Abstract
A filamentous benthic cyanobacteria strain isolated from a tropical man-made pond in Malaysia was characterised using combined phenotypic and genetic approaches. Morphological and ultrastructural observations were performed together with growth measurements. Cell dimensions, thylakoid arrangement and apical cell shape with aerotopes were consistent with the description of Pseudanabaena amphigranulata (Goor) Anagnostidis. Molecular characterisation of the16S rRNA gene gave 94% pairwise sequence identity with Pseudanabaena sp. PCC 6802,which corresponds to the genus identification threshold value while also suggesting that the strain is distinctly different to the species of Pseudanabaena currently represented in available databases. The strain showed identical 16S-23S ITS configuration with other strains of Pseudanabaena apart from having a larger spacer region. Cultures of the strain were exposed to various temperature and photoperiod treatments and harvested at exponential phase in order to examine phenotypic plasticity. Significant relationships between environmental conditions and morphological characteristics (cell dimensions and shape) were identified for the first time within the genus Pseudanabaena. The maximum cell length (5.7 ± 0.07 μm) was observed at 25 °C under 12:12 light to dark, while the greatest cell width (3.2 ± 0.11 μm) was also observed at 25 °C but under 16:8 light to dark. The strain showed high plasticity in cell dimensions and shape under different temperature and photoperiod treatments, with25 °C under 12:12 light to dark providing the optimal conditions for its growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The non-heterocystous cyanobacterial genus Pseudanabaena was first established by Lauterborn in 1915 (Geitler 1932). However, it has often been overlooked because of difficulties in observation related to relatively small cell/filament sizes. The genus includes a group of microscopic and simple members of Synechococcales, comprising more than 33 species (Kling and Watson 2003; Komárek and Anagnostidis 2005). The family Pseudanabaenaceae is characterised by simple trichomes with a width less than 4 μm (Acinas et al. 2009). Members of the genus play key roles in ecosystem dynamics (Bertos-Fortis et al. 2016) and have been reported to be sources of antibacterial and antifungal compounds (Oufdou et al. 2001). Therefore, increasing knowledge of the ecology, morphological variability and phylogenetic status of members of this key genus is of particular interest.
Pseudanabaena has undergone several taxonomic and systematic revisions (Yu et al. 2015), but current taxonomic knowledge remains unsatisfactory due to the typically superficial identification of strains of simple cyanobacteria (Komárek and Anagnostidis 2005). Additionally, the presence of apparently identical morphotypes under different ecological conditions, or different morphotypes under the same ecological conditions, leads to further taxonomic uncertainty (Kling and Watson 2003). While in classical taxonomy, morphological stability is often considered a key feature in taxonomic studies, it has been reported that filamentous cyanobacteria can show high morphological variability under different growth conditions, with only a few characters such as the shape of the apical cell being truly diagnostic (Lyra et al. 2001; Gugger et al. 2002; Gupta and Agrawal 2006). Temperature is one of the most important environmental variables affecting the growth and morphology of phytoplankton (Coles and Jones 2000), making identification of species based on morphology alone unreliable. Taxonomic information, such as ultrastructural features, pigment composition, DNA sequence data and robust phylogenetic analyses of most Pseudanabena species, remains incomplete (Yu et al. 2015).
Cyanobacterial classification has undergone important taxonomic and systematic changes in recent years (Komárek et al. 2014). Furthermore, sequence data in GenBank are unsatisfactory due to a general lack of integrated information on morphology that leads to incorrect recognition of strains. Komárek and Anagnostidis (2005) proposed a simplified generic classification based on morphological features. Advances in molecular technologies will increasingly play a critical role in prokaryote identification and characterisation (Emerson et al. 2008), especially when integrated with electron microscopy (EM) techniques and biochemical analyses (Castenholz 2001; Komárek 2003; Komárek and Anagnostidis 2005).
The species Pseudanabaena amphigranulata (Goor) Anagnostidisis is distributed worldwide and is often very common in fresh water (Anagnostidis 2001). It is benthic or planktic and has been reported from shallow eutrophic lakes and other water bodies with muddy sediments, together with purple and colourless sulphur bacteria (Anagnostidis 2001). Hence, we expected the species to occur in the freshwater pond Tasik Harapan (Penang Malaysia), which is shallow and polluted from the untreated effluent discharged from domestic sewage (Salih et al. 2013).
