Abstract—
The inflammatory interleukin (IL)-23/IL-17 axis plays an important role in the pathogenesis of ankylosing spondylitis (AS), but with an unknown regulatory mechanism. This study aimed to investigate the role of endoplasmic reticulum (ER) stress and autophagy pathway in the expression of IL-23 in peripheral blood–derived macrophages in AS patients. Peripheral blood samples were obtained from 15 AS and 15 healthy control subjects. MACS was used to isolate monocytes from PBMCs. Then, M-CSF was used to differentiate monocytes to M2 macrophages. IFN-γ and/or LPS were used to activate macrophages and M2 polarization towards M1 macrophages. Thapsigargin was used to induce ER stress and 3-MA to inhibit autophagy. The purity of extracted monocytes and macrophage markers was evaluated by flow cytometry. mRNA expression of HLA-B and-B27, ER stress–related genes, autophagy-related genes, and IL-23p19 was performed using RT-qPCR. Soluble levels of IL-23p19 were measured using ELISA. Significant increase in mRNA expression of HLA-B, HLA-B27, BiP, XBP1, CHOP, and PERK mRNAs was observed in macrophages of AS patients before and after stimulation with IFN-γ and LPS. No significant change in autophagy gene expression was detected. mRNA and soluble levels of IL-23p19 demonstrated a significant increase in macrophages of AS patients compared to healthy subjects. ER stress induction led to a significant increase in IL-23p19 in macrophages. Inhibition of autophagy did not affect IL-23 expression. ER stress, unlike autophagy, is associated with increased IL-23 levels in macrophages of AS patients.
Key Messages
-
ER stress in macrophages from AS patients plays a role in the increased production of IL-23.
-
The autophagy pathway is not involved in the modulation of IL-23 production by AS macrophages.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
BACKGROUND
Ankylosing spondylitis (AS), the most common type of spondyloarthritis (SpA), is a chronic inflammatory arthritis that predominantly involves the axial skeleton. Despite the significant contribution of the human leukocyte antigen (HLA)-B27 in conferring genetic risk of AS, several other genes have also been identified in association with the disease [1,2,3]. Alteration in the sequence of HLA-B27 is involved in the modulation of peptide-binding ability and stability of the antigen presentation [4, 5]. That notwithstanding, no certain “arthritogenic peptide” has been recognized in induced abnormal immune response in AS, and no differences have been found among peptides that are loaded onto AS-associated and non-associated HLA-B27 subtypes [6].
In the synovial tissues and gut of SpA patients as well as HLA-B27-transgenic rats, an accumulation of unconventional HLA-B27 forms, like the free heavy chain (FHC), has been observed [7]. The uncommon characteristic of HLA-B27 molecules is the slow rate of folding and misfolding during biosynthesis in the ER. Such unconventional HLA-B27 forms can be accumulated in the endoplasmic reticulum (ER), resulting in ER stress, which stimulates the unfolded protein response (UPR). On the other side, UPR has been associated with the promoted production of inflammatory IL-23 cytokine in AS patients [8].
Unconventional HLA-B27 molecules, like FHC dimers expressed on the surface of antigen-presenting cells (APCs), are able to induce IL-23 receptor (IL-23R) expressing T cells to generate IL-17 [9]. Killer cell immunoglobulin-like receptor (KIR) 3DL2 expressed on these T cells is bound to FHC of unconventional HLA-B27 molecules, leading to the upregulation of T helper (Th) 17 specific transcription factor, namely RAR-related orphan receptor-gamma t (RORγt). This, in turn, results in Th17 polarization and increased production of pro-inflammatory IL-17 in AS [10,11,12].
Autophagy is considered a protective cellular process that is involved in clearing the dying cellular organelles as well as misfolded proteins to enhance cell survival [13]. Autophagy has been suggested to be related to the pathogenesis of AS through digesting misfolded and unconventional HLA-B27 molecules and, hence, modulation of UPR. Modifications in the autophagy pathways have also been reported to be tissue-specific since dysregulated autophagy was not observed in the peripheral blood mononuclear cells (PBMCs) or synovium of AS patients [14].
Our previous investigations demonstrated that UPR is activated in the macrophage-colony stimulating factor (M-CSF)–derived macrophages of AS patients, along with the upregulation of HLA-B27 expression in these cells, which was linked to the overproduction of IL-23 in macrophages from AS [15]. Seeking for a broader insight, in the current study, the behavior of macrophages derived from peripheral blood monocytes of AS patients and healthy controls in establishing ER stress, induced either by treatment with IFN-γ (which triggers HLA-B27) or by a chemical ER stress inducer and, ultimately, production of IL-23 were assessed. In addition, the behavior of AS macrophages in response to the inhibition of autophagy was studied.
METHODS AND STUDY SUBJECTS
AS Patients and Controls
In this case–control study, 15 AS patients and 15 healthy controls were included. AS patients were recruited from the rheumatology outpatient clinic of Rheumatology Research Center, Shariati Hospital affiliated with Tehran University of Medical Sciences, Tehran, Iran. Patients fulfilled the classification index of the modified New York criteria for the diagnosis of AS. Patients were evaluated for functional capacity through examining the Bath Ankylosing Spondylitis Functional Index (BASFI), for spinal mobility by Bath Ankylosing Spondylitis Metrology Index (BASMI), and for disease activity via Bath Ankylosing Spondylitis Disease Activity Index (BASDAI). As the inclusion criteria, AS patients having a BASDAI score further than 4 were chosen. Healthy controls were age- and sex-matched with the patient group, had no familial relation with AS cases, and had no family history of AS as well as other autoimmune disorders (Supplementary Table 1). From all study subjects, fresh whole blood samples were obtained by venipuncture and collected into a 9-ml vacuum blood collection tube containing EDTA anticoagulant.
