Abstract
The human heart has a markedly low regenerative capacity, leaving patients who suffered from cardiac insults vulnerable to heart failure. The inability to regenerate lost myocardium is accompanied by extensive remodeling that leads to further deterioration in cardiac functions and structure. Although adult mammals seem to lack the ability to regenerate, some lower vertebrates have a cardio-regenerative potential. Emerging studies revealed that mammals do have the ability to undergo endogenous cardiac regeneration during development and shortly after birth. Later, it was proven that the source of the new cardiomyocytes is the proliferation of the pre-existing cardiomyocyte pool. Research is currently focused on finding suitable methods to restore this lost potential in adulthood and enhancing the proliferative capacity of cardiomyocytes. Long non-coding RNAs (lncRNAs) are critical functionally diverse epigenetic regulators capable of either activating or repressing gene expression. LncRNAs have been previously implicated in cardiac development, lineage commitment, and aging. Recent reports suggest that lncRNAs are capable of inducing endogenous cardiac regeneration through manipulating gene expression in cardiomyocytes. This review gives a concise overview of endogenous cardiac regeneration. It further summarizes and critically appraises the current literature on the roles of lncRNAs in endogenous cardiac regeneration and the challenges that face the field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
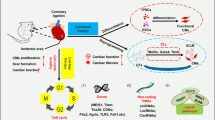
Coronary artery disease is the most common cardiovascular condition which often leads to myocardial infarction (MI), an ischemic shock that hits cardiomyocytes because of low blood supply, leaving nothing but necrosis and a poorly functioning myocardium [1]. Despite the extensive effort to fight acute MI, WHO estimates that its incidence will double by 2050 because of the increased cardiovascular risk factors including obesity and diabetes. MI causes significant disadvantageous remodeling of cardiac tissue and progresses later to heart failure (HF), which affects the lives of millions of people around the world and is a major cause of morbidity and mortality (WHO, June 2016). The current treatment regimens available for HF revolve around palliative therapy and heart transplants for end-stage HF [2]. Therefore, finding new treatments for HF is an urgent necessity. Recently, heart regeneration became an attractive therapeutic option for HF. Regenerating lost cardiomyocytes is expected to improve cardiac function and the quality of life of HF patients; however, extensive knowledge of the mechanisms and cellular pathways of cardiomyocytes is needed in order to pursue this hope. There are two possible ways of achieving cardiac regeneration, the first being supplying the myocardium exogenously with stem cells or cardiomyocytes originally reprogrammed from non-myocytes (cell therapy) and the second being the induction of endogenous regeneration from the proliferation of the pre-existing cardiomyocyte pool. However, the discouraging results of human cell therapy trials, the increasing evidence of the inability of stem cells to trans-differentiate into cardiomyocytes, and the concerns regarding the integrity of the data in the cardiac progenitor cells reports has made studying endogenous cardiac regeneration a priority in the field [3,4,5,6,7]. Several mechanisms regulating endogenous regeneration have been outlined; however, their therapeutic utilization is still a work in progress. Recently, long non-coding RNAs (lncRNAs), a class of RNA molecules that lack the ability to be translated into protein, have been shown to play an immense role in cardiac regeneration through their ability to push cardiomyocytes back into the cell cycle and enabling them to proliferate and divide. This review aims to discuss briefly the history of endogenous cardiac regeneration. It further discusses the roles of lncRNAs in promoting endogenous cardiac regeneration, their therapeutic implication, and the challenges facing the field.
Endogenous cardiac regeneration
It was previously assumed that cardiac regeneration is restricted to lower vertebrates and that the adult mammalian heart possesses no regenerative capacity. Recent studies harnessed data obtained from Carbon-14 dating generated by the nuclear bomb testing during the Cold War to identify the rate of turnover of human cardiomyocytes [8]. This study demonstrated that the turnover rate of cardiomyocytes is approximately 1% per year; however, this rate drops to 0.45% around the age of 75 [8]. Further studies indicated that the newly formed cardiomyocytes develop from the pre-existing pool of cardiomyocytes; however, this rate of regeneration is not sufficient to repair the myocardium after ischemic insults. Previous studies showed that the rate of DNA synthesis decreases and the cell cycle of cardiomyocyte arrests 5 to 7 days after birth [9, 10]. These findings suggested that cardiomyocytes might be able to proliferate in this narrow window of time. In 2011, Porrello et al. demonstrated that the hearts of neonatal mice were able to completely regenerate within 3 weeks after resecting 15% of the left ventricular apex at postnatal day 1 (P1); however, regeneration did not occur when the resection was done at postnatal day 7 (P7), suggesting that this window of regeneration is limited [11]. Further Fate mapping experiments using Cre recombinase technology confirmed that the newly formed myocardium originated from pre-existing cardiomyocytes. Although initially, one research group was not capable of reproducing the same results perhaps because of protocol and technique differences, further studies conducted by several other groups confirmed the authenticity of Porrello et al.’s findings [12]. Similar regenerative potential was also found after ligating the left anterior descending artery (LAD) in a neonatal mouse model of myocardial infarction.
These findings started a race between multiple research groups to identify the possible molecular mechanisms behind cardiac regeneration and to find methods that can restore this ability in adulthood. Interestingly, studies demonstrated that there is a marked increase in reactive oxygen species (ROS) in cardiomyocytes after birth. This increase in mitochondrial ROS promotes damage to various cell molecules including DNA which further results in cell-cycle arrest and cellular senescence. Furthermore, Puente et al. showed that 8-oxo-7,8-dihydroguanine (8-oxoG), an oxidized base, was present at P7 and not P1. This was also accompanied by an increase in the DNA damage response pathway as indicated by the increase of Ser1981 phosphorylated-Ataxia Telangiectasia Mutated (p-ATM) kinase, confirming that oxidative stress and DNA damage play a significant role in the loss of the regenerative capacity of mice hearts post-birth [13]. Additionally, Nakada et al. showed that adult mice are capable of regenerating injured myocardium when they were kept in a hypoxic environment (7% oxygen), suggesting that hypoxic cardiomyocytes are capable of cell-cycle re-entry [14]. These results raised questions regarding the existence of a rare hypoxic cardiomyocyte population that is responsible for the consistent turnover rate in the adult heart. Through pulse-chase experiments, studies identified the epicardium and subepicardium as a hypoxic niche (because of the low capillary and blood content) in which hypoxic cardiac progenitor cells/cardiomyocytes, expressing HIF-alpha (hypoxia-inducible factor), reside [15].
Signaling pathways such as the Hippo cascade have also been linked to cardiac regeneration. The pathway is characterized by a series of transductions that start by MST1/2 directly phosphorylating SALV and MOB1 and end up by the prevention of YAP and TAZ translocation into the nucleus and interact with several transcription factors. Studies have shown that mice cardiomyocytes lacking components of the Hippo pathway are capable of regenerating after insult, suggesting that this pathway reduces the regenerative ability of the heart [16]. Additionally, the enhancement of cardiac regenerative abilities upon Hippo inhibition can be heavily attributed to the upregulation of cell-cycle genes by YAP [17].