Here, we report the first application of apolyphasic phenotypic and genotypic approach to characterise a strain of Pseudanabaena obtained from Tasik Harapan using a combination of morphological characterisation, electron microscopy, 16SrRNA and 16S-23S internal transcribed spacer (ITS) sequencing. We also examined the morphological plasticity of this strain under different temperature and photoperiod treatments to examine the stability of the morphological features that are of highest taxonomic significance.
Materials and methods
Sample collection and isolation
Pond sediments were collected from Tasik Harapan, Universiti Sains Malaysia, Penang (5°21’ N 100°18′0″ E). A Pseudanabaena strain (strain USMAC18) was isolated by streaking and micropipette picking (Andersen and Kawachi 2005). The isolated strain was established in both BG-11 liquid and 1% agarised media (Rippka et al. 1979) at 25 ± 2 °C, under alight intensity of 27 μmol photons m−2 s−1and photoperiod of 12L:12D. Cycloheximide at a concentration of 200 μg mL−1 was added to both media to eliminate green algae contamination.
Morphological identification
The strain was examined morphologically using an Olympus light microscope (Model BX53F, Olympus, Japan) equipped with digital camera (Olympus, Japan). Individual trichomes were observed at 1000× magnification. The images were analysed concurrently with cell measurement software (Cell Sens Standard Version 1.4.1). Cell length and cell width of 30 individual mature trichomes were measured.
Morphological plasticity in response to environmental variation
To study morphological plasticity, 100 mL of BG-11 were added to 250-mL Erlenmeyer flasks, and then 10 mL aliquots containing 2 × 106 cells mL−1 density of cyanobacteria culture were inoculated into the flasks. Flasks were exposed to 12 different combinations of temperature and photoperiod in a cross-gradient design (Fig. 1) for 21 days. The temperature ranged from 4 to 25 °C, and photoperiod from 0L:24D to 24L:0D (L: light and D: dark), with illumination provided by a cool white fluorescent lamp (Philips), with a light intensity of 27 μmol photons m−2 s−1. Observations were made using an Olympus BX53F microscope at 1000× magnification, and length and width of apical cells from 30 mature trichomes were measured under each treatment combination. Growth rates were measured by the cell count method using a haemocytometer. Cell counts were completed under a light microscope each day until day 21.
Statistical analyses
The mean values of growth rates and measured morphological parameters were compared by using a 3 × 3 factorial experimental design with a two-way analysis of variance (ANOVA) in SPSS v20.0 software, with temperature and photoperiod as fixed factors. Duncan’s post hoc analysis was used to test multiple comparisons, when the main treatment effect was significant at P < 0.05. All of the experiments were conducted in triplicate. All data are presented as mean ± standard error.
TEM (transmission electron microscopy)
Cell ultrastructure was analysed using TEM. Samples were prepared from actively growing cultures and fixed at room temperature with McDowell-Trump fixative prepared in 0.1 M phosphate buffer (pH 7.2). After buffer washing, samples were fixed with osmium tetraoxide, dehydrated, embedded in Spurr’s resin and sectioned at < 0.1 μm thickness using an ultramicrotome (Power Tome XL, USA). Samples were stained with uranyl acetate and lead citrate before viewing under TEM (EFTEM Libra 120). Microphotographs were generated with a digital camera using Olympus, SIS iTem version 5.0, software.
DNA isolation
Genomic DNA was extracted from cells harvested during the exponential growth phase using the Tiangen DNA secure plant kit (China). The presence of DNA in the extracts was confirmed using 1% agarose gel electrophoresis. Extracted DNA was quantified with a Nanodrop spectrophotometer (Thermo Fisher Scientific, USA).