Monocyte Isolation and M1 Macrophage Polarization
To isolate PBMCs, blood samples were first diluted with phosphate-buffered saline (PBS; GIBCO Invitrogen, Carlsbad, CA) and then were separated using Ficoll/Hypaque 1.077 g/ml (Lymphodex, inno-Train, Kronberg, Germany) and gradient density centrifugation. The cells were then washed twice at 300 and 200 × g to reduce impurity. In order to isolate monocytes, magnetic-activated cell sorter columns (Miltenyi Biotec, San Diego, CA) were used for the positive selection of CD14+ cells from PBMCs according to the manufacturer’s guidelines. The monocytes were assessed for purity using flow cytometry that was approximately 90–94% (Supplementary Fig. 1). To further purify the samples, monocytes were cultured in serum-free RPMI 1640 media (GIBCO Invitrogen, Carlsbad, CA) in 12-well media at a density of 55 × 105 cells/well and then were kept in the humidified atmosphere under 5% CO2 at 37 °C for 2h. Afterward, non-adherent cells were removed, and differentiation of the adherent monocyte cells to M2 macrophages was implemented using 50 ng/ml of recombinant human M-CSF (eBioscience, San Diego, CA) in R10 media (containing RPMI-1640, 10% FBS, 4 mM l-glutamine, 1% penicillin/streptomycin antibiotic) for 7 days in the humidified atmosphere under 5% CO2 at 37 °C. Thereafter, interferon (IFN)-γ cytokine alone or in combination with LPS was used to polarize the M2 to M1 macrophage cells [16]. On the seventh day, the supernatant was removed from each well of culture media, and RPMI-1640 and 50 ng/ml IFN-γ (R&D Systems) alone or with 100 ng/ml lipopolysaccharides (LPS; Sigma-Aldrich) obtained from Salmonella enterica serotype Enteritidis were added to each well.
To confirm the polarization of M2 to M1 macrophages, expressions of specific mRNAs of CD36 and mannose receptor C-type 1 (MRC1) genes for M1 macrophages and CC-chemokine receptor 7 (CCR7) and indoleamine 2,3-dioxygenase 1 (IDO1) genes for M2 macrophages were evaluated (Supplementary Fig. 2) [16,17,18].
Flow Cytometry
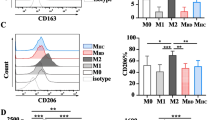
The flow cytometry was used for the phenotypic comparison of macrophages and monocytes. First, monocytes and macrophages collected from culture plates were washed twice with PBS. Cells were then incubated for 1h in PBS containing 5% FBS (GIBCO Invitrogen, Carlsbad, CA) at 4 °C to interrupt the non-specific monoclonal antibody binding. Afterward, cells were stained with APC-conjugated anti-human CD163, PE-Cy5-conjugated anti-human HLA-DR (Biolegend, San Diego, CA), and FITC-conjugated anti-human CD206 (Miltenyi Biotec, San Diego, CA) in the dark for 30 min at 4 °C. In parallel, irrelevant isotype-matched control labeling was also carried out. After that, the cells were rinsed and resuspended in 500 μl PBS, then chilled on ice, and evaluation of fluorescence was conducted using FACSCalibur (Becton Dickinson, Franklin Lakes, NJ) and analyzed using FlowJo software (Tree Star, Ashland, OR). At least 10,000 events were counted for each sample. Trypan blue staining was used to determine cell death.
Induction of ER Stress in M1 Macrophages
Thapsigargin (TPG) was used to induce ER stress in macrophages derived from blood monocytes of AS patients and then evaluate the effect of ER stress inhibition on IL-23 production. To this end, macrophages on the seventh day of culture were treated with 500 nM TPG (TOCRIS) along with or without IFN-γ and LPS for 12h.
Inhibition of Autophagy Pathway in M1 Macrophages
To investigate the inhibition of autophagy pathway in macrophages derived from blood monocytes of AS patients and determine the inhibition effect on IL-23 production, 3-methyladenine (3MA) was used. For this purpose, macrophages on the seventh day of culture were treated with 5 mM 3MA (TOCRIS) along with or without IFN-γ and LPS for 24h.
RNA Extraction, cDNA Synthesis, and Quantitative Real-time PCR
In order to extract the total RNA from M1 macrophages on day 7, the High Pure RNA Isolation Kit (Roche, Basel, Switzerland) was used, according to the company’s instructions. Synthesis of complementary DNA (cDNA) was conducted exerting the Transcriptor First Strand cDNA Synthesis Kit (Roche, Basel, Switzerland) using random hexamer primers, based on the manufacturer’s protocol. The quantitative mRNA expression levels of differentiation markers of macrophages, including CD36, MRC1, CCR7, and IDO1; HLA genes; ER stress–associated genes; autophagy-related genes; and IL-23p19 were determined to employ the real-time PCR performed by the StepOnePlus™ Real-Time PCR system (Applied Biosystems, Foster City, CA), SYBR Green master mix (Ampliqon, Odense, Denmark), and specific primers (Supplementary Table 2). The primers were designed through the Primer Express 3.0 tool (Applied Biosystems, Foster City, CA, USA) and checked for specificity by the Primer-BLAST Tool of the National Center for Biotechnology Information (NCBI) website (https://www.ncbi.nlm.nih.gov/tools/primer-blast/). Finally, the primers were generated by the custom oligonucleotide synthesis service (Metabion, Martinsried, Germany). Moreover, the specification of the HLA-B27-specific primers was obtained from the study by Zeng et al. [16]. The comparative CT approach, defined by Livak and Schmittgen [19], was used to determine the relative transcription levels. Normalization of target gene mRNAs was done corresponding to the expression level of 18S ribosomal RNA as the housekeeping gene to calculate the relative amounts of each gene transcription using the “Relative Expression = 2−ΔCt” equation.
ELISA
To measure the IL-23p19 cytokine concentration, the supernatant of each well was collected after different treatments of macrophages and then stored in − 70 °C for conducting the enzyme-linked immunosorbent assay (ELISA). The optical density in the samples was determined using the Human IL-23 ELISA development kit (HRP, Mabtech, Stockholm, Sweden).
Statistical Analysis
Data analysis and illustrations by graphs were both conducted using GraphPad Prism v.8 (GraphPad Software, San Diego, CA). Friedman test was used to compare three different macrophages (M2 macrophages, IFN-γ-induced M1 macrophages, and IFN-γ + LPS-induced M1 macrophages) from patients or controls. Additionally, before-after treatment analyses of comparing means were performed by Wilcoxon signed-rank test. The nonparametric spearman’s rho test was performed to assess the correlation between mRNA expression of different genes with mRNA and soluble levels of IL-23p19. Data were expressed as means ± standard deviation (SD), and a P value less than 0.05 was considered statistically significant level.
