Abstract
It has become apparent that natural hybridization is far more common and may play a much greater role in evolution than has historically been recognized. The skeletal morphology of hybrid primates is notoriously variable and difficult to predict. Indeed, before the advent of genetic sequencing techniques, many wild hybrid populations went undetected. Though many species of primates are now known to hybridize naturally and are likely to have done so for millions of years, anthropogenic alterations to the environment are increasingly restricting or altering primate species ranges and contact zones and driving hybridization between populations that may otherwise never have come into contact. The case of hybridizing Japanese and Taiwanese macaques (Macaca fuscata and Macaca cyclopis) is an excellent example of this, as these two island species could not have come into contact without human interference. Here we apply 3D geometric morphometrics and nonmetric trait analysis to the crania and dentition of hybrid macaques (N = 70) and their parental species, M. fuscata (N = 57) and M. cyclopis (N = 51). The exploration of 3D shape variation identifies mildly transgressive morphology in the hybrids and a general tendency toward the M. fuscata morphotype overall, but less variability in the hybrid morphotype than has been identified in previous studies of primate hybrids. We also identify a small number of nonmetric traits that differentiate the hybrids from the parental species, although the power of these traits to distinguish between groups is weak and their relationship with hybridity is unclear. We conclude that the relatively short divergence time between the parent species is likely to help explain the observed differences in hybrid morphotype, and that further exploration of the relationship between degree of evolutionary divergence and hybrid morphology may help us to better explain and predict hybrid morphology in other taxa.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Not so long ago, natural hybridization was thought to be rare, unnatural, and ultimately unimportant to evolution (e.g., Mallet 2005; Mayr 1963). It has since become apparent that hybridization is far more widespread than previously recognized and far more than just evolutionary “noise” (Arnold 1997); in fact, hybridization is likely to play an important creative role in evolution (Arnold 1997, 2006; Arnold and Meyer 2006; Arnold et al. 2008; Mallet 2005; Schwenk et al.2008; Seehausen 2004; Shurtliff 2013). Phenotypically, hybridization is generally considered to result in two main outcomes: heterosis (hybrid vigor), in which the hybrid offspring has some increase in fitness (e.g., growth, health, or fecundity), or dysgenesis (hybrid dysfunction or outbreeding depression), in which the hybrid offspring suffers a reduction in fitness (e.g., inviability, sterility) (e.g., Ackermann 2010; Burke and Arnold 2001). Hybridization is also thought to break down or disrupt developmental constraints that may lead to an increase in minor developmental abnormalities (Ackermann 2010) and fluctuating asymmetry (e.g., Tomkins and Kotiaho 2001). At present, though, our understanding of the underlying molecular processes (e.g., dominance, overdominance, pseudo-overdominance, epistasis, and epigenetics) remains contentious and ultimately superficial (Baranwal et al.2012).
Previously undiscovered hybrids and hybrid zones are being identified throughout all of the major primate radiations (Cortés-Ortiz 2017; Zinner et al.2011). Some of this is undoubtedly naturally occurring; ongoing zones of hybridization have been identified in minimally disturbed or protected habitats (e.g., Bicca-Marques et al.2008; Keller et al.2010; Tung et al.2008; Wyner et al.2002), and genetic evidence for ancient hybridization events has been discovered in several extant genera (e.g., Osada et al.2010; Tosi et al.2000; Zinner et al.2009). More frequently, though, isolated hybrids (or small groups of hybrids) and larger zones of hybridization have been identified in areas of substantial human impact (e.g., Aguiar et al.2007; Bynum et al.1997; Delmore et al.2011; Evans et al.2001; Fuzessy et al.2014; Gligor et al.2009; Hamada et al.2006; Kelaita and Cortés-Ortiz 2013; Malaivijitnond and Hamada 2008; Pastorini et al.2009). It is evident that anthropogenic influences are dramatically altering natural species interactions, whether this is by introducing species that had hitherto been separated by biogeographic barriers (e.g., Hamada et al.2012) or by creating barriers between groups that would otherwise have freely interacted (e.g., Malaivijitnond and Hamada 2008).
Genetic sequencing techniques have quickly become the dominant method for the detection and exploration of previously unidentified hybrid zones and events (e.g., Osada et al.2010) and are increasingly employed in studies alongside traditional phenotypic indicators of mixed parentage such as mixed or intermediate body size or proportions, and pelage color and patterning (e.g., Delmore et al.2013; Fuzessy et al.2014; Kelaita and Cortés-Ortiz 2013). Nevertheless, the relationship between molecular characteristics and hybrid phenotype remains poorly understood and difficult to predict (Cortés-Ortiz 2017). As far as size and shape variation is concerned, nearly any outcome seems possible; hybrids may be more like one parent (e.g., Tung et al.2008), intermediate to both (e.g., Hamada et al.2012), or different from either (Delmore et al.2011; Cortés-Ortiz 2017).
Craniofacial shape and nonmetric variants are commonly used to infer ancestry in living human populations (e.g., Bauchau 1988; Berry 1975; Hanihara and Ishida 2001a, b, c), and have been applied with some promise to baboon (Ackermann et al.2006, 2014; Eichel and Ackermann 2016) and macaque (Ito et al.2015) hybrids. Studies of craniofacial shape have used 2D point-to-point measurements to study craniofacial variation in three dimensions (Ackermann et al.2006, 2014), including targeted studies of nasal cavity (Eichel and Ackermann 2016) and the maxillary sinus (Ito et al.2015). The findings of these studies echo the broader literature on primate hybridization, indicating that hybrid craniofacial morphology is highly variable and that hybrids are very likely to display novel or transgressive phenotypes. A series of minor developmental anomalies were also identified in the hybrid baboons (Ackermann 2010; Ackermann et al.2006, 2014), a population of hybrid wildebeest (Ackermann et al.2010), and a proposed population of hybrid gorillas (Ackermann and Bishop 2010). These minor nonmetric variants (developmental abnormalities) have been suggested to be attributable to a breakdown in the coordination of early development caused by hybridization. How and whether these traits manifest in other species remains to be seen.
Macaques (genus Macaca) are an ideal study taxon for the further investigation of hybrid morphology. The genus has an exceptional biogeographical range and is represented by a large number of species that have the documented propensity to hybridize wherever their ranges meet (e.g., Bynum 2002; Bynum et al.1997; Evans et al.2001; Fooden 1964; Hamada et al.2006, 2008, 2012, 2016; Ito et al.2015; Jiang et al.2016; Malaivijitnond et al.2008; Schillaci et al.2005). An excellent example is a population of hybrids between of the Japanese macaque (M. fuscata) and Taiwanese macaque (M. cyclopis) (Hamada et al.2012). Contact between the two island-bound species first began in the 1970s, when the breakdown of a private zoo in the Ooike area of Wakayama led to the release or escape of several M. cyclopis individuals into the population of M. fuscata. The invasive M. cyclopis successfully adapted to the environment and interbred unchecked with M. fuscata (Hamada et al. 2012; Kawamoto 2005) until 2003, when the Wakayama prefectural government initiated an extermination project designed to remove M. cyclopis and their hybrids from the local population of M. fuscata. The Working Group of Hybrid Macaques was established to study the genetics, demography, and morphology of the hybrids (e.g., Hamada et al.2012; Ito et al.2015; Kawamoto 2005; Yamagiwa 2010). The resultant skeletal collection is likely one of the largest and best studied hybrid skeletal collections known, and a valuable resource for the study of hybrid morphology.
