Abstract
Downregulating miR-217-5p could protect cardiomyocytes against ischemia/reperfusion (I/R) injury, but its role in restoring mitochondrial function of I/R-injured cardiomyocytes remained unclear. H9C2 cardiomyocyte-derived cell line with I/R injury was established in vitro on the basis of hypoxia/reperfusion (H/R) model. Cell viability and apoptosis were respectively detected by MTT assay and flow cytometry. Contents of lactate dehydrogenase (LDH) and adenosine triphosphate (ATP) were determined. Flow cytometry was performed to measure the production of reactive oxygen species (ROS) and mitochondrial membrane potential (MMP). Target gene and potential binding sites between miR-217-5p and Sirtuin1 (SIRT1) were predicted by TargetScan and confirmed by dual-luciferase reporter assay. Relative SIRT1 and expressions of autophagy-related and apoptosis-related genes were measured by quantitative real-time polymerase chain reaction (qRT-PCR) and Western blot. After I/R treatment, the viability of H9C2 cardiomyocyte-derived cell line and ATP contents were reduced, but LDH and ROS contents were increased, at the same time, cell apoptosis and the expressions of miR-217-5p, p62 and cleaved caspase-3 were increased, whereas the expressions of SIRT1, LC3 (light chain 3), PINK1 (PTEN-induced kinase 1), Parkin, Bcl-2, and c-IAP (inhibitor of apoptosis protein) were reduced. However, downregulating miR-217-5p expression reversed the effects of I/R. SIRT1 was predicted and verified to be the target of miR-217-5p, and silencing SIRT1 reversed the effects of downregulating miR-217-5p on I/R-injured cells. Downregulating miR-217-5p could help restore mitochondrial function via targeting SIRT1, so as to protect cardiomyocytes against I/R-induced injury.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Myocardial ischemia/reperfusion (I/R) injury was a major challenge in myocardial infraction treatment, and reperfusion is currently the only effective therapy for treating myocardial infraction [1]. Factors such as oxidative stress, intracellular calcium overload, inflammation, and apoptosis have been found implicated in the pathogenesis of myocardial I/R [2]. However, the specific molecular mechanisms of myocardial I/R injury remained poorly understood, which requires a better and comprehensive study of I/R pathogenesis.
MicroRNAs (miRNAs) are single-strand RNAs with 18–25 nucleotides in length. MiRNAs have regulatory effects on gene expressions by binding to mRNA targets in the 3′-untranslated regions (3′-UTRs) [3], and they play pivotal roles in myocardial I/R injury through regulating multiple pathophysiological processes [4]. For example, miR-24-3p protects H9C2 cells against I/R injury via activating Nrf2-Keap1 pathway [5]; upregulation of miR-374a-5p also protects against I/R injury in vitro and in vivo [6]; miR-206 protects against myocardial I/R injury by targeting growth arrest DNA damage-inducible gene 45β (Gadd45β) [7].
The upregulation of miR-217-5p, which has been found in the myocardial tissues of patients with chronic heart failure, also promotes cardiac hypertrophy and dysfunction via targeting PTEN [8]. Xia et al. demonstrated that downregulating the expressions of miR-217 and miR-543 could mitigate inflammatory response and myocardial injury in children with viral myocarditis via regulating SIRT1/AMPK/NF-κB signaling pathway [9]. As for I/R injury, Li et al. revealed that inhibition of miR-217 could protect cardiomyocytes against I/R injury through suppressing the activation of NF-κB and MAPK pathways [10]. Moreover, dysfunction of mitochondria is closely related to I/R injury [11]; however, the role of miR-217-5p in the restoration of mitochondrial function to protect cardiomyocytes against I/R injury remained unclear. The present study established a hypoxia/reoxygenation (H/R) model as a simulation of I/R injury in vitro [12] to investigate the molecular mechanisms through which miR-217-5p restored mitochondrial function against myocardial I/R injury, hoping to develop a potential strategy for treating myocardial I/R injury.
MATERIAL AND METHODS
Cell Culture and Treatment
Rat H9C2 cardiomyocyte-derived cell lines were purchased from American Type Culture Collection (ATCC; CRL-1446; Rockville, MD, USA) and cultured in Dulbecco’s modified eagle medium (DMEM; 01–057-1; Biological Industries, Cromwell, CT, USA) containing 10% fetal bovine serum (FBS; S1620, Biowest, Nuaillé, France) and 100 U/mL penicillin-streptomycin (P1400; Solarbio, Beijing, China) at 37 °C with 5% CO2.
A H/R model was established as previously described for stimulating I/R injury in vitro [13]. Primary H9C2 cardiomyocyte-derived cell lines were first cultured in DMEM supplemented with 10% FBS and 100 U/mL penicillin-streptomycin at 37 °C with 5% CO2, and then incubated in a hypoxia incubator (#27310; StemCell, Beijing, China) with 95% N2 and 5% CO2 for 30 min. Finally, H9C2 cardiomyocyte-derived cell lines were cultured in fresh DMEM with 10% FBS at 37 °C with 5% CO2 for another 2 h.
Cell Transfection
MiR-217-5p inhibitor (I; B03001) and its control (IC; B04001) were purchased from Gene Pharma (Shanghai, China); small interfering RNA for SIRT1 (siSIRT1; SR30002) and negative control (siNC; SR3004) was synthesized by Origene (Beijing, China). The H9C2 cardiomyocyte-derived cell lines were cultured at a density of 5 × 104 cells/well in a 24-well plate to 80% confluence. Then, 2-μg inhibitor and its control, 30 pmol siSIRT1 and siNC were transfected into the cells using Lipofectamine 2000 reagent (11,668,027; Thermo Fisher Scientific, Waltham, MA, USA). After transfection for 48 h, the cells were harvested for subsequent studies. Transfection primer sequences are listed in Table 1.
MTT Assay
The transfected H9C2 cardiomyocyte-derived cell lines (5 × 103 cells/well) were seeded into 96-well plates. For the detection of cell viability, 20 μL MTT reagent (5 mg/mL; K017; Fine Biotech, Wuhan, China) was added into the cells and incubated together at 37 °C for 4 h, and then the visible formazan crystals were solubilized by 150 μL dimethyl sulfoxide (DMSO; 598–10, RJ Matthews, Massillon, OH, USA). OD values at an absorbance of 490 nm were recorded using a UV5 Spectrophotometer (Mettler Toledo, Columbus, OH, USA).
