Abstract
Abnormal expression and functioning of microRNAs in the immune system are frequently linked to various cancers. Our previous work demonstrated a universal, label-free, and reliable microRNA detection platform based on the nanochannels-based recognition interface and electrode-based sensing interface. Herein, the strategy of G-quadruplex DNAzyme-catalyzed deposition was introduced into the nanochannels to improve the detection sensitivity of the platform. G-quadruplex DNAzyme could catalyze the oxidation of 4-chloro-1-naphthol to produce insoluble precipitates into the nanochannels which amplified the effect of the hybridization reaction between the capture probe and the target to the mass transport of the redox probe through the recognition interface. Then, a porous carbon nanofibers-modified electrode was applied to further amplify the monitoring sensitivity of the sensing interface to the concentration change of the redox probe methylene blue. Thereby, sensitive electrochemical detection of microRNA-21 was acquired. The biosensing platform exhibited a broad linear range of 100 aM to 1 nM for microRNA-21, with a low detection limit of 40 aM. What is more, the selective and reproducible platform provided an alternative technique for the PCR-free determination of microRNA in serum and tumor cells.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
MicroRNAs (MiRs) are a kind of short nonprotein coding RNAs that serve as critical regulators of gene expression in plants, animals, and virus genomes [1]. As proto-oncogene or tumor suppressor, microRNA-21 (miR-21) plays an important role in controlling the expression of genes and therefore it has become a new biomarker for cancer diagnosis and prognosis [2]. MiR-21 is over-expressed in most tumor diseases such as breast cancer [3], cervical cancer [4], and glioblastoma [5]. However, the concentration of miR-21 in human serum or tumor cells is still on such a low level. So it is a rigorous challenge to establish a sensitive enough method for miR-21 detection [6]. The conventional golden nucleic acid detection method such as PCR or northern blotting is not amenable to detect short RNAs and distinguish miR’s family members with very similar sequences. In terms of sensitivity and specificity, surface-enhanced Raman spectroscopy, electrochemiluminescence, and electrochemical biosensing are more superior techniques for the highly sensitive determination of miR-21. Among them, electrochemical biosensing exhibits high sensitivity, good selectivity, easy operation, and are suitable for in vivo and in vitro applications [7,8,9,10,11,12]. In many cases, ultrasensitive detection of miR-21 was often connected with complicated probe labeling or/and time-consuming nucleic acid amplification [13].
Recently, biosensing platforms based on nanochannels have attracted great attention due to extensive, simple, and label-free detection mode [14,15,16,17]. Nanoporous membranes including anodic aluminum oxide (AAO) and polyethylene terephthalate (PET) are frequently applied to the development of biosensing platforms due to their ordered nanochannels, stable overall structure, and easy surface functionalization [18,19,20,21,22]. Usually, the structural transformation or specific capture of biomolecules will cause different occupied space in the nanochannels which can be reflexed by the mass transport of certain molecules or ions [23]. Therefore, electrochemical signal changes of these probe molecules or ions can be quantitatively connected with the formation of nanostructures or the existence of the target [24]. In the early years, Jiang’s fantastic group [16] reported a biomimetic K+ responsive nanochannel system. The conformational change of G-quadruplex DNA in a nanopore of track-etched PET membrane is positively correlated with the K+ concentration. However, the steric hindrance is too limited to miR detection if no any signal amplification strategy is applied. The early electrochemical biosensing platform for DNA detection only gives a detection limit of 0.1 nM [24]. Recently, a nanochannel-ion channel hybrid strategy was established for label-free detection of miR-10b using AAO nanochannels coupling with electrochemical detection. Using this strategy, a low detection limit of 15.4 aM for miR-10b is acquired [18]. By employing steric and electrostatic hindrance of the miRNA-initiated DNA-nanoflower blooming in the nanochannels of AAO, a detection limit of 4.53 fM for miR-21 has been obtained [19]. Therefore, the target-initiated increase of the steric hindrance is powerful to amplify the signal readout for miR detection to give excellent sensitivity.