Recently, the role of the Neuregulin 1 (NRG1) pathway was examined and linked to neonatal mouse heart regeneration as well [18, 19]. The authors found that NRG1, as well as the expression of ERBB2, an NRG1 coreceptor, was significantly reduced at P7. Interestingly, upon the activation of ERBB2 signaling in mice hearts at different age stages, an increase in the number of cardiomyocytes undergoing mitosis and proliferation was found [19]. Other inflammatory pathways, as well as transcription factors such as Meis1, have been implicated in the cardiac regeneration process. Detailed information about the mechanisms behind cardiac regeneration is extensively reviewed in the following articles [20, 21].
Are we chasing a mirage?
There is no doubt that animal studies have confirmed the neonatal myocardium’s ability to regenerate. But the question remains: does this apply to humans as well? It is hard to detect the human neonate’s heart ability to regenerate because of ethical considerations. Although direct experimental evidence is challenging to acquire from human infants, indirect observations from naturally occurring pathological conditions can be made. Anomalous left coronary artery from the pulmonary artery is a rare condition that affects infants resulting in decreased left ventricular function and myocardial scarring. The management of this condition is corrective surgery early in life. Fratz et al. reported long-term outcomes of several patients; four patients did not have any evidence of myocardial scarring during follow-up while the rest showed minimal scarring and normal LV function [22]. Functional recovery was also reported in other cardiovascular conditions. Neonatal myocardial infarction present at birth is a rare pathology caused by coronary artery occlusion. Haubner et al. reported a case of a newborn who suffered severe cardiac damage confirmed by serum markers for cardiomyocyte cell death, electrocardiograms, and echocardiography. After adequate therapeutic management, follow-up showed that the newborn’s heart restored normal functionality without any structural abnormalities; however, the authors could not exclude the possibility that hibernation of myocardium might have played a partial role in the heart’s functional recovery [23]. Several other similar studies reported neither structural nor severe functional abnormalities after incidents of neonatal myocardial infarction [24,25,26,27,28]. These reports support the notion of infantile cardiac plasticity and robust repair abilities that might be an indicator of the heart’s ability to regenerate. To further assess this ability, Mollova et al. directly examined postmortem human heart tissues obtained from donors of different ages. Strikingly, their study revealed the existence of cardiomyocytes positive for proliferation markers that persist long after birth. Positive cytokinesis markers were also present; however, these markers were not found after the age of 20 [29]. These interesting findings are suggestive of the presence of a regenerative window in humans but leave an important question unanswered. How can this regenerative potential be provoked and enhanced later in life?
An overview of the “junk”
Years ago, the concept of genes coding for mRNAs that are later translated into functional proteins was well established; however, the role of the rest of the non-coding DNA was unknown. It was thought that these non-coding pieces were more or less some evolutionary “junk” without an actual function [30]. The development of high-throughput sequencing and the completion of genome projects such as the ENCODE initiative have given us valuable insight into the human genome. They revealed that only 2% of the genome codes for proteins and that the non-coding parts of the genome are in fact transcribed into non-coding RNAs (ncRNAs) [31]. These RNA molecules were proven later to be major regulators of the protein-coding genes. ncRNAs are generally classified based on their length into short ncRNAs (less than 200 nucleotides) such as microRNAs (miRNAs) and long non-coding RNAs (lncRNAs) (more than 200 nucleotides). miRNAs are the most studied well-characterized class of ncRNAs. They are capable of regulating genes through binding to their mRNAs and hindering their translation or degrading them [32]. miRNAs play important roles in cardiac development, differentiation, and diseases. Moreover, studies have shown that miRNAs can induce direct endogenous regeneration through manipulating cell-cycle genes and other signaling pathways [33].
On the other hand, only a small portion of lncRNAs are well characterized such as the X inactivation-transcript (Xist) [34]; the functions of the rest are yet to be identified. LncRNAs are typically classified according to their genomic location into intergenic, intronic, bidirectional, enhancer, sense, and antisense lncRNAs (the biogenesis, regulation, and further characteristics of lncRNAs are thoroughly discussed in the following reviews [35, 36]). Unlike coding genes and miRNAs, lncRNAs are less conserved among species; however, this lack of conservation does not imply a lack of function but rather an evolutionary role associated with the complexity of the species [37]. In contrast to miRNAs who exert their function mainly through degrading or blocking the translation of mRNAs, the mechanisms of lncRNAs are much more diverse enabling them to both activate and repress genes (Fig. 1). Only a minority of lncRNAs are found in the cytoplasm while the rest are mainly localized in the nucleus [38], implying their ability to interact with DNA and directly regulate gene transcription. LncRNAs are capable of manipulating gene splicing, chromatin remodeling, and DNA methylation. They can also act as transcriptional signals, decoys, guides, and protein scaffolds. The fact that many lncRNAs appear to be cardiac specific or in some cases cardiac enriched makes studying their implication in heart disease and regeneration very tempting [39,40,41]. Recently, genome-wide association studies revealed an association between the loci of lncRNAs and the risk of cardiac conditions, confirming the biological relevance of these diverse RNA molecules in the human heart [42,43,44].
Mechanisms of action of long non-coding RNAs. LncRNAs are diverse molecules that are capable of manipulating gene expression through their cytoplasmic and/or nuclear action. Their actions can be classified into three main categories based on their level of regulation. They can regulate genes post-transcriptionally, epigenetically, or regulate factors responsible for gene transcription
LncRNAs in endogenous cardiac regeneration (an overview)
Although the function of the majority of lncRNAs remains uncharacterized, several transcripts have been shown to play critical roles in the heart. LncRNAs are linked to cardiac development, differentiation, and lineage commitment, proposing their use in reprogramming stem cells and mature differentiated cells (e.g., fibroblasts) into cardiomyocytes and introducing a new research avenue in cell therapy. Several other lncRNAs are also implicated in cardiovascular diseases including arrhythmias, myocardial infarction, and ventricular hypertrophy. The roles of lncRNAs in cardiovascular aging and disease have been extensively reviewed elsewhere [35, 45,46,47]. Instead, here, the focus is aimed towards a subset of the lncRNA studies that, to my knowledge, have only emerged recently and directly highlights the role of lncRNA-mediated regulation in endogenous cardiac regeneration defined by the ability of the pre-existing cardiomyocytes in the myocardium to undergo cell division and proliferation, restoring normal cardiac performance.
Although studies have profiled lncRNA expression at times of distress and managed to identify novel cardiac-specific lncRNAs and explore possible regenerative pathways, they do not provide concrete evidence of the exact role of lncRNAs in cardiac regeneration and need further confirmatory investigations. Better conclusions can be drawn from assessing lncRNA transcripts at times when cardiac regeneration is known to occur. An elegant example of this is the recent report by Chen et al. [48]. Using next-generation sequencing, they analyzed the expression of lncRNAs in the cardiac tissue of C57BL/6J mouse at P1 and P7, managing to identify 685 differentially expressed lncRNAs. Further, they constructed a CNC network between coding and non-coding genes which shed light on some highly correlated targets between dysregulated lncRNAs such as Igfbp3, Trnp1 Itgb6, and Pim3. Interestingly, KEGG pathway enrichment analysis revealed an association between differentially expressed lncRNAs and several signaling and metabolic pathways that have been linked to cardiac regeneration such as the Hippo pathway and Wnt signaling [48]. These findings not only reveal a strong association between lncRNAs and the ability of the heart to regenerate but also suggest a lncRNA-mediated regulation of several mechanisms that have been previously implicated in cardiac regeneration. As indicated earlier, conducting experiments on human hearts is ethically challenging; however, Wang et al. managed to conduct a microarray-based screening to identify differentially expressed lncRNAs between human fetal hearts obtained from unwanted terminated pregnancies and adult hearts [49]. As expected, unlike adult hearts, fetal myocardium exhibited a significant number of Ki-67-positive cells. Additionally, the authors identified 638 differentially expressed lncRNAs, indicating that lncRNAs might have a role in the proliferation and possibly regeneration of human cardiomyocytes. Strikingly, their enrichment analysis revealed that the Hippo pathway was among enriched down-regulated genes in the adult hearts, which contradicted the findings discovered in mice and indicated that Hippo inhibition induces regeneration. YAP1 was also found to be downregulated in adult hearts [49].