Molecular characterisation
The 16SrRNA gene sequences and 16S-23S rRNA internal transcribed spacer (ITS) sequences of strain USMAC18 were amplified from the genomic DNA with cyanobacteria-specific primers (Boyer et al. 2001). For PCR, 10 ng of extracted DNA was used in 20 μL reactions consisting of 1 μL of each forward and reverse primer, 2 μL of MgCl2 buffer, 2 μL dNTP mixture and 0.5 units Ex Taq DNA polymerase (all obtained from intron iTaq plus, Intron Biotechnology, South Korea). PCR was carried out in a Bio-Rad Thermal Cycler with parameters set as follows: 94 °C for 1 min, 56 °C for 1 min, 72 °C for 4 min (35 cycles), followed by 10 min extension at 72 °C. The products were confirmed on 1% agarose gels and sequenced commercially (Sanger sequencing, MyTACG Bioscience Enterprise, Malaysia). The 16s rRNA sequence (1285 bp) and 16S-23S ITS region (654 bp) were analysed by using the BLAST nucleotide search function of GenBank at the NCBI online site and the Seq Match tool of the Ribosomal Database Project II (http://www.ncbi.nlm.nih.gov/and http://rdp.cme.msu.edu/, respectively). Strain USMAC18 16SrRNA and 16S-23S ITS sequences are deposited in GenBank under accession numbers KU216231 and MF754077, respectively.
Phylogenetic analyses were conducted using the two sets of sequence data (16S rRNA and 16S-23S ITS) separately. The first set included all16S rRNA gene sequences of Pseudanabaena deposited in GenBank. Homologous sequences were identified using a MegaBlast search of the NCBI database in which only closely related sequences were selected to build the phylogenetic tree. These sequences were aligned using the CLUSTAL W program (European Bioinformatics Institute; http://www.ebi.ac.uk/). Phylogenetic analysis was carried out using MEGA version 6 (Tamura et al. 2013) and Mr. Bayes version 3.2 (Ronquist and Huelsenbeck 2003). Maximum likelihood (ML) analysis was implemented in MEGA version 6 (Tamura et al. 2013). The evolutionary model selected was the K2+gamma model, selected on the basis of the BIC (Bayesian Information Criterion) model using model test in MEGA version 6 (Tamura et al. 2013). Bootstrap re-sampling was performed using 1000 replications. Bayesian Inference was implemented in Mr. Bayes and analysis was performed using a mixed model with parameters set to two replicates of eight chains each for 1000,000 generations, with trees sampled every 100 generations. The first 1000 trees were discarded as burn-in. Parameter 220 stability was estimated by plotting log-likelihood values against generation time, and a consensus tree with posterior probabilities was then generated.
The second set included the16S-23S ITS sequence of strain USMAC18 and its closest match, identified through MegaBlast search and compared directly without tree building analysis. Presence and absence of tRNAs, sequence lengths and spacers were obtained by using the tRNA scan-SE Search Server (Lowe and Chan 2016). Limnothrix redekei was selected as an outgroup as it is often confused with Pseudanabaena due to close morphological resemblance making identification more difficult (Acinas et al. 2009).
Results
Morphology
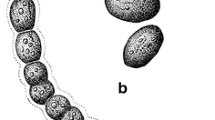
Trichomes were motile, straight, few slightly bent, without branching, 20–40 (50) μm long and 8–14 celled, rarely 20 celled. Cells dark blue-green, arranged in uniseriate row, cylindrical, up to 2 × longer than wide, 1.2–2 (2.5) μm wide and 2–5 (6) μm long, mostly slowly motile, distinctly constricted at cross walls with aerotopes on both sides of septa. Cell content was granulated and differentiated into centroplasm and chromatoplasm. The apical cells were rounded with polar aerotopes but without calyptras, no terminal attenuation, heterocytes and akinetes absent (Fig. 2). Reproduction was by hormogonia.
In liquid medium, the isolate grew as dark blue-green filaments suspended in the medium, and older cultures formed a thin layer on the walls and bottom of the culture vessel. On solid medium, the filaments aggregated into colonies and did not show any colonial gliding motility or phototaxis even after a long duration of light exposure.
Remarks
The specimens examined are consistent with the description of P. amphigranulata (Komarek and Anagnostidis 2005).