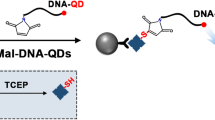
G-quadruplex/hemin DNAzyme is characterized by low cost, high stability, and peroxidase mimicking activity [25,26,27]. Using a capture probe containing G-rich sequence, our previous work [28] performed the G-quadruplex amplified biosensing in AAO nanochannels. As is well known, G-quadruplex/hemin DNAzyme can catalyze the redox reaction of H2O2 and 4-chloro-1-naphthol (4-CN), resulting in insoluble biocatalytic precipitation (IBP) [29]. When the formation of the insoluble products was introduced into the nanochannels, the steric hindrance of the biocomplexes toward the transport of molecules or ions would be greatly amplified. This strategy has been utilized to the photoelectrochemical bioanalysis of telomerase activity. IBP initiated by the sequence extending of telomerase blocked the nanochannels and inhibited the photocurrent signal of the photoelectrode, then realized the quantitative detection of telomerase [30].
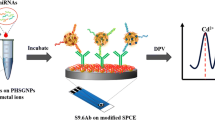
Herein, we constructed a miR-21 electrochemical sensing platform based on G-quadruplex/hemin DNAzyme-catalyzed IBP in nanochannels. As shown in Scheme 1, the nanochannels assumed the task of target recognition and signal amplification. Two DNA probes containing individually the complementary sequence for opposite ends of miR-21 were applied. After incubating miR-21 with recognition probe 2, the hybrids were captured by DNA probe 1 bound in the nanochannels. After sequentially introducing hemin and the raw materials for IBP into the nanochannels, the amount of electroactive methylene blue (MB) transported across the nanochannels was greatly reduced. A porous carbon nanofibers (PCNFs)-modified electrode was introduced into the permeating half-cell to amplify the monitoring sensitivity of the sensing interface. In this way, sensitive and label-free detection of miR-21 is achieved.
2 Experimental section
2.1 Materials and apparatus
AAO membranes (0.2 µm, 13 mm) were purchased from Whatman (UK). CNFs (D × L 100 nm × 20–200 µm) were purchased from Sigma-Aldrich (USA) 0.3-Glycidyloxypropyltrimethoxysilane (GPTMS), 4-CN were the products of Aladdin (China). Hemin was provided by Jingbo Biological Technology Co. Ltd. (Xi’an, China). All MiR and DNA sequences (Table S1) were prepared by Sangon Biotech (Shanghai, China).
All voltammetric experiments were carried out with a CHI440C electrochemical workstation (CH Instruments, China). The three-electrode electrochemical system consisted of a saturated calomel electrode (SCE) as the reference electrode, a platinum wire as the counter electrode, and a bare or modified glassy carbon electrode (GCE) as the working electrode. Other apparatuses applied for the characterization of the nanochannels, electrode, and system include scanning electron microscope (SEM, JSM-6390A, Japan), Fourier transform-infrared (FT-IR) spectroscopy (Tensor27, Germany), and Ultraviolet–visible (UV–Vis) spectrophotometer (N 5000, China).
2.2 Preparation of PCNFs-modified electrode
The CNFs and KOH were mixed in a 1:1 mass ratio and fully ground in a mortar, the mixture was calcined in a high-temperature furnace at 800 °C for 1 h. Subsequently, the product was washed repeatedly with 0.1 mol/L HCl until the pH was neutral, and then dried in an oven at 75 °C for 4 h to obtain PCNFs [28].
GCE is carefully polished with 0.05 μm Al2O3 powders, and then ultrasonicated with ethanol and distilled water to wash the electrode surface. With the aid of ultrasonic cleaner, 1.0 mg/mL PCNFs dispersion was obtained by dissolving 1 mg PCNFs in 1 mL N-methyl-2-pyrrolidone (NMP). Finally, 6 μL of porous PCNFs dispersion was dropped on the surface of the GCE. After the evaporation of the solvent, PCNFs/GCE was obtained.