Distinct lncRNA functions in endogenous cardiac regeneration
The screening for differentially expressed lncRNAs in regenerating tissues has hinted towards a putative role of lncRNAs in promoting cardiomyocyte proliferation and division; however, the “gold standard” evidence for lncRNA functional characterization, however, is the specific in vivo experimental studies that offer insight into the cells’ machinery and identify reliable therapeutic targets (Fig. 2). With this in mind, Li et al. aimed to identify a lncRNA that may contribute to cardiomyocyte proliferation [50]. Previously, they have demonstrated that Silent information regulator factor 2 related enzyme 1 (Sirt1) plays a role in cardiac regeneration and MI; hence, they suspected that its antisense lncRNA may be involved in the process as well. Analysis of Sirt1 antisense lncRNA in mice revealed high expression in embryonic 16.5 hearts, a downregulation at P1, and further reduction at P28, an expression profile that follows closely the time frame of the loss of cardiac regenerative ability in mice. To further assess its role in neonatal hearts, the authors knocked down the expression Sirt1 lncRNA using LNAs. Upon knockdown, a marked decrease in Ki-67-positive cells was observed, suggesting the lncRNA is necessary for cardiomyocyte proliferation and cell cycle. Overexpressing Sirt1 lncRNA through AAV9 vector injection in adult mice increased Ki-67 fluorescence and pH3, a mitosis marker [50]. To further assess the therapeutic potential of Sirt1 lncRNA, the authors induced its expression in an MI model and found reduced fibrotic area, improved left ventricular function, and decreased mortality rate in transfected mice. Mechanistically, Sirt1 lncRNA forms a lncRNA/mRNA complex with Sirt1 mRNA through interacting with its 3′UTR region, promoting the stabilization of the mRNA and increasing the levels of Sirt1 protein to a level that promotes regeneration [50]. Of note, excessive overexpression of Sirt1 is associated with cellular apoptosis and decreased cardiac function, raising questions about the other factors that regulate the functional shift of the Sirt1 axis from a possible regenerative state to a non-proliferative one [50].
Roles of various long non-coding RNAs in promoting cardiac regeneration. LncRNAs are capable of regulating the subcellular environment of cardiomyocytes, enhancing their proliferative capacity and promoting regeneration. LncRNAs sponge miRNAs, releasing mRNAs from the inhibitory threshold of miRNAs (e.g., AZIN2-sv, CAREL, and CRRL). Of note, all three lncRNAs negatively affect regeneration and their loss is what promotes cardiomyocyte proliferation. They may also interact with proteins and mRNAs directly and enhance their expression and stability (e.g., Sirt1 lncRNA and ECRAR)
Similarly, Li et al. characterized another lncRNA that is implicated in cardiac regeneration. Through comparing sequencing data from adult and fetal human hearts, they identified an AZIN2 splice variant (AZIN2-sv) that was differentially expressed between both tissues [51]. Analysis of the lncRNA revealed that it is conserved in rodents. AZIN2-sv expression was increased in mice at P7 and reduced at P1. Forced expression of AZIN2-SV at P1 reduced the number of proliferating cardiomyocytes while silencing its expression at P7 enhanced the proliferative and regenerative capacity of cardiomyocytes. Furthermore, loss of AZIN2-sv preserves cardiac performance and promotes angiogenesis post-MI. Functional analysis revealed that AZIN2-sv exhibits its function through manipulating the PTEN/PI3K/Akt pathway via two distinct mechanisms. First, it acts as a molecular sponge for miR-214, preventing its inhibitory action on PTEN. Second, it directly binds to PTEN protein and stabilizes its level. Therefore, loss of AZIN2-sv plays an important role in Akt pathway activation and promoting regeneration [51].
In another study, Chen et al. identified yet another differentially expressed lncRNA through analyzing human RNA-sequencing data. Cardiomyocyte regeneration-related lncRNA (CRRL) was found to be conserved among different mammals including humans, chimps, gorillas, mice, and rats [52]. Knocking down CRRL attenuates post-MI remodeling and improves cardiac function compared to controls. Silencing CRRL also increases the number of pH3, EdU, and Ki-67-positive cardiomyocytes at P1. Similar effects were observed upon CRRL knockdown at P7, suggesting that CRRL plays a role in cardiomyocyte proliferation and regeneration. Moreover, loss of CRRL did not increase the percentage of ANP-positive cardiomyocytes, indicating that the proliferation observed in CRRL-deficient cardiomyocytes is not accompanied by hypertrophy. Further analysis showed that CRRL harbors putative miR-199a-3p and miR-214 binding sites within its sequence; however, only the binding between miR-199a-3p and CRRL was confirmed by luciferase reporter assays. This binding relieves Hopx from the inhibitory action of miR-199a [52]. Interestingly, downregulation of Hopx does not entirely reproduce the efficacy of miR-199a in inducing cardiomyocyte proliferation which can be explained by the effect of miR-199a on various other targets [52].
Similar to the previously mentioned studies, Chen et al. identified a lncRNA that could be a viable therapeutic target for inducing cardiac regeneration. Endogenous cardiac regeneration-associated regulator (ECRAR) is a conserved fetal lncRNA that regulates postnatal cardiomyocyte proliferation and post-MI recovery without inducing hypertrophy [53]. The authors demonstrated that E2F1 regulates the expression of ECRAR through binding to its promoter. On the other hand, ECRAR directly interacts with ERK1/2, a crucial cell-cycle gene, increasing its phosphorylation and facilitating its translocation into the nucleus. This was accompanied by an increase in the expression of cyclin D1, cyclin E1, and E2F1 proteins. More importantly, this study is the first to directly visualize the effect of lncRNA therapy in the context of cardiac regeneration through time-lapse imaging. Cell division was observed in cardiomyocytes transfected with ECRAR at P7 [53]. Although this evidence is valuable and highlights the potential of lncRNAs in cardiac regeneration, it would be interesting to examine the effect of lncRNA transfection with time-lapse imaging on more senescent cardiomyocytes (e.g., P16 cardiomyocytes).
Another lncRNA, NR_045363, a human ortholog of LOC101927497, was identified in several microarray-based analyses; however, its function has not been explored. Recently, Wang et al. explored experimentally the role of NR_045363 in cardiac regeneration [54]. NR_045363 was found to be overexpressed in embryonic mice and human tissues. Interestingly, the expression of NR_045363 is threefold higher in cardiomyocytes than non-cardiomyocytes, suggesting that NR_045363 can act as a reliable therapeutic target without much off-target effect. Overexpression of NR_045363 in cardiomyocytes enhances cardiac proliferation and performance after MI. Mechanistically, NR_045363 acts as a decoy for miR-216a, elevating the activity of the JAK2–STAT3 pathway [54].