Lactate Dehydrogenase Leakage Assay
After I/R stimulation, H9C2 cardiomyocyte-derived cell lines (2 × 104 cells/well) were seeded into a 96-well plate. After collection of cell supernatant, 60 μL lactate dehydrogenase (LDH) detection reagent (A020–1-2; Nanjing Jiancheng Bioengineering Institute, Nanjing, China) was added to the supernatant for further incubation at 37 °C in the dark. Mass LDH leakage was measured at an absorbance at 490 nm using a fluorescence microplate reader (TriStar2 LB942; Berthold Technologies GmbH, Bad Wildbad, Germany).
Adenosine Triphosphate Measurement
The cardiomyocytes were washed by cold PBS twice, collected, and centrifuged at 13,000×g for 15 min at 4 °C. Proteins of H9C2 cardiomyocyte-derived cell lines were subsequently extracted and its concentration was determined by bicinchoninic acid (BCA) protein kit (J63283; Alfa Aesar, Shanghai, China). Two hundred microliters of protein supernatant was then mixed with adenosine triphosphate (ATP) assay kit (A095–1, Nanjing Jiancheng Bioengineering Institute, China), and the fluorescence was measured at 560-nm absorbance using a microplate reader (TriStar2 LB942; Berthold Technologies GmbH, Germany).
Detection of Cell Apoptosis by Flow Cytometry
Cell apoptosis was determined by Annexin V-FITC/Propidium iodide (PI) apoptosis kit (A211; Vazyme, Nanjing, China) according to the instructions. Forty-eight hours after the transfection, the H9C2 cardiomyocyte-derived cell lines were harvested, washed with cold phosphate buffered saline (PBS) twice, and subsequently treated by 5 μL Annexin V and PI together at room temperature for 20 min in the dark. Cell apoptosis was analyzed by Guava easyCyte Benchtop Flow Cytometer (BR168323; Luminex, Austin, TX, USA) with Kaluza C Analysis Software (Beckman Coulter, Indianapolis, IN, USA).
Reactive Oxygen Species Assay
The H9C2 cardiomyocyte-derived cell lines were cultured in a 24-well plate for 24 h. The cells were observed under a fluorescence microscope (BX51, Olympus, Tokyo, Japan), then collected, and resuspended in the flushing fluid at an adjusted density of 1 × 106 cells/well. Next, the cells were washed by PBS and incubated with 10 μM 2′, 7′-dichlorofluroscein diacetate (DCFH-DA; AAT Bioquest, Sunnyvale, CA, USA) in a serum-free DMEM at 37 °C for 30 min in the dark. The cells were finally analyzed using Guava easyCyte Benchtop Flow Cytometer (Luminex, USA) with an emission filter of 525 nm and the mean fluorescent intensity was measured.
Mitochondrial Membrane Potential Assessment
Mitochondrial membrane potential (MMP) assessment was performed using JC-1 Mitochondrial Membrane Potential Assay Kit (10,009,172, Cayman Chemical, Ann Arbor, MI, USA) as previously described [14]. The H9C2 cardiomyocyte-derived cell line samples were collected after centrifugation, then resuspended in JC-1 staining solution (10,009,908, Cayman Chemical), and further incubated for 15 min. Subsequently, the cells were resuspended in 500-μL pre-heated incubation buffer. MMP assessment was conducted using Guava easyCyte Benchtop Flow Cytometer (Luminex, USA).
Target Gene Prediction and Dual-Luciferase Reporter Assay
Target gene and potential binding sites between miR-217 and SIRT1 were predicted by TargetScan 7.2 and further confirmed using dual-luciferase reporter assay.
PMIR-REPORT Luciferase vector (AM5795; Thermo Fisher Scientific, USA) with wild-type or mutated SIRT1 sequences were cloned into the pMirGLO reporter vector (E1330; Promega, Madison, WI, USA) to generate SIRT1-WT (sequence: 5′-UGGCUACACUAAAGAAUGCAGUA-3′) and SIRT1-MUT (sequence: 5′-UGGCUACACUAAAGAGACUUUAAC-3′). The H9C2 cardiomyocyte-derived cell lines were cultured in 24-well plates at 2.5 × 104 cells/well and co-transfected with SIRT1-WT and SIRT1-MUT, miR-217-5p mimic (B02003; Gene Pharma, Shanghai), or mimic control (B04001; Gene pharma), using Lipofectamine 2000 reagent at 37 °c for 48 h. Finally, the firefly luciferase activity was detected and normalized to that of Renilla in a dual-luciferase reporter assay system (E1910; Promega, USA).
RNA Isolation and Quantitative Real-Time Polymerase Chain Reaction
Total RNAs were extracted from the cells using Trizol reagent (A33250, Invitrogen, Madison, MI, USA) according to the instructions and preserved in a − 80 °0 refrigerator. The RNA concentration was determined and quantified by Nano Drop 2000 biological spectrometer (catalog number: ND-LITE-PR; Thermo Fisher Scientific, USA). One microgram of the total RNA was reverse-transcribed into cDNA by First-strand cDNA Synthesis Kit (E6300L; New England Biolabs, Ipswich, MA, USA). Quantitative real-time polymerase chain reaction (qRT-PCR) experiment was conducted using Phusion® High-Fidelity PCR kit (New England BioLabs, Beijing, China) in AriaMx real-time PCR System (G8830A, Agilent, Santa Clara, CA, USA) under the following conditions: at 95 °C for 5 min, followed by 38 cycles at 95 °C for 15 s and at 60 °C for 30s. The primer sequences are listed in Table 2. GAPDH and U6 were used as internal references. Relative gene expressions were calculated by 2−ΔΔCT method [15].