Growth rates
Growth rates were evaluated under various culture temperatures and photoperiods over a period of 21 days (Fig. 3). Growth rates differed significantly between treatment combinations (P < 0.05, F11,24 = 281.7). The lowest growth rate (0.008 ± 0.03 day−1) was observed at 4 ± 2 °C and 0L:24D. Progressively higher growth rates were obtained at 15 ± 2 and 25 ± 2 °C with 12L:12D photoperiod, with the highest growth rate being 1.84 ± 0.04 day−1.
Morphological plasticity under varying growth conditions
Morphological plasticity was examined under various temperature and photoperiod treatments to investigate the stability of the measured diacritical characteristics. In stock culture under 25 ± 2 °C and 12L:12D, the cells were cylindrical, 2x longer than wide, 2–5 (6) μm long and 1.2–2 (2.5) μm in width, and apical cells were rounded with polar aerotopes and clear constrictions at the cross walls, consistent with the previous description of Anagnostidis (2001). Morphological features mainly related to size and appearance differed between treatments (Table 1). Individual effects of temperature and photoperiod on cell dimensions were insignificant, while significant effects of combined treatments were observed. Cells growing at 4 ± 2 °C with 0L:24D treatment had the smallest cell length (0.95 ± 0.05 μm, P < 0.05) and width (1.02 ± 0.08 μm, P < 0.05) amongst all the treatments, and were smaller than the range previously described by Anagnostidis (2001). Duncan’s test revealed that cell dimensions differed significantly between all photoperiods (P < 0.05) at 4 ± 2 °C. Cells were mostly isodiametric with aerotopes. Trichomes were narrower with unclear constrictions at the cross walls. Under the 25 ± 2 °C treatment with 12L:12D where tropical aquatic conditions were imitated, there were no differences apparent in cell shape and cell length from those reported by Anagnostidis (2001). The cell shape remained consistent throughout, which was cylindrical, longer than wide with polar aerotopes and distinctly constricted cross walls. Culture under this specific treatment led to cell length (5.7 ± 0.07 μm) in the range previously described by Anagnostidis (2001), and the longest amongst all treatments. However, the widest cells (3.2 ± 0.11 μm), observed at 25 °C under 16L:8D h photoperiod, exceeded the range (1.2–2.5 μm) reported by Anagnostidis (2001).
Transmission electron microscopy
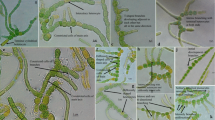
TEM analysis (Fig. 4) clearly showed the presence of cell wall constrictions and the uniseriate row of connected cells. Pseudanabaena amphigranulata has six to eight peripheral thylakoids (T) parietally arranged in the peripheral cytoplasm and concentrically arranged around the long axis of the trichome. The cell wall (Cw) and cross walls (Cr) are simple, similar to other Pseudanabaena species, without prominent mucilaginous envelopes or sheaths. Carboxylate (C) and cyanophycin granules (Cy) were present. TEM analysis was consistent with the description of P. amphigranulata ultrastructure as previously described by Komárek and Anagnostidis (2005).
Molecular characterisation
The newly sequenced P. amphigranulata strain USMAC18 had pairwise sequence identity in the range of 93–94% with 61 strains of Pseudanabaena currently available in GenBank. The best matches were 94% pairwise sequence identity with Pseudanabaena sp. PCC 6802 (accession no. AB039016.1) from Japan (query cover 79%) and P. mucicola KLL-C016 (KP726258.1) from Israel (query cover 71%). A sister relationship between P. amphigranulata USMAC18 and Pseudanabaena sp. PCC 6802 is supported by bootstrap percentage (81%) and BI (0.99). However, P. amphigranulata is clearly distant from P. mucicola KLL-C016 (Fig. 5).