2.3 Immobilization of probe 1 into nanochannels
Firstly, the blank AAO membrane was washed with distilled water and ethanol successively, and the membrane was immersed in 10 mL acetone solution containing 10% GPTMS and placed at 4 °C for 4 h. After the AAO membrane was washed with acetone and distilled water, 100 μL of DNA probe 1 solution (1 μM, dissolved in pH = 7.4 Tris–HCl buffer containing 100 mM NaCl) was dropped on the surface of the membrane. After waiting overnight at 4 °C, excessive probe 1 was rinsed with pH = 7.4 Tris–HCl buffer.
2.4 Electrochemical determination of miR-21
The probe 2 solution (2 μM) was mixed with different concentrations of miR-21 in equal volumes and the mixed solution was incubated at 37 °C for 1 h. Then, 100 μL of mixture containing different concentration of miR-21 was added to the membrane at 37 °C for 30 min. Next, the AAO membrane was incubated with 100 μL of hemin (0.1 mM in 25 mM KCl solution) at room temperature for 30 min. After that, 100 μL of the mixture of 4-CN (1.0 mM) and H2O2 (0.15 mM) was dropped on the AAO membrane at room temperature for 15 min. After each step, thorough rinsing was done with distilled water. Finally, the membrane was placed in a self-made double-cell device. Then, the left half-cell was filled with 2.0 mL of 0.1 M PBS solution, and the right half-cell was filled with 2.0 mL of 0.1 M PBS solution containing 100 μM MB. The working electrode (PCNFs/GCE) was inserted in the left half-cell, and the SCE and Pt counter electrode were put in the right half-cell. Differential pulse voltammogram (DPV) was carried out in the potential range of − 0.5 to 0.1 V after 45 min of mass diffusion.
3 Results and discussion
3.1 Characterization of PCNFs/GCE, AAO nanochannels, and the biosensing platform
Morphology of CNFs was a typical fibrous structure. After NaOH treatment, the obtained PCNFs have a loose porous or lotus-like structure (Fig. S1). The rough structure not only facilitates the dispersion of PCNFs in solvent, but also enlarges the surface area of the ultimate modified electrode. To confirm this, CV of the electrode was recorded in 0.1 mol∙L−1 KCl containing 5.0 mM K3[Fe(CN)6]. The peak current obtained at PCNFs/GCE is much bigger than it obtained at CNFs/GCE (Figure S2), and the effective surface area of PCNFs/GCE was calculated to be 0.092 cm2, which is bigger than the geometric area of the bare GCE, demonstrating the amplified surface area due to the modification of the material [31]. The voltammetric behavior of MB at the modified electrode was further investigated. Using PCNFs/GCE as the working electrode, multiple consecutive DPV scans was performed in 0.1 M PBS solution containing 20 μM MB. As shown in Fig. 1A, as the number of scan increased, the peak current increased gradually. It shows that MB adsorbs onto the electrode surface [27]. After immersing the electrode in MB solution for 20 min, the peak current acquired at PCNFs/GCE nearly doubled compared with it obtained at CNFs/GCE (Fig. 1B), indicating the applicability of the electrode to the sensitive MB monitoring. Therefore, PCNFs/GCE was chosen as the working electrode for subsequent experiments.
FT-IR and SEM were performed to characterize the AAO nanochannels. The amino-terminated recognition probe 1 was expected to be bound into the nanochannels via the epoxy group of silane. From FT-IR spectra, the peaks at 1627 cm−1 corresponding to the deformation vibration and 1174 cm−1 due to the C–N stretching vibration were observed after the incubation of probe 1 (Fig. S3), indicating that the DNA probe 1 was successfully immobilized in the AAO nanochannels. Representative SEM images of AAO membrane exhibited the uniformly distributed nanopores from the surface view and the vertically aligned nanochannel array from the side view (Fig. S4). Immobilization of probe 1 did not show distinctive variation of SEM image (not shown). After incubation with a mixture of G-rich sequence probe 2 and miR-21, hemin and mixture of 4-CN and H2O2 were further introduced into the nanochannels, then the G-quadruplex DNAzyme-catalyzed deposition of IBP was confirmed by SEM images (Fig. 2), demonstrating the feasibility of the signal amplification strategy.
