Abstract
Purpose
Gene expression analysis of the endometrium has been shown to be a useful approach for identifying the molecular signatures and pathways involved in recurrent implantation failure (RIF). Nevertheless, individual studies have limitations in terms of study design, methodology and analysis to detect minor changes in expression levels or identify novel gene signatures associated with RIF.
Method
To overcome this, we conducted an in silico meta-analysis of nine studies, the systematic collection and integration of gene expression data, utilizing rigorous selection criteria and statistical techniques to ensure the robustness of our findings.
Results
Our meta-analysis successfully unveiled a meta-signature of 49 genes closely associated with RIF. Of these genes, 38 were upregulated and 11 downregulated in RIF patients’ endometrium and believed to participate in key processes like cell differentiation, communication, and adhesion. GADD45A, IGF2, and LIF, known for their roles in implantation, were identified, along with lesser-studied genes like OPRK1, PSIP1, SMCHD1, and SOD2 related to female infertility. Many of these genes are involved in MAPK and PI3K-Akt pathways, indicating their role in inflammation. We also investigated to look for key miRNAs regulating these 49 dysregulated mRNAs as potential diagnostic biomarkers. Along with this, we went to associate protein–protein interactions of 49 genes, and we could recognize one cluster consisting of 11 genes (consisted of 22 nodes and 11 edges) with the highest score (p = 0.001). Finally, we validated some of the genes by qRT-PCR in our samples.
Conclusion
In summary, the meta-signature genes hold promise for improving RIF patient identification and facilitating the development of personalized treatment strategies, illuminating the multifaceted nature of this complex condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human endometrium is not receptive to embryonic implantation during most of the menstrual cycle; however, it becomes receptive for a period of 2 to 4 days within the mid-secretory phase of the menstrual cycle known as the window of implantation [1, 2]. Therefore, an embryo and endometrium must communicate with one another in synchrony and coordination for implantation to be successful [3]. With breakthroughs in laboratory techniques and ovarian stimulation over the last few decades, in vitro fertilization-embryo transfer (IVF-ET) has grown into an effective therapy for infertility. Nevertheless, it is estimated that 10% of women undergoing IVF will experience recurrent implantation failure (RIF) which is defined as the failure to achieve a clinical pregnancy after two or more IVF cycles with the transfer of at least four good-quality embryos [4,5,6]. RIF is a challenging problem in the field of reproductive medicine, as it is associated with significant emotional, psychological, and financial burden for patients [7, 8]. The causes of RIF can be multifactorial and include both maternal and embryonic factors. However, abnormalities in endometrial receptivity are increasingly recognized as one of the key contributors to RIF [9,10,11]. The molecular mechanisms underlying RIF are complex and not yet fully understood. We know that several factors affect endometrial receptivity, including hormonal imbalances, inflammation, and immune system dysregulation [12, 13]. Abnormalities in the endometrial thickness, pattern, and vascularization can also affect the success of implantation [14]. To improve endometrial receptivity and increase the success of IVF, various interventions have been proposed, such as endometrial scratching, hysteroscopy, immunomodulatory therapy, and transcriptomics.
One of the most promising approaches to identifying molecular signatures associated with RIF is the use of transcriptomics, a high-throughput technique that allows for the simultaneous analysis of thousands of genes [15, 16]. Studies have used microarray or RNA sequencing technology to profile the endometrial gene expression of patients with RIF [17] and identify differentially expressed genes and pathways for successful implantation [18, 19]. For instance, genes related to inflammation, immune response, and angiogenesis have been found to be upregulated in patients with RIF compared to controls [20,21,22,23]. Gene expression analysis of the endometrium is a useful approach for identifying the molecular pathways involved in RIF. However, individual studies may have limited power to detect insignificant changes in gene expression levels or identify novel gene signatures associated with RIF [24].
In this context, meta-analysis, which involves the integration of data from multiple studies, can provide a more comprehensive and robust analysis of gene expression patterns in RIF. Meta-analysis approaches have been widely used in genetic and genomic studies to identify gene expression signatures associated with various diseases and conditions [25, 26]. In recent years, meta-analysis approaches have also been employed to investigate the gene expression patterns associated with RIF [27]. These studies have combined gene expression data from multiple studies to identify common patterns of gene expression associated with RIF. In a more recent meta-analysis study, Zhao and co-workers conducted a comprehensive analysis of microarray gene expression data from 3 studies that investigated the endometrial gene expression patterns in women with RIF. The authors validated a set of 8 cellular senescence-associated differentially expressed genes that were consistently dysregulated in women with RIF [28]. Meta-analysis studies have also been used to investigate the effect of hormonal treatments on gene expression patterns in the endometrium of women with RIF [29,30,31]. For example, a meta-analysis study of endometrial gene expression data from women treated with gonadotropin-releasing hormone (GnRH) agonists identified several differentially expressed genes involved in cell cycle regulation and DNA damage response [32]. Although each study produces a set of genes, the overlap between different studies is limited. The limitations of this technology are widely recognized and arise from variations in experimental design, timing, and circumstances of endometrial sampling, as well as patient selection criteria. Additionally, variations in transcriptome array/sequencing platforms, genome annotation versions, and data processing pipelines contribute to these limitations [33,34,35].
While there are only a few meta-analysis studies investigating gene expression patterns, it’s important to highlight their limited sensitivity to refractory conditions such as repeated implantation failure (RIF). To overcome the limitations in endometrial transcriptome analyses, we employed a robust systematic analysis method, and subsequently conducted functional analysis to identify a meta-signature of highly probable biomarkers associated with RIF. This specific study compiles findings from nine research articles conducted globally by diverse groups, with a specific focus on recurrent implantation failure (RIF) on endometrial receptivity. Despite the extensive lists of genes provided by all the studies, our effort has been concentrated on narrowing down the gene numbers to gain a better understanding of the RIF pattern. We also analyzed potential microRNAs that could affect the genes/mRNAs associated with RIF.
In addition, our objective was to experimentally confirm the expression levels of the selected mRNA genes identified through our meta-analysis. To achieve this, we conducted experiments using our own set of samples to validate the findings from the in silico analysis. This experimental validation step adds a crucial layer of confidence to our results, ensuring the dependability and reliability of the gene expression patterns observed in our study.
Materials and methods
Systematic search of the literature
A systematic review of the literature in PubMed, Scopus, Google Scholar, MEDLINE and Embase was independently conducted from January 2018 up to December 2022. The terms ‘embryo implantation’, ‘endometrium’, ‘gene expression’ and ‘Recurrent implantation failure’ were used individually and combined with the Boolean operator ‘AND’. The reference lists of review articles and relevant original studies were explored in-literature to include other appropriate studies. We followed steps as described in the PRISMA 2020 [36] flow chart for new systematic reviews which included searches of databases, registers, and other sources. The study protocol was registered in PROSPERO under the registration number CRD42023445555.