Cai et al. also discovered cardiac regeneration-related lncRNA (CAREL), another lncRNA implicated in cardiac generation. CAREL acts as a decoy for miR-296, relieving the expression of Trp53inp1 and Itm2a from miR-296 inhibitory action; however, the low number of miR-296 binding sites in the sequence CAREL makes the efficiency of the mechanism questionable [55]. Additionally, unlike the other studies, the authors did not identify it from analyzing human sequencing data. Instead, CAREL was recognized through its differential expression between P1 and P7 in mice, which made conservation analysis a necessity for translational purposes. Only a short sequence was found to be conserved in humans. Although this sequence contains the miR-296 binding site, it is only a small part of the long functional fragment of CAREL and is present in multiple parts of the human genome; thus, it cannot indicate a specific non-coding RNA [56]. Further, although an increase in cardiomyocyte size was observed in CAREL transgenic mice, the authors did not assess the possible effects of CAREL on hypertrophy.
Other lncRNAs in cardiac development, cell fate, and regeneration
Although activating an endogenous cardiac regenerative response might be of clinical benefit to patients with HF, other curative strategies are also being explored. For example, reprogramming stem cells into a cardiomyocyte fate and injecting these cells into the failing myocardium or direct in vivo injection of reprogramming factors into the heart to mediate differentiation of non-myocytes into functional cardiomyocytes represent other viable approaches for regenerating the heart. However, these approaches require extensive knowledge of cardiac developmental programs and the factors that regulate the life cycle stages of cardiomyocytes in order to improve the effectiveness and efficiency of these strategies. For example, it has been shown that genomic enhancers play crucial roles in controlling spatial and temporal gene expression during development and embryogenesis. Long non-coding RNAs transcribed from enhancer regions termed “enhancer-associated lncRNA” (elncRNAs) aid in regulating the enhancer’s activity and the expression of their neighboring genes during development and differentiation including cardiac fate determination. In a recent study, Ounzain et al. managed to identify hundreds of elncRNAs expressed during embryonic stem cell transitioning into cardiomyocytes using deep RNA sequencing and ab initio reconstruction [57]. These cardiac-specific transcripts undergo chromatin state transitions throughout cardiac differentiation. The study also revealed an increase in fetal enhancer transcripts during the reactivation of fetal gene program, a distinct event that accompanies stress response in adult hearts [57].
Using loss-of-function strategies, the function of distinct lncRNAs in cardiac commitment and development has also been studied. For example, Braveheart (BVHT), a lncRNA expressed in cardiac mesoderm, is essential for cardiac lineage commitment in mice and embryonic stem cell differentiation. BVHT functions upstream of mesoderm posterior 1 (MesP1) and portrays its role through interacting with SUZ12, a component of polycomb-repressive complex 2 (PRC2), during cardiac differentiation [40]. CARMEN, an enhancer lncRNA, is found upstream of cardiac mesoderm-specifying genes. Like BVHT, CARMEN exerts its function through interacting with components of PRC2, SUZ12, and EZH2 in cis. Although CARMEN is closely related to miR-143 and -145, the authors demonstrated that CARMEN functions independently of these miRNAs. Owing to its regulatory action, CARMEN is crucial for cardiac differentiation and maintaining cardiac identity [58]. Fetal-lethal non-coding developmental regulatory RNA (Fendrr) is another example of lncRNAs controlling cardiac fate [59]. Loss of Fendrr during embryonic life leads to major cardiac wall defects due to decreased cardiomyocyte proliferation. Fendrr interacts with PRC2 and TrxG/MLL complexes, controlling the methylation status of the promoter of lateral plate mesoderm genes either through increasing or decreasing their expression during cardiac development. Additionally, increased expression of two cardiac commitment regulators, Nkx-2.5 and GATA-6, can be noted in Fendrr-mutated mice [59]. These results initiated a parade of experimental studies examining the roles of lncRNAs in cardiac development, differentiation, and commitment leading to the identification of several lncRNAs implicated in these mechanisms as well as the exploration of possible regenerative pathways regulated by lncRNAs (Table 1).
Challenges and concerns
In cancer, lncRNAs are mostly studied in human tissues and cell lines; hence, each identified lncRNA directly represents a therapeutic target and is biologically relevant. Conversely, in cardiovascular research, animal models are necessary to study the role of lncRNAs in cardiac regeneration and other mechanisms; however, their poor conservation affects the translation and interpretation of any findings discovered in animals if a human ortholog cannot be found. The lack of conservation of Braveheart (Bvht) is an example of this [40]. This issue can be bypassed through analyzing human RNA-sequencing data then studying the identified RNAs in animals, making use of the fact that human lncRNAs are more conserved. Although this method was proven successful (from the studies reviewed), it cannot completely replace the need for animal-based lncRNA characterization. Hence, finding new ways to identify human orthologs is due. Instead of completely relying on detecting primary linear sequence conservation, other methods have been proposed based on several observations. First, Ulitsky et al. demonstrated that conserved short sequences within the lncRNA transcripts, and not the entire Transcript sequence, are sufficient for their function to be conserved [72]. Second, it has been proposed that functional conservation can be detected through examining lncRNAs’ secondary and tertiary structures, rather than the primary sequence. However, this approach is harder to apply when analyzing excessively long transcripts [73]. Third, functional orthologs may be found through analyzing persevered genomic localization relative to conserved coding genes (often related to development) as well as the conservation of lncRNA promoter [74]. Lastly, Kirk et al. proposed a functional classification of lncRNAs based on their K-mers content. They managed to identify marked similarity between specific mouse and human lncRNA communities through their analysis [75]. In the future, these discoveries may help the field blast through the translational obstacles of lncRNAs.
It has become evident that the binding of lncRNAs to miRNAs is a mechanism that constitutes a major portion of lncRNAs’ regulatory action in cardiomyocytes. The mechanism commonly referred to as the competing endogenous RNA (ceRNA) effect relies on the existence of short sequences in the lncRNA transcript complementary to miRNAs through which lncRNAs are capable of titrating the expression of miRNAs and relieving mRNAs from the inhibitory threshold of miRNAs [76]. The biological relevance of this mechanism has been previously questioned with some studies nullifying its effectiveness to induce observable regulatory effect under normal physiological conditions [77]. Nevertheless, experimental lncRNA studies, in the context of cancer particularly, continue to emerge and confirm the relevance of the mechanism in contributing to disease progression. However, several factors such as the abundance of miRNAs and lncRNAs, number of binding sites on the lncRNA, the spacing between these binding sites, and the target abundance control the efficacy of the mechanism [78]; thus, all these factors need to be characterized before relying on this mechanism solely to explain the action of a lncRNA. This concern can be noted in the cases of CAREL and CRRL as they both have a low number of miRNA binding sites, which makes the efficacy of the mechanism questionable and begs for a stoichiometric analysis to be conducted. And although identifying a miRNA interaction is a straightforward way to explain a lncRNA function, the other potential mechanisms should not be left unattended. AZIN2-sv’s ability to interact directly with PTEN protein on top of binding to miR-214 is an elegant example of this. Conversely, other lncRNA–miRNA interactions should not be ignored. To further elaborate, cytoplasmic localization of lncRNA hints toward a lncRNA-sponging action; however, recent studies showed that both cytoplasmic and nuclear lncRNAs are capable of manipulating miRNAs through regulating their promoters, maturation and miRNA precursor production [79, 80]. Additionally, none of the reports I reviewed examined the effect of miRNA binding on the expression of the lncRNA itself as it has been reported that miRNAs may degrade lncRNAs in an AGO2-dependent manner.