Western Blot
The expressions of SIRT1 and autophagy-related proteins (Light Chain 3, LC3; p62; PTEN-induced kinase 1, PINK1; Parkin; Cleaved caspase-3; Bcl-2 and c-inhibitor of apoptosis, c-IAP) were determined by Western blot as previously described [16]. The proteins were lysed and extracted by RIPA buffer (MB-077-0015; Rockland, Limerick, PA, USA) shortly after the cell collection. The protein concentrations were measured by bicinchoninic acid (BCA) protein kit (Alfa Aesar, China). Sample protein lysates (30 μg) were electrophoresed by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE; P1200; Solarbio, Beijing, China) and then transferred into Amersham Hybond polyvinylidene fluoride (PVDF) membranes (1,060,069, GE Health Care, Pittsburgh, PA, USA), which were blocked by 5% non-fat milk for 2 h. The membranes were incubated with the following primary antibodies: anti-SIRT1 antibody (mouse, 1:1000, ab110304, Abcam, Cambridge, UK), anti-LC3B antibody (rabbit, 1:2000, ab192890, Abcam, UK), anti-p62 antibody (rabbit, ab109012, 1:10000, Abcam, UK), anti-PINK1 antibody (rabbit, 1:1000, ab23707, Abcam, UK), anti-Parkin antibody (rabbit, 1:1000, ab109199, Abcam, UK), anti-cleaved caspase-3 antibody (rabbit, 1:500, ab49822, Abcam, UK), anti-Bcl-2 antibody (rabbit, 1:1000, ab59348, Abcam, UK), anti-c-IAP antibody (rabbit, ab2399, 1:200, Abcam, UK), and anti-β-actin antibody (rabbit, ab8226, 1:2000, Abcam, UK) at 4 °C overnight, with β-actin as an internal control. Then the membranes were incubated with secondary horseradish peroxidase (HRP)-combined antibodies goat anti-rabbit IgG H&L (HRP) (NBP2-30348H, 1:2000, Bio-Techne, Shanghai, China) and goat anti-mouse IgG H&L (HRP) (NB7539, 1:2000, Bio-Techne, China) at room temperature for 1 h, and then washed by tris-buffer saline tween (TBST) for three times. Next, the protein bands were collected and analyzed by an enhanced chemiluminescence (ECL) kit (SW2020; Solarbio, China). The gray values of the strips were calculated by ImageJ (version 5.0; Bio-Rad, Hercules, CA, USA).
Statistical Analysis
All the experiments were independently performed at least three times. The data were expressed as mean ± standard deviation (SD). Statistical analysis was conducted using SPSS 19.0 software (Chicago, IL, USA). Student’s t test and one-way ANOVA followed by Dunnett’s post hoc test were applied to analyze statistical significance, which was defined as a p < 0.05.
RESULTS
Downregulating miR-217-5p Reversed the Effects of I/R on the Viability, Apoptosis, and Contents of LDH, ATP, and ROS of H9C2 Cardiomyocyte-Derived Cell Lines
After I/R treatment, H9C2 cardiomyocyte-derived cell lines were transfected miR-217-5p inhibitor to explore the effects of I/R and miR-217-5p. We found that H9C2 cardiomyocyte-derived cell line viability was decreased after I/R treatment (Fig. 1a, P < 0.001) but was increased after the transfection with miR-217-5p inhibitor (Fig. 1a, P < 0.01), suggesting that downregulating miR-217-5p could reverse the effects of I/R on the cell viability of H9C2.
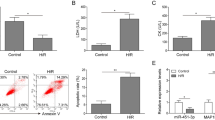
Downregulation of miR-217-5p reversed the effects of I/R on viability, apoptosis, and contents of LDH, ATP, and ROS of H9C2 cardiomyocyte-derived cell line. A, The viability of H9C2 cardiomyocyte-derived cell line after I/R treatment and miR-217-5p inhibitor (I) transfection was measured by MTT assay. B, LDH content of H9C2 cardiomyocyte-derived cell line after I/R treatment and miR-217-5p inhibitor (I) transfection was measured by LDH detection kit. C, ATP content in H9C2 cardiomyocyte-derived cell line after I/R treatment and miR-217-5p inhibitor (I) transfection were detected by ATP detection kit. D, e apoptosis of H9C2 cardiomyocyte-derived cell line after I/R treatment and miR-217-5p inhibitor (I) transfection was detected by flow cytometry. F, g ROS content of H9C2 cardiomyocyte-derived cell line after I/R treatment and miR-217-5p inhibitor (I) transfection were quantified by DCFH-DA assay and flow cytometry. All experiments were performed in triplicate and the data were expressed as mean ± standard deviation (SD). ***P < 0.001, vs. control; ^^P < 0.01, ^^^P < 0.001, vs. I/R + IC. LDH, lactate dehydrogenase; ATP, adenosine triphosphate; ROS, reactive oxygen species; I/R, ischemia/reperfusion; IC, inhibitor control.
As shown in Fig. 1b and c, after I/R treatment, LDH content was significantly increased and ATP content was reduced, whereas the opposite results were observed after downregulation of miR-217-5p (Fig. 1b and c, P < 0.01), indicating that downregulating miR-217-5p could reversed the effects of I/R on LDH and ATP contents of the H9C2 cardiomyocyte-derived cell lines.
The data of flow cytometry demonstrated that the apoptosis of H9C2 cardiomyocyte-derived cell lines was high after I/R treatment (Fig. 1d and e, P < 0.001) but was reduced after downregulating miR-217-5p (Fig. 1d and e, P < 0.001). This demonstrated that downregulating miR-217-5p could reverse the effects of I/R on H9C2 cardiomyocyte-derived cell line apoptosis.
Reactive oxygen species (ROS) content was determined by DCFH-DA fluorescence assay and flow cytometry. In Fig. 1f and g, it could be observed that ROS content of H9C2 cardiomyocyte-derived cell lines was evidently increased after I/R treatment, but downregulating miR-217-5p reduced ROS content of H9C2 cardiomyocyte-derived cell lines (Fig. 1f and g, P < 0.001). These two findings showed that downregulating miR-217-5p could reverse the effects of I/R on ROS content in H9C2 cardiomyocyte-derived cell lines.
Downregulating miR-217-5p Reversed the Effects of I/R on MMP and Expressions of miR-217-5p and SIRT1 of H9C2 Cardiomyocyte-Derived Cell Lines
Flow cytometry was performed to examine the effects of I/R and downregulation of miR-217-5p on MMP. As shown in Fig. 2a and b, MMP was reduced after I/R treatment but was restored by downregulation of miR-217-5p (Fig. 2a and b, P < 0.001), indicating that the downregulation of miR-217-5p could reverse the effects of I/R on MMP of H9C2 cardiomyocyte-derived cell lines.