Study review and eligibility criteria
The search retrieved all identified abstracts, which were all examined to determine which studies were eligible. The entire text of each pertinent article was meticulously evaluated. For the final analysis, only unique experimental papers published in English that addressed the endometrial transcriptome in women undergoing Assisted Reproductive Treatment (ART) in the mid-secretory phase were considered. The following inclusion and exclusion criteria were employed for selecting studies for meta-analysis: research involving patients undergoing Assisted Reproductive Treatment (ART) cycles who experienced at least two implantation failures; investigations on the relationship between control-pregnancy positive results and outcomes in patients with Recurrent Implantation Failure (RIF). No limitation was set on the minimum number of patients in each study. If multiple articles were using the same patient dataset by the same research group, only the recent relevant article was considered. Endometrial transcriptome analysis in connection with any pathological condition, such as, endometriosis, adenomyosis, fibroids, hydrosalpinx and cancer was excluded. Additionally, gene expression analyses focusing on different endometrial tissue-sections of normal individuals were excluded in this meta-analysis.
Data extraction and quality assessment
After full text screening, the quality of each included study was assessed for data availability on databases and primary sources. Available raw data was retrieved from ArrayExpress database (https://www.ebi.ac.uk/arrayexpress/) and the Gene Expression Omnibus database (https://www.ncbi.nlm.nih.gov/geo/). Additionally, the lists of genes differentially expressed in control and RIF in mid-secretory endometrium were extracted from the selected publications.
Data analysis settings
The acquired.CEL files of Affymetrix platform and.TXT files of Agilent platform were imported into GeneSpring version 14.9.1 GX-PA software (Agilent technologies). Data from both the platforms were analyzed separately. For 3 of the studies, gene lists were considered for final compilation of data. A differentially regulated gene list common between the studies was pooled for additional analysis. The final acquired gene list was converted to ENTREZ IDs by using the DAVID Gene ID converter Tool. The default statistical analysis options for all the studies for gene list acquisition (false discovery rate, FDR < 0.05; Fold Change, FC > 2.0).
Enrichment analysis
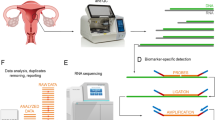
Enrichment analyses for Gene Ontology (GO) terms and Kyoto Encyclopedia of genes and genomes (KEGG) for biological pathways were carried out by using two tools, g:Profiler web tool (biit.cs.ut.ee/gprofiler/) and GeneSpring. We also used miRNA that targets our mRNA genes by easy-to-use web tool MIENTURNET (MicroRNA ENrichment TURned NETwork- http://userver.bio.uniroma1.it/apps/mienturnet/) [37]. They provide a graphical and tabular output. Additionally, both these platforms enable the user to view each detailed pathway diagram highlighting the number of entities in a particular pathway. The obtained results were corrected by the default setting provided by these platforms, unless mentioned in detail in the results. STRING, Protein–Protein Interaction Networks Functional Enrichment Analysis (https://string-db.org/) was used to assess protein–protein interaction (PPI) information with its basic settings and particularly focused on 3 clusters using k-means clustering options.
Validation of meta-analysis genes by RT-qPCR
Out of 49 genes, representative genes of major biological processes controlling endometrial receptivity like immune response, response to stress, defense response, response to external stimulus, cell cycle, cell adhesion, anatomical structure development, cell–cell signaling, and receptor binding were selected to check them in our RIF patients. We selected five genes (3 downregulated and 2 upregulated), CTNNA2↑, GADD45A↓, LIF↓, PPP1R1A↑ and SMCHD1↓ and a housekeeping gene GAPDH (Primer sequence in Table 1). Regulation of selected genes in RIF patients was performed using qRT-PCR. For this study, a total of 10 samples were selected, including 5 individuals with RIF and 5 control (Pregnancy Positive) samples. The aim was to investigate the regulation of these specific genes in our samples and determine if there are any differences in gene expression identified in meta-analysis between the RIF patients and the control group.
Results
Systematic search of the literature
A literature search turned up 1381 items, but 386 were disregarded since they were duplicates in different databases. From the remaining 995 articles, 896 articles were excluded after screening their titles and abstracts due to research carried out on non-human samples. The full text of the remaining 99 articles was assessed for eligibility, resulting in 9 of them being included in the meta-analyses based on criteria like a study on RIF patients, availability of full text articles, retrieval of full gene list and accessibility of file formats for Genespring software (Fig. 1). Most selected studies on endometrial receptivity markers in the context of IVF between the RIF and Control were assessed using microarray. These studies involved 492 women with mid-secretary phase endometrium from various countries (Table 2). Overall, the study quality was moderate, with excellent scores for participant selection and follow-up but low scores for cohort comparability. Almost in every study, the RIF group consisted of women who had more than three good quality embryos that fail repeatedly, whereas the control group consisted of women who had a successful pregnancy.
PRISMA 2020 Flow diagram showing the process to obtain information for the meta-analysis [36]. *RIF-Recurrent Implantation Failure
Data analysis
Due to the computational limitations of functional enrichment analyses of all studies from raw data, our analysis was restricted to 6 studies from raw data and 3 studies pooled gene list. We identified a statistically significant meta-signature of 49 genes of which 38 up-regulated and 11 down-regulated genes in mid-secretory endometrium between control and RIF patients (Table 3). Most significantly, differentially expressed genes identified by meta-analysis were statistically computed to p-value < 0.05 and standardized fold change > 2. The thirty-eight up-regulated genes in RIF were ABLIM3, ANK3, BIRC3, BTNL9, CPT1A, CTNNA2, FLT4, GDF15, GNAT1, GPR52, GPRC5C, IGFN1, IL2RA, KCNMA1, KLRC1, MC3R, MUC17, MUC22, NANOS1, NNMT, NTRK2, PAX7, PDPR, PHF8, PLXNA4, PPP1R1A, RANBP17, SAMD12, SGSM1, SH3D21, SLC22A12, SORBS1, SPAG11B, SRSF6, SYT2, TRAPPC8, TUBAL3 and ZNF90. The eleven down-regulated genes identified in RIF were BTN2A1, CYBRD1, FOLR3, GADD45A, GBP2, IGF2, LIF, OPRK1, PSIP1, SMCHD1 and SOD2 (Table 3).