There is no doubt that single-cell RNA-seq provides a new paradigm for studying organ development and cell characteristics. Delaughter el al. used spatial–temporal cardiac transcriptomic analysis to provide insight into cardiac development from E9.5 to P21 [81]. They managed to identify a sub-population of cardiomyocytes with proliferative capacity, which expressed cell cycle and division-associated genes. Interestingly, this sub-population represented 60% of cardiomyocytes at E9.5 and E14.5 and only 20% at P0 and P7. At P21, this sub-population vanished [81]. In another study, See et al. analyzed single nuclear transcriptomes of failing and healthy cardiomyocytes [82]. They found marked transcriptomic heterogeneity between different cardiomyocyte sub-populations. Moreover, they identified a group of novel lnRNAs that were not detected in bulk tissue, highlighting the importance of individual cell analysis. The authors attributed the inability to detect the lncRNAs to the large number of cytoplasmic mRNA that might have diluted their expression. Additionally, they demonstrated that lncRNAs regulate dedifferentiation, cell-cycle genes, and cardiomyocytes’ regenerative potential [82]. Furthermore, it has been noted that functional analysis of lncRNAs in single cell setting produces more accurate results and may prevent the imprecise findings of bulk tissue analysis. Together, these findings suggest that relying on single cell analysis in different developmental stages and after various stimuli may help us discover previously unannotated lncRNAs as well as enhance our understanding of the association between distinct lncRNAs and cardiac regeneration.
Overall, better experimental methodology is still needed (Fig. 3). After birth, cardiomyocytes shift from proliferative phase into a hypertrophic phase where they lose the ability to undergo cytokinesis and become binucleated [83]. A hallmark of a successful stimulation of cardiac regeneration is the detection of cell division and successful cytokines. Most studies rely on Aurora B as a marker of cytokinesis. Recently, Hesse et al. demonstrated that Aurora B stains both binucleated and dividing cells, confirming that Aurora B lacks sufficient specificity to distinguish between binucleating and proliferating cardiomyocytes [84]. Instead, they proposed the use of midbody positioning and the distance between the two daughter nuclei as predictors of successful cell division [84]. Careful interpretation of Ki-67, PCNA (proliferating cell nuclear antigen), and pH3 markers in the context of cardiomyocyte division and proliferation is warranted as well [84]. Direct visualization of cell division by time-lapse imaging can also be conducted to confirm the robustness of experimental conclusions. Additionally, cells should be closely monitored even after the occurrence of cell division. In their experiment, Mohamed et al. noticed that although cardiomyocytes divided upon overexpressing CDK1, CCNB, and AURKB within 48–72 h, this division was shortly followed by cell death because of marked DNA damage [85]. Without full characterization of lncRNAs’ mechanistic pathways, it is possible that the promising results of lncRNA transfection might be accompanied by other catastrophic unknown confounders. Furthermore, although costly, lncRNA experiments and other cardiac regeneration-related studies, in general, should be upsized to larger mammals and tissues that closely resemble humans and not just mice. Recently, two independent research groups demonstrated that porcine hearts are capable of initiating a regenerative response after cardiac insult shortly after birth; however, this regenerative window seems to be relatively shorter than that observed in mice [86, 87]. On the other hand, another group managed to develop a human fetal cardiac organoid model through harnessing the advancements in the fields of tissue engineering and stem cell differentiation, providing a new research tool for studying cardiac regeneration [88].
Although cardiac regeneration is a relatively unexplored field with lots of knowledge to unravel, one must think about its future therapeutic potential. LncRNAs are diverse molecules capable of inducing cardiac regeneration; however, targeting them remains a hurdle. Several methods have been proposed to knock down their expression in the preclinical reports such as shRNAs, gapmeRs, and siRNAs. Of them, gapmeRs seem to be most suitable for therapeutic utilization due to their ability to target lncRNAs in the nucleus [89]. Additionally, ASOs have been shown to be clinically feasible as several ASO-based drugs have been recently approved [90]. However, several factors should be taken into consideration when considering such drugs such as tissue specificity of the drug, pharmacological profile, their stability, and off-targets. When considering overexpressing lncRNAs, the therapeutic options are even more limited to viruses and viral-associated vectors. Despite the current effort to improve these methods of delivery, their immunogenicity and specificity remain a major concern. With that being said, several clinical trials have used viral-based delivery methods without reporting significant side effects [91,92,93]. Crispr/cas9 system may also be used to activate the expression of lncRNAs through regulating their promoter or knock their expression out through targeting their genomic locus [94]. Delivery of non-coding RNAs through exosomes seems promising as well [95]. However, the most appropriate therapeutic vehicle for targeting lncRNAs is yet to be identified and further research is required.
Conclusion and future directions
Cardiac regeneration is a new exciting field that holds the promise for healing failing hearts. Until this date, several molecular mechanisms and pathways implicated in cardiac regeneration have been identified. LncRNAs have the ability to induce a regenerative response in cardiomyocytes; however, several aspects of the field remain uncharted. Although cardiac regeneration originates mainly from cardiomyocytes, it is not a one-man show. Several other mechanisms play crucial roles in determining cardiomyocytes’ proliferative response and promoting regeneration. For example, neovascularization has been shown to improve the regenerative capacity after injury [96]. Additionally, Mahmoud et al. observed impaired regeneration in neonatal mice upon interrupting the left vagus nerve mechanically, suggesting a role for nerves in promoting cardiac regeneration [97]. The immune system was also proposed as an important regulator of the process. For example, Aurora et al. found differences between immune responses in 1-day-old mice and 14-day-old mice. Further, they demonstrated that macrophages are key players in the regeneration process of neonatal mice; however, they concluded that macrophages associated with regeneration are transcriptionally and functionally different from adult macrophages [98]. Recently, Boeckel et al. found that lncRNA Heat2 is upregulated in patients with heart failure and heavily regulates immune cells responses [99]. Other studies have shown that lncRNAs regulate angiogenesis and endothelial cell functions [100, 101]. Therefore, exploring the interplay between lncRNA-mediated regulation and these mechanisms may improve our understanding of cardiac regeneration.
ElncRNAs transcribed from genomic enhancer regions have been implicated in cardiac development; however, their role in cardiovascular medicine is yet to be fully understood. Because of their cis and trans regulatory actions, elncRNAs may have a role in regulating endogenous cardiac regeneration gene networks [102]. Other non-coding RNAs such as circular RNAs have been implicated in cardiovascular diseases; however, their possible functions have not been studied in the context of heart regeneration [103]. Finally, the evolving genome editing tools such as Crisp/cas9 system can be harnessed to manipulate lncRNA expression and screen for regeneration-relevant lncRNAs, providing a new paradigm for lncRNA research.
Although at the moment several obstacles may interfere with the translational potential of lncRNAs such as their poor conservation and the lack of appropriate therapeutic tools, investigating their functions and roles in cardiomyocytes should be pursued to further enhance our understanding of the epigenetic control laid upon the genome. Characterizing their functions may also act as a gate for revealing other targetable mechanisms. For example, a lncRNA that acts mainly through sponging a certain miRNA provides insight into the relevance and function of that particular miRNA, which will most probably be conserved and targetable. The same applies to genes and signaling cascades regulated by lncRNAs.