Downregulation of miR-217-5p reversed the effects of I/R on mitochondrial membrane potential (MMP) and expressions of miR-217-5p and SIRT1 of H9C2 cardiomyocyte-derived cell line. A, b MMP of H9C2 cardiomyocyte-derived cell line after I/R treatment and miR-217-5p inhibitor (I) transfection was assessed by flow cytometry. C, d relative expressions of miR-217-5p and SIRT1 in H9C2 cardiomyocyte-derived cell line after I/R treatment and miR-217-5p inhibitor (I) transfection were measured by quantitative real-time polymerase chain reaction (qRT-PCR). U6 (for miR-217-5p) and β-actin (for SIRT1) were internal controls. All experiments have been performed in triplicate and the data were expressed as mean ± standard deviation (SD). ***P < 0.001, vs. control; ^P < 0.05, ^^^P < 0.001, vs. I/R + IC. SIRT1, Sirtuin 1.
In Fig. 2c and d, after I/R treatment, miR-217-5p expression was increased and SIRT1 expression was reduced, while the miR-217-5p inhibitor inhibited the miR-217-5p expression and promoted SIRT1 expression in the I/R + I group compared with I/R group (Fig. 2c and d, P < 0.01). The results suggested that downregulation of miR-217-5p could reverse the effects of I/R on the expressions of miR-217-5p and SIRT1 of H9C2 cardiomyocyte-derived cell lines.
Downregulating miR-217-5p Reversed the Effects of I/R on Protein Expressions of SIRT1 and Autophagy-Related and Apoptosis-Related Genes
After I/R treatment and miR-217-5p inhibitor transfection, the expressions of SIRT1 and autophagy-related and apoptosis-related genes were determined by conducting Western blot. In Fig. 3a–c, we found that the expressions of SIRT1, LC3-II, Mito-LC3-II, PINK1, Parkin, Bcl-2, and c-IAP were reduced but those of LC3-I, p62, and C Caspase-3 were increased after I/R treatment; however, these effects were reversed by downregulation of miR-217-5p (Fig. 3a–c, P < 0.001). Also, as shown in Fig. 3d, we found that LC3-II/LC3-I ratio was reduced after I/R treatment, while downregulating miR-217-5p reversed such an effect (Fig. 3d, P < 0.001).
Downregulation of miR-217-5p reversed the effects of I/R on expressions of SIRT1 and autophagy-related and apoptosis-related genes. A–c relative protein/β-actin expressions of SIRT1, LC3-I, LC3-II, Mito-LC3-II, p62, PINK1, Parkin, cleaved (C) caspase-3, Bcl-2, and c-IAP after I/R treatment and miR-217-5p inhibitor (I) transfection were measured by Western blot. β-actin was an internal control. D LC3-II/LC3-I ratio after I/R treatment and miR-217-5p inhibitor (I) transfection was quantified. All experiments have been performed in triplicate and the data were expressed as mean ± standard deviation (SD). ***P < 0.001, vs. control; ^^^P < 0.001, vs. I/R + IC. LC3, light chain 3; PINK1, PTEN-induced kinase 1; Bcl-2, B cell lymphoma 2; c-IAP, cellular inhibitor of apoptosis protein.
SIRT1 Was the Target of miR-217-5p, and Silencing SIRT1 Reversed the Effects of Downregulation of miR-217-5p on SIRT1 Expression of I/R-Treated H9C2 Cardiomyocyte-Derived Cell Lines
TargetScan predicted that SIRT1 was the target of miR-217-5p because it contained the complementary binding sites at 3′-UTRs (Fig. 4a). The prediction was further verified by dual-luciferase reporter vectors containing their 3′-UTRs, and we observed that the relative luciferase activity of SIRT1-WT-mimic (M) group was reduced as compared with SIRT-WT-mimic control (MC) group (Fig. 4b, P < 0.001), whereas that in SIRT-MUT-M group was not affected as compared with SIRT-MUT-MC group. Thus, SIRT1 was confirmed to be the target of miR-217-5p.
SIRT1 was the target of miR-217-5p, and silencing SIRT1 reversed the effects of downregulation of miR-217-5p on SIRT1 expression, cell viability, and LDH and ATP contents in I/R-treated H9C2 cardiomyocyte-derived cell line. A predicted target gene and binding sites of miR-217-5p and SIRT1 at 3′-UTR and SIRT1 wild-type and mutated 3′-UTR (SIRT1 3′-UTR WT; SIRT1 3′-UTR MUT) were listed. B dual-luciferase reporter assay confirmed that SIRT1 was the target gene of miR-217-5p. C–e relative mRNA (c) and protein/β-actin expressions (d, e) of SIRT1 in I/R-injured H9C2 cardiomyocyte-derived cell line after transfection of miR-217-5p inhibitor and siSIRT1 were measured by qRT-PCR and Western blot. β-actin was an internal control. F H9C2 cardiomyocyte-derived cell line viability after transfection of miR-217-5p inhibitor and siSIRT1 were measured by MTT assay. G LDH content in H9C2 cardiomyocyte-derived cell line after transfection of miR-217-5p inhibitor and siSIRT1 was detected. H ATP content in H9C2 cardiomyocyte-derived cell line after transfection of miR-217-5p inhibitor and siSIRT1 was measured. All experiments have been performed in triplicate and the experimental data were expressed as mean ± standard deviation (SD). +++P < 0.001, vs. MC; **P < 0.01, ***P < 0.001, vs. I/R; ^^P < 0.01, ^^^P < 0.001, vs. I/R + I + siNC. siSIRT1, small interfering RNA for SIRT1; NC, negative control; MC, mimic control.
The mRNA and protein expressions of SIRT1 were measured after transfecting miR-217-5p inhibitor and siSIRT1 into the I/R-treated H9C2 cardiomyocyte-derived cell lines. We observed that after transfection of miR-217-5p inhibitor into I/R-injured H9C2 cardiomyocyte-derived cell lines, SIRT1 mRNA and protein expressions were both increased (Fig. 4c–e, P < 0.001). Silencing SIRT1, however, reduced both mRNA and protein expressions of SIRT1 of I/R-treated H9C2 cardiomyocyte-derived cell lines (Fig. 4c–e, P < 0.001).
Silencing SIRT1 Reversed the Effects of Downregulation of miR-217-5p on Viability, Apoptosis, and LDH, ATP, and ROS Contents of I/R-Injured H9C2 Cardiomyocyte-Derived Cell Lines
After the transfection of miR-217-5p inhibitor and siSIRT1 into I/R-treated H9C2 cardiomyocyte-derived cell lines, the viability, apoptosis, and contents of LDH, ATP, and ROS of the cells were determined. As shown in Fig. 4f, the viability of I/R-injured cells was increased after downregulation of miR-217-5p but was then reduced after silencing SIRT1 (Fig. 4f, P < 0.01). This indicated that silencing SIRT1 reversed the effects of downregulating miR-217-5p on the viability of I/R-injured H9C2 cardiomyocyte-derived cell lines.