Enrichment analysis to identify GO terms
To investigate the molecular mechanisms and pathways underlying the meta-signature of the mid-secretory endometrium of the RIF group, we utilized a range of contemporary enrichment analysis methods. Specifically, we employed g:GOSt, a tool for functional enrichment analysis (also known as gene set enrichment analysis), which was applied to a set of 49 genes. This tool associates’ genes with well-established functional information sources and identifies functional terms that show significant enrichment through statistical analysis. As depicted in Fig. 2, out of the 49 genes, 16 were associated with GO-Molecular Functions (depicted in red), 101 with GO-Biological Processes (depicted in orange), 30 with GO-Cellular Components (depicted in green), and 8 with KEGG Pathways (depicted in pink).
The identified genes were predominantly associated with molecular functions related to ion binding, chemical binding, and catalytic activity, as shown in Fig. 3A. In terms of biological processes (BP), most of the genes were involved in the regulation of cellular and metabolic processes, cell communication, signaling, and signal transduction, as depicted in Fig. 3B. Furthermore, essential genes in cellular components were found in the lumen of the intracellular membrane, the nucleus, and the plasma membrane as in Fig. 3C. Genes from the meta-signature gene list, namely FLT4, GADD45A, IGF2, NTRK2, IL2RA, TUBAL3, GDF15, LIF, PAX7 have been revealed to primarily belong to MAPK signaling pathway, PI3K-Akt signaling pathway, Apoptosis, Cytokine-cytokine receptor interaction, Ras signaling pathway and Transcriptional misregulation in cancer as shown in Table 4.
Gene Ontology (GO) analysis of genes. Genes that were found to be differentially expressed with a fold change greater than 2 in patients with implantation failure compared to control. A false discovery rate (FDR) of less than 5.0 was considered significant. Panel A presents Molecular function in implantation failure patients with the count of genes involved in each function. Panel B shows the dysregulated biological process in implantation failure patients with the count of genes involved in each process. Panel C illustrates the dysregulated cellular components in implantation failure patients with the count of genes involved in each component
microRNA target prediction
Using the go-profiler miRNA scan to predict their putative regulatory microRNAs, we assessed the possible regulation of the 49 meta-signature genes. Table 5 lists the top 15 human miRNAs that regulate most genes. In parallel, we evaluated our gene list in MIENTURNET, an in silico target prediction algorithm [37] that employs Targetscan and miRTarBase enrichment analysis. To further enhance bioinformatic predictions, we implemented an extra filter by developing a network algorithm that focused on a small group of genes. The resulting network was visualized using charts and display networks, representing miRNA and its corresponding predicted genes, shown in Fig. 4.
Integrated miRNA-mRNA analysis by MIENTURNET web tool. The top 9 correlated putative miRNA-mRNA pairs (p < 0.05). Light blue square indicates miRNAs with their interacting partners’ mRNAs, blue circles [37]
Protein–protein interaction prediction
In the STRING website, a total of 49 differentially expressed genes (DEGs) were filtered and included in the PPI network complex and some extra genes with protein homology. The network comprised of 49 nodes and 20 edges, representing protein–protein interactions (enrichment p-value 0.001) among the DEGs (Fig. 5A). To identify clusters within the PPI network, we performed a k-core analysis with a threshold of 2, resulting in the identification of three distinct clusters. Among these clusters, cluster 1 had the highest score, consisting of 22 nodes and 11 edges, as shown in Fig. 5B. These findings suggest that the 22 DEGs (ANK3, BTNL9, CYBRD1, FLT4, GBP2, GSG1, IGF2, IGFN1, IL2RA, LIF, MC3R, NOS1, NTRK2, OPRK1, PAX7, PLXNA4, PSIP1, SAMD12, SLC22A12, SMCHD1, SPAG118, and TUBAL3) within this cluster may have a pivotal role in mid-secretory endometrium.
String Protein–Protein Interaction Output. (A) Cluster analysis of the 49 DEGs were filtered into PPI network complex that contained 49 nodes and 20 edges with PPI enrichment p-value: 0.001. (B) Module analysis of Protein–Protein Interaction network cluster 1. This cluster consists of 22 nodes and 11 edges and has the highest score in those clusters with a PPI enrichment p-value: 0.001
Validation of meta-analysis genes by RT-qPCR
Since the attachment between the embryo and the endometrium during implantation relies on interactions, CTNNA2 is believed to be involved in maintaining the structural integrity of endometrial tissue. GADD45A regulates decidualization, LIF promotes endometrial receptivity and differentiation of stromal cells, SMCHD1 may be important for endometrial development during pregnancy, and PPP1R1A is expressed during the menstrual cycle and may regulate endometrial cell proliferation and differentiation. Hence, the expression levels of the listed genes were analyzed in few patient samples (RIF and Control). The meta-signature gene analysis comparing RIF versus Control indicated significance (p < 0.05), as illustrated in Fig. 6. The results showed that the expression levels of the selected genes followed the expected trend. Specifically, CTNNA2 and PP1R1A were upregulated, as seen in our meta-analysis. Gene CTNNA2 exhibited a cumulative fold change of 1.5, which was comparable to the control samples. Additionally, other genes, namely GADD45A, LIF, and SMCHD1, were downregulated in our selected samples.
Discussion
The findings of this study reveal a meta-signature, comprising 49 identified genes, which holds potential as an indicator for RIF. The approach involved using data from diverse transcriptomic studies. However, a limitation arose as only data from six studies were examined directly from raw data, while for three studies, a compiled gene list was utilized due to its incompatible file format for use in Genespring. Hence, this report provides stronger evidence for the role of these genes in endometrial functions and their potential clinical implications and understanding these genes can provide valuable insights into the mechanisms that underlie successful implantation and may have implications for the diagnosis and treatment of infertility.
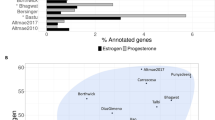
Despite advances in system biology approaches in recent years, there are very few meta-analysis studies comparing RIF transcriptome data to fertile controls. One of the extensive meta-analysis studies including RIF was conducted by Devesa-Peiro and co-workers in 2020 [46]. The authors conducted a meta-analysis of microarray gene expression data in 119 who had endometrial adenocarcinoma (ADC), recurrent implantation failure (RIF), recurrent pregnancy loss (RPL), or stage II–IV endometriosis. They identified 12 functional groups which were significantly dysregulated for RIF; the functional group with the most up-regulated genes was chromosomal and DNA binding, followed by phosphorylation. Genes related to membranes accounted for the downregulated functional group, but they have failed to provide corresponding genes highly involved in these functional groups. Other available studies talk of meta-analysis of endometrial receptivity concept, Altmäe and co-workers 2017 [24] have identified a group of 57 mRNA genes as potential markers of receptivity in the endometrium, these underscore the significance of immune system reactions, the complement cascade pathway, and the role of exosomes in the functions of mid-secretory endometrium. Notably, only three genes, i.e.—GADD45A, GBP2, and NNMT are—overlap between their study and the present study. This difference may be attributed to the distinct focus and selected studies, as their primary emphasis was on endometrial receptivity concepts at mid-secretory endometrium.