In conclusion, several challenges face lncRNA research in endogenous cardiac regeneration; however, overcoming these challenges seems to be an attainable goal. Further, lots of lncRNAs and their associated mechanisms are yet to be identified; however, studying this “dark matter” of the genome is definitely worthy as the loss- and gain-of-function studies have clearly demonstrated their potential in inducing cardiomyocyte regeneration, declaring lncRNAs a new hope for healing failing hearts.
References
Anderson JL, Morrow DA (2017) Acute myocardial infarction. N Engl J Med 376(21):2053–2064. https://doi.org/10.1056/NEJMra1606915
Inamdar AA, Inamdar AC (2016) Heart failure: diagnosis, management and utilization. J Clin Med 5(7). https://doi.org/10.3390/jcm5070062
Banerjee MN, Bolli R, Hare JM (2018) Clinical studies of cell therapy in cardiovascular medicine: recent developments and future directions. Circ Res 123(2):266–287. https://doi.org/10.1161/CIRCRESAHA.118.311217
Eschenhagen T, Bolli R, Braun T, Field LJ, Fleischmann BK, Frisen J, Giacca M, Hare JM, Houser S, Lee RT, Marban E, Martin JF, Molkentin JD, Murry CE, Riley PR, Ruiz-Lozano P, Sadek HA, Sussman MA, Hill JA (2017) Cardiomyocyte regeneration: a consensus statement. Circulation 136(7):680–686. https://doi.org/10.1161/CIRCULATIONAHA.117.029343
The Lancet E (2014) Expression of concern: the SCIPIO trial. Lancet 383(9925):1279. https://doi.org/10.1016/S0140-6736(14)60608-5
Li Y, He L, Huang X, Bhaloo SI, Zhao H, Zhang S, Pu W, Tian X, Li Y, Liu Q, Yu W, Zhang L, Liu X, Liu K, Tang J, Zhang H, Cai D, Ralf AH, Xu Q, Lui KO, Zhou B (2018) Genetic lineage tracing of nonmyocyte population by dual recombinases. Circulation 138(8):793–805. https://doi.org/10.1161/CIRCULATIONAHA.118.034250
Menasche P (2018) Cell therapy trials for heart regeneration—lessons learned and future directions. Nat Rev Cardiol 15(11):659–671. https://doi.org/10.1038/s41569-018-0013-0
Bergmann O, Bhardwaj RD, Bernard S, Zdunek S, Barnabe-Heider F, Walsh S, Zupicich J, Alkass K, Buchholz BA, Druid H, Jovinge S, Frisen J (2009) Evidence for cardiomyocyte renewal in humans. Science 324(5923):98–102. https://doi.org/10.1126/science.1164680
Soonpaa MH, Field LJ (1997) Assessment of cardiomyocyte DNA synthesis in normal and injured adult mouse hearts. Am J Phys 272(1 Pt 2):H220–H226. https://doi.org/10.1152/ajpheart.1997.272.1.H220
Soonpaa MH, Field LJ (1998) Survey of studies examining mammalian cardiomyocyte DNA synthesis. Circ Res 83(1):15–26
Porrello ER, Mahmoud AI, Simpson E, Hill JA, Richardson JA, Olson EN, Sadek HA (2011) Transient regenerative potential of the neonatal mouse heart. Science 331(6020):1078–1080. https://doi.org/10.1126/science.1200708
Lam N, Sadek H (2018) Neonatal heart regeneration: comprehensive literature review, vol 138. doi:https://doi.org/10.1161/CIRCULATIONAHA.118.033648
Puente BN, Kimura W, Muralidhar SA, Moon J, Amatruda JF, Phelps KL, Grinsfelder D, Rothermel BA, Chen R, Garcia JA, Santos CX, Thet S, Mori E, Kinter MT, Rindler PM, Zacchigna S, Mukherjee S, Chen DJ, Mahmoud AI, Giacca M, Rabinovitch PS, Aroumougame A, Shah AM, Szweda LI, Sadek HA (2014) The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 157(3):565–579. https://doi.org/10.1016/j.cell.2014.03.032
Nakada Y, Canseco DC, Thet S, Abdisalaam S, Asaithamby A, Santos CX, Shah AM, Zhang H, Faber JE, Kinter MT, Szweda LI, Xing C, Hu Z, Deberardinis RJ, Schiattarella G, Hill JA, Oz O, Lu Z, Zhang CC, Kimura W, Sadek HA (2017) Hypoxia induces heart regeneration in adult mice. Nature 541(7636):222–227. https://doi.org/10.1038/nature20173
Kimura W, Sadek HA (2012) The cardiac hypoxic niche: emerging role of hypoxic microenvironment in cardiac progenitors. Cardiovasc Diagn Ther 2(4):278–289. https://doi.org/10.3978/j.issn.2223-3652.2012.12.02
Heallen T, Morikawa Y, Leach J, Tao G, Willerson JT, Johnson RL, Martin JF (2013) Hippo signaling impedes adult heart regeneration. Development 140(23):4683–4690. https://doi.org/10.1242/dev.102798
Lin Z, von Gise A, Zhou P, Gu F, Ma Q, Jiang J, Yau AL, Buck JN, Gouin KA, van Gorp PR, Zhou B, Chen J, Seidman JG, Wang DZ, Pu WT (2014) Cardiac-specific YAP activation improves cardiac function and survival in an experimental murine MI model. Circ Res 115(3):354–363. https://doi.org/10.1161/CIRCRESAHA.115.303632
Bersell K, Arab S, Haring B, Kuhn B (2009) Neuregulin1/ErbB4 signaling induces cardiomyocyte proliferation and repair of heart injury. Cell 138(2):257–270. https://doi.org/10.1016/j.cell.2009.04.060
D'Uva G, Aharonov A, Lauriola M, Kain D, Yahalom-Ronen Y, Carvalho S, Weisinger K, Bassat E, Rajchman D, Yifa O, Lysenko M, Konfino T, Hegesh J, Brenner O, Neeman M, Yarden Y, Leor J, Sarig R, Harvey RP, Tzahor E (2015) ERBB2 triggers mammalian heart regeneration by promoting cardiomyocyte dedifferentiation and proliferation. Nat Cell Biol 17(5):627–638. https://doi.org/10.1038/ncb3149
Porrello ER, Olson EN (2014) A neonatal blueprint for cardiac regeneration. Stem Cell Res 13(3 Pt B):556–570. https://doi.org/10.1016/j.scr.2014.06.003
Uygur A, Lee RT (2016) Mechanisms of cardiac regeneration. Dev Cell 36(4):362–374. https://doi.org/10.1016/j.devcel.2016.01.018
Fratz S, Hager A, Schreiber C, Schwaiger M, Hess J, Stern HC (2011) Long-term myocardial scarring after operation for anomalous left coronary artery from the pulmonary artery. Ann Thorac Surg 92(5):1761–1765. https://doi.org/10.1016/j.athoracsur.2011.06.021
Haubner BJ, Schneider J, Schweigmann U, Schuetz T, Dichtl W, Velik-Salchner C, Stein JI, Penninger JM (2016) Functional recovery of a human neonatal heart after severe myocardial infarction. Circ Res 118(2):216–221. https://doi.org/10.1161/CIRCRESAHA.115.307017