As shown in the Scheme 1, after incubating miR-21 with recognition probe 2, the hybrids were captured by DNA probe 1 bound in the nanochannels and sequentially introducing hemin and the raw materials for IBP into the nanochannels, the amount of electroactive methylene blue (MB) transported across the nanochannels was greatly reduced. G-quadruplex DNAzyme-catalyzed formation of IBP was quantitatively related with the concentration of miR-21 introduced into the nanochannels. The steric hindrance caused by IBP reduced the amount of MB that crosses the channels from the left pool into the right pool. Therefore, the peak current in DPVs should show a downward trend after target incubation. As displayed in Fig. 3A, sequentially introducing probe 2 and the necessary reagents for catalyzed deposition, the peak current (curve b) decreased by 0.545 μA compared with it for the probe 1 functionalized AAO nanochannels (curve a). The low amplitude decrease was contributed to the steric hindrance of nonspecifically deposited IBP. However, replacing probe 2 with the mixture of miR-21 and probe 2, the peak current reduced further by 1.09 μA (curve c), demonstrating the amplified spatial baffle effect of G-quadruplex DNAzyme-catalyzed IBP. Figure 3B shows the UV–Vis spectrum of MB in the left cell. The absorbance variation of the characteristic peak of MB at 666 nm is consistent with the change of peak current in DPV. These results proved the feasibility of the biosensing platform.
DPV response (A) and UV–Vis spectrum (B) of MB transmitted across channels (curves a, b, and c correspond to the AAO empty nanochannels, DNA recognition probe 1 functionalized nanochannels with hemin & 4-CN & H2O2, and DNA recognition probe 2 + 1 × 10−13 M miR-21 immobilized DNA recognition probe 1 nanochannels with hemin & 4-CN & H2O2, respectively)
3.2 Optimization of experimental conditions
To achieve the best sensing performance, the effect of several experimental parameters including the pore size of the AAO membrane, incubation time for the target, and the concentration of the necessary reagents on the change of the DPV response (ΔI, ΔI = I0–I1, where I0 and I1 are the peak current obtained without and with the target, respectively) was investigated.
The pore size of the AAO membrane has an important influence on the analytical performance of the sensing platform. As shown in Fig. 4A, AAO nanochannels with average channels diameter of 100 nm and 200 nm were assessed. It shows that 100 nm is a proper size. For the incubation time of target, ΔI increased rapidly until the miR-21 incubation reached 45 min (Fig. 4B). As observed in Fig. 4C, the maximum ΔI were obtained when the concentration of hemin was 0.06 mM.
In addition, the concentration of 4-CN and H2O2 as well as the reaction time for the G-quadruplex/hemin DNAzyme-catalyzed IBP are also important in the analysis system. The change of the DPV response increased with the increase in concentration of 4-CN and there was almost a plateau trend after 0.06 mM (Fig. 4D). Meanwhile, ΔI is the largest when the concentration of H2O2 was 0.1 mM (Fig. 4E). Furthermore, Our results also showed that 15 min was the optimized reaction time for DNAzyme-catalyzed IBP (Fig. 4F). Prolonging the reaction time decreased the signal sensitivity, probably because of increased I0 caused by the nonspecific BCP into the channels.
3.3 Analytical performance
Under optimal conditions, the DPV responses of the sensing platform for different concentrations of miR-21 were monitored. As shown in Fig. 5A, the peak current gradually decreased with the concentration of miR-21 from 100 aM to 1 nM. The linear-regression equation is I = 2.85 − 0.26 lgC, the linear-correlation coefficient is 0.997 (Fig. 5B), and the detection limit was calculated to be 40 aM. It can be seen from Table 1 that the constructed biosensing platform has a wider linear range and lower detection limit compared with some other methods.
