Abstract
Comparing fragment based molecular fingerprints of drug-like molecules is one of the most robust and frequently used approaches in computer-assisted drug discovery. Molprint2D, a popular atom environment (AE) descriptor, yielded the best enrichment of active compounds across a diverse set of targets in a recent large-scale study. We present here BCL::Mol2D descriptors that outperformed Molprint2D on nine PubChem datasets spanning a wide range of protein classes. Because BCL::Mol2D records the number of AEs from a universal AE library, a novel aspect of BCL::Mol2D over the Molprint2D is its reversibility. This property enables decomposition of prediction from machine learning models to particular molecular substructures. Artificial neural networks with dropout, when trained on BCL::Mol2D descriptors outperform those trained on Molprint2D descriptors by up to 26% in logAUC metric. When combined with the Reduced Short Range descriptor set, our previously published set of descriptors optimized for QSARs, BCL::Mol2D yields a modest improvement. Finally, we demonstrate how the reversibility of BCL::Mol2D enables visualization of a ‘pharmacophore map’ that could guide lead optimization for serine/threonine kinase 33 inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ligand-based computer aided drug design (LB-CADD) relies on the observation that small molecule ligands often share a defined set of molecular features that promote molecular recognition of a ligand by a target protein—the so-called pharmacophore [1, 2]. While structurally unrelated chemotypes can represent the same pharmacophore, it is also correct that often molecules of similar structure share the pharmacophore required for targeting a protein. One advantage of LB-CADD methods is that the comparison of small molecule structures is independent of the knowledge of the three-dimensional structures of the target protein and its dynamics [3]. Two fundamental approaches of LB-CADD include similarity search and quantitative structure activity relationship (QSAR) models. While the former selects molecules that have similar structures to known actives, the latter infers a relationship between physicochemical properties of molecules and the bioactivity of interest and uses this relationship to select for molecules with high predicted output [4]. Due to its ability to rapidly screen libraries of compounds and significantly improve the discovery rate of actives, LB-CADD has become an increasingly popular in silico approach. A typical LB-CADD model is comprised of two major components: (1) a quantitative representation of chemical structures (descriptors) and (2) a similarity metric or, in the case of QSAR models, a mathematical function to compute bioactivity from these descriptors, often a machine-learning algorithm. While the former quantifies the similarity between input descriptors, the latter predicts bioactivity of compounds from the molecular descriptors [4].
Molprint2D is a 2D similarity search method based on atom environments (AE), which encode atomic properties, such as element types and bond types, of surrounding atoms within two bonds distance from the atom of interest (height = 2, Fig. 1) [5]. A largescale benchmark study of eight different 2D fingerprint methods has shown that Molprint2D fingerprint generated by the CANVAS software package yielded the best enrichments of active compounds on a diverse set of targets [6]. Each binary bit in a Molprint2D fingerprint only documents the presence or absence of a unique AE [5]. In the current work, we test the hypothesis that in addition to presence also the number of AEs could be important to distinguish substructures with similar AE composition (e.g., six-member rings vs. five-member rings).
The Molprint2D defines different AEs based on the element type of atoms bound up to two bonds away from the central atom of interest (height = 2). This description is highly overlapping as every atom will be represented in many AEs. We hypothesized that a more fine-grained list of AEs that includes hybridization state, i.e. electron configuration [7], in addition to element type but ventures only one bond around the atom of interest (height = 1) would provide a more information dense description of the AE. We set out to test this idea in the present work. Furthermore, Molprint2D generates AE set from the training dataset. We also hypothesized, that focusing on the most likely AEs in drug-like molecules can remove any bias from the training data. This AE library enables the model to be readily applicable to scaffold-hop into new chemical space and reduce the length of the descriptor vector. Thus, we generated a list of common AEs (i.e. the AE library) from a large database of over 900,000 drug-like compounds.
Artificial Neural Networks (ANN) are one of the most commonly used non-linear classifiers in QSAR models for LB-CADD due to their strong predictive power [8, 9]. We have previously shown that ANN-QSAR models outperformed fingerprint-similarity searches on Molprint2D descriptors [9]. However, their advantage in predictive capacity comes with the pitfall of their “black-box” nature [10]; it is difficult to map which structural features contribute to the activity. Previous efforts at interpreting QSAR models to aid molecular design used sensitivity analysis to rank importance of each descriptor on ANN training [11,12,13,14,15]. Yet the success of characterizing an ANNs’ internal function also depends on the nature of the descriptors used in the QSAR studies [10]. We hypothesize that fingerprint descriptors are particularly well-suited to interpretation when used for training an ANN as they are reversible; each input number refers to one specific structural motive. It is one goal of the present study to test this hypothesis.