Boulton J, Henry R, Roddick LG, Rogers D, Thompson L, Warner G (1991) Survival after neonatal myocardial infarction. Pediatrics 88(1):145–150
Cesna S, Eicken A, Juenger H, Hess J (2013) Successful treatment of a newborn with acute myocardial infarction on the first day of life. Pediatr Cardiol 34(8):1868–1870. https://doi.org/10.1007/s00246-012-0417-2
Deutsch MA, Cleuziou J, Noebauer C, Eicken A, Vogt M, Hoerer J, Lange R, Schreiber C (2014) Successful management of neonatal myocardial infarction with ECMO and intracoronary r-tPA lysis. Congenit Heart Dis 9(5):E169–E174. https://doi.org/10.1111/chd.12117
Farooqi KM, Sutton N, Weinstein S, Menegus M, Spindola-Franco H, Pass RH (2012) Neonatal myocardial infarction: case report and review of the literature. Congenit Heart Dis 7(6):E97–E102. https://doi.org/10.1111/j.1747-0803.2012.00660.x
Saker DM, Walsh-Sukys M, Spector M, Zahka KG (1997) Cardiac recovery and survival after neonatal myocardial infarction. Pediatr Cardiol 18(2):139–142. https://doi.org/10.1007/s002469900133
Mollova M, Bersell K, Walsh S, Savla J, Das LT, Park SY, Silberstein LE, Dos Remedios CG, Graham D, Colan S, Kuhn B (2013) Cardiomyocyte proliferation contributes to heart growth in young humans. Proc Natl Acad Sci U S A 110(4):1446–1451. https://doi.org/10.1073/pnas.1214608110
Ohno S (1972) So much "junk" DNA in our genome. Brookhaven Symp Biol 23:366–370
Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, Hilton JA, Jain K, Baymuradov UK, Narayanan AK, Onate KC, Graham K, Miyasato SR, Dreszer TR, Strattan JS, Jolanki O, Tanaka FY, Cherry JM (2018) The encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res 46(D1):D794–D801. https://doi.org/10.1093/nar/gkx1081
O'Brien J, Hayder H, Zayed Y, Peng C (2018) Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol (Lausanne) 9:402. https://doi.org/10.3389/fendo.2018.00402
Gnecchi M, Pisano F, Bariani R (2015) microRNA and cardiac regeneration. Adv Exp Med Biol 887:119–141. https://doi.org/10.1007/978-3-319-22380-3_7
Sahakyan A, Yang Y, Plath K (2018) The role of Xist in X-chromosome dosage compensation. Trends Cell Biol 28(12):999–1013. https://doi.org/10.1016/j.tcb.2018.05.005
Devaux Y, Zangrando J, Schroen B, Creemers EE, Pedrazzini T, Chang CP, Dorn GW, 2nd, Thum T, Heymans S, Cardiolinc n (2015) Long noncoding RNAs in cardiac development and ageing. Nat Rev Cardiol 12 (7):415–425. doi:https://doi.org/10.1038/nrcardio.2015.55
Quinn JJ, Chang HY (2016) Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet 17(1):47–62. https://doi.org/10.1038/nrg.2015.10
Pang KC, Frith MC, Mattick JS (2006) Rapid evolution of noncoding RNAs: lack of conservation does not mean lack of function. Trends Genet 22(1):1–5. https://doi.org/10.1016/j.tig.2005.10.003
Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, Bell I, Cheung E, Drenkow J, Dumais E, Patel S, Helt G, Ganesh M, Ghosh S, Piccolboni A, Sementchenko V, Tammana H, Gingeras TR (2007) RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316(5830):1484–1488. https://doi.org/10.1126/science.1138341
Han P, Li W, Lin CH, Yang J, Shang C, Nuernberg ST, Jin KK, Xu W, Lin CY, Lin CJ, Xiong Y, Chien H, Zhou B, Ashley E, Bernstein D, Chen PS, Chen HV, Quertermous T, Chang CP (2014) A long noncoding RNA protects the heart from pathological hypertrophy. Nature 514(7520):102–106. https://doi.org/10.1038/nature13596
Klattenhoff CA, Scheuermann JC, Surface LE, Bradley RK, Fields PA, Steinhauser ML, Ding H, Butty VL, Torrey L, Haas S, Abo R, Tabebordbar M, Lee RT, Burge CB, Boyer LA (2013) Braveheart, a long noncoding RNA required for cardiovascular lineage commitment. Cell 152(3):570–583. https://doi.org/10.1016/j.cell.2013.01.003
Ounzain S, Micheletti R, Beckmann T, Schroen B, Alexanian M, Pezzuto I, Crippa S, Nemir M, Sarre A, Johnson R, Dauvillier J, Burdet F, Ibberson M, Guigo R, Xenarios I, Heymans S, Pedrazzini T (2015) Genome-wide profiling of the cardiac transcriptome after myocardial infarction identifies novel heart-specific long non-coding RNAs. Eur Heart J 36(6):353–368a. https://doi.org/10.1093/eurheartj/ehu180
Tragante V, Barnes MR, Ganesh SK, Lanktree MB, Guo W, Franceschini N, Smith EN, Johnson T, Holmes MV, Padmanabhan S, Karczewski KJ, Almoguera B, Barnard J, Baumert J, Chang YP, Elbers CC, Farrall M, Fischer ME, Gaunt TR, Gho JM, Gieger C, Goel A, Gong Y, Isaacs A, Kleber ME, Mateo Leach I, McDonough CW, Meijs MF, Melander O, Nelson CP, Nolte IM, Pankratz N, Price TS, Shaffer J, Shah S, Tomaszewski M, van der Most PJ, Van Iperen EP, Vonk JM, Witkowska K, Wong CO, Zhang L, Beitelshees AL, Berenson GS, Bhatt DL, Brown M, Burt A, Cooper-DeHoff RM, Connell JM, Cruickshanks KJ, Curtis SP, Davey-Smith G, Delles C, Gansevoort RT, Guo X, Haiqing S, Hastie CE, Hofker MH, Hovingh GK, Kim DS, Kirkland SA, Klein BE, Klein R, Li YR, Maiwald S, Newton-Cheh C, O'Brien ET, Onland-Moret NC, Palmas W, Parsa A, Penninx BW, Pettinger M, Vasan RS, Ranchalis JE, MR P, Rose LM, Sever P, Shimbo D, Steele L, Stolk RP, Thorand B, Trip MD, van Duijn CM, Verschuren WM, Wijmenga C, Wyatt S, Young JH, Zwinderman AH, Bezzina CR, Boerwinkle E, Casas JP, Caulfield MJ, Chakravarti A, Chasman DI, Davidson KW, Doevendans PA, Dominiczak AF, FitzGerald GA, Gums JG, Fornage M, Hakonarson H, Halder I, Hillege HL, Illig T, Jarvik GP, Johnson JA, Kastelein JJ, Koenig W, Kumari M, Marz W, Murray SS, O'Connell JR, Oldehinkel AJ, Pankow JS, Rader DJ, Redline S, Reilly MP, Schadt EE, Kottke-Marchant K, Snieder H, Snyder M, Stanton AV, Tobin MD, Uitterlinden AG, van der Harst P, van der Schouw YT, Samani NJ, Watkins H, Johnson AD, Reiner AP, Zhu X, de Bakker PI, Levy D, Asselbergs FW, Munroe PB, Keating BJ (2014) Gene-centric meta-analysis in 87,736 individuals of European ancestry identifies multiple blood-pressure-related loci. Am J Hum Genet 94(3):349–360. https://doi.org/10.1016/j.ajhg.2013.12.016