We have applied 3D geometric morphometric methods to the sample of hybrid Macaca fuscata and M. cyclopis to explore the craniofacial morphology of the hybrids and parental species. We also assessed the presence and significance of minor developmental abnormalities as observed in other primate hybrids (Ackermann 2010). The first aim of the study was to document quantitative and qualitative aspects of the craniofacial morphology and to explore their relationships to degree of hybridity. Though this part of the research was intended to be exploratory rather than hypothesis driven, observed broad trends in hybrid phenotype guided our expectations. It has been hypothesized (e.g., Ackermann et al.2006; Kelaita and Cortés-Ortiz 2013) that the high degree of variability and sometimes novel or transgressive phenotypes observed in primate hybrids results from a complex suite of heterotic, dysgenetic, and neutral interactions between the parental genotypes, and that the effect of these is reduced in later generation hybrids. Based on this, we made three broad predictions about the presentation in our sample:
-
1.
Hybrid morphology would be highly variable and possibly transgressive but would largely fall within the range of the parental morphotypes.
-
2.
Genetically intermediate hybrids were likely to be more morphologically variable than any other group.
-
3.
Later generation hybrids were likely to be most similar to the parent species with which they share the most DNA.
The second aim of the study was to test the hypothesis that the breakdown of coordination in early development would result in a range of minor developmental abnormalities in the craniofacial skeleton and dentition (Ackermann 2010). If the hypothesis is correct, we predicted that we would identify minor developmental abnormalities in higher frequency in the hybrids than in the parent species. For the same reasons we might also expect to see a greater degree of fluctuating asymmetry in the hybrid sample, though this has yet to be discussed in this context.
Methods
Samples
We collected 3D geometric morphometric (3D GMM) and nonmetric data from 178 adult primates. One hundred and fifty-five of the specimens are housed at the Kyoto University Primate Research Institute (KUPRI) and a further 23 specimens are housed at the Smithsonian Museum of Natural History in Washington, DC. We also examined CT scans of 177 individuals (from both parental species and their hybrids) for the presence of other dental nonmetric variants. We took 51 new CT scans from a mix of hybrid and parental individuals. Forty-five CT scans of hybrid individuals were provided by Dr. T. Ito from other research undertaken at KUPRI (Ito et al.2015), and we gathered 81 from the KUPRI Digital Morphology Museum. All scans that we included in the study were of sufficient quality to easily observe nonmetric dental morphology. There was partial overlap between the CT scan and physical (3D GMM and nonmetric) data sets; we included 105 individuals in both data sets, 72 only in the CT scan data set, and 73 only in the physical data set. In total we included 250 individuals in the study (Table I).
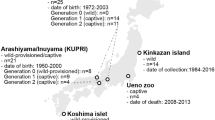
Species assignment in the sample is fairly robust; we obtained most pure-bred samples from collections in which interaction with other species was impossible, due to either geographic isolation or controlled breeding. We identified hybrids based on live observation data (group membership) and tail length, which has been determined to be a reliable indicator of hybridity in this population (Hamada et al.2012). Of the 70 hybrids we included in the sample, genetic data from other research indicating degree of hybridization are available for 39. Detailed reporting of the methodology and results are available in a series of publications (Hamada et al.2012; Kawamoto 2005; Kawamoto et al.1999, 2001). In brief, degree of hybridization (hybrid index) was estimated from eight alleles of four marker gene loci that were typed as either Macaca cyclopis or M. fuscata and is given on a scale from 0/8 (M. cyclopis) to 8/8 (M. fuscata) (Table II). Three diagnostic blood proteins (plasma transferrin [TF], blood cell adenosine deaminase [ADA] and blood cell diaphorase [DIA]) were tested using starch gel electrophoresis and staining (amido black for TF and formazan for ADA and DIA) (Kawamoto et al.1999, 2001). The fourth, a single nucleotide polymorphism (SNP) of the NRAMP 1 protein, was tested using polymerase chain reaction (PCR) and direct sequencing from whole blood (Kawamoto 2005). In addition, the natal origin of each individual was assessed using the Y-linked testis-specific protein (TSPY) (paternal origin) and hypervariable region II of the mitochondrial D-loop (maternal origin), using PCR restriction fragment length polymorphism analysis (PCR-RFLP) to classify each of the regions as either M. cyclopis or M. fuscata type (Kawamoto et al.2001; Hamada et al.2012).
Ideally we would have assigned hybrids to categories according to whether they are first generation (F1) hybrids, later generation hybrids (Fn), or single or multigenerational backcrosses (Bn). Lacking pedigree data, however, this was impossible. For example, one male individual has a hybrid index of 6/8, placing it genetically closest to Macaca fuscata (M. cyclopis = 0/8, M. fuscata = 8/8) and appearing to indicate that it is a first-generation backcross (B1) between an F1 hybrid (4/8) and a pure-bred M. fuscata. However, both the TSPY (testis-specific protein encoded on the Y chromosome) and mtDNA of this individual are M. cyclopis type, a pattern that would be impossible to obtain if this individual had not had both a male and a female M. cyclopis ancestor. This is likely the result of multigenerational hybridization and backcrossing. As such, we made no assumptions regarding the generational status or parental composition of hybrids. Of the 39 hybrids for which genetic data are available, 7 either have minimal hybrid ancestry or are genetically indistinguishable from M. cyclopis. Of the remaining hybrids, 7 are genetically intermediate, 15 are closer to M. cyclopis (M. cyclopis type), and 10 are closer to M. fuscata (M. fuscata type).
Data Collection
Metric Data
We collected landmark data from all crania for 3D geometric morphometric analyses of shape and shape variance between individuals and groups. A single observer collected all landmark data using a MicroScribe™ G2X desktop digitizer (Immersion Corporation, San Jose, CA, USA), with MicroScribe Utility Software v4.0 and Microsoft Office Excel 2010. We collected landmarks in two object orientations that were subsequently “stitched” together using FileConverter (http://www.flywings.org.uk).
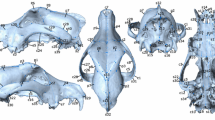
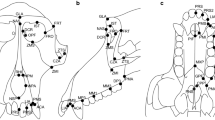
We selected basic landmarks from existing sets of standard osteological landmarks for both human and nonhuman primates (Howells 1973; White and Folkens 2000) and added further purpose designed landmarks as necessary to ensure that meaningful structures were captured, and that overall craniofacial shape was well represented. Of the 67 cranial landmarks originally selected, we used 66 in the final analyses (Table III); we removed one landmark because of consistent difficulties with identification and recording accuracy.