To investigate the selectivity of the sensing platform, DPV response of the platform for the same concentration of mismatched MiRNAs, miR-122 as well as mixture of 100 fM miR-21 and interfering substances was recorded. The results showed that the peak current for the mismatched sequences is similar with it for the blank control, and the peak current for the mixture of 100 fM miR-21 and interfering substances is only 1.1% higher than it for 100 fM miR-21 (Fig. S5A). These results proved the excellent selectivity of the sensing platform. The reproducibility of the sensing platform was further studied. The platforms were reconstructed by repeatedly modifying the electrode with PCNFs and immobilizing probe 1 into the new nanochannels. The relative standard deviation of five responses obtained on the reconstructed platform for 100 fM miR-21 was 2.7% (Fig. S5B). The stability of the platform was also evaluated by storing the bio-functionalized nanochannels and the PCNFs/GCE at 4℃ for one week. The electrochemical response only decreased 7.6% compared with it obtained from the freshly prepared platform.
3.4 Sample analysis
In order to further determine the feasibility of the developed biosensing platform, recovery tests were performed in human serum of healthy volunteers. The peak current for the serum sample diluted with PBS buffer solution (1:100) is very close to it for the blank control (Fig. S6), hinting that content of miR-21 in the tested serum sample is below the detection limit. The recoveries of miR-21 in spiked serum ranged from 82.7% to 107.3% (Table S2), indicating the application probability of the method [39, 40].
To demonstrate the applicability of the proposed biosensing platform for miR-21 detection in cancer cells, human breast cancer cells (MCF-7) and cervical cancer cells (Hela) were analyzed. The total RNA extraction from MCF-7 and HeLa was performed according to the instructions of the M5 Universal RNA Mini Kit. As shown in Fig. 6, the response for both HeLa and MCF-7 increased with the increase of the number of cancer cells, and the response for the same number of MCF-7 is larger than that for HeLa. It shows that MCF-7 contains more miR-21, which is consistent with the results of the previous literature studies [41].
4 Conclusion
A dual-interface biosensing platform with AAO nanochannels as the recognition interface and PCNFs/GCE as the detection interface was constructed. The application of the porous carbon nanofibers-modified electrode enhanced the monitoring sensitivity of the electrode to methylene blue. The introduction of insoluble precipitate catalytically produced by the G-quadruplex DNAzyme into the nanochannels amplified the steric hindrance of the interaction occurred into the nanochannels. Based on this, highly sensitive electrochemical detection of miR-21 was achieved with a linear range of 100 aM to 1 nM and a detection limit of 40 aM. What is more, this platform avoided the electrochemical labeling of recognition probe and the use of expensive nuclease, and thus provided a promise platform for the early diagnosis of related diseases.