RESULTS
Expression of HLA-B and HLA-B27 Molecules
There was a statistically significant difference in mRNA expression of HLA-B among IFN-γ-induced M1 macrophages; IFN-γ + LPS-induced M1 macrophages; and M2 macrophages in AS-B27+, AS-B27−, Con-B27+, and Con-B27− (Fig. 1a). There was a statistically significant difference in mRNA expression of HLA-B27 among IFN-γ-induced M1 macrophages, IFN-γ + LPS-induced M1 macrophages, and M2 macrophages in both AS patients and the control group (Fig. 1b). Refer to Supplementary Table 3 for fold change values and P values with more details.
mRNA expression levels of HLA-B (a) and HLA-B27 (b) in AS patients and healthy subjects. Monocytes were isolated from 10 HLA-B27-positive AS patients (AS-B27+), 5 HLA-B27-negative AS patients (AS-B27−), 10 HLA-B27-positive healthy controls (Con-B27+), and 5 HLA-B27-negative healthy controls (Con-B27−) and were differentiated into M2 macrophages upon treating with M-CSF in 7 days. M2 macrophages were then polarized into M1 macrophages after treating with IFN-γ alone or in combination with LPS for 24h. The expression levels of HLA-B mRNA (a) and HLA-B27 mRNA (b) in these three cell groups were compared between patients and controls. Results are presented as mean ± standard deviation (SD). Significance within each cell group was indicated with “*” sign, between M1 (IFN-γ) and M1 (IFN-γ + LPS) group compared to M2 with “#” sign, and between M1 (IFN-γ) and M1 (IFN-γ + LPS) group by “$” sign.
mRNA Expression of ER Stress–Associated Genes
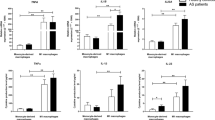
A statistically significant difference was observed in mRNA expression of BiP (Fig. 2a), XBP1 (Fig. 2b), and PERK (Fig. 2c) between different groups of AS patients and controls as well as between IFN-γ-induced M1 macrophages, IFN-γ + LPS-induced M1 macrophages, and M2 macrophages. In the IFN-γ + LPS-induced M1 macrophages, upregulated levels of BiP mRNA in the AS-B27+ patients compared with the Con-B27− and Con-B27+ groups were observed. Besides, there was a similar pattern in BiP mRNA expression in IFN-γ-induced M1 macrophages and M2 macrophages. The highest expression of XBP1 mRNA was seen in IFN-γ + LPS-induced M1 macrophages from B27 + individuals, which was significantly higher than both IFN-γ-induced M1 macrophages and M2 macrophages. PERK mRNA expression was also higher in IFN-γ + LPS-induced M1 macrophages compared to IFN-γ-induced M1 macrophages in AS-B27+ patients. However, the mRNA expression of ATF6 (Fig. 2d) and CHOP (Fig. 2e) did not show a significant difference between different cells. Refer to Supplementary Table 3 for fold change values and P values with more details.
mRNA expression levels of ER stress–related genes, including BiP (a), XBP1 (b), PERK (c), ATF6 (d), and CHOP (e) in AS patients and healthy subjects. Monocytes were isolated from 10 HLA-B27-positive AS patients (AS-B27+), 5 HLA-B27-negative AS patients (AS-B27−), 10 HLA-B27-positive healthy controls (Con-B27+), and 5 HLA-B27-negative healthy controls (Con-B27−) and were differentiated into M2 macrophages upon treating with M-CSF in 7 days. M2 macrophages were then polarized into M1 macrophages after treating with IFN-γ alone or in combination with LPS for 24h. The mRNA expression levels of the mentioned genes in these three cell groups were compared between patients and controls. Results are presented as mean ± standard deviation (SD). Significance within each cell group was indicated with “*” sign, between M1 (IFN-γ) and M1 (IFN-γ + LPS) group compared to M2 with “#” sign, and between M1 (IFN-γ) and M1 (IFN-γ + LPS) group by “$” sign.
mRNA Expression of Autophagy-Related Genes
In the IFN-γ + LPS-induced M1 macrophages, there were upregulated levels of ATG5 mRNA in the AS-B27+ patients compared with the AS-B27− patients and Con-B27+ group (Fig. 3a). Significantly altered expression of ATG12 mRNA was detected in the M2 macrophages between patients and controls. AS-B27+ patients expressed increased levels of ATG12 mRNA in the M2 macrophages in comparison to the Con-B27− group. Moreover, mRNA expression of ATG12 was significantly upregulated in the Con-B27+ in relation to the Con-B27− group (Fig. 3b). Analysis indicated a statistically significant difference in mRNA expression of ATG16L among IFN-γ-induced M1 macrophages; IFN-γ + LPS-induced M1 macrophages; and M2 macrophages in AS-B27+, Con-B27+, and Con-B27− groups (Fig. 3c). Refer to Supplementary Table 3 for fold change values and P values with more details.
mRNA expression levels of autophagy-related genes, including Atg5 (a), Atg12 (b), and Atg16L (c) in AS patients and healthy subjects. Monocytes were isolated from 10 HLA-B27-positive AS patients (AS-B27+), 5 HLA-B27-negative AS patients (AS-B27−), 10 HLA-B27 positive healthy controls (Con-B27+), and 5 HLA-B27-negative healthy controls (Con-B27−) and were differentiated into M2 macrophages upon treating with M-CSF in 7 days. M2 macrophages were then polarized into M1 macrophages after treating with IFN-γ alone or in combination with LPS for 24h. The mRNA expression levels of the mentioned genes in these three cell groups were compared between patients and controls. Results are presented as mean ± standard deviation (SD). Significance within each cell group was indicated with “*” sign, between M1 (IFN-γ) and M1 (IFN-γ + LPS) group compared to M2 with “#” sign, and between M1 (IFN-γ) and M1 (IFN-γ + LPS) group by “$” sign.