In this paper, we introduce BCL::Mol2D, which significantly outperforms Molprint2D in predictive capacity and ANN interpretability. BCL::Mol2D documents the counts of common AEs, in which atoms are classified based on their element types and hybridization states (Fig. 2). There are two atomic encoding schemes for BCL::Mol2D descriptors: the ‘Element type’ enciphers atoms based on their atomic numbers and bond orders, while the ‘Atom type’ further distinguishes between elements with different orbital configurations [7]. ANNs, with drop-outs [16], trained on BCL::Mol2D descriptors perform significantly better than ANNs trained on Molprint2D descriptors. Moreover, we demonstrate the potential of BCL::Mol2D in the interpretation of ANN-QSAR models. The BCL::Mol2D descriptors are reversible from their numerical representation to the original chemical structures. Therefore, they allow extraction of the AEs that are crucial for optimization of ANN prediction of compound candidates. BCL::Mol2D descriptor method has been added to the BCL::Cheminfo package [17], which is free for non-commercial users. Finally, BCL::Mol2D are also combined with our previously-described best performing reduced short-range 3D descriptor set (BCL::3D-RSR) [9]. The resulting hybrid descriptor set modestly, albeit consistently, improve the performance of QSAR models.
Illustration of BCL::Mol2D element and atom type fingerprints of Phenol. a A Phenol molecule with atoms are colored based on their corresponding AEs: 5 carbon atoms inside the benzene ring (blue), 1 carbon in the benzene ring that also connect to the hydroxyl group (green), and 1 oxygen in the hydroxyl group (red). b The non-zero entries in BCL::Mol2D fingerprint with Element type (yellow-upper) or Atom type (grey-under) represent the counts of their corresponding unique AEs. Each entry stores the count of a unique AE, whose center atom is shown in bold. The location of an AE on the BCL::Mol2D fingerprint (shown in black number next to the black arrows) is determined by its prevalence in the AE library
Results
This study is comprised of a benchmark and sensitivity analysis to evaluate performance and functionality of BCL::Mol2D descriptors. BCL::Mol2D (height = 2) was first benchmarked against Molprin2D to examine the effects of changing the descriptor value from presence/absence to count of unique AEs. Then, we verified that decreasing the height of BCL::Mol2D from 2 to 1 would not significantly alter performance. The sensitivity analysis on BCL::Mol2D (height = 1) aims at estimating how alteration of a certain substituent affects its corresponding molecular prediction output. Finite differences of AEs were computed and mapped on the pharmacophore to signify potential impacts of adding or removing their corresponding substituents. In the final stage of the benchmark, we determined if adding BCL::Mol2D descriptor into the BCL::3D-RSR set improves its performance. Different descriptor configurations were evaluated through logAUC scores of trained QSAR-ANN models.
ANN-QSAR benchmarks on BCL::Mol2D in comparison to Molprint2D
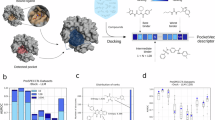
ANN-QSAR models were trained to compare the performance of BCL::Mol2D vs. Molprint2D [5, 6] across nine HTS PubChem datasets [18]. Table 1 summarizes the details of descriptor configurations and their average logAUC scores, and Fig. 3 illustrates the performance of those descriptors broken down into individual datasets. Following the design of Molprint2D, our initial implementation of BCL::Mol2D with the AE height of 2. While Molprint2D (height = 2) contains binary bits that record the presence/absence of Element type AEs, BCL::Mol2D documents counts of either Element or Atom type AEs. The logAUC scores of ANNs trained with either of those two descriptors were measured across nine HTS PubChem datasets. Compared to the performance of ANNs with Molprint2D, BCL::Mol2D significantly improved ANN predictive power up 23% for Atom type, and 26% for Element type (p-values < 0.01).
BCL::Mol2D (grey bars) outperforms Molprint2D (yellow bars) by 26.7%, and improves RSR (orange bars)’s performance by 6.8% when combined with RSR (blue bars) on average across nine PubChem HTS datasets. BCL::Mol2D descriptors are atom typed with height = 1. RSR + BCL::Mol2D are hybrid fingerprints from combining BCL::Mol2D (atom type, height = 1) and the BCL::3D-RSR descriptor set. Error bars represent SDs. Datasets are referred to by their PubChem assay IDs along the x-axis
Reducing the AE height from two bonds to one shrinks the size of the BCL::Mol2D fingerprint without reducing the QSAR performance
Since each BCL::Mol2D descriptors documents count of a unique AE, length of BCL::Mol2D fingerprint equals the size of the AE library. The AE library was built by collecting AEs that appeared more than 100 times among 900,000 drug-like small molecules. However, this common AE list contains several thousand AEs if the height is set to 2. The resulting fingerprints were likewise very sparse—less than 0.7% of all descriptor values were non-zero. We hypothesized that this fingerprint is unnecessarily large to encode even the most complex, drug-like, molecules with less than 100 unique AEs. Reducing the AE height to 1 reduces the length of the BCL::Mol2D fingerprint. It is 14-fold for Atom type and 20-fold for Element type (Table 1). The sparsity of the descriptors is also reduced—now up to 25% of all descriptor values are non-zero.