Nonmetric Data
We conducted nonmetric data collection in two parts: we visually examined all samples on which metric data had been collected, and observed CT scans for abnormalities that were not externally visible. Our selection of characters was heavily influenced by those that had been identified in previous studies of hybrid developmental abnormalities (Ackermann 2010; Ackermann et al.2006, 2014); these include supernumerary teeth, rotated teeth, supernumerary sutures or ossicles in the zygomaxillary region, remnant metopic sutures, dental crowding, ossicles at lambda and asterion, bregmatic bones, coronal ossicles, epipteric bones, and parietal notch bones. We also considered similar nonmetric traits that have been observed in other (nonhybrid) human and primate studies, such as infraorbital suture persistence and supraorbital notches (Manzi et al.2000), particularly where these traits are considered as potential indicators of developmental disturbance (e.g., Gruneberg 1963; Kaur et al.2012; Korey 1980; Manzi 1996; Manzi et al.2000; Trinkaus 1978). We refined the final character list (Table IV) to conform to actual manifestation on the macaque cranium; we discounted traits that were not identified in the sample and included some previously unidentified traits (such as a supernumerary suture in the premaxilla). We scored the characters as present or absent and added scores for variation in appearance where relevant—for example, full or partial metopism, or the location or number or teeth affected.
The KUPRI Digital Morphology Museum houses CT and MRI scans from several institutions. Details of scanning methodologies for each individual sample are available on the KUPRI DMM website (http://dmm.pri.kyoto-u.ac.jp/dmm/WebGallery/index.html). For consistency within this study, we selected CT scans taken at the KUPRI using the same or similar methodology.
We collected further CT scans (N = 51) using the same methodology as for the scans provided by Dr. T. Ito (N = 45) (Ito et al.2015). We used a helical scanner (Asterion Premium 4 Edition; Toshiba Medical Systems Co. Ltd., Otawara, Japan) at the KUPRI in Inuyama (Aichi, Japan), with slice thickness of 0.5 mm and a slice interval of 0.2 or 0.3 mm, and pixel size ranging from 0.166 mm × 0.166 mm to 0.314 mm × 0.314 mm. We viewed the scans using Sante DICOM viewer (64-bit) version 3.0.11 (©1998–2014 Santesoft).
Statistical Analysis
Metric Data
We conducted morphometric analyses using MorphoJ (Klingenberg 2011). First we subjected the data to a generalized Procrustes analysis (GPA), which centers all shapes at the origin (0.0.0), optimally translating and rotating the specimens, while removing the size component with uniform scaling (Dryden and Mardia 1998; Klingenberg 2011; Rohlf 1990, 1999). MorphoJ employs a full Procrustes fit, which uses orthogonal projection to cast the data in a tangent space (Dryden and Mardia 1998), and is advantageous for data sets with a large amount of variation. Less weight is placed on observations that are far away from the average shape, meaning the results are more robust against the influence of outliers (Klingenberg 2011). The program allows the user to specify the type of symmetry of the samples (e.g., object symmetry vs. matching symmetry [Klingenberg 2015]) and takes this into account.
We generated covariance matrices from the symmetric component of the data and used these for principal components analysis (PCA) and canonical variates analysis (CVA) within MorphoJ. PCA extracts the major features of shape variation in the entire data set and arranges data points along the major axes of variation. Variance is measured in units of Procrustes distance, which is approximately the square root of the sum of squared differences between the positions of the two landmarks in optimally aligned configurations (Klingenberg and McIntyre 1998; Slice et al.1996). Principal components are largely statistical artefacts rather than representing biologically meaningful phenomena (Mitteroecker et al.2005; Mitteroecker and Gunz 2009) and should not be interpreted as such, although in a single species (and possibly in very closely related species), PC1 is likely to represent allometric shape variation (Klingenberg 1998). CVA performed on these principal components examines the separation between groups of observations and maximizes the separation between groups by scaling for within group variation. In the CVA, Mahalanobis distances are preferred over Procrustes distances because they account for the nonisotropic nature of the data (Klingenberg and Monteiro 2005).
We used the asymmetric component of the data to quantify fluctuating asymmetry via Procrustes ANOVA using MorphoJ (Klingenberg 2011, 2015; Klingenberg and McIntyre 1998) and compared the population measure of fluctuating asymmetry to the error effect to assess whether error was problematic in this analysis (Klingenberg 2015). We assessed measurement error via repeated digitization of a single specimen before, at intervals throughout, and at the completion of the data collection process (a total of 10 measurements) and compared the total variance for all samples to the variance of the repeated measurements. We then conducted pairwise F tests for equality of variances between the two parental species (Macaca fuscata and M. cyclopis) and between the hybrids and each of the parent species.
Nonmetric Data
To facilitate comparison with relevant prior studies (e.g., Ackermann 2010; Ackermann et al.2006, 2014), we applied χ2 analyses to test the significance of trait frequency variations between groups. Wherever multiple pairwise comparisons are made there is a risk of introducing type 1 error through alpha-inflation, so we applied a Bonferroni correction to the resulting P values (new significance level = α/m, where α = 0.05 and m = the number of hypotheses). Where sample size for any given trait was inappropriately small for χ2 (N < 5), we applied Fisher’s Exact test instead.
Pairwise χ2 tests, though widely used, are poorly suited to the data and to the research questions; they cannot address covariation between traits or to assess which factors may be significant in explaining the pattern of trait frequencies. As such, we conducted further statistical analyses using a model-based approach in the program R (R Core Team 2015), using the mvabund package (Wang et al.2012). Using this model-based approach, we fitted abundance (presence/absence) data to generalized linear models using the manyglm function, which fits generalized linear models to multivariate abundance data. There is an assumed relationship between the means and variances of the variables and an assumed relationship between mean abundance and environmental variables. The mean–variance relationship can be assessed by plotting it directly; a linear relationship was achieved, indicating that the data are appropriate. The second assumption is unimportant here because the model includes orthogonal factors only (Wang et al.2012). The manyglm function is designed to handle this type of nonnormal multivariate abundance count or binary data for which a distribution can be specified, which offers a reasonable model for the data (Wang et al.2018). We constructed a series of models for each of the effects being tested for—in this case, species membership, hybridity, and sex. We then compared each of these models (Table V) to a null model in which there was no effect using analysis of deviance (α = 0.05). In this case, deviance is a measure of unexplained variance and quantifies the discrepancy between a null model (no effect) and the applied model. Deviance is usually not interpreted directly, but rather in comparison with other models fitted to the same data. Similarly, we assessed trait covariance by comparing a model in which there is a positive correlation between traits to a null model with no correlation between traits (1000 bootstrap iterations). Owing to the difference in sample composition (physical specimens vs. CT scans; see Table I), we treated surface nonmetric characters and CT results separately.
Results
Metric Data
Error Assessment
Error accounted for 1.37% of overall variance (repeated measure variance = 8.22E-05, total sample variance = 5.99E-03).
Principal Components Analysis
Approximately half of the variation in the sample was explained by the first four principal components, of which PC1 accounts for 29.83% (Fig. 1). Overall there was very little differentiation between the groups except in PC2, where Macaca cyclopis occupied the negative extreme and M. fuscata occupied the positive extreme (Figs. 1 and 2). In PCs 1 and 2 the hybrids were more closely clustered with M. fuscata and in PC3 they exceeded the range of either parent. Both parental species exhibited a greater amount of variance than the hybrids (Table VI).
Principal component scatterplots and percentage variance explained in 3D shape analyses of hybrid macaques (M. fuscata × M. cyclopis) and parental species. Equal frequency ellipses are drawn to a probability of 0.95. Shape changes associated with each of the PCs (eigenvectors) are plotted as Procrustes distances to scale factor of 0.1.