mRNA Expression of IL-23
There was a statistically significant difference in the mRNA expression of IL-23p19 among IFN-γ-induced M1 macrophages; IFN-γ + LPS-induced M1 macrophages; and M2 macrophages in AS-B27+, AS-B27−, Con-B27+, and Con-B27−. In comparison to M2 macrophages, the mRNA expression of IL-23p19 was overexpressed in the IFN-γ + LPS-induced M1 macrophages in AS-B27+, AS-B27−, Con-B27+, and Con-B27−. In both M2 macrophages and IFN-γ-induced M1 macrophages, the highest expression of mRNA of IL-23p19 was observed in the AS-B27+ patients, which was significantly higher than those in the AS-B27− patients and the control groups. Although this pattern is not observed in IFN-γ + LPS-induced M1 macrophages, the mRNA expression of IL-23p19 in all AS-B27+, AS-B27−, Con-B27+, and Con-B27− individuals was significantly higher than those in M2 macrophages and IFN-γ-induced M1 macrophages (Fig. 4a). Refer to Supplementary Table 3 for fold change values and P values with more details.
Bar graphs indicate the mRNA expression and soluble protein levels of IL-23p19 in the patients and healthy controls before and after TPG treatment (to induce ER stress). Monocytes were isolated from 10 HLA-B27-positive AS patients (AS-B27+), 5 HLA-B27-negative AS patients (AS-B27−), 10 HLA-B27-positive healthy controls (Con-B27+), and 5 HLA-B27-negative healthy controls (Con-B27−) and were differentiated into M2 macrophages upon treating with M-CSF in 7 days. M2 macrophages were then polarized into M1 macrophages after treating with IFN-γ alone or in combination with LPS for 24h. To induce ER stress, TPG at a concentration of 500 nM was added into M1 macrophages and incubated for 12h. The mRNA expression and serum levels of IL-23p19 were compared between patients and controls. Soluble levels of IL-23p19 cytokine in the supernatant of the IFN-γ + LPS-induced M1 macrophages were measured by ELISA and then were compared between patients and controls. a Baseline mRNA expression of IL-23p19 in the IFN-γ + LPS-induced M1 macrophages, IFN-γ-induced M1 macrophages, and M2 macrophages. b mRNA expression of IL-23p19 in the IFN-γ-induced M1 macrophages before and after TPG treatment. c mRNA expression of IL-23p19 in the IFN-γ + LPS-induced M1 macrophages before and after TPG treatment. d Soluble levels of IL-23p19 in the supernatant of the IFN-γ + LPS-induced M1 macrophages before and after TPG treatment. Results are presented as mean ± standard deviation (SD). Significance within each cell group is marked with the “*” sign, and between different cell groups with “#” sign.
Effect of ER Stress Induction on mRNA Expression of IL-23
After induction of ER stress with TPG in the IFN-γ-induced M1 macrophages, the mRNA expression of IL-23p19 was upregulated in AS-B27+, AS-B27−, Con-B27+, and Con-B27− in comparison to non-ER stress–induced IFN-γ-induced M1 macrophages (Fig. 4b). Similar results were observed in the IFN-γ + LPS-induced M1 macrophages.
Furthermore, there was an overexpression of the mRNA of IL-23p19 in the ER stress–induced IFN-γ + LPS-induced M1 macrophages from AS-B27+ patients in comparison to the Con-B27+ group, as well as in the AS-B27− patients compared with the Con-B27+ group and Con-B27− group (Fig. 4c). Refer to Supplementary Table 4 for fold change values and P values with more details.
Effect of ER Stress Induction on Soluble Level of IL-23
The measurement range of the kit used (Mabtech, Stockholm, Sweden) in the detection of soluble levels of IL-23p19 was between 7 and 700 pg/ml. Levels of IL-23p19 were not detectable in the M2 macrophages and IFN-γ-induced M1 macrophages due to very low levels. However, the cytokine levels were measured in the IFN-γ + LPS-induced M1 macrophages before and after induction of ER stress.
Analysis indicated that in comparison to the non-ER stress–induced IFN-γ + LPS-induced M1 macrophages, the soluble levels of IL-23p19 were higher in the supernatant of ER stress–induced IFN-γ + LPS-induced M1 macrophages in AS-B27+, AS-B27− Con-B27+, and Con-B27−. In the supernatant of ER stress–induced IFN-γ + LPS-induced M1 macrophages, soluble levels of IL-23p19 were higher in the AS-B27+ patients in comparison to the Con-B27+ group, as well as in the AS-B27− patients compared to the Con-B27+ and Con-B27− groups (Fig. 4d).
Effect of Autophagy Pathway Inhibition on mRNA Expression and Soluble Levels of IL-23
To inhibit the autophagy pathway in macrophages derived from blood monocytes of AS patients and determine its consequence on IL-23 production, cells were treated with 3MA along with or without IFN-γ and LPS. The mRNA expression of IL-23p19 did not change significantly in either IFN-γ (Fig. 5a) or IFN-γ + LPS-induced M1 macrophages (Fig. 5b) in any of the study groups after treatment. Besides, no altered levels of soluble IL-23p19 were detected between patients and controls in the IFN-γ + LPS-induced M1 macrophages (Fig. 5c).
Bar graphs indicate the mRNA expression and soluble protein levels of IL-23p19 in the patients and healthy controls before and after 3MA treatment (to inhibit autophagy pathway). Monocytes were isolated from 10 HLA-B27-positive AS patients (AS-B27+), 5 HLA-B27-negative AS patients (AS-B27−), 10 HLA-B27-positive healthy controls (Con-B27+), and 5 HLA-B27-negative healthy controls (Con-B27−) and were differentiated into M2 macrophages upon treating with M-CSF in 7 days. M2 macrophages were then polarized into M1 macrophages after treating with IFN-γ alone or in combination with LPS for 24h. For inhibiting the autophagy pathway, 3MA at a concentration of 5 mM was added into M1 macrophages and incubated for 24h. The expression levels of IL-23p19 mRNA in these cell groups were compared between patients and controls. Soluble levels of IL-23p19 cytokine in the supernatant of the IFN-γ + LPS-induced M1 macrophages were measured by ELISA and then were compared between patients and controls. a mRNA expression of IL-23p19 in the IFN-γ-induced M1 macrophages. b mRNA expression of IL-23p19 in the IFN-γ + LPS-induced M1 macrophages. c Soluble levels of IL-23p19 in the supernatant of the IFN-γ + LPS-induced M1 macrophages. Results are presented as mean ± standard deviation (SD).