Specifically, our investigation concentrated on the RIF group, incorporating nine carefully chosen studies with notable variations in study designs, analytical approaches, and data processing. Furthermore, chosen studies have extensive lists of genes that are expressed differentially.
We examined huge datasets to find common and promising meta-signatures that characterize the endometrium of the RIF group. Eleven genes were found to be significantly downregulated in the endometrium of RIF patients when compared to fertile controls. These genes included BTN2A1, CYBRD1, FOLR3, GADD45A, GBP2, IGF2, LIF, OPRK1, PSIP1, SMCHD1 and SOD2. These genes are involved in a wide range of biological processes, including cell proliferation, DNA repair, and oxidative stress response [47,48,49]. Among them, GADD45A, IGF2, and LIF are known to play important roles in embryo implantation and placentation [50, 51]. The downregulation of these genes may contribute to the impaired implantation and decreased endometrial receptivity in RIF patients [21, 52]. For instance, GADD45A has been shown to be involved in DNA repair and cell cycle regulation, and its downregulation may lead to decreased cell proliferation and impaired endometrial development [53]. IGF2, on the other hand, plays a critical role in embryonic growth and development, and its decreased expression may impair embryo development and implantation [52]. Similarly, LIF, a cytokine essential for embryo implantation and placentation, was also found to be downregulated in RIF patients [21, 54].
Despite garnering less attention in research on female infertility, the four genes OPRK1, PSIP1, SMCHD1, and SOD2 still appear to have an important role. OPRK1 is a gene encoding the opioid receptor kappa 1, which is expressed in the human endometrium and plays a crucial role in implantation and pregnancy maintenance by regulating the immune response and angiogenesis [55, 56]. PSIP1 (PC4 and SFRS1-interacting protein 1) encodes a protein involved in transcriptional regulation and DNA repair processes. It is also involved in the regulation of the endometrial decidualization process, which is essential for successful implantation. SMCHD1 (Structural maintenance of chromosomes flexible hinge domain-containing protein 1) is a recently identified maternal effect gene that functions in the oocyte and is essential for genomic imprinting in the mouse placenta [57]. It is involved in the maintenance of chromatin structure and gene expression regulation. It plays an essential role in the initial stages of embryonic development and implantation [58, 59]. SOD2 (superoxide dismutase 2) encodes an enzyme that scavenges reactive oxygen species (ROS), which can be toxic to cells and tissues. ROS accumulation can cause oxidative stress, leading to DNA damage and cellular dysfunction. Increased SOD2 expression has been reported in steroid producing follicle granulosa and theca internal cells, functional corpus luteum luteinized granulosa and theca cells, and degenerating corpus luteum luteinized theca cells in humans [60, 61]. A study reported that the decreased expression of SOD2, a key antioxidant enzyme, may increase the levels of oxidative stress in the endometrium, leading to impaired endometrial receptivity and decreased implantation success [62, 63].
Asymmetry in the expression of these genes, either upregulation or downregulation, may contribute to RIF pathogenesis by altering the intricate interplay between the embryo and the maternal environment during implantation. Consequently, 38 genes were found to be significantly elevated in the RIF group. These genes are involved in a variety of biological activities, including cell signaling, metabolism, and immunological function, and they may be implicated in endometrial receptivity and implantation. FLT4 (Fms Related Receptor Tyrosine Kinase 4), an upregulated gene involved in angiogenesis and lymphangiogenesis, was found to play a role in endometrial growth and function [64]. Another gene involved is IGFN1 (Immunoglobulin Like And Fibronectin Type III Domain Containing 1), which affects cell migration and adhesion and may control endometrial receptivity [65]. PAX7 (Paired Box 7) is a transcription factor that plays a role in muscle development and has been shown to be upregulated in the endometrium during the implantation window [66]. Endometrial cancer has been connected to the genes SLC22A12 (Solute Carrier Family 22 Member 12) and ANK3 (Ankyrin 3), which are also involved in the regulation of uric acid levels, cytoskeleton organization, and membrane trafficking [67, 68]. CTNNA2 (Catenin Alpha 2), which is involved in cell adhesion and has been shown to be elevated in the endometrium during the implantation window [69] and BIRC3 (Baculoviral IAP Repeat Containing 3), which is involved in apoptosis and immune modulation and has been linked in endometriosis [70]. These findings imply that RIF may be caused by the dysregulation of numerous biological systems, and more research is required to completely understand the molecular mechanisms behind this condition.
In 2002, the endometrium and its receptivity entered the transcriptomic era. Carson and co-workers [71] were the first to address their view on transcriptomics in the endometrium, comparing the early luteal phase with the mid-luteal phase in samples taken from fertile patients. Therefore, when impacted pathways are identified, the importance of differentially expressed genes (DEGs) is more clearly recognized. It is important to note that genes eventually affect pathway functioning via up- or downregulation [20]. Consequently, the present study also intended to identify the dysregulated pathways involved in the pathophysiology of RIF of mentioned 49 meta-signature genes. The results showed significant dysregulation in several pathways, including the MAPK signaling pathway, PI3K-Akt signaling pathway, Apoptosis, Cytokine-cytokine receptor interaction, Ras signaling pathway and Transcriptional misregulation in cancer. The most significantly dysregulated pathways are MAPK and PI3K-Akt signaling pathways, in which FLT4, GADD45A, IGF2, and NTRK2 were the common genes found to be involved in both pathways. The PI3K-Akt signaling pathway plays an essential role in the regulation of the cell cycle, cell proliferation, and apoptosis [72,73,74] while the MAPK signaling pathway plays a crucial role in cellular processes such as cell differentiation, proliferation, and survival [75, 76]. These pathways have also been implicated in endometrial development and implantation [20, 77, 78]. Furthermore, the results showed dysregulation in the apoptosis pathway, with BIRC3, GADD45A, and TUBAL3 genes found to be downregulated. This pathway plays a crucial role in maintaining tissue homeostasis by regulating cell death and is essential for proper embryo implantation [79, 80]. The present study also found dysregulation in the cytokine-cytokine receptor interaction pathway, with GDF15, IL2RA, and LIF genes found to be involved which tend to play a role in the regulation of immune and inflammatory responses essential for successful implantation and pregnancy [81]. Overall, the dysregulated pathways and genes identified in this study provide insight into the molecular mechanisms underlying RIF. The results suggest that dysregulation of genes involved in the MAPK signaling pathway, PI3K-Akt signaling pathway, apoptosis pathway, and cytokine-cytokine receptor interaction pathway may contribute to recurrent implantation failure (RIF) and could potentially enable earlier diagnosis of infertility [82]. These discoveries could aid in the creation of novel diagnostic and therapeutic approaches for females experiencing RIF.