LDH and ATP contents of I/R-injured H9C2 cardiomyocyte-derived cell lines after the transfection of miR-217-5p inhibitor and siSIRT1 were subsequently measured. We found that LDH content was reduced and ATP content was increased after downregulating miR-217-5p (Fig. 4g and h, P < 0.01). However, silencing SIRT1 increased LDH content and decreased ATP content (Fig. 4g and h, P < 0.01), showing that silencing SIRT1 reversed the effects of downregulating miR-217-5p on LDH and ATP contents of I/R-injured H9C2 cardiomyocyte-derived cell lines.
Flow cytometry was performed to detect apoptosis of I/R-injured H9C2 cardiomyocyte-derived cell lines after the transfection of miR-217-5p inhibitor and siSIRT1. In Fig. 5a, the cell apoptosis was reduced after downregulation of miR-217-5p, whereas silencing SIRT1 increased the cell apoptosis (Fig. 5a, P < 0.001), indicating that silencing SIRT1 reversed the effects of downregulation of miR-217-5p on the apoptosis of I/R-injured H9C2 cardiomyocyte-derived cell lines.
Silencing SIRT1 reversed the effects of downregulation of miR-217-5p on cell apoptosis, ROS content, and MMP in I/R-injured H9C2 cardiomyocyte-derived cell line. A H9C2 cardiomyocyte-derived cell line apoptosis after transfection of miR-217-5p inhibitor and siSIRT1 was detected by flow cytometry. B ROS content in H9C2 cardiomyocyte-derived cell line after transfection of miR-217-5p inhibitor and siSIRT1 was measured by DCFH-DA assay and flow cytometry. C mitochondrial membrane potential in H9C2 cardiomyocyte-derived cell line after transfection of miR-217-5p inhibitor and siSIRT1 was assessed by flow cytometry. All experiments have been performed in triplicate and the experimental data were expressed as mean ± standard deviation (SD). ***P < 0.001, vs. I/R; ^^^P < 0.001, vs. I/R + I + siNC. DCFH-DA: 2’, 7’-dichlorofluroscein diacetate.
After the transfection of miR-217-5p inhibitor and siSIRT1, ROS content of I/R-injured H9C2 cardiomyocyte-derived cell lines was decreased after downregulation of miR-217-5p (Fig. 5b, P < 0.001), while silencing SIRT1 increased ROS content of the cells (Fig. 5b, P < 0.001). This showed that silencing SIRT1 reversed the effects of downregulating miR-217-5p on ROS content of I/R-injured H9C2 cardiomyocyte-derived cell lines.
Silencing SIRT1 Reversed the Effects of Downregulating miR-217-5p on MMP of I/R-Treated H9C2 Cardiomyocyte-Derived Cell Lines
Flow cytometry was performed to evaluate the effects of downregulating miR-217-5p and silencing SIRT1 on mitochondrial membrane potential of I/R-injured H9C2 cardiomyocyte-derived cell lines. In Fig. 5c mitochondrial membrane potential of I/R-treated H9C2 cardiomyocyte-derived cell lines was increased but then decreased after silencing SIRT1 in the I/R-treated H9C2 cardiomyocyte-derived cell lines (Fig. 5c, P < 0.001). Therefore, it could be concluded that silencing SIRT1 reversed the effects of downregulation of miR-217-5p on mitochondrial membrane potential of I/R-injured H9C2 cardiomyocyte-derived cell lines.
Silencing SIRT1 Reversed the Effects of Downregulation of miR-217-5p on the Expressions of Autophagy-Related and Apoptosis-Related Genes of I/R-Injured H9C2 Cardiomyocyte-Derived Cell Lines
The expressions of autophagy-related and apoptosis-related genes of I/R-treated H9C2 cardiomyocyte-derived cell lines after the transfection of miR-217-5p inhibitor and siSIRT1 were determined. As shown in Fig. 6a–c, after the transfection of miR-217-5p inhibitor into I/R-treated cells, the expressions of LC3-II, Mito-LC3-II, PINK1, Parkin, Bcl-2, and c-IAP were increased but those of LC3-I, p62, and C Caspase-3 were reduced, whereas silencing SIRT1 resulted in opposite effects (Fig. 6a–c, P < 0.001). In addition, as shown in Fig. 6d, LC3-II/LC3-I ratio of I/R-injured H9C2 cardiomyocyte-derived cell lines was significantly increased after downregulation of miR-217-5p, while silencing SIRT1 reduced the LC3-II/LC3-I ratio (Fig. 6d, P < 0.001).
Silencing SIRT1 reversed the effects of downregulation of miR-217-5p on expressions of autophagy-related and apoptosis-related genes in I/R-injured H9C2 cardiomyocyte-derived cell line. A–c relative protein/β-actin expressions of LC3-I, LC3-II, Mito-LC3-II, p62, PINK1, Parkin, cleaved (C) caspase-3, Bcl-2, and c-IAP after transfection of miR-217-5p inhibitor and siSIRT1 were measured by Western blot. β-actin was an internal control. D LC3-II/LC3-I ratio after transfection of miR-217-5p inhibitor and siSIRT1 was quantified. All experiments have been performed in triplicate and the data were expressed as mean ± standard deviation (SD). ***P < 0.001, vs. I/R; ^^^P < 0.001, vs. I/R + I + siNC.
DISCUSSION
In the present study, the H/R-treated H9C2 cardiomyocyte-derived cell lines were cultured in vitro to simulate myocardial I/R injury in vivo. The viability and apoptosis of H/R-injured H9C2 cardiomyocyte-derived cell lines were detected to explore the effects of I/R injury on miR-217-5p expression and the effect of downregulation miR-217-5p on the H9C2 cardiomyocyte-derived cell lines. The results showed that the viability of H9C2 cardiomyocyte-derived cell lines was reduced by H/R treatment, whereas the apoptosis of H9C2 cardiomyocyte-derived cell lines was increased, but downregulation of miR-217-5p caused opposite effects, suggesting that downregulation miR-217-5p might increase viability and reduce apoptosis of the H9C2 cardiomyocyte-derived cell lines after I/R injury.