Correlation Analysis
In order to evaluate the relation between the ER stress–related genes and autophagy genes with IL-23p19, correlation analyses were conducted. In IFN-γ + LPS-induced M1 macrophages from AS-B27+ patients, the mRNA expression of HLA-B27 and PERK were negatively associated with IL-23p19 expression. Moreover, the mRNA expression of BiP and ATF6 in IFN-γ + LPS-induced M1 macrophages obtained from AS-B27+ patients was significantly associated with soluble levels of IL-23p19. In IFN-γ-induced M1 macrophages from AS-B27+ patients, only the mRNA expression of CHOP was significantly correlated with IL-23p19 expression. In IFN-γ or IFN-γ + LPS-induced M1 macrophages from AS-B27− patients, no significant relationship was found between ER stress/autophagy genes and IL-23p19 mRNA expression (Supplementary Tables 5 and 6).
DISCUSSION
In the current study, we tried to evaluate the role of UPR in the modulation of AS macrophages to establish an inflammatory setting by the production of IL-23. The behavior of macrophages derived from peripheral blood monocytes isolated from AS patients and healthy controls in stimulating the ER stress induced either by treatment with IFN-γ (HLA-B27 enhancer stimulant) or by ER stress inducer (TPG) and IL-23 production was assessed. Moreover, the macrophage response against the inhibition of autophagy (by 3MA agent) was evaluated.
A bulk of investigations have shown increased IL-23 in tissue, serum, and peripheral blood cells in AS patients [20, 21]. However, the specific pathway that increases IL-23 in this disease has not yet been clearly identified. Among the different hypotheses presented for AS pathogenesis, the theory of misfolding and accumulation of HLA-B27 in the ER and induction of ER stress has been suggested. According to this hypothesis, stress induced in the ER by misfolding of the HLA-B27 molecule can induce the UPR cellular response, leading to increased inflammatory cytokines like IL-23 [22, 23].
Here in this study, IFN-γ was used to differentiate M2 to M1 macrophages and increase HLA-B27 expression in these cells. It was observed that M2 macrophages, IFN-γ-induced M1 macrophages, and IFN-γ + LPS-induced M1 macrophages obtained from AS-B27+ patients had upregulated mRNA expression of HLA-B in comparison to the AS-B27− patients and control group. All these macrophage cells also overexpressed mRNA of HLA-B in AS-B27+ patients compared with Con-B27+ individuals. Previous studies using immunofluorescence, flow cytometry [24], and quantitative PCR [25] showed increased HLA-B27 expression in the PBMCs from HLA-B27+ AS patients compared to healthy HLA-B27+ subjects.
Considering increased expression of the HLA-B27 in the IFN-γ-induced M1 macrophages and IFN-γ + LPS-induced M1 macrophages, the behavior of macrophages with respect to ER stress and mRNA expression of related genes was investigated. The results showed that M2 macrophages, as well as IFN-γ-induced M1 macrophages and IFN-γ + LPS-induced M1 macrophages of AS-B27+ patients compared to AS-B27− patients and control group, overexpressed ER stress–associated genes of BiP, XBP1, PERK, and CHOP, implying to ER stress in these cells. It should be noted that measuring the spliced XBP1 along with total XBP1 could be a better marker for XBP1 activation, since upon UPR activation, XBP1 gets spliced [26]. Besides, the differences in the mRNA expression of BiP and CHOP were not different between the three macrophages. PERK mRNA expression in the IFN-γ + LPS-induced M1 macrophages was higher than IFN-γ-induced M1 macrophages in AS-B27+ patients. Only mRNA expression of XBP1 showed a significant increase under IFN-γ treatment alone or with LPS. Feng et al. also indicated increased expression of UPR genes, such as BiP, CHOP, and XBP1, in the peripheral blood as well as the synovial fluid of SpA patients [27].
Similarly, using microarray and gene expression profile analysis, Gu et al. reported the increase of BiP in the synovial fluid mononuclear cells (SFMC) in SpA patients [28]. However, lamina propria mononuclear cells (LPMC) isolated from the gut of AS patients did not show significant changes in the expression of BiP, XBP1, PERK, and ATF6 genes [29]. Zeng et al. also could not show a significant increase in UPR-associated genes in macrophages of AS patients after treatment with IFN-γ and LPS [16].
Our experiments indicated that macrophages from AS-B27+ patients produced higher levels of IL-23p19 in comparison to AS-B27− patients and healthy individuals. It should be noted that after treatment with IFN-γ, there was no significant difference in the level of IL-23p19 expression between M2 macrophages and IFN-γ-induced M1 macrophages. The highest mRNA expression of IL-23p19 was observed in IFN-γ + LPS-induced M1 macrophages obtained from AS-B27+ patients, indicating that HLA-B27 levels in combination with LPS might further increase IL23p19 expression. Therefore, it should be noted that although there is no difference in the expression of HLA-B27 between IFN-γ and IFN-γ + LPS-induced M1 macrophages, the effect of HLA-B27 is likely to be the “straw that broke the camel’s back” in the background of LPS association with higher mRNA expression of IL-23p19. This is consistent with a study by DeLay et al., which showed that IFN-γ alone did not affect IL-23p19 expression [30].