miRNAs are a type of non-coding RNA that acts as a regulator of mRNA and primarily targets the 3′ untranslated region (UTR) of gene transcripts [83, 84]. These are important regulators of cellular processes involved in embryo implantation, as they play a vital role in controlling gene expression post-transcriptionally [85, 86]. About 2500 mature miRNAs have been found so far, with many of those implicated in reproduction and pregnancy [87, 88]. Our analyzed data shows the list of miRNAs most common among the dysregulated mRNAs involved in RIF. miR-335-5p is the most prevalent miRNA, with 11 genes identified, including GDF15, LIF, NTRK2, and PPP1R1A, involved in cellular processes such as cell proliferation, differentiation, and apoptosis [89, 90]. Similarly, miR-26b-5p targets seven genes, including GADD45A and SRSF6, believed to be involved in the regulation of the MAPK and Ras signaling pathways [91]. In addition, miR-17-5p, miR-20a-5p, miR-20b-5p, and miR-106b-5p are predictive to target CPT1A and CYBRD1, earlier studies have shown its involvement in myoblast differentiation [92, 93]. These miRNAs are also involved in the regulation of other genes, such as MUC17, SAMD12, and OPRK1, which are involved in cellular processes, including cell adhesion, proliferation, and differentiation. Interestingly, miR-124-3p targets IGFN1, which is involved in the regulation of the PI3K-Akt signaling pathway [94] and NNMT, which is involved in the regulation of methylation status of histones and DNA [68, 95]. Hence, the dysregulation of miRNAs and their target genes can lead to defects in endometrial receptivity, resulting in the failure of embryo implantation in RIF patients [96, 97]. These findings highlight the potential miRNAs which can also be used as biomarkers for RIF diagnosis.
In conclusion, the molecular signatures identified in the endometrium could provide valuable insights into the pathogenesis of RIF and guide personalized treatment. Transcriptomics, proteomics, and metabolomics are promising techniques that could help identify differentially expressed genes, proteins, and metabolites associated with RIF. Further studies are needed to confirm these findings. Nevertheless, it is crucial to recognize a limitation in this study: the selected samples, while representative of Recurrent Implantation Failure (RIF), were not compared to gene expression profiles associated with other pathologies causing infertility, such as endometritis or endometrial polyps. Despite this limitation, our study paves the way to create tailored medicines that could improve the chances of success in RIF patient.
References
Harper MJK. 10 The implantation window. Baillieres Clin Obstet Gynaecol. 1992;6:351–71.
Wilcox AJ, Baird DD, Weinberg CR. Time of implantation of the conceptus and loss of pregnancy. N Engl J Med. 1999;340:1796–9.
Diedrich K, Fauser BCJM, Devroey P, Griesinger G. The role of the endometrium and embryo in human implantation. Hum Reprod Update. 2007;13:365–77.
Coughlan C, Ledger W, Wang Q, Liu F, Demirol A, Gurgan T, Cutting R, Ong K, Sallam H, Li TC. Recurrent implantation failure: definition and management. Reprod Biomed Online. 2014;28:14–38.
Lai ZZ, Wang Y, Zhou WJ, Liang Z, Shi JW, Yang HL, Xie F, Chen WD, Zhu R, Zhang C, Mei J. Single-cell transcriptome profiling of the human endometrium of patients with recurrent implantation failure. Theranostics. 2022;12:6527–47.
Polanski LT, Baumgarten MN, Quenby S, Brosens J, Campbell BK, Raine-Fenning NJ. What exactly do we mean by ‘recurrent implantation failure’? A systematic review and opinion. Reprod Biomed Online. 2014;28:409–23.
Ma J, Gao W, Li D. Recurrent implantation failure: a comprehensive summary from etiology to treatment. Front Endocrinol. 2023;13:1061766.
Simon A, Laufer N. Repeated implantation failure: clinical approach. Fertil Steril. 2012;97:1039–43.
Fatemi HM, Popovic-Todorovic B. Implantation in assisted reproduction: a look at endometrial receptivity. Reprod Biomed Online. 2013;27:530–8.
Salker MS, Nautiyal J, Steel JH, Webster Z, Šućurović S, Nicou M, Singh Y, Lucas ES, Murakami K, Chan YW, James S. Disordered IL-33/ST2 activation in decidualizing stromal cells prolongs uterine receptivity in women with recurrent pregnancy loss. PLoS ONE. 2012;7:e52252.
Yu Ng EH, Chi Wai Chan C, Tang OS, Shu Biu Yeung W, Ho PC. Endometrial and subendometrial blood flow measured by three-dimensional power Doppler ultrasound in patients with small intramural uterine fibroids during IVF treatment. Hum Reprod. 2005;20:501–6.
Gargett CE, Chan RW, Schwab KE. Hormone and growth factor signaling in endometrial renewal: role of stem/progenitor cells. Mol Cell Endocrinol. 2008;288:22–9.
Maybin JA, Critchley HO, Jabbour HN. Inflammatory pathways in endometrial disorders. Mol Cell Endocrinol. 2011;335:42–51.
Hickey M, Fraser IS. Clinical implications of disturbances of uterine vascular morphology and function. Best Pract Res Clin Obstet Gynaecol. 2000;14:937–51.
Díaz-Gimeno P, Ruíz-Alonso M, Blesa D, Simón C. Transcriptomics of the human endometrium. Int J Dev Biol. 2014;58(2–3–4):127–37.
Dwivedi S, Purohit P, Misra R, Pareek P, Goel A, Khattri S, Pant KK, Misra S, Sharma P. Diseases and molecular diagnostics: a step closer to precision medicine. Indian J Clin Biochem. 2017;32:374–98.
Ou J, Wang W, Feng T, Liao L, Meng Q, Zou Q, Ding J, Zheng A, Duan C, Li P, Liu Q. Identification of small segmental translocations in patients with repeated implantation failure and recurrent miscarriage using next generation sequencing after in vitro fertilization/intracytoplasmic sperm injection. Mol Cytogenet. 2015;8:1–7.
Altmäe S, Reimand J, Hovatta O, Zhang P, Kere J, Laisk T, Saare M, Peters M, Vilo J, Stavreus-Evers A, Salumets A. Research resource: interactome of human embryo implantation: identification of gene expression pathways, regulation, and integrated regulatory networks. Mol Endocrinol. 2012;26:203–17.