References
Cissell KA, Rahimi Y, Shrestha S, Hunt EA, Deo SK (2008) Bioluminescence-based detection of microRNA, miR21 in breast cancer cells. Anal Chem 80(7):2319–2325. https://doi.org/10.1021/ac702577a
Zhang X, Wang Y, Fricke BL, Gu L-Q (2014) Programming nanopore ion flow for encoded multiplex microRNA detection. ACS Nano 8(4):3444–3450. https://doi.org/10.1021/nn406339n
Yan L-X, Huang X-F, Shao Q, Huang M-Y, Deng L, Wu Q-L, Zeng Y-X, Shao J-Y (2008) MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA 14:2348–2360. https://doi.org/10.1261/rna.1034808
Lee J-W, Choi CH, Choi J-J, Zhang L, Zhu A, Tian Y (2008) Altered microRNA expression in cervical carcinomas. Clin Cancer Res 14:2535–2542. https://doi.org/10.1158/1078-0432.CCR-07-1231
Chan JA, Krichevsky AM, Kosik KS (2005) MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65:6029–6033. https://doi.org/10.1158/0008-5472.CAN-05-0137
Nossier AI, Abdelzaher H, Matboli ES (2018) Dual approach for the colorimetric determination of unamplified microRNAs by using citrate capped gold nanoparticles. Microchim Acta 185:236. https://doi.org/10.1007/s00604-018-2767-9
Mohammed YE, Mohamed AS, Hiromi M, Sherif AE-S (2018) One-step selective screening of bioactive molecules in living cells using sulfur-doped microporous carbon. Biosens Bioelectron 109:237–245. https://doi.org/10.1016/j.bios.2018.03.026
Wang J, Lu J, Dong S, Zhu N, Gyimah E, Wang K, Li Y, Zhang Z (2019) An ultrasensitive electrochemical biosensor for detection of microRNA-21 based on redox reaction of ascorbic acid/iodine and duplex-specific nuclease assisted target recycling. Biosens Bioelectron 130:81–87. https://doi.org/10.1016/j.bios.2019.01.031
Wang D, Wang J (2021) A sensitive and label-free electrochemical microRNA biosensor based on Polyamidoamine Dendrimer functionalized Polypyrrole nanowires hybrid. Microchim Acta 188:173. https://doi.org/10.1007/s00604-021-04824-y
Yang J, Tang M, Diao W, Cheng W, Zhang Y, Yan Y (2016) Electrochemical strategy for ultrasensitive detection of microRNA based on MNAzyme-mediated rolling circle amplification on a gold electrode. Microchim Acta 183:3061–3067. https://doi.org/10.1007/s00604-016-1958-5
Liang Y-R, Zhang Z-M, Liu Z-J, Wang K, Wu X-Y, Zeng K, Meng H, Zhang Z (2017) A highly sensitive signal-amplified gold nanoparticle-based electrochemical immunosensor for dibutyl phthalate detection. Biosens Bioelectron 91:199–202. https://doi.org/10.1016/j.bios.2016.12.007
Zhou L, Wang J, Chen Z, Li J, Wang T, Zhang Z, Xie G (2017) A universal electrochemical biosensor for the highly sensitive determination of microRNAs based on isothermal target recycling amplification and a DNA signal transducer triggered reaction. Microchim Acta 184:1305–1313. https://doi.org/10.1007/s00604-017-2129-z
Tian L, Qi J, Ma X, Wang X, Yao C, Song W, Wang Y (2018) A facile DNA strand displacement reaction sensing strategy of electrochemical biosensor based on N-carboxymethyl chitosan/molybdenum carbide nanocomposite for microRNA-21 detection. Biosens Bioelectron 122:43–50. https://doi.org/10.1016/j.bios.2018.09.037
Hou X, Guo W, Xia F, Nie F-Q, Dong H, Tian Y, Wen L, Wang L, Cao L, Yang Y, Xue J, Song Y, Wang Y, Liu D, Jiang L (2009) A biomimetic potassium responsive nanochannel: G-quadruplex DNA conformational switching in a synthetic nanopore. J Am Chem Soc 131(22):7800–7805. https://doi.org/10.1021/ja901574c