Maximum likelihood (ML) tree showing phylogenetic relationships between Pseudanabaena amphigranulata USMAC18 based on 16S rRNA gene sequences with other species of Pseudanabaena. The newly isolated strain in this study is shown with a filled diamond. Numbers associated with nodes are maximum likelihood bootstrap percentages/Bayesian posterior probability
We obtained a single band of 16S-23S ITS PCR from P. amphigranulata USMAC18, indicating the presence of a single rRNA operon. The ITS sequence of P. amphigranulata strain USMAC18 was compared with other ITS sequences of members of Pseudanabaena retrieved from GenBank. There was variability in the size of the ITS region across the species examined. This took the form of variation in the ITS sequence length, with P. amphigranulata having two inserts, a tRNAIle and tRNAAla, otherwise showing similar ITS composition to the other strains of Pseudanabaena retrieved from GenBank except Pseudanabaena sp. CCY9705, which lacked tRNAAla (Table 2). Pseudanabaena. amphigranulata had the longest ITS length (603 bp) and also a significantly larger spacer region than other close relatives retrieved from GenBank. Limnothrix redekei CCAP 1443/1, used as an outgroup, had a slightly different ITS configuration with a longer tRNAAla gene sequence (77 bp).
A summary of diacritical features of morphospecies closely related to P. amphigranulata USMAC18, derived from Komárek and Anagnostidis (2005), is presented in Table 3. Cell lengths and widths for P. amphigranulata and strains from Genbank with similar ITS composition were mostly in the same ranges, except for P. minima which has a somewhat shorter cell length. All these strains of Pseudanabaena have isodiametric/longer than wide cells. Apical cells were rounded, but conical rounded for P. mucicola.
Discussion
Taxonomic revisions of cyanobacteria at the genus level have been achieved by integrating polyphasic studies during the last decade (Yu et al. 2015). However, considering the complexity of Pseudanabaena, Komárek and Anagnostidis (2005) concluded that the genus still needs further taxonomic revision. Modern taxonomic studies on the genus are limited by the lack of reference polyphasic characteristics from the type strain. This background supported the use of microscopy in combination with molecular studies here, since the description of microbial species without genetic support has been criticised by numerous authors (e.g., Comte et al. 2007; Bruno et al. 2009). Polyphasic approaches are now recognised as crucial to identifying cyanobacterial strains at genus as well as at species level (Marquardt and Palinska 2007).
Growth and morphological plasticity under varying experimental conditions were documented in this study for the first time in the genus Pseudanabaena. Temperature changes between 4 ± 2 and 25 ± 2 °C affected growth, with growth rates increasing significantly with temperature and the highest values observed at 25 ± 2 °C. Growth rates were also affected by photoperiod. The highest growth rate was achieved with 12:12 L:D while the lowest was observed in the absence of light. In the absence of light, the ability of a phototroph to grow will clearly be reduced, and cells will eventually die (Dehning and Tilzer 1989). However, growth of the study strain in the absence of light indicates possession of some mixotrophic capacity. Strains cultured under 24:0 L:D experience photoinhibition due to continuous irradiance, which results in a reduction in photosynthetic rate and leads to reduced growth rates (Tyystjarvi and Aro 1996). Previous studies have emphasised the importance of average temperature and light exposure for active metabolic process (Winder and Sommer 2012; Singh and Singh 2015). Temperatures around 25 ± 2 °C with 12:12 L:D photoperiod, which typify many tropical systems, are favourable for the growth of this strain of P. amphigranulata.
Temperature and light had substantial effects on morphological features of the strain studied. Under various treatments, P. amphigranulata USMAC18 showed significant changes in diacritical characteristics. These changes corroborate the widely accepted observation that cell size changes with temperature (Montagnes and Franklin 2001). Additionally, growth and metabolism are clearly limited at low temperature (sensu Graumann and Marahiel 1996), therefore, at 4 ± 2 °C, we assume that metabolic deficiencies were expressed as shorter and thinner cells in trichomes. At 25 ± 2 °C, longer cells were observed with increased metabolic activities. Changing photoperiod alone did not elicit significant effects on the morphological features of strain USMAC18, but in combination with temperature significant responses were obtained. Intermediate photoperiods (12:12 L:D) in the cross-gradient experiments can be considered optimal for maximising cell length and width. An analogous conclusion was drawn by Zapomelova et al. (2008), who reported morphological variation occurring in two strains of cyanobacteria, Anabaena circinalis and Anabaena crassa, cultivated under various temperature and light conditions.
An important finding of our study is the stability of apical cell shape with polar aerotopes under varied experimental treatments, as originally described by Anagnostidis (2001), supporting the morphological identity of this strain. Thus, the presence of polar aerotopes appears to be a reliable criterion for the identification of P. amphigranulata (Anagnostidis 2001), discriminating it from morphospecies with similar cell dimensions such as P. mucicola (Schwabe 1964) and P. catenata (Lauterborn 1915).