Munch EM, Sparks AE, Gonzalez Bosquet J, Christenson LK, Devor EJ, Van Voorhis BJ. Differentially expressed genes in preimplantation human embryos: potential candidate genes for blastocyst formation and implantation. J Assist Reprod Genet. 2016;33:1017–25.
Bastu E, Demiral I, Gunel T, Ulgen E, Gumusoglu E, Hosseini MK, Sezerman U, Buyru F, Yeh J. Potential marker pathways in the endometrium that may cause recurrent implantation failure. Reprod Sci. 2019;26:879–90.
Mrozikiewicz AE, Ożarowski M, Jędrzejczak P. Biomolecular markers of recurrent implantation failure—a review. Int J Mol Sci. 2021;22:10082.
Pathare AD, Zaveri K, Hinduja I. Downregulation of genes related to immune and inflammatory response in IVF implantation failure cases under controlled ovarian stimulation. Am J Reprod Immunol. 2017;78:e12679.
Sheikhansari G, Soltani-Zangbar MS, Pourmoghadam Z, Kamrani A, Azizi R, Aghebati-Maleki L, Danaii S, Koushaeian L, Hojat-Farsangi M, Yousefi M. Oxidative stress, inflammatory settings, and microRNA regulation in the recurrent implantation failure patients with metabolic syndrome. Am J Reprod Immunol. 2019;82:e13170.
Altmäe S, Koel M, Võsa U, Adler P, Suhorutšenko M, Laisk-Podar T, Kukushkina V, Saare M, Velthut-Meikas A, Krjutškov K, Aghajanova L. Meta-signature of human endometrial receptivity: a meta-analysis and validation study of transcriptomic biomarkers. Sci Rep. 2017;7(1):10077.
Hamid JS, Hu P, Roslin NM, Ling V, Greenwood CM, Beyene J. Data integration in genetics and genomics: methods and challenges. Human genomics and proteomics HGP. 2009; 2009.
Toro-Domínguez D, Carmona-Sáez P, Alarcón-Riquelme ME. Shared signatures between rheumatoid arthritis, systemic lupus erythematosus and Sjögren’s syndrome uncovered through gene expression meta-analysis. Arthritis Res Ther. 2014;16:1–8.
Wang C, Guan D, Li Z, Yang Y, Yang K. Emerging trends and frontier research on recurrent implantation failure: a bibliometric analysis. Annals of Translational Medicine. 2022;10(6).
Zhao X, Zhao Y, Jiang Y, Zhang Q. Deciphering the endometrial immune landscape of RIF during the window of implantation from cellular senescence by integrated bioinformatics analysis and machine learning. Front Immunol. 2022;13:952708.
Potdar N, Gelbaya T, Nardo LG. Endometrial injury to overcome recurrent embryo implantation failure: a systematic review and meta-analysis. Reprod Biomed Online. 2012;25:561–71.
Valdes CT, Schutt A, Simon C. Implantation failure of endometrial origin: it is not pathology, but our failure to synchronize the developing embryo with a receptive endometrium. Fertil Steril. 2017;108:15–8.
Woon EV, Greer O, Shah N, Nikolaou D, Johnson M, Male V. Number and function of uterine natural killer cells in recurrent miscarriage and implantation failure: a systematic review and meta-analysis. Hum Reprod Update. 2022;28:548–82.
Bourdon M, Peigné M, Solignac C, Darné B, Languille S, Pocate-Cheriet K, Santulli P. F&S Reviews. 2021;2:353–70.
Soini S, Ibarreta D, Anastasiadou V, Aymé S, Braga S, Cornel M, Coviello DA, Evers-Kiebooms G, Geraedts J, Gianaroli L, Harper J. The interface between assisted reproductive technologies and genetics: technical, social, ethical, and legal issues. Eur J Hum Genet. 2006;14:588–645.
Altmäe S, Esteban FJ, Stavreus-Evers A, Simon C, Giudice L, Lessey BA, Horcajadas JA, Macklon NS, D’Hooghe T, Campoy C, Fauser BC. Guidelines for the design, analysis and interpretation of ‘omics’ data: focus on human endometrium. Hum Reprod Update. 2014;20:12–28.
Dai X, Shen L. Advances and trends in omics technology development. Front Med. 2022;9:911861.
Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. Int J Surg. 2021;88:105906.
Licursi V, Conte F, Fiscon G, Paci P. MIENTURNET: an interactive web tool for microRNA-target enrichment and network-based analysis. BMC Bioinformatics. 2019;20:1–10.
Díaz-Gimeno P, Horcajadas JA, Martínez-Conejero JA, Esteban FJ, Alamá P, Pellicer A, Simón C. A genomic diagnostic tool for human endometrial receptivity based on the transcriptomic signature. Fertil Steril. 2011;95:50–60.
Lédée N, Munaut C, Aubert J, Sérazin V, Rahmati M, Chaouat G, Sandra O, Foidart JM. Specific and extensive endometrial deregulation is present before conception in IVF/ICSI repeated implantation failures (IF) or recurrent miscarriages. J Pathol. 2011;225:554–64.
Altmäe S, Tamm-Rosenstein K, Esteban FJ, Simm J, Kolberg L, Peterson H, Metsis M, Haldre K, Horcajadas JA, Salumets A, Stavreus-Evers A. Endometrial transcriptome analysis indicates superiority of natural over artificial cycles in recurrent implantation failure patients undergoing frozen embryo transfer. Reprod Biomed Online. 2016;32:597–613.
Shi C, Han HJ, Fan LJ, Guan J, Zheng XB, Chen X, Liang R, Zhang XW, Sun KK, Cui QH, Shen H. Diverse endometrial mRNA signatures during the window of implantation in patients with repeated implantation failure. Hum Fertil. 2018;21:183–94.
Zhang WB, Li Q, Liu H, Chen WJ, Zhang CL, Li H, Lu X, Chen JL, Li L, Wu H, Sun XX. Transcriptomic analysis of endometrial receptivity for a genomic diagnostics model of Chinese women. Fertil Steril. 2021;116:157–64.
He A, Zou Y, Wan C, Zhao J, Zhang Q, Yao Z, Tian F, Wu H, Huang X, Fu J, Hu C. The role of transcriptomic biomarkers of endometrial receptivity in personalized embryo transfer for patients with repeated implantation failure. J Transl Med. 2021;19:176.