ANN-QSAR models were trained on BCL::Mol2D descriptors with either Atom or Element atom encoding scheme, which built AEs of either one or two-bond limit. The results (Table 1) suggested that reducing the bond limit in BCL::Mol2D had no significant effects on predictive power of the QSAR models with Atom type (+ 3.4%, p-value > 0.05), although doing so would moderately lower performance of BCL::Mol2D Element type fingerprints across nine HTS datasets (− 7.5%, p-value < 0.05). Compared to the logAUC scores of ANNs with Molprint2D, BCL::Mol2D (height = 1) still significantly improved ANN predictive power up 26.7% for Atom type (Fig. 3), and 16.5% for Element type (p-values < 0.01). Moreover, when combining the Atom and Element type, the resulting descriptor, BCL::Mol2D (Element + Atom, height = 1), show almost no performance difference when compared to that of Atom type counterpart, BCL::Mol2D (Atom, height = 1). A plausible explanation for this observation is that adding Element type to Atom type would not increase the useful information contents in the fingerprints because Atom type contains all the information of the Element type.
BCL::Mol2D moderately improves logAUC when combined with the BCL::3D-RSR set
The BCL::Mol2D (height = 1) descriptor was also tested in conjunction with a previously optimized descriptor set, BCL::3D-RSR, that utilizes a mix of 2D and 3D auto-correlation functions, along with scalar molecular descriptors [9]. More specifically, the combined sets modestly, though consistently, performed better than just the BCL::3D-RSR set alone. Combining the BCL::Mol2D (atom type, height = 1) with the BCL::3D-RSR descriptor set improves the logAUC from 0.385 to 0.406 (+ 6.8%, p-value < 0.01) with Atom type AEs, and to 0.411 (+ 5.5%, p-value < 0.01) with Element type AEs (Table 1). Additionally, combining the BCL::3D-RSR set with BCL::Mol2D consistently performed better than BCL::Mol2D alone. In particular, adding BCL::3D-RSR set improved the average logAUC score by 20.3% for Element type, and 12.0% for Atom type (p-values < 0.01) (Table 1; Fig. 3). Means, standard deviations, and 95% confident intervals of logAUC scores from ANN-QSAR models trained on all descriptor configurations mentioned above are summarized in the supplementary Table S1.
Detection of hot spots on scaffold through sensitivity analysis of the ANNs
Unlike Molprint2D descriptors, BCL::Mol2D descriptor values correspond to the counts of molecular substructures that they represent. Hence, we can estimate the effects of adding or removing a certain substituent based on sensitivity analysis of the AEs that the substituent encompasses. Discrete derivatives of molecular ANN prediction output were computed for each unique AE when adding (increment derivative) or removing (decrement derivative) that AE. Eight pairs of an active and an inactive with less than 10% difference in structure (according to Tanimoto index [19]) were selected for this analysis as described in the “Methods” section. We investigated whether the ANN model can predict which structural differences between those active and inactive compounds cause significant differences in their bioactivity.
The decrement derivatives of AEs are mapped onto their corresponding atom for each of 16 compounds in the analysis (Fig. 4, first column). The results show that removal of substructures with positive decrement derivatives (blue region) often boost the ANN predicted activity, while addition of the ones with negative decrement derivatives (red regions) leads to a decreased expected activity of the compound. Atoms colored white indicate close to 0 values of their corresponding AE derivatives (− 0.01 to 0.01).The corresponding figures for seven of the eight actives illustrate that altering the blue regions while keeping red regions intact lead to higher ANN prediction output, with only the 3rd scaffold appearing to contradict this conclusion. The transformation from inactive 3 to active 3 is the only one that involved removal of an AE with negative decrement derivative. The change to the hydroxyl indeed made a relatively small change in ANN prediction output, merely increasing the compound ANN prediction by 0.27. However, this increase was enough to move it from a strongly ANN prediction output inactive to a weakly ANN prediction output active, because 99% of inactives have an ANN prediction output below 0.1.