Shape changes associated with each of the first four principal components in 3D shape analyses of hybrid macaques (M. fuscata × M. cyclopis) and parental species. The wireframe reflects shape at the negative end of each PC axis (see Fig. 2) and bold lines indicate shape change progressing positively along the axis. Shape change associated with each PC is presented in superior, lateral, and anterior views (respectively).
When subdivided according to degree of hybridity (Table II), Macaca fuscata–type and M. cyclopis–type hybrids clustered most closely with the more closely related parental species. Intermediate hybrids were entirely subsumed within the range of M. fuscata but still overlapping with M. cyclopis, especially in PC1 (Fig. 3). Some sample sizes were so small as to be uninformative at this point, and so this division is impractical for gaining meaningful or statistically significant results.
Principal component scatterplot and percentage variance explained in 3D shape analyses of hybrid macaques (M. fuscata × M. cyclopis) and parental species, with hybrids subdivided according to degree of hybridity. Equal frequency ellipses are drawn to a probability of 0.95. Shape changes associated with each of the PCs (eigenvectors) are plotted as Procrustes distances to scale factor of 0.1.
Canonical Variates Analysis
More than two-thirds of the variance between groups was accounted for by CV1 (Fig. 4), in which the hybrids sat clearly with Macaca fuscata at the positive end of the axis, to the exclusion of M. cyclopis. In CV2, accounting for the remaining third of the variance, the three groups were more distinct but retained a small degree of overlap at the margins. Mahalanobis distances between groups indicated that the hybrids are most similar to M. fuscata, and that both M. cyclopis and M. fuscata are more similar to the hybrids than they are to one another (Table VII).
Canonical variate scatterplots and percentage variance explained in 3D shape analyses of hybrid macaques (M. fuscata × M. cyclopis) and parental species (a) and with hybrids subdivided according to degree of hybridity (b). Equal frequency ellipses are drawn to a probability of 0.95. Shape changes associated with each of the PCs (eigenvectors) are plotted as Procrustes distances to scale factor of 0.1.
When subdivided according to degree of hybridity the hybrids remain clustered with one another rather than associating with the parental species with which they share the most ancestry (Fig. 4). As in the PCA, sample sizes subdivided according to degree of hybridity are too small for statistically significant results.
Fluctuating Asymmetry
The hybrids had the lowest mean FA score and the lowest individual FA score over all (Fig. 5). Kruskal–Wallis (H(χ2): 14.43, P = 0.0007, N = 171) indicated that the hybrids can be distinguished statistically from Macaca cyclopis (Dunn’s post hoc P = 0.0002) and M. fuscata (Dunn’s post hoc P = 0.023), but that M. cyclopis and M. fuscata cannot be distinguished from one another (Dunn’s post hoc P = 0.144).
Mean squares of the FA effect (3.29E-06) divided by the associated error effect (7.98E-07) resulted in a ratio of 4.12, indicating that the effect of fluctuating asymmetry is somewhat larger than the error effect but that the difference is not large enough for the effect of error to be considered negligible; FA results should therefore be interpreted with caution.
Nonmetric Data
Pairwise χ2 testing identified several statistically significant differences in trait frequency between the groups (Tables VIII and IX). Hybrids compared to a composite of both parental species had a significantly greater frequency of anomalous premaxillary sutures (Fig. 6a and b), the tuberculum intermedium (Fig. 6h), and the absence of one or more cusps on the lower third molars (M3) (Fig. 6f and g). When compared to each parental species individually, only the anomalous premaxillary suture and the presence of the tuberculum intermedium were significant compared to both parents. The absence of a cusp on the M3 was significantly more frequent only when compared to Macaca fuscata, and statistically significant differences appeared in the presence of interconuli (Fig. 6k and l) and anomalous supranasal sutures (Fig. 6e) (vs. M. cyclopis), and rotated lower fourth premolars (P4) (Fig. 6i and j) and anomalous sutures in the temporal fossae (Fig. 6c and d) (vs. M. fuscata). Compared to one another, the parental species differed significantly in the appearance of anomalous sutures in the temporal fossae, anomalous supranasal sutures, and P4 rotation. Differences between the sexes were minimal; neither parental species had a significant difference between males and females, but hybrid females were more likely to have cusp number variations in the M3 than hybrid males. Females were generally more likely than males to differ significantly from the parental composite.
We consistently found group membership and degree of hybridity to be significant explanatory factors in the model-based analyses conducted in R (P < 0.05), though the magnitude of the effect (deviance) differed between sets (Table X). An exception is the CT data in set 8, where group membership was not found to be a significant explanatory factor. Sex also did not have a significant effect on trait frequency.
Hybrids differ significantly from each parent species separately and from the composite of the parent species (sets 1 and 2, Table X). The parent species also differ significantly from one another. The amount of deviance explained (magnitude of the effect) is greater between the parent species (77.48) and between the hybrids and Macaca fuscata (72.81) than it is between the hybrids and M. cyclopis (47.52), indicating that the hybrids are more like M. cyclopis than M. fuscata (set 3). Degree of hybridity is a significant explanatory factor whether group membership is considered or not (sets 4 and 5), and both when compared to the parental composite and when hybrids are considered alone (sets 6 and 7).
Discussion
3D analyses of shape indicated that the hybrids fall largely within the range of the parental morphotypes, with some slightly transgressive morphological components. There appears to be a slight tendency for nonintermediate hybrids to more closely resemble the parent with which they share more DNA, but overall the hybrids tended toward the Macaca fuscata morphotype regardless of genetic ancestry. The nonmetric data similarly identified a small number of transgressive features in the hybrids, but the analysis of deviance appeared to suggest a closer affinity to M. cyclopis.
Craniofacial Shape
The large degree of overlap and complexity of shape variability prohibit generalizations about the hybrid morphotype. We could not clearly distinguish hybrids from either parental species, nor were the parental species adequately distinguishable from one another using PCA. The hybrids sit further toward the positive end of the PC3 axis than either parent, indicating the presence of some slightly transgressive morphological components, but it is impossible from this to identify a particular feature or character state that distinguishes hybrids from either parent.
Despite the general overlap in the three group morphotypes, the CVA discriminated effectively between Macaca fuscata, M. cyclopis, and the hybrids. CV1, which described more than two-thirds of the total variance between the groups, very clearly aligned the hybrids with the M. fuscata morphotype. CV2, describing the other third of the variation, separated all three groups to a good degree—there was overlap only between the outliers of the M. fuscata and M. cyclopis groups, and only a small degree of overlap between the M. cyclopis and hybrid groups. The corresponding Mahalanobis distances between groups supported the observation that the hybrids are morphologically more similar to M. fuscata than to M. cyclopis; this observation is particularly interesting in light of the fact that the available genetic data indicate that the hybrid sample tends to be more closely related to M. cyclopis. Despite this tendency toward the M. fuscata morphotype, all hybrids, regardless of the degree of hybridity, clustered with one another rather than with either of the parental species when subjected to CVA. This suggested that they exhibit a unique hybrid morphology that is distinct from both parental species.
We predicted that hybrids would be more variable than either parent sample and that intermediate hybrids would be more variable than backcrossed hybrids. Owing to the limited availability of genetic data, it is impossible to determine relative degrees of variability within the hybrid sample. Interestingly, though, data for the hybrid group as a whole indicated that they show markedly less total sample variance than either parent, even with their greater overall sample size. This contradicts the results from other studies (e.g., Ackermann et al.2006 and Kelaita and Cortés-Ortiz 2013), which have indicated that hybrids, particularly genetically intermediate hybrids, are highly variable with respect to parental species.