Keleş ID, Günel T, Özgör BY, Ülgen E, Gümüşoğlu E, Hosseini MK, Sezerman U, Buyru F, Yeh J, Baştu E. Gene pathway analysis of the endometrium at the start of the window of implantation in women with unexplained infertility and unexplained recurrent pregnancy loss: is unexplained recurrent pregnancy loss a subset of unexplained infertility? Human Fertility. 2023;26(5):1129–41.
Zhao F, Chen T, Zhao X, Wang Q, Lan Y, Liang Y, Li Y, Wang S, Yang Y, Yang X. LINC02190 inhibits the embryo–endometrial attachment by decreasing ITGAD expression. Reproduction. 2022;163:107–18.
Devesa-Peiro A, Sebastian-Leon P, Garcia-Garcia F, Arnau V, Aleman A, Pellicer A, Diaz-Gimeno P. Uterine disorders affecting female fertility: what are the molecular functions altered in endometrium? Fertil Steril. 2020;113:1261–74.
Burova E, Borodkina A, Shatrova A, Nikolsky N. Sublethal. Oxidative stress induces the premature senescence of human mesenchymal stem cells derived from endometrium. Oxidative Medicine and Cellular Longevity. 2013.
Fung JN, Mortlock S, Girling JE, Holdsworth-Carson SJ, Teh WT, Zhu Z, Lukowski SW, McKinnon BD, McRae A, Yang J, Healey M. Genetic regulation of disease risk and endometrial gene expression highlights potential target genes for endometriosis and polycystic ovarian syndrome. Scientific reports. 2018;8(1):11424.
Mierzejewski K, Paukszto Ł, Kurzyńska A, Kunicka Z, Jastrzębski JP, Makowczenko KG, Golubska M, Bogacka I. PPARγ regulates the expression of genes involved in the DNA damage response in an inflamed endometrium. Sci Rep. 2022;12(1):4026.
Ochoa-Bernal MA, Fazleabas AT. Physiologic events of embryo implantation and decidualization in human and non-human primates. Int J Mol Sci. 2020;21(6):1973.
Silva JF, Serakides R. Intrauterine trophoblast migration: a comparative view of humans and rodents. Cell Adh Migr. 2016;10:88–110.
Negrón-Pérez VM, Echevarría FD, Huffman SR, Rivera RM. Determination of allelic expression of H19 in pre- and peri-implantation mouse embryos. Biol Reprod. 2013;88(4):97–101.
Salvador JM, Brown-Clay JD, Fornace AJ. Gadd45 in stress signaling, cell cycle control, and apoptosis. Adv Exp Med Biol. 2013;793:1–19.
Massimiani V, Weiland B, Chatelet E, Cornuault P-H, Faucheu J, Massi F. The role of mechanical stimuli on hedonistic and topographical discrimination of textures. Tribol Int. 2020;143:106082.
Cemerikic B, Cheng J, Agbas A, Ahmed MS. Opioids regulate the release of human chorionic gonadotropin hormone from trophoblast tissue. Life Sci. 1991;49:813–24.
Maekawa R, Taketani T, Mihara Y, Sato S, Okada M, Tamura I, Jozaki K, Kajimura T, Asada H, Tamura H, Takasaki A. Thin endometrium transcriptome analysis reveals a potential mechanism of implantation failure. Reproductive Medicine and Biology. 2017;16:206–27.
Wanigasuriya I, Gouil Q, Kinkel SA, Tapia del Fierro A, , Beck T, , Roper EA, , Breslin K, Stringer J, Hutt K, Lee HJ, Keniry A. Smchd1 is a maternal effect gene required for genomic imprinting. eLife. 2020;9:e55529.
Benetti N, Gouil Q, Tapia del Fierro A, Beck T, Breslin K, Keniry A, McGlinn E, Blewitt ME. Maternal SMCHD1 regulates Hox gene expression and patterning in the mouse embryo. Nat Commun. 2022;13(1):4295.
Jansz N, Keniry A, Trussart M, Bildsoe H, Beck T, Tonks ID, Mould AW, Hickey P, Breslin K, Iminitoff M, Ritchie ME. Smchd1 regulates long-range chromatin interactions on the inactive X chromosome and at Hox clusters. Nat Struct Mol Biol. 2018;25:766–77.
Suzuki T, Sugino N, Fukaya T, Sugiyama S, Uda T, Takaya R, Yajima A, Sasano H. Superoxide dismutase in normal cycling human ovaries: immunohistochemical localization and characterization. Fertil Steril. 1999;72:720–6.
Tamate K, Sengoku K, Ishikawa M. The role of superoxide dismutase in the human ovary and fallopian tube. J Obstet Gynaecol. 1995;21:401–9.
Chen C, Zhou Y, Hu C, Wang Y, Yan Z, Li Z, Wu R. Mitochondria and oxidative stress in ovarian endometriosis. Free Radical Biol Med. 2019;136:22–34.
Zaidi SK, Shen WJ, Cortez Y, Bittner S, Bittner A, Arshad S, Huang TT, Kraemer FB, Azhar S. SOD2 deficiency-induced oxidative stress attenuates steroidogenesis in mouse ovarian granulosa cells. Mol Cell Endocrinol. 2021;519:110888.
Guo X, Yi H, Li TC, Wang Y, Wang H, Chen X. Role of vascular endothelial growth factor (Vegf) in human embryo implantation: clinical implications. Biomolecules. 2021;11(2):253.
Li X, Baker J, Cracknell T, Haynes AR, Blanco G. IGFN1_v1 is required for myoblast fusion and differentiation. PLoS ONE. 2017;12:e0180217.
Zong L, Zheng S, Meng Y, Tang W, Li D, Wang Z, Tong X, Xu B. Integrated transcriptomic analysis of the miRNA–mRNA interaction network in thin endometrium. Front Genet. 2021;12:589408.
Matsubayashi M, Sakaguchi YM, Sahara Y, Nanaura H, Kikuchi S, Asghari A, Bui L, Kobashigawa S, Nakanishi M, Nagata R, Matsui TK. 27-Hydroxycholesterol regulates human SLC22A12 gene expression through estrogen receptor action. FASEB. 2021;35:e21262.
Tapia A, Gangi LM, Zegers-Hochschild F, Balmaceda J, Pommer R, Trejo L, Pacheco IM, Salvatierra AM, Henríquez S, Quezada M, Vargas M. Differences in the endometrial transcript profile during the receptive period between women who were refractory to implantation and those who achieved pregnancy. Hum Reprod. 2008;23:340–51.
Akbar R, Ullah K, Rahman TU, Cheng Y, Pang HY, Jin LY, Wang QJ, Huang HF, Sheng JZ. miR-183-5p regulates uterine receptivity and enhances embryo implantation. J Mol Endocrinol. 2020;64:43–52.