Experiments by induction of ER stress using TPG implied the different behavior of macrophages from patients compared to the control group. TPG treatment of macrophages significantly increased both mRNA expression and protein levels of IL-23p19 in both patients and controls. This increase was significant for macrophages that were not treated with TPG. Additionally, TPG-treated macrophages of patients showed higher IL-23p19 cytokine levels than healthy controls. The results of the present study were in line with increased IL-23 levels in macrophages of AS patients and demonstrated the previously reported involvement of the IL-23/IL17 inflammatory pathway in the pathogenesis of AS. Many genome-wide association studies (GWASs) have reported the association of IL23 receptor gene polymorphisms with AS susceptibility [31, 32]. In a study by Goodall et al., it was shown that the ER stress–related gene CHOP directly regulated IL-23. In fact, CHOP is an ER stress–induced transcription factor and possesses a specific binding site to the IL23 gene regulatory domain [33]. Animal studies have also shown that TPG-induced ER stress can increase IL-23 production in rat bone marrow–derived macrophages [23, 34]. This increase by ER stress has also been detected in human dendritic cells (DCs) [33].
Experiments on the autophagy genes indicated no significant difference in expression of ATG5 and ATG12 in the three macrophage groups between patients and healthy individuals. Only IFN-γ + LPS-induced M1 macrophages from AS-B27+ patients expressed higher levels of ATG16L than M2 macrophages and IFN-γ-induced M1 macrophages. In addition, mRNA expression of ATG12 in M2 macrophages from patients was upregulated compared to controls. In a 2013 study by Ciccia and colleagues on LPMCs isolated from the gut of AS patients, it was found that the autophagy pathway in these cells was more active than that in the control group. They showed that the expression levels of ATG5, ATG12, and LC3II were higher in the gut of AS patients than in healthy individuals and that the autophagy pathway was involved in regulating IL-23 production in the gut of these patients [29]. Data from autophagy inhibition in M1 macrophages by 3MA and IL-23p19 cytokine analysis showed that inhibition of the autophagy pathway could not have a significant effect on IL-23p19 expression and production. Contrary to the results of the present study, a study by Castro et al. on the autophagy pathway showed that both mouse bone marrow–derived macrophages and human macrophages and DCs were more susceptible to autophagy inhibition either by chemical inhibitors or by interfering RNA. Autophagy pathway inhibition increased the production of IL-23 in the cells mentioned [35]. The possible reason for not statistically significant differences in the expression of IL-23p19 in this study after 3MA treatment could be the mechanism of action of this drug in inhibiting the autophagy pathway. A recent study has shown that 3MA can induce massive DNA damage and an abrupt decrease in cell viability compared to other autophagy inhibitors such as spautin-1, Cpd18, bafilomycin A1, and chloroquine [36]. In our observation, the morphology of treated cells was changed during the treatment, and the viability is also affected by 3MA especially in IFN-γ + LPS-induced M1 macrophages (Supplementary Figs. 3 and 4).
All of the mentioned information suggests that targeting the IL-23 axis may be useful in treating AS patients. However, a recent randomized, placebo-controlled phase II clinical trial using an anti-p19 antibody, risankizumab, for AS treatment failed to show any evidence for clinically significant improvement of the primary and secondary endpoints [37, 38]. This may be because the IL-23 axis is not a linear cascade. Due to the autoimmune nature of the disease, there are several other factors that can greatly affect the pathogenesis of AS.
CONCLUSION
Overall, this study revealed that AS patients express higher levels of HLA-B27 than healthy controls. Macrophages in these patients can also produce higher levels of IL-23, especially in the presence of LPS and IFN-γ. The increased expression of several genes involved in the UPR response indicated an ER stress state in macrophages of AS patients. Evaluation of the ER stress pathway revealed that AS macrophages behave differently towards ER stress compared to healthy subjects. Hence, ER stress can be proposed as a regulatory pathway for IL-23 production in macrophages of AS patients. Inhibition of autophagy pathway did not alter IL-23 production and probably could not be considered the regulatory pathway of this cytokine in macrophages of AS patients. Considering the results of the risankizumab phase II clinical trial, it cannot be concluded that targeting IL-23 alone can improve the patients’ outcome, but IL-23 inhibitors will certainly be one of the important arms of future combination therapies for the treatment of AS patients, which will require more molecular and preclinical studies.
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- 3MA:
-
3-Methyladenine
- APC:
-
Antigen-presenting cell
- AS:
-
Ankylosing spondylitis
- BASDAI:
-
Bath Ankylosing Spondylitis Disease Activity Index
- BASFI:
-
Bath Ankylosing Spondylitis Functional Index
- BASMI:
-
Bath Ankylosing Spondylitis Metrology Index
- CCR7:
-
CC-chemokine receptor 7
- DC:
-
Dendritic cells
- ER:
-
Endoplasmic reticulum
- FHC:
-
Free heavy chain
- GWAS:
-
Genome-wide association study
- HLA-B27:
-
Human leukocyte antigen B27
- IDO1:
-
Indoleamine 2,3-dioxygenase 1
- IFN-γ:
-
Interferon gamma
- IL-23:
-
Interleukin-23
- IL-23R:
-
IL-23 receptor
- KIR:
-
Killer cell immunoglobulin-like receptor
- LPMC:
-
Lamina propria mononuclear cells
- LPS:
-
Lipopolysaccharide
- M-CSF:
-
Macrophage-colony-stimulating factor
- MRC1:
-
Mannose receptor C-type 1
- PBMC:
-
Peripheral blood mononuclear cell
- PBS:
-
Phosphate-buffered saline
- RORγt:
-
RAR-related orphan receptor-gamma t
- SFMC:
-
Synovial fluid mononuclear cells
- SpA:
-
Spondyloarthritis
- Th17:
-
T helper 17
- TPG:
-
Thapsigargin
- UPR:
-
Unfolded protein response
References
Ranganathan, V., E. Gracey, M.A. Brown, R.D. Inman, and N. Haroon. 2017. Pathogenesis of ankylosing spondylitis—recent advances and future directions. Nature Reviews Rheumatology 13: 359.
Mahmoudi, M., S. Aslani, M.H. Nicknam, J. Karami, and A.R. Jamshidi. 2017. New insights toward the pathogenesis of ankylosing spondylitis; genetic variations and epigenetic modifications. Modern rheumatology 27: 198–209.
Mohammadi H, F., Babaie, M., Hemmatzadeh, G., Azizi, M., Hajaliloo, A. A., Ebrahimi, T., Kazemi, M., Yousefi, A., Rezaiemanesh, E., afarzadeh, E., Baghbani. 2018 Oct 20. Evaluation of ERAP1 gene single nucleotide polymorphism in impressing the inflammatory cytokine profile of ankylosing spondylitis patients. Iranian Journal of Allergy, Asthma and Immunology 464–474.