We also predicted that hybrids would demonstrate a greater degree of fluctuating asymmetry than the parental species, owing to the breakdown or disruption of developmental constraints which may be associated with hybridization (e.g., Tomkins and Kotiaho 2001). The macaque hybrids appeared to be significantly less affected by fluctuating asymmetry than either parental species; however, digitizing error is not sufficiently low in comparison to the degree of fluctuating asymmetry for the effect to be considered negligible, and so these results should be interpreted with caution.
Nonmetric Variation
Several anomalous nonmetric traits were identified in all three sample groups. Multivariate analyses indicated that all three groups (hybrids, Macaca cyclopis, and M. fuscata) differed significantly in trait frequencies. Hybrids could be distinguished from each parental species and from an amalgamated sample of the parental species together (notionally “pure-bred”), but the parental species were also found to differ significantly from one another. Trait frequency patterning did vary significantly by degree of hybridity, appearing to suggest an effect of hybridization, but it was impossible to establish whether this was a true effect or whether it might be due to affinities between the backcrossed specimens and the more closely related parental species. There was no clear hybrid/pure-bred dichotomy in the overall appearance and frequency of anomalous nonmetric traits. This method of analysis considered the pattern of trait frequencies and did not isolate particular traits or sets of traits that distinguish groups. Pairwise significance testing of individual trait frequency identified a small number of traits that differed significantly from one parental species to the other and three traits that differed from the “pure-bred” composite. Only an increased frequency of anomalous sutures in the premaxilla and the presence of the tuberculum intermedium consistently differ between hybrids and parental/“pure-bred” samples. Though the hybrids differed by sex in the frequency of cusp number variations in the M3, there were no significant sex differences within either parental species or the “pure-bred” composite. Similarly, the multivariate analyses found no significant effect of sex in any of the scenarios. The effect of sex, though not rejected entirely, was contextually minimal.
Understanding the Hybrid Morphotype
So far as general shape is concerned, the results of this study are largely compatible with findings from prior research. The hybrids were largely within the range of the parental species, were mildly transgressive in some respects, and there was a tendency for the hybrids to cluster more closely with one parent (Macaca fuscata) than the other. It is somewhat unexpected that the hybrids appeared to cluster more closely with M. fuscata considering the available genetic data, which seemed to indicate that the hybrid sample were genetically more closely related to M. cyclopis. However, genetic information is available for only a little more than half of the hybrid sample and may not be representative of the true sample composition.
It is possible that the hybrid tendency toward the Macaca fuscata morphotype is related to sex. No specific hypotheses were made regarding the effect of sex on hybrid morphotype, owing largely to the fact that differences associated with sex have in the past been highly unpredictable. Prior studies have identified a distinct or different effect of hybridization in one sex (Charpentier et al.2008, 2012; Delmore et al.2011; Eichel and Ackermann 2016), and a preference for each sex to appear more like different parental species (e.g., Kelaita and Cortés-Ortiz 2013); it appears that there may be sexual differences in the expression of hybridity that extend beyond normal primate sexual dimorphism. Furthermore, asymmetric gene flow and sexually biased introgression are frequently observed among primate hybrids (e.g., Charpentier et al.2012; Evans et al.2001; Tung et al.2008), often associated with an apparent mismatch between phenotype and genetic affiliation. Though the physical differences between the sexes were found to be statistically negligible in this sample, these prior findings suggest that sex should not be discounted as a possible contributor to the observed morphological bias toward M. fuscata. On the other hand, it is not necessarily required to invoke sexual factors to explain the observed morphology; expression level dominance of one parent over the others is not an unusual occurrence in interspecies hybridization (e.g., Combes et al.2015), regardless of sex.
Though truly intermediate morphology appears to be a relatively rare outcome of hybridization, some traits have been identified in which phenotype does correspond well with the degree of hybridity. A clear relationship between tail length and degree of hybridity has been identified in this same hybrid population (Hamada et al.2012). There are two reasons why tail length and craniofacial shape may offer dissimilar results in this case. The craniofacial complex is considered to be composed of two (e.g., Engleka et al.2008) or three (e.g., Sperber 1989) main components or developmental modules: the neurocranium (1), which comprises the bones that encase the brain, and the viscerocranium, which may be further divided into the facial skeleton (2) and masticatory apparatus (3). These major components are highly integrated with one another, yet each has different patterns of growth, development, maturation, and function, determined by the morphogenetic fields in which they develop. In contrast, control of tail length appears to be much simpler. Mammalian vertebrae are determined by the Hox gene complex, with the development of the caudal vertebrae thought to be controlled by Hoxd 10–13 gene expression (Wolpert et al.2007). Hybrids between Macaca cyclopis and M. fuscata have been found to have anywhere from 9 to 24 caudal vertebrae, largely encompassing the ranges of both parental species. Relative tail length (percent crown–rump length) has been found to correlate well with the number of caudal vertebrae, and also with degree of hybridization (Hamada et al.2012); in this case, the combination of the two parental genotypes may interact under a simple additive model with minimal epistatic effects (e.g., Cheverud and Routman 1995; Falconer 1989).
It is possible that the parental populations used in this study may not accurately represent the local parental populations in Wakayama. Actual specimens used were a combination of captive bred, traded, and wild caught specimens from source populations all over Japan. In light of the relatively minor nature of the craniofacial shape differences between the two species and large degree of overlap, it is unlikely that local population variation would have had much effect on these results. It is more likely that this could affect the nonmetric trait frequencies, many of which are known to vary considerably between human populations (e.g., Bauchau 1988; Berry 1975; Hanihara and Ishida 2001a, b, and c). Furthermore, there are poorly understood relationships between relevant trait frequencies and variables such as age and sex (e.g., Epstein 2008; Hauser and DeStefano 1989; Perizonius 1979). Allowing for local variation in the parental populations, it remains unlikely that the Wakayama macaques would vary significantly from the larger parental species samples in any more than a few of the observed traits. The significance of local population variation in the frequency and manifestation of nonmetric variants is an avenue worth considering for further study.
Finally, the unexpected relative reduction in variability and fluctuating asymmetry, as well as the general lack of increase in developmental abnormalities with respect to the parental populations, may be at least partially explained by the relatively small degree of evolutionary divergence between the parental populations. In contrast to many other hybridizing primates, Macaca fuscata and M. cyclopis are very closely related, having diverged as recently as 0.17 million years ago (Chu et al.2007). Though this seems intuitive, a general relationship between the degree of evolutionary divergence between species and the morphological consequences of hybridization has yet to be established; the bulk of the literature describing the outcomes of hybridization refers to intraspecific strain crosses, rather than true interspecific hybridization (e.g., Baranwal et al.2012; Chen 2010). It is often cited as fact that the degree of heterosis in hybrids increases proportionally with the degree of genetic divergence between the two lineages, (e.g., Chen 2010; Groszmann et al.2013), but this generalization is contingent on the outdated (i.e., Muller 1925) assumption that mammalian interspecific hybrids are extremely rare and generally sterile (e.g., Chen 2010). Interspecific hybrids simply are not included in the models, and so we have little basis on which to draw inferences. Fortunately, with the current rate of accumulation of new data on interspecific and even intergeneric (e.g., Chiarelli 1973; Dunbar and Dunbar 1974; Jolly and Disotell 1997; Markarjan et al.1974; Moore et al.1999) hybrids, this need no longer be the case. Further research capitalizing on this growing and varied database would go a long way toward building a clearer and more nuanced understanding of the consequences of hybridization across all taxa, including primates..