Mapping partial contributions of AEs to the ANN prediction output of STK33 inhibitors using BCL::Mol2D (Atom type, height = 1). The first column contains general structures of two pairs of compounds (one active and one inactive) with their corresponding ANN predicted activities. The atoms that are different between active and inactive compounds are colored in green (green rectangles). The second and third columns illustrate the transformation from active to inactive and from inactive to active, respectively. The directions of the transformation are shown in black arrows. The atoms that are highlighted in green are colored based on the finite differences of their corresponding AEs. Red circles mean negative values, and blue circles have positive values. The decrement derivatives (marked with minus signs) are represented on the deleted substructures, and the increment derivatives (marked with plus signs) are represented on added substructures in each transformation. Additional examples are reported in Fig. S5
To identify the type of changes in molecular structures that significantly affect their ANN prediction output, sums of increment and derivatives were computed for added and removed AEs, respectively for transformation from inactive to active, and from active to inactive. Generally, transformation from inactive to active compounds replaced AEs, whose sum of decrement derivatives is positive, by AEs with positive sum of increment derivatives (Fig. 4, second column). Again, in the case of the scaffold 5, the benefit of adding the chloride groups on the inactive 5 (with the sum of increment derivatives of 0.727) might outweigh the effect of the removed the carbons (decrement derivative of − 0.043). A similar trend is shown in the transformation from active to inactive compounds. If the relation between structure and activity were linear, one could expect decrement and increment derivatives to always have opposite signs and similar values. To test this hypothesis, we correlated increment and decrement derivatives of the 16 STK inhibitors used in the sensitivity analysis (Fig. S9). With a low \({R}^{2}\) value (0.13), this result suggests a non-linear dependency.
A case study of applying lead optimization through derivatization
We illustrate here an example of applying knowledge from the sensitivity analysis of the BCL::Mol2D descriptors on derivatization to improve the ANN prediction output (Fig. 5). From an inactive STK inhibitor (the inactive compound of the compound pair 5 from the sensitivity analysis, Fig. S5), we have created a new compound with improvement in ANN prediction output from 0.42 to 1.0 after two steps of modification. In each step, we manually select the added and removed functional group with positive decrement and increment derivatives, respectively. One exception is the added in aromatic carbon atom the second step, which has a negligible increment derivative. However, the transformation in step 2 still improves the ANN prediction output of the compound because the effect of removing the chloride group (decrement derivative sum = 0.12) outweighs the impact of adding the aromatic carbon (increment derivative = − 0.01). The values of removed and added AEs for each step are listed in the Table S3.
Applying sensitivity analysis of BCL::Mol2D descriptor to lead optimization through derivatization. Starting from the inactive compound from the STK inhibitor HTS, we remove functional groups with favorable decrement (marked with black minus signs) and add functional groups with favorable increment (mark with black plus signs) derivatives. The process results in a known active compound much higher ANN prediction output (denoted by black numbers on the most left side). The substructures that are modified in each step are labeled and framed in green, and colored based on the decrement derivatives of their corresponding AEs (positive: blue; negative: red). Between molecular structures: added or removed substructures in each step are framed according to the sum of increment and decrement derivatives, respectively (blue: positive value, red: negative value)
Discussion
In this study, BCL::Mol2D descriptors were tested using an established QSAR benchmark of nine large high-throughput screens to ensure general applicability of the method. We observe consistent improvements in logAUC scores over all datasets when the QSAR models when trained with BCL::Mol2D instead of Molprint2D, even though the height of AEs of BCL::Mol2D was reduced from two to one. This observation suggests that 2D information of an additional layer of neighboring atoms fails to improve the useful informational content to the ANNs. Interestingly, performance of Element type BCL::Mol2D fingerprints, although with fewer AEs, is significantly higher than performance of Molprint2D. This improvement suggests that counts of unique AEs, even with a height of one, provide more information than just their presence/absence with height of two, like Molprint2D, perhaps by helping distinguish substructures with similar AE composition. Adding electron configuration of atoms, which further differentiates AEs with the same element type, was also shown to improve performance of the fingerprints.
Comparing the QSAR-ANN performance of BCL::Mol2D descriptors at height of one and two, we notice that reducing height from two to one improves performance of Atom type BCL:: Mol2D descriptor but worsens the performance of the Element type counterpart. One explanation for those two contrast behaviors is that the electron configuration encoded in the Atom hash already contains valuable information regarding bond types and electron hybridization of the neighboring atoms that are two bonds away from the center atom. In contrast, Element type AEs at a height of one lacks this hybridization information. As a result performance of the corresponding Element type AE BCL::Mol2D descriptors suffers.
Combining BCL::Mol2D and the BCL::3D-RSR descriptors sets yields a modest, but consistent performance improvement over the optimized BCL::3D-RSR set alone. This suggests possible partial information overlap between two descriptor sets. Future studies could consider performing descriptor selection analysis on the hybrid fingerprints to prune out the descriptors that do not provide meaningful information to the model.