García-Medel, N., A. Sanz-Bravo, C. Alvarez-Navarro, P. Gómez-Molina, E. Barnea, M. Marcilla, A. Admon, and J.A.L. de Castro. 2014. Peptide handling by HLA-B27 subtypes influences their biological behavior association with ankylosing spondylitis and susceptibility to endoplasmic reticulum aminopeptidase 1 (ERAP1). Molecular & Cellular Proteomics 13: 3367–3380.
Dashti, N., M. Mahmoudi, S. Aslani, and A. Jamshidi. 2018. HLA-B* 27 subtypes and their implications in the pathogenesis of ankylosing spondylitis. Gene 670: 15–21.
Schittenhelm, R.B., T.C.L.K. Sian, P.G. Wilmann, N.L. Dudek, and A.W. Purcell. 2015. Revisiting the arthritogenic peptide theory: Quantitative not qualitative changes in the peptide repertoire of HLA–B27 allotypes. Arthritis & rheumatology 67: 702–713.
Rysnik, O., K. McHugh, L. van Duivenvoorde, M. van Tok, G. Guggino, J. Taurog, S. Kollnberger, F. Ciccia, D. Baeten, and P. Bowness. 2016. Non-conventional forms of HLA-B27 are expressed in spondyloarthritis joints and gut tissue. Journal of autoimmunity 70: 12–21.
Colbert, R.A., M.L. DeLay, E.I. Klenk, and G. Layh-Schmitt. 2010. From HLA-B27 to spondyloarthritis: A journey through the ER. Immunological reviews 233: 181–202.
Bowness, P., A. Ridley, J. Shaw, A.T. Chan, I. Wong-Baeza, M. Fleming, F. Cummings, A. McMichael, and S. Kollnberger. 2011. Th17 cells expressing KIR3DL2+ and responsive to HLA-B27 homodimers are increased in ankylosing spondylitis. The Journal of Immunology 186: 2672–2680.
Babaie, F., M. Hasankhani, H. Mohammadi, E. Safarzadeh, A. Rezaiemanesh, R. Salimi, B. Baradaran, and Z. Babaloo. 2018. The role of gut microbiota and IL-23/IL-17 pathway in ankylosing spondylitis immunopathogenesis: New insights and updates. Immunology letters 196: 52–62.
Rezaiemanesh, A., M. Abdolmaleki, K. Abdolmohammadi, H. Aghaei, F.D. Pakdel, Y. Fatahi, N. Soleimanifar, M. Zavvar, and M.H. Nicknam. 2018. Immune cells involved in the pathogenesis of ankylosing spondylitis. Biomedicine & Pharmacotherapy 100: 198–204.
Ridley, A., H. Hatano, I. Wong-Baeza, J. Shaw, K.K. Matthews, H. Al-Mossawi, K. Ladell, D.A. Price, P. Bowness, and S. Kollnberger. 2016. Activation-induced killer cell immunoglobulin-like receptor 3DL2 binding to HLA–B27 licenses pathogenic t cell differentiation in spondyloarthritis. Arthritis & rheumatology 68: 901–914.
Ciccia, F., and N. Haroon. 2016. Autophagy in the pathogenesis of ankylosing spondylitis. Clinical rheumatology 35: 1433–1436.
Neerinckx, B., S. Carter, and R. . 2014. Lories. IL-23 expression and activation of autophagy in synovium and PBMCs of HLA-B27 positive patients with ankylosing spondylitis. Arthritis & Rheumatology 66: S844–S844.
Rezaiemanesh, A., M. Mahmoudi, A.A. Amirzargar, M. Vojdanian, A.R. Jamshidi, and M.H. Nicknam. 2017. Ankylosing spondylitis M-CSF-derived macrophages are undergoing unfolded protein response (UPR) and express higher levels of interleukin-23. Modern rheumatology 27: 862–867.
Zeng, L., M.J. Lindstrom, and J.A. Smith. 2011. Ankylosing spondylitis macrophage production of higher levels of interleukin-23 in response to lipopolysaccharide without induction of a significant unfolded protein response. Arthritis & Rheumatism 63: 3807–3817.
Svensson, J., M.C. Jenmalm, A. Matussek, R. Geffers, G. Berg, and J. Ernerudh. 2011. Macrophages at the fetal–maternal interface express markers of alternative activation and are induced by M-CSF and IL-10. The Journal of Immunology 187: 3671–3682.
Brown, M.B., M. von Chamier, A.B. Allam, and L. Reyes. 2014. M1/M2 macrophage polarity in normal and complicated pregnancy. Frontiers in immunology 5: 606.
Schmittgen, T.D., and K.J. Livak. 2008. Analyzing real-time PCR data by the comparative CT method. Nature protocols 3: 1101–1108.
Ciccia, F., M. Bombardieri, A. Principato, A. Giardina, C. Tripodo, R. Porcasi, S. Peralta, V. Franco, E. Giardina, and A. Craxi. 2009. Overexpression of interleukin-23, but not interleukin-17, as an immunologic signature of subclinical intestinal inflammation in ankylosing spondylitis. Arthritis & Rheumatism: Official Journal of the American College of Rheumatology 60: 955–965.
Mei, Y., F. Pan, J. Gao, R. Ge, Z. Duan, Z. Zeng, F. Liao, G. Xia, S. Wang, and S. Xu. 2011. Increased serum IL-17 and IL-23 in the patient with ankylosing spondylitis. Clinical rheumatology 30: 269–273.
Appel, H., R. Maier, J. Bleil, A. Hempfing, C. Loddenkemper, U. Schlichting, U. Syrbe, and J. Sieper. 2013. In situ analysis of interleukin–23–and interleukin-12–positive cells in the spine of patients with ankylosing spondylitis. Arthritis & Rheumatism 65: 1522–1529.
DeLay, M.L., M.J. Turner, E.I. Klenk, J.A. Smith, D.P. Sowders, and R.A. Colbert. 2009. HLA–B27 misfolding and the unfolded protein response augment interleukin-23 production and are associated with Th17 activation in transgenic rats. Arthritis & Rheumatism 60: 2633–2643.