References
Ackermann, R. R. (2010). Phenotypic traits of primate hybrids: Recognizing admixture in the fossil record. Evolutionary Anthropology, 19(6), 258–270.
Ackermann, R. R., & Bishop, J. M. (2010). Morphological and molecular evidence reveals recent hybridization between gorilla taxa. Evolution, 64(1), 271–290.
Ackermann, R. R., Rogers, J., & Cheverud, J. M. (2006). Identifying the morphological signatures of hybridization in primate and human evolution. Journal of Human Evolution, 51(6), 632–645.
Ackermann, R. R., Brink, J. S., Vrahimis, S., De Klerk, B., & Klerk, B. (2010). Hybrid wildebeest (Artiodactyla: Bovidae) provide further evidence for shared signatures of admixture in mammalian crania. South African Journal of Science, 106(11–12). https://doi.org/10.4102/sajs.v106i11/12.423.
Ackermann, R. R., Schroeder, L., Rogers, J., & Cheverud, J. M. (2014). Further evidence for phenotypic signatures of hybridization in descendant baboon populations. Journal of Human Evolution, 76(C), 54–62.
Aguiar, L. M., Mellek, D. M., Abreu, K. C., Boscarato, T. G., Bernardi, I. P., et al (2007). Sympatry between Alouatta caraya and Alouatta clamitans and the rediscovery of free-ranging potential hybrids in southern Brazil. Primates, 48(3), 245–248.
Arnold, M. L. (1997). Natural hybridization and evolution. Oxford: Oxford University Press.
Arnold, M. L., & Meyer, A. (2006). Natural hybridization in primates: one evolutionary mechanism. Zoology, 109(4), 261–276.
Arnold, M. L., Sapir, Y., & Martin, N. H. (2008). Genetic exchange and the origin of adaptations: prokaryotes to primates. Philosophical Transactions of the Royal Society B: Biological Sciences, 363(1505), 2813–2820.
Baranwal, V. K., Mikkilineni, V., Zehr, U. B., Tyagi, A. K., & Kapoor, S. (2012). Heterosis: Emerging ideas about hybrid vigour. Journal of Experimental Botany, 63(18), 6309–6314.
Bauchau, V. (1988). Non-metrical variation in wild mammals: A bibliography. Mammal Review, 18(4), 195–200.
Berry, A. C. (1975). Factors affecting the incidence of non-metrical skeletal variants. Journal of Anatomy, 120(3), 519–535.
Bicca-Marques, J. C., Prates, H. M., Aguiar, F. R. C., & Jones, C. B. (2008). Survey of Alouatta caraya, the black-and-gold howler monkey, and Alouatta guariba clamitans, the brown howler monkey, in a contact zone, State of Rio Grande do Sul, Brazil: Evidence for hybridization. Primates, 49(4), 246–252.
Bookstein, F. L. (1991). Morphometric tools for landmark data: geometry and biology. Cambridge: Cambridge University Press.
Burke, J., & Arnold, M. (2001). Genetics and the fitness of hybrids. Annual Review of Genetics, 35, 31–52.
Bynum, N. (2002). Morphological variation within a macaque hybrid zone. American Journal of Physical Anthropology, 118(1), 45–49.
Bynum, E. L., Bynum, D. Z., & Supriatna, J. (1997). Confirmation and location of the hybrid zone between wild populations of Macaca tonkeana and Macaca hecki in Central Sulawesi, Indonesia. American Journal of Primatology, 43(3), 181–209.
Charpentier, M. J. E., Tung, J., Altmann, J., & Alberts, S. C. (2008). Age at maturity in wild baboons: Genetic, environmental and demographic influences. Molecular Ecology, 17, 2026–2040.
Charpentier, M. J. E., Fontaine, M. C., Cherel, E., Renoult, J. P., Jenkins, T., et al (2012). Genetic structure in a dynamic baboon hybrid zone corroborates behavioural observations in a hybrid population. Molecular Ecology, 21(3), 715–731.
Chen, Z. J. (2010). Molecular mechanisms of polyploidy and hybrid vigor. Trends in Plant Science, 15(2), 57–71.
Cheverud, J. M., & Routman, E. J. (1995). Epistasis and its contribution to genetic variance components. Genetics, 139(3), 1455–1461.
Chiarelli, B. (1973). Check-list of catarrhina primate hybrids. Journal of Human Evolution, 2(4), 259–355.
Chu, J. H., Lin, Y. S., & Wu, H. Y. (2007). Evolution and dispersal of three closely related macaque species, Macaca mulatta, M. cyclopis, and M. fuscata, in the eastern Asia. Molecular Phylogenetics and Evolution, 43(2), 418–429.
Combes, M. C., Hueber, Y., Dereeper, A., Rialle, S., Herrera, J. C., & Lashermes, P. (2015). Regulatory divergence between parental alleles determines gene expression patterns in hybrids. Genome Biology and Evolution, 7(4), 1110–1121.
Cortés-Ortiz, L. (2017). Hybridization and hybrid zones. In The international encyclopedia of primatology. Wiley Online Library.
Delmore, K. E., Louis, E. E., & Johnson, S. E. (2011). Morphological characterization of a brown lemur hybrid zone (Eulemur rufifrons × E. cinereiceps). American Journal of Physical Anthropology, 145(1), 55–66.
Delmore, K. E., Brenneman, R. A., Lei, R., Bailey, C. A., Brelsford, A., et al (2013). Clinal variation in a brown lemur (Eulemur spp.) hybrid zone: Combining morphological, genetic and climatic data to examine stability. Journal of Evolutionary Biology, 26(8), 1677–1690.
Dryden, L., & Mardia, K. V. (1998). Statistical shape analysis. Hoboken: Wiley.
Dunbar, R. I. M., & Dunbar, P. (1974). On hybridization between Theropithecus gelada and Papio anubis in the wild. Journal of Human Evolution, 3(3), 187–192.
Eichel, K. A., & Ackermann, R. R. (2016). Variation in the nasal cavity of baboon hybrids with implications for late Pleistocene hominins. Journal of Human Evolution, 94, 134–145.
Engleka, K. A., Lang, D., Brown, C. B., Antonucci, N. B., & Epstein, J. A. (2008). Neural crest formation and craniofacial development. In C. J. Epstein, R. P. Erickson, & A. Wynshaw-Boris (Eds.), Inborn errors of development: The molecular basis of clinical disorders of morphogenesis (pp. 69–78). Oxford Monographs on Medical Genetics. Oxford: Oxford University Press.
Epstein, C. J. (2008). Human malformations and their genetic basis. In C. J. Epstein, R. P. Erickson, & A. Wynshaw-Boris (Eds.), Inborn errors of development: The molecular basis of clinical disorders of morphogenesis (pp. 3–8). Oxford Monographs on Medical Genetics. Oxford: Oxford University Press.