Since the values of BCL::Mol2D descriptors directly relates to atomic fragments of the molecular structures, derivatives of individual AEs can be used to estimate the effects of removal/addition of functional groups on the scaffolds. Previous studies [2, 15, 20] have attempted to estimate/rank the global importance of descriptors based on their partial derivatives. However, since we are interested in extracting information from the ANNs to optimize specific scaffolds, we only focused on effects of changes in local substructures to a specific prediction. Furthermore, as values BCL::Mol2D descriptors are discrete integers, their decrement and increment “finite differences” are likely to be different than those computed using a traditional continuous derivative calculation. Hence, using these two types of finite differences would distinguish the effects of removing and adding a particular AE in scaffold optimization. Carlson et al. suggested that changes of important regions (with high sensitivity scores) would alter the ANN prediction output of that molecules [2]. However, equivocating descriptor importance with their partial first derivatives for a molecule is not always meaningful because descriptor values could have a partial first derivative of zero while retaining a large second derivative when they are at local optima. Likewise, we did not measure the centered first derivatives of the ANN with respect to descriptors in order to rank their importance, instead looking at discrete increments or decrements of the descriptor.
We propose that AEs with positive decrement derivatives should be replaced by AEs with positive increment derivatives to improve the ANN prediction output. We tested this proposal by applying the sensitivity analysis of BCL::Mol2D on transform an inactive STK inhibitor to a novel compound with more than substantial improvement in ANN prediction output. We hence demonstrated that we can use BCL::2D descriptors to leverage knowledge from the QSAR-ANN models to optimize lead compounds. Although in this study, we only focus on the derivatization aspect of the lead optimization, performing more central and dramatic modifications on the scaffolds should be possible, though the success of such an approach will depend heavily on whether the training data contained molecules with similar structures.
Conclusion
We present the BCL::Mol2D molecular descriptor that significantly improves both predictive power and interpretability of the ANN-QSARs compared to Molprint2D and our previous-best descriptor set. We further illustrate how BCL::Mol2D can be used to identify potential modification of a given inactive molecule to improve its ANN prediction output. Therefore, ANN-QSAR models trained on BCL::Mol2D could be employed in conjunction with a Monte Carlo or genetic algorithm as a structure generator [4, 21] to automate the process of rational combinatorial drug-like molecule design. The sensitivity analysis on BCL::Mol2D can guide medicinal chemists in the design of focused libraries [22] to optimize new derivatives by filtering out unfruitful scaffold modification. This will potentially reduce the number of compounds for synthesis and testing in drug discovery campaigns.
Methods
Data curation
A previously established QSAR benchmark of nine HTS datasets (Table 2) was used to evaluate performance of ANNs. These datasets are comprised of compounds from HTS scans on eight protein targets: two class A G-protein coupled receptor (GPCRs), three ion channels, one transporter, one kinase, and one enzyme. To address the concern regarding the quality of PubChem data, as previously detailed [18, 23, 24], molecules were labeled as active compounds only if their activity was verified in follow up confirmatory assays, selectivity assays, and dose response experiments, reducing significant amount of false positives in the datasets. Each data set contained more than 170 active and 61,000 inactive compounds. Three-dimensional conformations were generated with Corina version 3.60 [25], with the driver options of adding hydrogens (wh) and removing molecules from which the software could not generate 3D structures from (r2d).
Generation of descriptors
BCL::Mol2D
To generate each AE, one of two atomic encoding schemes (Element or Atom) is assigned. All neighbor heavy atoms that are up to either two bonds (e.g. AE height = 2) or one bond away (AE height = 1) from the central atom are included in each AE. The shorter AE height of one was tested to see whether an increase in information density was beneficial for ANN training. Atom type and Element type AE libraries contain AEs were generated from in-house database of 900,000 drug-like small molecules, and with more than 100 counts. There are 574 AEs in the Atom type AE library and 240 in the Element type counterpart when AE height is set to 1. The BCL::Mol2D fingerprints are then generated to document counts of AEs in the AE library (Fig. 2).
Molprint2D
The descriptors were generated with ElemRC atom types using the Schrodinger Canvas software suit, consistent with the optimal settings in the 2D fingerprint benchmark [6]. The AEs that appear from 1 to 90% molecules of each PubChem dataset were selected. The length of the Molprint2D ranges from 300 to 334 across the nine datasets.