In cardiac I/R injury, excessive mitochondrial fission, together with mitophagy defection, oxidative stress, dyshomeostasis on calcium, and programmed cell death are modulated by ER-mitochondria microdomains [17]. Study found that mitochondrial dysfunction, which is characterized by the loss of mitochondrial membrane potential and altered mitochondrial biogenesis and clearance, plays a key role in multiple cardiovascular diseases and could reduce ATP production [18]. Production of mitochondrial ROS is related to the oxidative damage caused by I/R injury in several pathological situations such as heart attack and stroke [19], and LDH is indicative of the degree of myocardial injury [20]. Previous study demonstrated that I/R consist of several distinct phases of cellular injury including ATP depletion, accumulation of LDH during ischemia, and production of ROS during reperfusion [21]. In the present study, we found that LDH and ROS contents were increased but ATP and mitochondrial membrane potential were decreased, whereas downregulation of miR-217-5p caused opposite effects, suggesting that I/R treatment could aggravate injury to H9C2 cardiomyocyte-derived cell lines, but downregulation of miR-217-5p could ameliorate myocardial I/R injury by the restoration of mitochondrial functions.
Mitophagy refers to an autophagic response specifically targeting damaged and cytotoxic mitochondria, and is involved with several cardiovascular disorders [22]. Several proteins, such as LC3, p62, PINK-1, and Parkin, have been found to play pivotal roles in mitophagy by increasing p62 expression and reducing the expressions of LC3, PINK-1, and Parkin [23, 24]. Autophagosomes could further damage mitochondria through interaction with LC3 or LC3-related proteins and autophagy adaptors or receptors [25]. P62, a selective cargo autophagy receptor that degenerates misfolded proteins, affects the balance between mitophagy and recognize and binds ubiquitinated proteins through binding to LC3 [26, 27]. Several types of mitophagy will be activated to damage mitochondria by reducing membrane potential dependent on PTEN-induced putative kinase-1 (PINK-1) [28]. Parkin, together with PINK-1, is indispensable for mitophagy, and PINK-1-Parkin axis is a core mechanism of neurons mitophagy [29]. In our study, after I/R treatment, the p62 expression was significantly increased and those of expressions of LC3, PINK1, and Parkin were decreased, whereas downregulation of miR-217-5p produced opposite results. To our best knowledge, we were the first to demonstrate that downregulation of miR-217-5p could reverse the effects of I/R on mitophagy.
Apoptosis plays an important role in I/R pathogenesis, and generally prolonged ischemia will increase necrosis and reperfusion will promote cell apoptosis [30]. Bcl-2, caspase-3, and c-IAP play key roles in cell apoptosis [31]. Bcl-2 and its family of proteins could regulate cell apoptosis through regulating mitochondrial permeability [32]. Caspase-3 is a marker of apoptosis because its activity is required for major apoptosis-related morphological and biochemical events, and increasing the expression of caspase-3 will induce cell apoptosis [33]. C-inhibitor of apoptosis (c-IAP) protein dose not binds to caspases, but it indirectly regulates caspase activation [34]. The present study found that after I/R treatment, cleaved caspase-3 expression was increased, but those of Bcl-2 and c-IAP were reduced, whereas downregulation of miR-217-5p caused opposite effects, suggesting that downregulation of miR-217-5p could reversed the effects of I/R on the apoptosis of cardiomyocytes, which was also consistent with a previous study [35].
Sirtuin 1 (SIRT1), which is a NAD-dependent deacetylase, plays a protective role in several diseases including in cardiovascular diseases [36]. SIRT1 cardiomyocyte-specific deletion could sensitize myocardium to ischemia and reperfusion injury [37], and SIRT1 can increase autophagy and reduce apoptosis of cardiomyocytes against hypoxic stress [38]. For its effects on mitochondrial function of cardiomyocytes, Wu et al. found that SIRT1 plays a key role in protecting Icariin-pretreated cardiomyocytes against I/R injury [39]. In addition, as Rao et al. showed that SIRT1 is the target of miR-217, and silencing SIRT1 attenuated the effects of downregulation of miR-217 on the neurons treated by oxygen-glucose deprivation and reoxygenation (OGD/R) [40], the present study found that SIRT1 was the target of miR-217-5p, and that silencing SIRT1 reversed the effects of downregulating miR-217-5p on I/R-treated cells through the restoration of mitochondrial function in I/R-injured cells.
There were also some limitations in our present study, as we only explored the effects of downregulation of miR-217-5p on reversing I/R-induced injury in vitro, but its efficacy in vivo remained unclear. Thus, further studies were required to validate our findings in vivo.
In conclusion, downregulation of miR-217-5p could protect cardiomyocytes against I/R injury in vitro though restoring mitochondrial function via targeting SIRT1, indicating that miR-217-5p could be a therapeutic target for the diagnosis and prognosis of I/R injury.
Data Availability
The analyzed data sets generated during the study are available from the corresponding author on reasonable request.
References
Li, Z., Y. Zhang, N. Ding, Y. Zhao, Z. Ye, L. Shen, H. Yi, and Y. Zhu. 2019. Inhibition of lncRNA XIST improves myocardial I/R injury by targeting miR-133a through inhibition of autophagy and regulation of SOCS2. Molecular Therapy--Nucleic Acids 18: 764–773. https://doi.org/10.1016/j.omtn.2019.10.004.
Wu, Q., R. Wang, Y. Shi, W. Li, M. Li, P. Chen, B. Pan, Q. Wang, C. Li, J. Wang, G. Sun, X. Sun, and H. Fu. 2020. Synthesis and biological evaluation of panaxatriol derivatives against myocardial ischemia/reperfusion injury in the rat. European Journal of Medicinal Chemistry 185: 111729. https://doi.org/10.1016/j.ejmech.2019.111729.
Krol, J., I. Loedige, and W. Filipowicz. 2010. The widespread regulation of microRNA biogenesis, function and decay. Nature Reviews. Genetics 11 (9): 597–610. https://doi.org/10.1038/nrg2843.
Fan, Zhi-Xing, and Jian Yang. 2015. The role of microRNAs in regulating myocardial ischemia reperfusion injury. Saudi Medical Journal 36 (7): 787–793. https://doi.org/10.15537/smj.2015.7.11089.