Evans, B. J., Supriatna, J., & Melnick, D. J. (2001). Hybridization and population genetics of two macaque species in Sulawesi, Indonesia. Evolution, 55(8), 1686–1702.
Falconer, D. S. (1989). Introduction to quantitative genetics (3rd ed.). New York: Longman.
Fooden, J. (1964). Rhesus and crab-eating macaques: Intergradation in Thailand. Science, 143(3604), 363–364.
Fuzessy, L. F., Silva, I. D. O., Malukiewicz, J., Silva, F. F. R., Pônzio, M. D. C., et al (2014). Morphological variation in wild marmosets (Callithrix penicillata and C. geoffroyi) and their hybrids. Evolutionary Biology, 41(3), 480–493.
Gligor, M., Ganzhorn, J. U., Rakotondravony, D., Ramilijaona, O. R., Razafimahatratra, E., et al (2009). Hybridization between mouse lemurs in an ecological transition zone in southern Madagascar. Molecular Ecology, 18(3), 520–533.
Groszmann, M., Greaves, I. K., Fujimoto, R., James Peacock, W., & Dennis, E. S. (2013). The role of epigenetics in hybrid vigour. Trends in Genetics, 29(12), 684–690.
Gruneberg, H. (1963). The pathology of development: A study of inherited skeletal disorders in animals. New York: Wiley.
Hamada, Y., Urasopon, N., Hadi, I., & Malaivijitnond, S. (2006). Body size and proportions and pelage color of free-ranging Macaca mulatta from a zone of hybridization in northeastern Thailand. International Journal of Primatology, 27(2), 497–513.
Hamada, Y., Suryobroto, B., Goto, S., & Malaivijitnond, S. (2008). Morphological and body color variation in Thai Macaca fascicularis fascicularis north and south of the Isthmus of Kra. International Journal of Primatology, 29(5), 1271–1294.
Hamada, Y., Yamamoto, A., Kunimatsu, Y., Tojima, S., Mouri, T., & Kawamoto, Y. (2012). Variability of tail length in hybrids of the Japanese macaque (Macaca fuscata) and the Taiwanese macaque (Macaca cyclopis). Primates, 53(4), 397–411.
Hamada, Y., San, A. M., & Malaivijitnond, S. (2016). Assessment of the hybridization between rhesus (Macaca mulatta) and long-tailed macaques (M. fascicularis) based on morphological characters. American Journal of Physical Anthropology, 159(2), 189–198.
Hanihara, T., & Ishida, H. (2001a). Frequency variations of discrete cranial traits in major human populations. I. Supernumerary ossicle variations. Journal of Anatomy, 198(6), 689–706.
Hanihara, T., & Ishida, H. (2001b). Frequency variations of discrete cranial traits in major human populations. III. Hyperostotic variations. Journal of Anatomy, 199(3), 251–272.
Hanihara, T., & Ishida, H. (2001c). Frequency variations of discrete cranial traits in major human populations. IV. Vessel and nerve related variations. Journal of Anatomy, 199(3), 273–287.
Hauser, G., & DeStefano, G. F. (1989). Epigenetic variants of the human skull. Stuttgart: Schweizerbart.
Howells, W. W. (1973). Cranial variation in man: A study by multivariate anaysis of patterns of difference among recent human populations. Papers of the Peabody Museum of Archaeology and Ethnology, Vol. 67 (pp. 1–259). Cambridge: Harvard University Press.
Ito, T., Kawamoto, Y., Hamada, Y., & Nishimura, T. D. (2015). Maxillary sinus variation in hybrid macaques: Implications for the genetic basis of craniofacial pneumatization. Biological Journal of the Linnean Society, 115(2), 333–347.
Jiang, J., Yu, J., Li, J., Li, P., Fan, Z., Niu, L., Deng, J., Yue, B., Li, J., & Stanyon, R. (2016). Mitochondrial genome and nuclear markers provide new insight into the evolutionary history of macaques. PLoS One, 11(5), e0154665.
Jolly, C. J., & Disotell, T. R. (1997). Intergeneric hybrid baboons. International Journal of Primatology, 18(4), 597–627.
Kaur, J., Choudhry, R., Raheja, S., & Dhissa, N. C. (2012). Non-metric traits of the skull and their role in anthropological studies. Journal of Morphological Sciences, 29(4), 189–194.
Kawamoto, Y. (2005). NRAMP1 polymorphism in a hybrid population between Japanese and Taiwanese macaques in Wakayama, Japan. Primates, 46(3), 203–206.
Kawamoto, Y., Shirai, H., Araki, S., & Kyoko, M. (1999). A case of mixed typing of Japanese monkey and Taiwan monkey in Wakayama Prefecture (translated). Primate Research, 15, 53–60.
Kawamoto, Y., Ohsawa, H., Nigi, H., Maruhashi, T., Maekawa, S., et al (2001). Genetic assessment of a hybrid population between Japanese and Taiwan macaques in Wakayama Prefecture. Primate Research, 17(1), 13–24.
Kelaita, M. A., & Cortés-Ortiz, L. (2013). Morphological variation of genetically confirmed Alouatta pigra × A. palliata hybrids from a natural hybrid zone in Tabasco, Mexico. American Journal of Physical Anthropology, 150(2), 223–234.
Keller, C., Roos, C., Groeneveld, L. F., Fischer, J., & Zinner, D. (2010). Introgressive hybridization in southern African baboons shapes patterns of mtDNA variation. American Journal of Physical Anthropology, 142(1), 125–136.
Klingenberg, C. P. (1998). Heterochrony and allometry: The analysis of evolutionary change in ontogeny. Biological Reviews, 73, 79–123.
Klingenberg, C. P. (2011). MorphoJ: An integrated software package for geometric morphometrics. Molecular Ecology Resources, 11(2), 353–357.
Klingenberg, C. P. (2015). Analyzing fluctuating asymmetry with geometric morphometrics: Concepts, methods, and applications. Symmetry, 7(2), 843–934.
Klingenberg, C. P., & McIntyre, G. S. (1998). Geometric morphometrics of developmental instability: Analyzing patterns of fluctuating asymmetry with procrustes methods. Evolution, 52(5), 1363.
Klingenberg, C. P., & Monteiro, L. R. (2005). Distances and directions in multidimensional shape spaces: Implications for morphometric applications. Systematic Biology, 54(4), 678–688.
Korey, K. A. (1980). The incidence of bilateral nonmetric skeletal traits: A reanalysis of sampling procedures. American Journal of Physical Anthropology, 53(1), 19–23.
Mallet, J. (2005). Hybridization as an invasion of the genome. Trends in Ecology & Evolution, 20(5), 229–237.
Malaivijitnond, S., & Hamada, Y. (2008). Current situation and status of long-tailed macaques (Macaca fascicularis) in Thailand. The Natural History Journal of Chulalongkorn University, 8(2), 185–204.
Malaivijitnond, S., Sae-Low, W., & Hamada, Y. (2008). The human-ABO blood groups of free-ranging long-tailed macaques (Macaca fascicularis) and parapatric rhesus macaques (M. mulatta) in Thailand. Journal of Medical Primatology, 37(1), 31–37.