Reduced SR (BCL::3D-RSR) descriptor set
This is a shortened version of the short range (SR) descriptor set introduced in a precedent study [9]. The SR, containing 1315 descriptors in total, calculates six atomic properties for both signed and unsigned 2D/3D autocorrelation descriptors [23, 26]. The BCL::3D-RSR set reduces the number of descriptors down to 391: 23 scalar, 132 short range 2DA_Sign and 240 3DA_Sign descriptors (Supplementary Table S1). Most of reduction is a result of using only signed versions of 2D and 3D autocorrelation (2DA_Signed and 3DA_Signed) [23] and only four atomic properties are calculated.
Hybrid fingerprint of BCL::3D-RSR set and BCL::Mol2D descriptor
Descriptors from the BCL::3D-RSR set and BCL::Mol2D descriptors (height = 1) were combined to create hybrid fingerprints. BCL::Mol2D(Atom) + BCL::3D-RSR hybrid fingerprints comprise of 965 descriptor values, while BCL::Mol2D(Element) + BCL::3D-RSR hybrid fingerprints contain 631 descriptor values.
ANN-QSAR model training and evaluation
The performance of artificial neural network (ANN)—QSAR models using BCL::Mol2D descriptors was compared with those with the Molprint2D on each of the nine datasets. All ANN-QSAR models were trained with simple back propagation using a sigmoid transfer function with η = 0.05 and α = 0.5. The architecture of the ANNs consisted of a single hidden layer of 32 neurons and drop-out rates [9, 16] of 0.05 for visible (input-layer) neurons, and 0.25 for hidden neurons, as previously optimized, with full connectivity to the input and output layer of the ANN The reported results were the evaluation of ANN predictions on independent test sets, which have no overlap with the training sets. Each ANN training was trained for 100 iterations without early stopping, which we previously found unnecessary when dropout is used [9].
QSAR models were evaluated with logAUC [27], which is area under the curve of the logarithmic receiver operating characteristic curve (logROC) between false positive rates of 0.001 to 0.1 [9]. We also computed full AUCs of the ROC curves. Each QSAR experiment was bootstrapped with replacement 2000 times to obtain logAUC and AUC mean and confident intervals using BCL v3.5, model:ComputeStatistics application. Average logAUC and AUC, and their standard deviations (SDs) were computed across the nine HTS datasets for each descriptor condition. The SDs of mean metric values across nine PubChem datasets were computed as \({\sigma }_{\mu }=\sqrt{\frac{{\sum }_{i=1}^{9}{\sigma }_{i}^{2}}{{9}^{2}}}\), where \({\sigma }_{i}^{2}\) is the variance of the metric of the dataset \(i\) [28].
Two tailed two-sample t-test was then conducted to compare the average logAUC of different descriptor configurations. The p-values were computed from the Student’s paired t-test to compare pairwise average logAUC scores across nine different PubChem datasets for each pair of descriptor configurations. The standard deviation of the logAUC for each dataset was not used in these calculations. In each dataset, the active compounds were duplicated during ANN training such that for each ten inactive compounds that are in the training set and presented to the ANN, an active is presented (A:Iratio = 0.1).
Cross-validation
Five-fold cross-validation was used throughout the evaluation of the QSAR models. After each of the nine PubChem datasets was randomized, it is split into five parts. The ANN was trained on four of the parts (e.g. the training set), and independently tested on the last part (e.g. the test set). Subsequent ANNs are then trained holding out a disparate fifth of the dataset as the test set from the training. This process is repeated (5×) until each fifth of the dataset has been used as the independent test set for one model. The final performance metrics are averaged across the resulting predictions on the five test sets, which covers the complete dataset.
Sensitivity analysis
We used the QSAR model trained on the dataset 2689 [18], which contains compounds from bioassays scanning for inhibitors of serine/threonine kinase 33 (STK33) [29] because this model yielded highest logAUC score. Each BCL::Mol2D descriptor, which corresponds to a specific substructure of the molecule, was evaluated for output sensitivity, i.e. the change in ANN output resulting from of adding or removing each individual atom environment. Sensitivity score, \({ S}_{o,{d}_{i}}\), of a descriptor \({d}_{i}\) for a molecule m, was defined to be the discrete derivative of the output \(f\) with respect to the value of that descriptor, and was calculated as
where \(\delta\) is the change applied to \({d}_{i}\). \({S}_{f\left({d}_{i}\right),{d}_{i}}^{m}\) is referred to as the decrement derivative when \(\delta\)is − 1 and increment derivative when \(\delta\) is 1, so as to approximate the change in ANN output from adding or removing discrete AEs.To investigate the effects of removing and adding different AEs on altering the ANN prediction output of a structure, we selected eight actives for which there was a corresponding inactive molecule in the dataset with at least 90% of substructure in common. ANN prediction output (ANN output) of the active compounds are greater than 0.95 and that of the inactive compounds is lower than 0.60. PubChem IDs and ANN prediction output of 16 compounds used in the sensitivity analysis are included in Table S4.