Neubauer NL, Ward EC, Patel P, Lu Z, Lee I, Blok LJ, Hanifi-Moghaddam P, Schink J, Kim JJ. Progesterone receptor-B induction of BIRC3 protects endometrial cancer cells from AP1-59-mediated apoptosis. Hormones and Cancer. 2011;2:170–81.
Carson DD, Lagow E, Thathiah A, Al-Shami R, Farach-Carson MC, Vernon M, Yuan L, Fritz MA, Lessey B. Changes in gene expression during the early to mid-luteal (receptive phase) transition in human endometrium detected by high-density microarray screening. Mol Hum Reprod. 2002;8:871–9.
Zhang J, Yang Y, Zhang Z, He Y, Liu Z, Yu Y, Wu S, Cai B, Feng Y. Gankyrin plays an essential role in estrogen-driven and GPR30-mediated endometrial carcinoma cell proliferation via the PTEN/PI3K/AKT signaling pathway. Cancer Lett. 2013;339:279–87.
Ma LI, Chang Y, Yu L, He W, Liu Y. Pro-apoptotic and anti-proliferative effects of mitofusin-2 via PI3K/Akt signaling in breast cancer cells. Oncol Lett. 2015;10:3816–22.
Liu C, Wang M, Zhang H, Sui C. Altered microRNA profiles of extracellular vesicles secreted by endometrial cells from women with recurrent implantation failure. Reprod Sci. 2021;28:1945–55.
Hong K, Choi Y. Role of estrogen and RAS signaling in repeated implantation failure. BMB Rep. 2018;51:225–9.
Luo J, Zhu L, Zhou N, Zhang Y, Zhang L, Zhang R. Construction of circular RNA–MicroRNA–Messenger RNA regulatory network of recurrent implantation failure to explore its potential pathogenesis. Front Genet. 2021;11: 627459.
Tabibzadeh S, Babaknia A. The signals and molecular pathways involved in implantation, a symbiotic interaction between blastocyst and endometrium involving adhesion and tissue invasion. Mol Hum Reprod. 1995;1:179–202.
Dey SK, Lim H, Das SK, Reese J, Paria BC, Daikoku T, et al. Molecular cues to implantation. Endocr Rev. 2004;25:341–73.
Danial NN, Korsmeyer SJ. Cell Death. Cell. 2004;116:205–19.
Wu H, Che X, Zheng Q, Wu A, Pan K, Shao A, et al. Caspases: a molecular switch node in the crosstalk between autophagy and apoptosis. Int J Biol Sci. 2014;10:1072–83.
Mor G, Cardenas I, Abrahams V, Guller S. Inflammation, and pregnancy: the role of the immune system at the implantation site. Ann N Y Acad Sci. 2011;1221:80–7.
Parikh FR, Panpalia M, Mehta T, Agarwal S, Khandeparker M, Chettiar SS, et al. Dysfunctional regulation of pivotal and key inflammatory pathways in infertile Indian women with genital tuberculosis. Am J Reprod Immunol. 2022;88:e13624.
Martinez NJ, Walhout AJM. The interplay between transcription factors and microRNAs in genome-scale regulatory networks. BioEssays. 2009;31:435–45.
Ebert MS, Sharp PA. Emerging roles for natural MicroRNA sponges. Curr Biol. 2010;20:R858–61.
Rarani FZ, Borhani F, Rashidi B. Endometrial pinopode biomarkers: molecules and microRNAs. J Cell Physiol. 2018;233:9145–58.
Li R, Qiao J, Wang L, Li L, Zhen X, Liu P, et al. MicroRNA array and microarray evaluation of endometrial receptivity in patients with high serum progesterone levels on the day of hCG administration. Reprod Biol Endocrinol. 2011;9(1):1–9.
Shekibi M, Heng S, Nie G. MicroRNAs in the regulation of endometrial receptivity for embryo implantation. Int J Mol Sci. 2022;23(11):6210.
Wang E. An overview of microRNA. RNA Technologies in Cardiovascular Medicine and Research. 2008;3–15.
Bahmyari S, Jamali Z, Khatami SH, Vakili O, Roozitalab M, Savardashtaki A, Solati A, Mousavi P, Shabaninejad Z, Vakili S, Behrouj H. microRNAs in female infertility: an overview. Cell Biochem Funct. 2021;39:955–69.
Butler AE, Cunningham TK, Ramachandran V, Diboun I, Halama A, Sathyapalan T, Najafi-Shoushtari SH, Atkin SL. Association of microRNAs with embryo development and fertilization in women undergoing subfertility treatments: a pilot study. Front Reprod Health. 2021;3:719326.
Wu Y, Yuan W, Ding H, Wu X. Serum exosomal miRNA from endometriosis patients correlates with disease severity. Arch Gynecol Obstet. 2022;305:117–27.
Luo W, Li G, Yi Z, Nie Q, Zhang X. E2F1-miR-20a-5p/20b-5p auto-regulatory feedback loop involved in myoblast proliferation and differentiation. Sci Rep. 2016;6:27904.
Wang Y, Hussein AM, Somasundaram L, Sankar R, Detraux D, Mathieu J, Ruohola-Baker H. MicroRNAs regulating human and mouse naïve pluripotency. Int J Mol Sci. 2019;20(23):5864.
Lu Y, Yang J, Sun J, Lu W, Wang J-H. mRNA and miRNA profiles in the nucleus accumbens are associated with psychological stress-induced susceptible and resilient mice. Pharmacol Biochem Behav. 2020;199:173062.
Su X, Wellen KE, Rabinowitz JD. Metabolic control of methylation and acetylation. Curr Opin Chem Biol. 2016;30:52–60.
von Grothusen C, Frisendahl C, Modhukur V, Lalitkumar PG, Peters M, Faridani OR, et al. Uterine fluid microRNAs are dysregulated in women with recurrent implantation failure. Hum Reprod. 2022;37:734–46.
Revel A, Achache H, Stevens J, Smith Y, Reich R. MicroRNAs are associated with human embryo implantation defects. Hum Reprod. 2011;26:2830–40.
Acknowledgements
We thank Shreya Johnson, Research Fellow, Gujarat Biotechnology Research Centre (GBRC), Gandhinagar, India for helping with data analysis and for her useful comments in preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
All authors were involved in the conception, design, analysis, and interpretation of the data, as well as drafting, revising, and final approval of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chettiar, V., Patel, A., Chettiar, S.S. et al. Meta-analysis of endometrial transcriptome data reveals novel molecular targets for recurrent implantation failure. J Assist Reprod Genet 41, 1417–1431 (2024). https://doi.org/10.1007/s10815-024-03077-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-024-03077-x