Manzi, G. (1996). Developmental stress and cranial hypostosis by epigenetic trait occurrence and distribution: An exploratory study on the Italian Neandertals. Journal of Human Evolution, 30(6), 511–527.
Manzi, G., Gracia, A., & Arsuaga, J. L. (2000). Cranial discrete traits in the Middle Pleistocene humans from Sima de los Huesos (Sierra de Atapuerca, Spain). Does hypostosis represent any increase in “ontogenetic stress” along the Neanderthal lineage? Journal of Human Evolution, 38, 425–446.
Markarjan, D. S., Isakov, E. P., & Kondakov, G. I. (1974). Intergeneric hybrids of the lower (42-chromosome) monkey species of the Sukhumi monkey colony. Journal of Human Evolution, 3(3), 247–250.
Mayr, E. (1963). Animal species and evolution. Cambridge: Belknap Press of Harvard University Press.
Mitteroecker, P., & Gunz, P. (2009). Advances in geometric morphometrics. Evolutionary Biology, 36, 235–247.
Mitteroecker, P., Gunz, P., & Bookstein, F. L. (2005). Heterochrony and geometric morphometrics: A comparison of cranial growth in Pan paniscus versus Pan troglodytes. Evolution & Development, 7(3), 244–258.
Moore, C. M., Janish, C., Eddy, C. A., Hubbard, G. B., Leland, M. M., & Rogers, J. (1999). Cytogenetic and fertility studies of a rheboon, rhesus macaque (Macaca mulatta) × baboon (Papio hamadryas) cross: Further support for a single karyotype nomenclature. American Journal of Physical Anthropology, 110(2), 119–127.
Osada, N., Uno, Y., Mineta, K., Kameoka, Y., Takahashi, I., & Terao, K. (2010). Ancient genome-wide admixture extends beyond the current hybrid zone between Macaca fascicularis and M. mulatta. Molecular Ecology, 19(14), 2884–2895.
Pastorini, J., Zaramody, A., Curtis, D. J., Nievergelt, C. M., & Mundy, N. I. (2009). Genetic analysis of hybridization and introgression between wild mongoose and brown lemurs. BMC Evolutionary Biology, 9, 32.
Perizonius, W. R. K. (1979). Non-metric cranial traits: Sex difference and age dependence. Journal of Human Evolution, 8(7), 679–684.
R Development Core Team (2015). R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing.
Rohlf, F. J. (1990). Morphometrics. Annual Review of Ecology and Systematics, 21, 299–316.
Rohlf, F. J. (1999). Shape statistics: procrustes superimpositions and tangent spaces. Journal of Classification, 16(2), 197–223.
Schillaci, M. A., Froehlich, J. W., Supriatna, J., & Jones-Engel, L. (2005). The effects of hybridization on growth allometry and craniofacial form in Sulawesi macaques. Journal of Human Evolution, 49(3), 335–369.
Schwenk, K., Brede, N., & Streit, B. (2008). Introduction. Extent, processes and evolutionary impact of interspecific hybridization in animals. Philosophical Transactions of the Royal Society B: Biological Sciences, 363(1505), 2805–2811.
Seehausen, O. (2004). Hybridization and adaptive radiation. Trends in Ecology & Evolution, 19(4), 198–207.
Shurtliff, Q. R. (2013). Mammalian hybrid zones: a review. Mammal Review, 43(1), 1–21.
Slice, D. E., Bookstein, F. L., Marcus, L. F., Rohlf, F. J. (1996). Appendix I: a glossary for geometric morphometrics. Advances in Morphometrics, (part 1), 531–552.
Sperber, G. H. (1989). Craniofacial Embryology. London: John Wright.
Tomkins, J. L., & Kotiaho, J. S. (2001). Fluctuating asymmetry. In eLS. Wiley Online Library.
Tosi, A. J., Morales, J. C., & Melnick, D. J. (2000). Comparison of Y chromosome and mtDNA phylogenies leads to unique inferences of macaque evolutionary history. Molecular Phylogenetics and Evolution, 17(2), 133–144.
Trinkaus, E. (1978). Bilateral asymmetry of human skeletal non-metric traits. American Journal of Physical Anthropology, 49(3), 315–318.
Tung, J., Charpentier, M. J. E., Garfield, D. A., Altmann, J., & Alberts, S. C. (2008). Genetic evidence reveals temporal change in hybridization patterns in a wild baboon population. Molecular Ecology, 17(8), 1998–2011.
Wang, Y., Naumann, U., Wright, S. T., & Warton, D. I. (2012). Mvabund: An R package for model-based analysis of multivariate abundance data. Methods in Ecology and Evolution, 3(3), 471–474.
Wang, Y., Naumann, U., Eddelbuettel, D. & Warton, D. (2018). mvabund: Statistical Methods for Analysing Multivariate Abundance Data. R package version 3.13.1. https://CRAN.R-project.org/package=mvabund.
White, T. D., & Folkens, P. (2000). Human osteology (2nd ed.). San Diego: Academic Press.
Wolpert, L., Jessel, T., Lawrence, P., Meyerowitz, E., Robertson, E., & Smith, J. (2007). Principles of development (3rd ed.). New York: Oxford University Press.
Wyner, Y. M., Johnson, S. E., Stumpf, R. M., & Desalle, R. (2002). Genetic assessment of a white-collared × red-fronted lemur hybrid zone at Andringitra, Madagascar. American Journal of Primatology, 57(2), 51–66.
Yamagiwa, J. (2010). Research history of Japanese macaques in Japan. In N. Nakagawa, M. Nakamichi, & H. Sigiura (Eds.), The Japanese macaques. New York: Springer Science+Business Media.
Zinner, D., Groeneveld, L. F., Keller, C., & Roos, C. (2009). Mitochondrial phylogeography of baboons (Papio spp.): Indication for introgressive hybridization? BMC Evolutionary Biology, 9, 83.
Zinner, D., Arnold, M. L., & Roos, C. (2011). The strange blood: Natural hybridization in primates. Evolutionary Anthropology, 20(3), 96–103.
Acknowledgments
This research was conducted as part of PhD research undertaken at the University of New South Wales, supported by an Australian Postgraduate Award and Research Excellence Award (UNSW School of Biological, Earth and Environmental Sciences). This research was chiefly funded by the Kyoto University Primate Research Institute Cooperative Research Program (awarded to C. Boel). We would like to thank Associate Professor Yoshi Kawamoto and Dr. Tsuyoshi Ito of the Kyoto University Primate Research Institute (KU-PRI) for providing access to genetic data and CT scans relevant to the hybrid sample, Professor Toshio Mouri for his generous guidance and assistance during data collection at the KU-PRI, and the Working Group of Hybrid Macaques who studied the demography and caught hybrids and Taiwanese macaques. We would also like to acknowledge the effort of the editor and reviewers, and thank them for the valuable contributions they have made to this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Joanna M. Setchell
Electronic supplementary material
ESM 1
(DOCX 219 kb)
Rights and permissions
About this article
Cite this article
Boel, C., Curnoe, D. & Hamada, Y. Craniofacial Shape and Nonmetric Trait Variation in Hybrids of the Japanese Macaque (Macaca fuscata) and the Taiwanese Macaque (Macaca cyclopis). Int J Primatol 40, 214–243 (2019). https://doi.org/10.1007/s10764-019-00081-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10764-019-00081-2