The genus Pseudanabaena is further divided into the subgenera Skujanema (species with pointed apical cell), Pseudanabaena (species with cylindrical and rounded apical cell, with or without polar aerotopes) and Ilyonema (species with polar aerotopes in cells) (Komárek and Anagnostidis 2005). The cell shape and dimensions of the strain USMAC18 closely resemble those of the group of species with aerotopes classified in the subgenus Pseudanabaena.
Molecular analysis demonstrated that the strain P. amphigranulata USMAC18 had only 94% 16S rRNA sequence similarity with Pseudanabaena sp. PCC 6802. Additionally, sequence match analysis of other Pseudanabaena 16S rRNA gene sequences of significant length (> 1000 pb) available in GenBank indicated that these strains generally showed similarities close to the limit for genus delimitation (Stackebrandt and Ebers 2006). The best match (94%) sequence similarity of strain USMAC18 with P. mucicola KLL-C016 is also close to the limit of genus delimitation, and well below the molecular limits (98.5%) for species identification indicated by Stackebrandt and Ebers (2006). However, P. amphigranulata USMAC18 is placed in a single clade with Pseudanabaena sp. PCC 6802, with high bootstrap and BI values. Furthermore, P. mucicola KLL-C016 is distant from strain USMAC18 and positioned with other P. mucicola. The current lack of available sequence data for other strains of P. amphigranulata, or other more closely related species, limits further speculation on the taxonomic position and relationships of this strain.
Limnothrix redekei strains (NIVA CYA 227/1 and CCAP 1459/29), as shown in the phylogenetic tree, are embedded in a cluster with type species of Pseudanabaena, suggesting close genetic similarities between these two genera. A similar conclusion was also drawn by Yu et al. (2015) who suggested to transfer the stains of L. redekei positioned inside the Pseudanabaena cluster to the latter genus. The result is also consistent with Whitton (2011), who used Pseudanabaena redeckei (Goor 1918) as the synonym of Limnothrix redekei. However, differences in the morphology of Pseudanabaena and Limnothrix are still not resolved. Morphologically, the type species differ by the absence or presence of polar aerotopes. L. redeki shows similarities with P. amphigranulata in possessing polar aerotopes, but cell dimensions differ widely (Komárek and Anagnostidis 2005).
Pseudanabaena amphigranulata showed nearly identical ITS configuration with other representatives of Pseudanabaena available in GenBank (except Pseudanabaena sp. CCY9705), apart from possessing a larger spacer region. Multiple rRNA operons have been reported in cyanobacteria, but in most cases the tRNA configuration is identical in both operons. The nucleotide sequences of tRNAAla and tRNAIle were identical and highly conserved between species, but the spacer regions surrounding these two tRNA gene sequences were variable (Chen et al. 2000), signifying that ITS regions in the rRNA operon can be helpful in discrimination between species (Boyer et al. 2001). Both the presence and absence of tRNA sequences between multiple copies of the ITS region in the cyanobacterial genome have been reported previously (Sciuto et al. 2012).
The strain USMAC18 differs from P. mucicola KLL-C016 and other members of this genus in terms of morphology and molecular sequence data. There are no sequences for this species currently available in GenBank against which to compare and support a precise taxonomic identification of this strain. Therefore, according to the ecological and morphological differences documented here we conclude that strain USMAC 18 is a representative of P. amphigranulata. Further molecular studies are necessary to clarify the taxonomic status of this strain.
Conclusions
A polyphasic approach was used to characterise a strain of the cyanobacterial genus Pseudanabaena isolated from a tropical pond. Our data indicate that environmental features (temperature, day length) have a clear influence on cell morphology, with longer day length leading to reduced width and length of the cell. A light-dark cycle of 12:12 light to dark allowed the strain to produce the longest and widest cells. The consistent presence of a rounded apical cell under various culture conditions is a reliable criterion for the identification of this strain, which was morphologically consistent with the description of P. amphigranulata. 16S rRNA and ITS sequence data confirm the strain is highly distinct at the species level from more than 60 members of the genus Pseudanabaena currently available in GenBank.