Ying Y-L, Yu R-J, Hu Y-X, Gao R, Long Y-T (2017) Single antibody-antigen interactions monitored via transient ionic current recording using nanopore sensors. Chem Commun 53:8620–8623. https://doi.org/10.1039/c7cc03927a
Gao R, Ying Y-L, Hu Y-X, Li Y-J, Long Y-T (2017) Wireless bipolar nanopore electrode for single small molecule detection. Anal Chem 89(14):7382–7387. https://doi.org/10.1021/acs.analchem.7b00729
Zhang H, Tian Y, Hou J, Hou X, Hou G, Ou R, Wang H, Jiang L (2015) Bioinspired smart gate-location-controllable single nanochannels: experiment and theoretical simulation. ACS Nano 9(12):12264–12273. https://doi.org/10.1021/acsnano.5b05542
Zhao X-P, Liu F-F, Hu W-C, Younis MR, Wang C, Xia X-H (2019) Biomimetic nanochannel-ionchannel hybrid for ultrasensitive and label-free detection of microRNA in cells. Anal Chem 91(5):3582–3589. https://doi.org/10.1021/acs.analchem.8b05536
Shi L, Mu C, Gao T, Chen T, Hei S, Yang J, Li G (2018) DNA nanoflower blooms in nanochannels: a new strategy for miRNA detection. Chem Commun 54:11391–11394. https://doi.org/10.1039/c8cc05690k
Liu X, Wei M, Liu Y, Lv B, Wei W, Zhang Y, Liu S (2016) Label-free detection of telomerase activity in urine using telomerase-responsive porous anodic alumina nanochannels. Anal Chem 88(16):8107–8114. https://doi.org/10.1021/acs.analchem.6b01817
Xia X, Li H, Zhou G, Ge L, Li F (2020) In situ growth of nano-gold on anodized aluminum oxide with tandem nanozyme activities towards sensitive electrochemical nanochannel sensing. Analyst 145:6617–6624. https://doi.org/10.1039/d0an01271h
Zhao X-P, Cao J, Nie X-G, Wang S-S, Wang C, Xia X-H (2017) Label-free monitoring of the thrombin–aptamer recognition reaction using an array of nanochannels coupled with electrochemical detection. Electrochem Commun 81:5–9. https://doi.org/10.1016/j.elecom.2017.05.018
Han C, Hou X, Zhang H, Guo W, Li H, Jiang L (2011) Enantioselective recognition in biomimetic single artificial nanochannels. J Am Chem Soc 133(20):7644–7643. https://doi.org/10.1021/ja2004939
Li S-J, Li J, Wang K, Wang C, Xu J-J, Chen H-Y, Xia X-H, Huo Q (2010) A nanochannel array-based electrochemical device for quantitative label-free DNA analysis. ACS Nano 4(11):6417–6424. https://doi.org/10.1021/nn101050r
Chen JH, Zhang J, Guo Y, Li J, Fu FF, Yang H-H, Chen G (2011) An ultrasensitive electrochemical biosensor for detection of DNA species related to oral cancer based on nuclease-assisted target recycling and amplification of DNAzyme. Chem Commun 47:8004–8006. https://doi.org/10.1039/c1cc11929j
Zhang J, Gao QL, Chen PP, Chen JH, Chen GN, Fu FF (2011) A novel Tb3+-promoted G-quadruplex-hemin DNAzyme for the development of label-free visual biosensors. Biosens Bioelectron 26:4053–4057. https://doi.org/10.1016/j.bios.2011.03.029
Willner I, Shlyahovsky B, Zayats M, Willne B (2008) DNAzymes for sensing, nanobiotechnology and logic gate applications. Chem Soc Rev 37:1153–1165. https://doi.org/10.1039/b718428j
Zhao F, Zhang H, Zheng J (2021) Novel electrochemical biosensing platform for microRNA detection based on G-quadruplex formation in nanochannels. Sens Actuators B Chem 327:128898. https://doi.org/10.1016/j.snb.2020.128898
Hou L, Gao Z, Xu M, Cao X, Wu X, Chen G, Tang D (2014) DNAzyme-functionalized gold–palladium hybrid nanostructures for triple signal amplification of impedimetric immunosensor. Biosens Bioelectron 54:365–371. https://doi.org/10.1016/j.bios.2013.11.014