Xiao, X., Z. Lu, V. Lin, A. May, D.H. Shaw, Z. Wang, B. Che, K. Tran, H. Du, and P.X. Shaw. 2018. MicroRNA miR-24-3p reduces apoptosis and regulates Keap1-Nrf2 pathway in mouse cardiomyocytes responding to ischemia/reperfusion injury. Oxidative Medicine and Cellular Longevity 2018: 7042105–7042109. https://doi.org/10.1155/2018/7042105.
Huang, Z.Q., W. Xu, J.L. Wu, X. Lu, and X.M. Chen. 2019. MicroRNA-374a protects against myocardial ischemia-reperfusion injury in mice by targeting the MAPK6 pathway. Life Sciences 232: 116619. https://doi.org/10.1016/j.lfs.2019.116619.
Zhai, C., Q. Qian, G. Tang, B. Han, H. Hu, D. Yin, H. Pan, and S. Zhang. 2017. MicroRNA-206 protects against myocardial ischaemia-reperfusion injury in rats by targeting Gadd45beta. Molecules and Cells 40 (12): 916–924. https://doi.org/10.14348/molcells.2017.0164.
Nie, Xiang, Jiahui Fan, Huaping Li, Zhongwei Yin, Yanru Zhao, Beibei Dai, Nianguo Dong, Chen Chen, and Dao Wen Wang. 2018. miR-217 promotes cardiac hypertrophy and dysfunction by targeting PTEN. Molecular Therapy--Nucleic Acids 12: 254–266. https://doi.org/10.1016/j.omtn.2018.05.013.
Xia, K., Y. Zhang, and D. Sun. 2020. miR217 and miR543 downregulation mitigates inflammatory response and myocardial injury in children with viral myocarditis by regulating the SIRT1/AMPK/NFkappaB signaling pathway. International Journal of Molecular Medicine 45 (2): 634–646. https://doi.org/10.3892/ijmm.2019.4442.
Li, Y., L. Fei, J. Wang, and Q. Niu. 2020. Inhibition of miR-217 protects against myocardial ischemia-reperfusion injury through inactivating NF-kappaB and MAPK pathways. Cardiovascular Engineering and Technology 11: 219–227. https://doi.org/10.1007/s13239-019-00452-z.
Herr, D. J., T. Singh, T. Dhammu, and D. R. Menick. 2020. Regulation of metabolism by mitochondrial enzyme acetylation in cardiac ischemia-reperfusion injury. Biochimica et biophysica acta. Molecular basis of disease 1866 (6):165728. https://doi.org/10.1016/j.bbadis.2020.165728.
Chen, Hui, Ri-Sheng Huang, Xian-Xian Yu, Qiong Ye, Lu-Lu Pan, Guo-Jian Shao, and Jing Pan. 2017. Emodin protects against oxidative stress and apoptosis in HK-2 renal tubular epithelial cells after hypoxia/reoxygenation. Experimental and Therapeutic Medicine 14 (1): 447–452. https://doi.org/10.3892/etm.2017.4473.
Yu, W., M. Xu, T. Zhang, Q. Zhang, and C. Zou. 2019. Mst1 promotes cardiac ischemia-reperfusion injury by inhibiting the ERK-CREB pathway and repressing FUNDC1-mediated mitophagy. The journal of physiological sciences : JPS 69 (1): 113–127. https://doi.org/10.1007/s12576-018-0627-3.
Yang, Y.Y., D.J. Gong, J.J. Zhang, X.H. Liu, and L. Wang. 2019. Diabetes aggravates renal ischemia-reperfusion injury by repressing mitochondrial function and PINK1/Parkin-mediated mitophagy. American Journal of Physiology. Renal Physiology 317 (4): F852–F864. https://doi.org/10.1152/ajprenal.00181.2019.
Livak, K.J., and T.D. Schmittgen. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−delta delta C(T)) method. Methods 25 (4): 402–408. https://doi.org/10.1006/meth.2001.1262.
Lin, Q., S. Li, N. Jiang, X. Shao, M. Zhang, H. Jin, Z. Zhang, J. Shen, Y. Zhou, W. Zhou, L. Gu, R. Lu, and Z. Ni. 2019. PINK1-parkin pathway of mitophagy protects against contrast-induced acute kidney injury via decreasing mitochondrial ROS and NLRP3 inflammasome activation. Redox Biology 26: 101254. https://doi.org/10.1016/j.redox.2019.101254.
Zhou, Hao, Shuyi Wang, Shunying Hu, Yundai Chen, and Jun Ren. 2018. ER-mitochondria microdomains in cardiac ischemia-reperfusion injury: A fresh perspective. Frontiers in Physiology 9: 755. https://doi.org/10.3389/fphys.2018.00755.
Chistiakov, D.A., T.P. Shkurat, A.A. Melnichenko, A.V. Grechko, and A.N. Orekhov. 2018. The role of mitochondrial dysfunction in cardiovascular disease: A brief review. Annals of Medicine 50 (2): 121–127. https://doi.org/10.1080/07853890.2017.1417631.
Prag, Hiran A., Duvaraka Kula-Alwar, Timothy E. Beach, Anja V. Gruszczyk, Nils Burger, and Michael P. Murphy. 2020. Mitochondrial ROS production during ischemia-reperfusion injury. In Oxidative Stress, ed. Helmut Sies, 513–538. Academic Press.
Liu, Z.Y., S.P. Hu, Q.R. Ji, H.B. Yang, D.H. Zhou, and F.F. Wu. 2017. Sevoflurane pretreatment inhibits the myocardial apoptosis caused by hypoxia reoxygenation through AMPK pathway: An experimental study. Asian Pacific Journal of Tropical Medicine 10 (2): 148–151. https://doi.org/10.1016/j.apjtm.2017.01.006.
Yu, H., Q. Guan, L. Guo, H. Zhang, X. Pang, Y. Cheng, X. Zhang, and Y. Sun. 2016. Gypenosides alleviate myocardial ischemia-reperfusion injury via attenuation of oxidative stress and preservation of mitochondrial function in rat heart. Cell Stress & Chaperones 21 (3): 429–437. https://doi.org/10.1007/s12192-016-0669-5.
Bravo-San Pedro, J.M., G. Kroemer, and L. Galluzzi. 2017. Autophagy and mitophagy in cardiovascular disease. Circulation Research 120 (11): 1812–1824. https://doi.org/10.1161/CIRCRESAHA.117.311082.