References
Acinas SG, Haverkamp TA, Huisman J, Stal LJ (2009) Phenotypic and genetic diversification of Pseudanabaena spp. (cyanobacteria). ISME J 3(1):31–46
Anagnostidis K (2001) Nomenclatural changes in cyanoprokaryotic order Oscillatoriales. Preslia 73:359–375
Andersen RA, Kawachi M (2005) Traditional microalgae isolation techniques. In: Anderson RA (ed) Algal culturing techniques. Elsevier, Amsterdam, pp 83-100.
Bertos-Fortis M, Farnelid HM, Lindh MV, Casini M, Andersson A, Pinhassi J, Legrand C (2016) Unscrambling cyanobacteria community dynamics related to environmental factors. Front Microbiol 7:625
Boyer SL, Flechtner VR, Johansen JR (2001) Is the 16S-23S rRNA internal transcribed spacer region a good tool for use in molecular systematics and population genetics? A case study in cyanobacteria. Mol Biol Evol 18:1057–1069
Bruno L, Billi D, Bellezza S, Albertano P (2009) Cytomorphological and genetic characterization of troglobitic Leptolyngbya strains isolated from Roman hypogea. Appl Environ Microbiol 75:608–617
Castenholz R W (2001) Phylum BX. Cyanobacteria. Oxygenic photosynthetic bacteria. In: Bergey’s manual of systematic bacteriology. Volume 1: the Archaea and the deeply branching and phototropic Bacteria. Springer, New York, pp. 413-439
Chen J, Banks D, Jarret RL, Jones JB (2000) Evidence for conserved tRNA genes in the 16S-23S rDNA spacer sequence and two rrn operons of Xylella fastidiosa. Can J Microbiol 46:1171–1175
Comte K, Sabacka M, Carre-Mlouka A, Elster J, Komárek J (2007) Relationships between the Arctic and the Antarctic cyanobacteria; three Phormidium–like strains evaluated by a polyphasic approach. FEMS Microbiol Ecol 59:366–376
Dehning I, Tilzer M (1989) Survival of Scenedesmus acuminatus (Chlorophyceae) in darkness. J Phycol 25:509–515
Emerson D, Agulto L, Liu H, Liu L (2008) Identifying and characterizing bacteria in an era of genomics and proteomics. Bioscience 58:925–936
Geitler L (1932) Cyanophyceae In: Rabenhorst, L. (Ed.) Kryptogamen Flora von Deutschland, Österreich und der Schweiz 14. Akademische Verlagsgesellschaft, Leipzig, pp. 130–159
van Goor ACJ (1918) Zur Kenntnis der Oscillatoriaceen. Reçueil des Travaux Botaniques Néerlandais 15:255–262
Graumann P, Marahiel MA (1996) Some like it cold: response of microorganisms to cold shock. Arch Microbiol 166:293–300
Gupta S, Agrawal SC (2006) Survival of blue-green and green algae under stress conditions. Folia Microbiol 51(2):121–128
Gugger M, Lyra C, Suominen I, Tsitko I, Humbert JF, Salkinoja-Salonen MS, Sivonen K (2002) Cellular fatty acids as chemotaxonomic markers of the genera Anabaena, Aphanizomenon, Microcystis, Nostoc and Planktothrix (cyanobacteria). Int J Syst Evol Microbiol 52:1007–1015
James F. Coles, R. Christian Jones, (2000) Effect of temperature on photosynthesis-light response and growth of four phytoplankton species isolated from a tidal freshwater river. J Phycol 36 :7-16
Kling H, Watson S (2003) A new planktic species of Pseudanabaena (Cyanoprokaryota, Oscillatoriales) from North American large lakes. Hydrobiologia 502:383–388
Komarek J (2003) Problem of the taxonomic category “species” in cyanobacteria. Algol Stud 109:281–297
Komárek J, Anagnostidis K (2005) Süsswasserflora von Mitteleuropa. Cyanoprokaryota: 2.Teil/2nd Part: Oscillatoriales. (Vol. 19): Elsevier Spektrum Akademischer Verlag, Munich, pp.86
Komarek J, Kastovsky J, Mares J, Johansen RJ (2014) Taxonomic classification of cyanoprokaryotes (cyanobacterial genera) 2014, using a polyphasic approach. Preslia 86:295–335