Cauli, A., G. Dessole, M. Fiorillo, A. Vacca, A. Mameli, P. Bitti, G. Passiu, R. Sorrentino, and A. Mathieu. 2002. Increased level of HLA-B27 expression in ankylosing spondylitis patients compared with healthy HLA-B27-positive subjects: A possible further susceptibility factor for the development of disease. Rheumatology 41: 1375–1379.
Liu, S.-Q., H.-C. Yu, Y.-Z. Gong, and N.-S. Lai. 2006. Quantitative measurement of HLA-B27 mRNA in patients with ankylosing spondylitis–correlation with clinical activity. The Journal of rheumatology 33: 1128–1132.
Colbert, R.A., T.M. Tran, and G. Layh-Schmitt. 2014. HLA-B27 misfolding and ankylosing spondylitis. Molecular Immunology 57: 44–51.
Feng, Y., J. Ding, C.M. Fan, and P. Zhu. 2012. Interferon-γ contributes to HLA-B27-associated unfolded protein response in spondyloarthropathies. The Journal of rheumatology 39: 574–582.
Gu, J., M. Rihl, E. Märker-Hermann, D. Baeten, J.G. Kuipers, Y.W. Song, W.P. Maksymowych, R. Burgos-Vargas, E.M. Veys, and F. De Keyser. 2002. Clues to pathogenesis of spondyloarthropathy derived from synovial fluid mononuclear cell gene expression profiles. The Journal of Rheumatology 29: 2159–2164.
Ciccia, F., A. Accardo-Palumbo, A. Rizzo, G. Guggino, S. Raimondo, A. Giardina, A. Cannizzaro, R.A. Colbert, R. Alessandro, and G. Triolo. 2014. Evidence that autophagy, but not the unfolded protein response, regulates the expression of IL-23 in the gut of patients with ankylosing spondylitis and subclinical gut inflammation. Annals of the rheumatic diseases 73: 1566–1574.
DeLay, M.L., M.J. Turner, E.I. Klenk, J.A. Smith, D.P. Sowders, and R.A. Colbert. 2009. HLA-B27 misfolding and the unfolded protein response augment interleukin-23 production and are associated with Th17 activation in transgenic rats. Arthritis & Rheumatism 60: 2633–2643.
Rahman, P., R.D. Inman, D.D. Gladman, J.P. Reeve, L. Peddle, and W.P. Maksymowych. 2008. Association of interleukin-23 receptor variants with ankylosing spondylitis. Arthritis & Rheumatism 58: 1020–1025.
Reveille, J.D., A.-M. Sims, P. Danoy, D.M. Evans, P. Leo, J.J. Pointon, R. Jin, X. Zhou, L.A. Bradbury, and L.H. Appleton. 2010. Genome-wide association study of ankylosing spondylitis identifies non-MHC susceptibility loci. Nature genetics 42: 123.
Goodall, J.C., C. Wu, Y. Zhang, L. McNeill, L. Ellis, V. Saudek, and J.H. Gaston. 2010. Endoplasmic reticulum stress-induced transcription factor, CHOP, is crucial for dendritic cell IL-23 expression. Proceedings of the national academy of Sciences 107: 17698–17703.
Turner, M.J., D.P. Sowders, M.L. DeLay, R. Mohapatra, S. Bai, J.A. Smith, J.R. Brandewie, J.D. Taurog, and R.A. Colbert. 2005. HLA-B27 misfolding in transgenic rats is associated with activation of the unfolded protein response. The Journal of Immunology 175: 2438–2448.
de Castro, C.P., S.A. Jones, C.N. Cheallaigh, C.A. Hearnden, L. Williams, J. Winter, E.C. Lavelle, K.H. Mills, and J. Harris. 2012. Autophagy regulates IL-23 secretion and innate T cell responses through effects on IL-1 secretion. The Journal of Immunology 189: 4144–4153.
Chicote, J., V.J. Yuste, J. Boix, and J. Ribas. 2020. Cell death triggered by the autophagy inhibitory drug 3-methyladenine in growing conditions proceeds with DNA damage. Frontiers in Pharmacology 11: 580343.
Siebert, S., N.L. Millar, and I.B. McInnes. 2019. Why did IL-23p19 inhibition fail in AS: A tale of tissues, trials or translation? Annals of the Rheumatic Diseases 78: 1015–1018.
Baeten, D., M. Østergaard, J.C. Wei, J. Sieper, P. Järvinen, L.S. Tam, C. Salvarani, T.H. Kim, A. Solinger, Y. Datsenko, et al. 2018. Risankizumab, an IL-23 inhibitor for ankylosing spondylitis: Results of a randomised double-blind placebo-controlled proof-of-concept dose-finding phase 2 study. Annals of the Rheumatic Diseases 77: 1295–1302.
Acknowledgements
We thank all the patients and medical staff who generously contributed to this study.
Funding
This research has been supported by grants from Tehran University of Medical Sciences (TUMS), Grant No. 92–02-30–23095.
Author information
Authors and Affiliations
Contributions
A.R. conceived the study and wrote the manuscript. M.V. and A.R.J. contributed to sample preparation. F.B., J.M., and M.R. took the lead in writing the manuscript. M.M. and A.A.A. contributed to the interpretation of the results. M.H.N. critically revised the manuscript and provided the final approval. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
The protocol of the study was endorsed by the ethical committee of Tehran University of Medical Sciences (Ethical code: IR.TUMS.REC.1394.671) for use of human participants. All study subjects signed informed consent forms before sampling. All research was performed in accordance with relevant guidelines and regulations of the Tehran University of Medical Sciences.
Consent for Publication
Written informed consent was obtained from all participating individuals.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rezaiemanesh, A., Mahmoudi, M., Amirzargar, A.A. et al. Upregulation of Unfolded Protein Response and ER Stress–Related IL-23 Production in M1 Macrophages from Ankylosing Spondylitis Patients. Inflammation 45, 665–676 (2022). https://doi.org/10.1007/s10753-021-01575-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10753-021-01575-z