Electronic supplementary material
Protocols of performing benchmark and sensitivity analysis are included in this GitHub repository: https://github.com/vuoanh/BCL_Mol2D_benchmark.
Supplementary tables S1–S4. List of descriptors in the BCL::3D-RSR set (Table S1); Average, standard deviation (SD), and 95% confidence interval (CI) of logAUC scores of different descriptor configurations (atom hash and height) across nine PubChem datasets (Table S2); Average, standard deviation (SD), and 95% confidence interval (CI) of logAUC scores of different descriptor configurations across nine PubChem datasets. Atom-1bond is BCL::Mol2D descriptors with atom type and height of 1. (Element + Atom)-1bond is BCL::Mol2D descriptors with combination of atom and element type with height of 1 (Table S3); PubChem IDs and ANN prediction output of 16 compounds used in the sensitivity analysis (Table S4). (PDF)
Supplementary files list of unique AEs that are sorted based on their prevalence in the AE library (Atom_AE_1_bond_sorted, Element_AE_1_bond_sorted).
Software
Descriptor generation and QSAR model training were performed using the BCL::Cheminfo package, which is free of charge for non-commercial use. For more information of BCL::Cheminfo, please visit its webpage: http://meilerlab.org/qsar_benchmark_2015. Different applications of the package can be downloaded at: http://meilerlab.org/index.php/servers/bcl-academic-license.
Abbreviations
- AE:
-
Atom environment
- ANN:
-
Artificial neural network
- AUC:
-
Area under curve
- BCL:
-
BioChemical Library
- CADD:
-
Computer aided drug discovery
- LB-CADD:
-
Ligand based computer aided drug discovery
- QSAR:
-
Quantitative structure–activity relationship
- RSR:
-
Reduced short range
References
Kim KH, Kim ND, Seong BL (2010) Pharmacophore-based virtual screening: a review of recent applications. Expert Opin Drug Discov 5(3):205–222
Carlsson L, Helgee EA, Boyer S (2009) Interpretation of nonlinear QSAR models applied to ames mutagenicity data. J Chem Inf Model 49(11):2551–2558
Cramer RD (2012) The inevitable QSAR renaissance. J Comput Aided Mol Des 26(1):35–38
Sliwoski G, Kothiwale S, Meiler J, Lowe EW Jr. (2014) Computational methods in drug discovery. Pharmacol Rev 66(1):334–395
Bender A, Mussa HY, Glen RC, Reiling S (2004) Similarity searching of chemical databases using atom environment descriptors (MOLPRINT 2D): evaluation of performance. J Chem Inf Comput Sci 44(5):1708–1718
Sastry M, Lowrie JF, Dixon SL, Sherman W (2010) Large-scale systematic analysis of 2D fingerprint methods and parameters to improve virtual screening enrichments. J Chem Inf Model 50(5):771–784
Gasteiger J, Marsili M (1980) Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron 36(22):3219–3228
Montañez-Godínez N, Martínez-Olguín AC, Deeb O, Garduño-Juárez R, Ramírez-Galicia G (2015) QSAR/QSPR as an application of artificial neural networks. In: Cartwright H (ed) Artificial neural networks. Springer, New York, pp 319–333
Mendenhall J, Meiler J (2016) Improving quantitative structure–activity relationship models using Artificial Neural Networks trained with dropout. J Comput Aided Mol Des 30(2):177–189
Cherkasov A, Muratov EN, Fourches D, Varnek A, Baskin II, Cronin M et al (2014) QSAR modeling: where have you been? Where are you going to? J Med Chem 57(12):4977–5010
Tetko IV, Tanchuk VY, Chentsova NP, Antonenko SV, Poda GI, Kukhar VP et al (1994) HIV-1 reverse transcriptase inhibitor design using artificial neural networks. J Med Chem 37(16):2520–2526
Tetko IV, Villa AE, Livingstone DJ (1996) Neural network studies. 2. Variable selection. J Chem Inform Comput Sci 36(4):794–803
Guha R, Stanton DT, Jurs PC (2005) Interpreting computational neural network quantitative structure-activity relationship models: a detailed interpretation of the weights and biases. J Chem Inform Model 45(4):1109–1121
Guha R, Jurs PC (2005) Interpreting computational neural network QSAR models: a measure of descriptor importance. J Chem Inform Model 45(3):800–806
Marcou G, Horvath D, Solov’ev V, Arrault A, Vayer P, Varnek A (2012) Interpretability of SAR/QSAR models of any complexity by atomic contributions. Mol Inform 31(9):639–642