Lauterborn R (1915) Die sapropelische Lebewelt. Ein Beitragzur Biologie des Faulschlammesnatürlicher Gewässer. Verhs Naturhist Med Vereins Heidelberg. Neue Folge 13:395–481
Lyra C, Suomalainen S, Gugger M, Vezie C, Sundman P, Paulin L, Sivonen K (2001) Molecular characterization of planktic cyanobacteria of Anabaena, Aphanizomenon, Microcystis and Planktothrixgenera. Int J Syst Evol Microbiol 51:513–526
Marquardt J, Palinska KA (2007) Genotypic and phenotypic diversity of cyanobacteria assigned to the genus Phormidium (Oscillatoriales) from different habitats and geographical sites. Arch Microbiol 187:397–413
Montagnes DS, Franklin DJ (2001) Effect of temperature on diatom volume, growth rate, and carbon and nitrogen content: reconsidering some paradigms. Limnol Oceanogr 46:2008–2018
Oufdou K, Mezrioui N, Oudra B, Loudiki M, Barakate M, Sbiyyaa B (2001) Bioactive compounds from Pseudanabaena species (cyanobacteria). Microbios 106(Suppl 1):21–29
Rippka R, Deruelles J, Waterbury JB, Herdman M, Stanier RY (1979) Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol 111:1–61
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3—Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Salih S, Alkarkhi A, Lalung J, Ismail N (2013) Water quality of river, lake and drinking water supply in Penang State by means of multivariate analysis. World Appl Sci J 26:75–82
Schwabe GH (1964) Grundprobleme der Cyanophytentaxonomie. Gewässer und Abwässer 36:7–39
Sciuto K, Andreoli C, Rascio N, La Rocca N, Moro I (2012) Polyphasic approach and typification of selected Phormidium strains (cyanobacteria). Cladistics 28:357–374
Singh SP, Singh P (2015) Effect of temperature and light on the growth of algae species: a review. Renew Sust Energy Rev 50:431–444
Stackebrandt E, Ebers J (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33:152–155
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6, molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tyystjarvi E, Aro EM (1996) The rate constant of photoinhibition, measured in lincomycin-treated leaves, is directly proportional to light intensity. Plant Biol 93:2213–2218
Whitton B A (2011) Cyanobacteria (Cyanophyta) In: John, D.M., Whitton, B.A. and Brook, A.J. (Eds.) The freshwater algal flora of the British Isles. An identification guide to freshwater and terrestrial algae. Cambridge University Press, Cambridge, pp. 31–158
Winder M, Sommer U (2012) Phytoplankton response to a changing climate. Hydrobiologia 698:5–16
Yu G, Zhu M, Chen Y, Pan Q, Chai W, Li R (2015) Polyphasic characterization of four species of Pseudanabaena (Oscillatoriales,Cyanobacteria) from China and insights into polyphyletic divergence within the Pseudanabaena genus. Phytotaxa 192:1–12
Zapomelova E, Hisem D, Rehakova K, Hrouzek P, Jezberova J, Komarkova J, Korelusova J, Znachor P (2008) Experimental comparison of phenotypic plasticity and growth (cyanobacteria). J Plankton Res 30:1257–1269
Acknowledgements
We thank Mohammed BasriEshak for assistance with statistical analysis.
Funding
This study was funded and supported by Flagship grant (304/PBIOLOGI/650723/P131) under Ministry of Science, Technology and Innovation, Malaysia. P. Convey is supported by NERC core funding to the BAS ‘Biodiversity, Evolution and Adaptation’ Team.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Khan, Z., Wan Omar, W., Mohd Sidik Merican, F. et al. Characterisation of Pseudanabaena amphigranulata (Synechococcales) isolated from a man-made pond, Malaysia: a polyphasic approach. J Appl Phycol 30, 3187–3196 (2018). https://doi.org/10.1007/s10811-018-1392-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-018-1392-7