Geisler, S., K.M. Holmstrom, D. Skujat, F.C. Fiesel, O.C. Rothfuss, P.J. Kahle, and W. Springer. 2010. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nature Cell Biology 12 (2): 119–131. https://doi.org/10.1038/ncb2012.
Lahuerta, M., C. Aguado, P. Sanchez-Martin, P. Sanz, and E. Knecht. 2018. Degradation of altered mitochondria by autophagy is impaired in Lafora disease. The FEBS Journal 285 (11): 2071–2090. https://doi.org/10.1111/febs.14468.
Tong, M., and J. Sadoshima. 2016. Mitochondrial autophagy in cardiomyopathy. Current Opinion in Genetics & Development 38: 8–15. https://doi.org/10.1016/j.gde.2016.02.006.
Chen, S., L. Zhou, Y. Zhang, Y. Leng, X.Y. Pei, H. Lin, R. Jones, R.Z. Orlowski, Y. Dai, and S. Grant. 2014. Targeting SQSTM1/p62 induces cargo loading failure and converts autophagy to apoptosis via NBK/Bik. Molecular and Cellular Biology 34 (18): 3435–3449. https://doi.org/10.1128/MCB.01383-13.
Liu, H., C. Dai, Y. Fan, B. Guo, K. Ren, T. Sun, and W. Wang. 2017. From autophagy to mitophagy: The roles of P62 in neurodegenerative diseases. Journal of Bioenergetics and Biomembranes 49 (5): 413–422. https://doi.org/10.1007/s10863-017-9727-7.
Hibshman, J.D., T.C. Leuthner, C. Shoben, D.F. Mello, D.R. Sherwood, J.N. Meyer, and L.R. Baugh. 2018. Nonselective autophagy reduces mitochondrial content during starvation in Caenorhabditis elegans. American Journal of Physiology. Cell Physiology 315 (6): C781–C792. https://doi.org/10.1152/ajpcell.00109.2018.
Tanaka, Keiji. 2020. The PINK1-Parkin axis: an overview. Neuroscience research. https://doi.org/10.1016/j.neures.2020.01.006.
Favaloro, B., N. Allocati, V. Graziano, C. Di Ilio, and V. De Laurenzi. 2012. Role of apoptosis in disease. Aging (Albany NY) 4 (5): 330–349. https://doi.org/10.18632/aging.100459.
Moon, D.O., S.Y. Park, M.S. Heo, K.C. Kim, C. Park, W.S. Ko, Y.H. Choi, and G.Y. Kim. 2006. Key regulators in bee venom-induced apoptosis are Bcl-2 and caspase-3 in human leukemic U937 cells through downregulation of ERK and Akt. International Immunopharmacology 6 (12): 1796–1807. https://doi.org/10.1016/j.intimp.2006.07.027.
Brunelle, J.K., and A. Letai. 2009. Control of mitochondrial apoptosis by the Bcl-2 family. Journal of Cell Science 122 (Pt 4): 437–441. https://doi.org/10.1242/jcs.031682.
Kuo, W.T., L. Shen, L. Zuo, N. Shashikanth, Ong Mldm, L. Wu, J. Zha, et al. 2019. Inflammation-induced occludin downregulation limits epithelial apoptosis by suppressing caspase-3 expression. Gastroenterology 157 (5): 1323–1337. https://doi.org/10.1053/j.gastro.2019.07.058.
de Almagro, M.C., and D. Vucic. 2012. The inhibitor of apoptosis (IAP) proteins are critical regulators of signaling pathways and targets for anti-cancer therapy. Experimental Oncology 34 (3): 200–211.
Yang, C.L., X.L. Zheng, K. Ye, Y.N. Sun, Y.F. Lu, H. Ge, and H. Liu. 2019. Effects of microRNA-217 on proliferation, apoptosis, and autophagy of hepatocytes in rat models of CCL4-induced liver injury by targeting NAT2. Journal of Cellular Physiology 234 (4): 3410–3424. https://doi.org/10.1002/jcp.26748.
Saini, A., N. Al-Shanti, A.P. Sharples, and C.E. Stewart. 2012. Sirtuin 1 regulates skeletal myoblast survival and enhances differentiation in the presence of resveratrol. Experimental Physiology 97 (3): 400–418. https://doi.org/10.1113/expphysiol.2011.061028.
Wang, L., N. Quan, W. Sun, X. Chen, C. Cates, T. Rousselle, X. Zhou, X. Zhao, and J. Li. 2018. Cardiomyocyte-specific deletion of Sirt1 gene sensitizes myocardium to ischaemia and reperfusion injury. Cardiovascular Research 114 (6): 805–821. https://doi.org/10.1093/cvr/cvy033.
Luo, G., Z. Jian, Y. Zhu, Y. Zhu, B. Chen, R. Ma, F. Tang, and Y. Xiao. 2019. Sirt1 promotes autophagy and inhibits apoptosis to protect cardiomyocytes from hypoxic stress. International Journal of Molecular Medicine 43 (5): 2033–2043. https://doi.org/10.3892/ijmm.2019.4125.
Wu, B., J.Y. Feng, L.M. Yu, Y.C. Wang, Y.Q. Chen, Y. Wei, J.S. Han, X. Feng, Y. Zhang, S.Y. di, Z.Q. Ma, C.X. Fan, and X.Q. Ha. 2018. Icariin protects cardiomyocytes against ischaemia/reperfusion injury by attenuating sirtuin 1-dependent mitochondrial oxidative damage. British Journal of Pharmacology 175 (21): 4137–4153. https://doi.org/10.1111/bph.14457.
Rao, G., W. Zhang, and S. Song. 2019. MicroRNA217 inhibition relieves cerebral ischemia/reperfusion injury by targeting SIRT1. Molecular Medicine Reports 20 (2): 1221–1229. https://doi.org/10.3892/mmr.2019.10317.
Funding
This work was financially supported by the Science and Technology Foundation of Tianjin Health Bureau (Grant Number 2015ky31).
Author information
Authors and Affiliations
Contributions
YQ designed the research study. YQ and PL performed the research. PL and ZW analyzed the data. YQ wrote the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Qi, Y., Zhang, K., Li, P. et al. Down–regulating miR-217-5p Protects Cardiomyocytes against Ischemia/Reperfusion Injury by Restoring Mitochondrial Function via Targeting SIRT1. Inflammation 44, 383–396 (2021). https://doi.org/10.1007/s10753-020-01343-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10753-020-01343-5