Fan G-C, Lu Y, Ma L, Song Z-L, Luo X, Zhao W-W (2019) Target-induced formation of multiple DNAzymes in solid-state nanochannels: Toward innovative photoelectrochemical probing of telomerase activity. Biosens Bioelectron 142:111564. https://doi.org/10.1016/j.bios.2019.111564
Mohammed YE, Sherif A-S, Mahmoud MS, Takashi M, Ahmed E, Mohamed AS (2017) Non-metal sensory electrode design and protocol of DNA-nucleobases in living cells exposed to oxidative stresses. Anal Chim Acta 1142:143–156. https://doi.org/10.1016/j.aca.2020.11.004
Zhou W, Tian Y-F, Yin B-C, Ye B-C (2017) Simultaneous surface-enhanced raman spectroscopy detection of multiplexed microRNA biomarkers. Anal Chem 89(11):6120–6128. https://doi.org/10.1021/acs.analchem.7b00902
Xia Y, Wang L, Li J, Chen X, Lan J, Yan A, Lei Y, Yang S, Yang H, Chen J (2018) A ratiometric fluorescent bioprobe based on carbon dots and acridone derivate for signal amplification detection exosomal microRNA. Anal Chem 90(15):8969–8976. https://doi.org/10.1021/acs.analchem.8b01143
Peng L, Yuan Y, Fu X, Fu A, Zhang P, Chai Y, Gan X, Yuan R (2019) Reversible and distance-controllable DNA scissor: a regenerated electrochemiluminescence biosensing platform for ultrasensitive detection of microRNA. Anal Chem 91(5):3239–3245. https://doi.org/10.1021/acs.analchem.8b02757
Zhang Y, Li Z, Cheng Y, Lv X (2009) Colorimetric detection of microRNA and RNase H activity in homogeneous solution with cationic polythiophene derivative. Chem Commun. https://doi.org/10.1039/b904579a
Miao P, Zhang T, Xu J, Tang Y (2018) Electrochemical detection of miRNA combining T7 exonuclease-assisted cascade signal amplification and DNA-templated copper nanoparticles. Anal Chem 90(18):11154–11160. https://doi.org/10.1021/acs.analchem.8b03425
Tian L, Qi J, Ma X, Wang X, Yao C, Song W, Wang Y (2018) A facile DNA strand displacement reaction sensing strategy of electrochemical biosensor based on N-carboxymethyl chitosan/ molybdenum carbide nanocomposite for microRNA-21 detection. Biosens Bioelectron 122:4350. https://doi.org/10.1016/j.bios.2018.09.037
Guo W-J, Wu Z, Yang X-Y, Pang D-W, Zhang Z-L (2019) Ultrasensitive electrochemical detection of microRNA-21 with wide linear dynamic range based on dual signal amplification. Biosens Bioelectron 131:267–273. https://doi.org/10.1016/j.bios.2019.02.026
Mohammed YE, Mohamed AS, Sherif AE-S, Abdullah R, Mahmoud MS (2021) Microporous P-doped carbon spheres sensory electrode for voltammetry and amperometry adrenaline screening in human fluids. Microchim Acta 188:138. https://doi.org/10.1007/s00604-021-04782-5
Jiang Z, Jiao J, Li J, Zhang H, Zheng J (2021) Novel electrochemical biosensing platform for microRNA: Bivalent recognition-induced nanoparticle amplification occurred in nanochannels. Sens Actuators, B Chem 344:0925–4005. https://doi.org/10.1016/j.snb.2021.130209
Fu P, Xing S, Xu M, Zhao Y, Zhao C (2020) Peptide nucleic acid-based electrochemical biosensor for simultaneous detection of multiple microRNAs from cancer cells with catalytic hairpin assembly amplification. Sens Actuators B Chem 305:127545. https://doi.org/10.1016/j.snb.2019.127545
Funding
This work is financially supported by the National Natural Science Foundation of China (Grant No. 21775120).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, F., Xue, W. & Zhang, H. Electrochemical microRNA detection based on catalytic deposition of G-quadruplex DNAzyme in nanochannels. J Appl Electrochem 52, 885–893 (2022). https://doi.org/10.1007/s10800-022-01673-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10800-022-01673-2