Nitish Srivastava GH, Krizhevsky A, Sutskever I, Salakhutdinov R (2014) Dropout: a simple way to prevent neural networks from overfitting. J Mach Learn Res 15(1):1929–1958
Butkiewicz M, Lowe EW, Meiler J, Bcl∷ChemInfo—Qualitative analysis of machine learning models for activation of HSD involved in Alzheimer’s Disease. 2012 IEEE Symposium on Computational Intelligence in Bioinformatics and Computational Biology (CIBCB); 9–12 May 2012
Butkiewicz M, Lowe EW Jr, Mueller R, Mendenhall JL, Teixeira PL, Weaver CD et al (2013) Benchmarking ligand-based virtual high-throughput screening with the PubChem database. Molecules 18(1):735–756
Rogers DJ, Tanimoto TT (1960) A computer program for classifying plants. Science 132(3434):1115–1118
Baskin II, Ait AO, Halberstam NM, Palyulin VA, Zefirov NS (2002) An approach to the interpretation of backpropagation neural network models in QSAR studies. SAR QSAR Environ Res 13(1):35–41
Meiler J, Will M. Genius (2002) A genetic algorithm for automated structure elucidation from 13C NMR Spectra. J Am Chem Soc 124(9):1868–1870
Zheng W, Cho SJ, Tropsha A (1998) Rational combinatorial library design. 1. Focus-2D: a new approach to the design of targeted combinatorial chemical libraries. J Chem Inform Comput Sci 38(2):251–258
Sliwoski G, Mendenhall J, Meiler J (2016) Autocorrelation descriptor improvements for QSAR: 2DA_Sign and 3DA_Sign. J Comput Aided Mol Des 30(3):209–217
Butkiewicz M, Bryant SH, Lowe EW Jr., David C, Meiler J (2017) High-throughput screening assay datasets from the PubChem database. Chem Inform 3(1):1
Gasteiger J, Teckentrup A, Terfloth L, Spycher S (2003) Neural networks as data mining tools in drug design. J Phys Org Chem 16(4):232–245
Pierre Broto GM, Vandycke C (1984) Molecular structures: perception, autocorrelation descriptor and SAR studies. Autocorrelation descriptor. Eur J Med Chem 19(1):66–70
Mysinger MM, Shoichet BK (2010) Rapid context-dependent ligand desolvation in molecular docking. J Chem Inf Model 50(9):1561–1573
Weisstein E (2000) Normal sum distribution: Wolfram Research, Inc. http://mathworld.wolfram.com/NormalSumDistribution.html
Liao Z, Thibaut L, Jobson A, Pommier Y (2006) Inhibition of human tyrosyl-DNA phosphodiesterase by aminoglycoside antibiotics and ribosome inhibitors. Mol Pharmacol 70(1):366
Krylov A, Windus TL, Barnes T, Marin-Rimoldi E, Nash JA, Pritchard B et al (2018) Perspective: computational chemistry software and its advancement as illustrated through three grand challenge cases for molecular science. J Chem Phys 149(18):180901
Wilkins-Diehr N, Crawford TD, NSF’s Inaugural Software Institutes (2018) The science gateways community institute and the molecular sciences software institute. Comput Sci Eng 20(5):26–38
Acknowledgements
This study is funded by Molecular Science Software Institute (MolSSI) [30, 31] Fellowship and NIH. MolSSI is funded by the NSF Grant (ACI-1547580). Work in the Meiler laboratory is supported through NIH (R01 GM099842, R01 DK097376) and NSF (CHE 1305874). The author would like to thank Dr. Francois Berenger for discussion regarding descriptor design.
Author information
Authors and Affiliations
Contributions
OV, JM and JM designed the study. OV implemented the descriptor, performed the benchmark and analysis, and wrote the manuscript. JM and DA supervised the project. JM, DA and JM edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10822_2019_199_MOESM4_ESM.pdf
Supplementary material 4: Mapping partial contributions of AEs to the ANN prediction output of STK33 inhibitors using BCL::Mol2D (Atom type, height = 1) (Fig. S5). (PDF 481 KB)
10822_2019_199_MOESM5_ESM.pdf
Supplementary material 5: Linear regression between increment and decrement derivatives of 16 STK inhibitors used in the sensitivity analysis. The formula and the value are denoted in red next to the red trend line (Fig. S6). (PDF 14 KB)
Rights and permissions
About this article
Cite this article
Vu, O., Mendenhall, J., Altarawy, D. et al. BCL::Mol2D—a robust atom environment descriptor for QSAR modeling and lead optimization. J Comput Aided Mol Des 33, 477–486 (2019). https://doi.org/10.1007/s10822-019-00199-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-019-00199-8