Helgadottir A, Thorleifsson G, Manolescu A, Gretarsdottir S, Blondal T, Jonasdottir A, Jonasdottir A, Sigurdsson A, Baker A, Palsson A, Masson G, Gudbjartsson DF, Magnusson KP, Andersen K, Levey AI, Backman VM, Matthiasdottir S, Jonsdottir T, Palsson S, Einarsdottir H, Gunnarsdottir S, Gylfason A, Vaccarino V, Hooper WC, Reilly MP, Granger CB, Austin H, Rader DJ, Shah SH, Quyyumi AA, Gulcher JR, Thorgeirsson G, Thorsteinsdottir U, Kong A, Stefansson K (2007) A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 316(5830):1491–1493. https://doi.org/10.1126/science.1142842
Ishii N, Ozaki K, Sato H, Mizuno H, Saito S, Takahashi A, Miyamoto Y, Ikegawa S, Kamatani N, Hori M, Saito S, Nakamura Y, Tanaka T (2006) Identification of a novel non-coding RNA, MIAT, that confers risk of myocardial infarction. J Hum Genet 51(12):1087–1099. https://doi.org/10.1007/s10038-006-0070-9
Frank S, Aguirre A, Hescheler J, Kurian L (2016) A lncRNA perspective into (re)building the heart. Front Cell Dev Biol 4:128. https://doi.org/10.3389/fcell.2016.00128
Ounzain S, Crippa S, Pedrazzini T (2013) Small and long non-coding RNAs in cardiac homeostasis and regeneration. Biochim Biophys Acta 1833(4):923–933. https://doi.org/10.1016/j.bbamcr.2012.08.010
Uchida S, Dimmeler S (2015) Long noncoding RNAs in cardiovascular diseases. Circ Res 116(4):737–750. https://doi.org/10.1161/CIRCRESAHA.116.302521
Chen YM, Li H, Fan Y, Zhang QJ, Li X, Wu LJ, Chen ZJ, Zhu C, Qian LM (2017) Identification of differentially expressed lncRNAs involved in transient regeneration of the neonatal C57BL/6J mouse heart by next-generation high-throughput RNA sequencing. Oncotarget 8(17):28052–28062. https://doi.org/10.18632/oncotarget.15887
Wang J, Geng Z, Weng J, Shen L, Li M, Cai X, Sun C, Chu M (2016) Microarray analysis reveals a potential role of lncRNAs expression in cardiac cell proliferation. BMC Dev Biol 16(1):41. https://doi.org/10.1186/s12861-016-0139-4
Li B, Hu Y, Li X, Jin G, Chen X, Chen G, Yanmei C, Huang S, Liao W, Liao Y, Teng Z, Bin J (2018) Sirt1 antisense long noncoding RNA promotes cardiomyocyte proliferation by enhancing the stability of Sirt1, vol 7. doi:https://doi.org/10.1161/JAHA.118.009700
Li X, He X, Wang H, Li M, Huang S, Chen G, Jing Y, Wang S, Chen Y, Liao W, Liao Y, Bin J (2018) Loss of AZIN2 splice variant facilitates endogenous cardiac regeneration. Cardiovasc Res 114(12):1642–1655. https://doi.org/10.1093/cvr/cvy075
Chen G, Li H, Li X, Li B, Zhong L, Huang S, Zheng H, Li M, Jin G, Liao W, Liao Y, Chen Y, Bin J (2018) Loss of long non-coding RNA CRRL promotes cardiomyocyte regeneration and improves cardiac repair by functioning as a competing endogenous RNA. J Mol Cell Cardiol 122:152–164. https://doi.org/10.1016/j.yjmcc.2018.08.013
Yanmei C, Li X, Li B, Wang H, Li M, Huang S, Sun Y, Chen G, Si X, Huang C, Liao W, Liao Y, Bin J (2018) Long non-coding RNA ECRAR triggers postnatal myocardial regeneration by activating ERK1/2 signaling. https://doi.org/10.1016/j.ymthe.2018.10.021
Wang J, Chen X, Shen D, Ge D, Chen J, Pei J, Li Y, Yue Z, Feng J, Chu M, Nie Y (2018) A long noncoding RNA NR_045363 controls cardiomyocyte proliferation and cardiac repair. J Mol Cell Cardiol 127:105–114. https://doi.org/10.1016/j.yjmcc.2018.12.005
Cai B, Ma W, Ding F, Zhang L, Huang Q, Wang X, Hua B, Xu J, Li J, Bi C, Guo S, Yang F, Han Z, Li Y, Yan G, Yu Y, Bao Z, Yu M, Li F, Tian Y, Pan Z, Yang B (2018) The long noncoding RNA CAREL controls cardiac regeneration. J Am Coll Cardiol 72(5):534–550. https://doi.org/10.1016/j.jacc.2018.04.085
Thum T (2018) Translational opportunities and challenges of long noncoding RNAs in cardiac regeneration. J Am Coll Cardiol 72(5):551–552. https://doi.org/10.1016/j.jacc.2018.05.039
Ounzain S, Pezzuto I, Micheletti R, Burdet F, Sheta R, Nemir M, Gonzales C, Sarre A, Alexanian M, Blow MJ, May D, Johnson R, Dauvillier J, Pennacchio LA, Pedrazzini T (2014) Functional importance of cardiac enhancer-associated noncoding RNAs in heart development and disease. J Mol Cell Cardiol 76:55–70. https://doi.org/10.1016/j.yjmcc.2014.08.009
Ounzain S, Micheletti R, Arnan C, Plaisance I, Cecchi D, Schroen B, Reverter F, Alexanian M, Gonzales C, Ng SY, Bussotti G, Pezzuto I, Notredame C, Heymans S, Guigo R, Johnson R, Pedrazzini T (2015) CARMEN, a human super enhancer-associated long noncoding RNA controlling cardiac specification, differentiation and homeostasis. J Mol Cell Cardiol 89 (Pt A:98–112. https://doi.org/10.1016/j.yjmcc.2015.09.016
Grote P, Wittler L, Hendrix D, Koch F, Wahrisch S, Beisaw A, Macura K, Blass G, Kellis M, Werber M, Herrmann BG (2013) The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell 24(2):206–214. https://doi.org/10.1016/j.devcel.2012.12.012
Plaisance I, Perruchoud S, Fernandez-Tenorio M, Gonzales C, Ounzain S, Ruchat P, Nemir M, Niggli E, Pedrazzini T (2016) Cardiomyocyte lineage specification in adult human cardiac precursor cells via modulation of enhancer-associated long noncoding RNA expression. JACC Basic Transl Sci 1(6):472–493. https://doi.org/10.1016/j.jacbts.2016.06.008
Kurian L, Aguirre A, Sancho-Martinez I, Benner C, Hishida T, Nguyen TB, Reddy P, Nivet E, Krause MN, Nelles DA, Esteban CR, Campistol JM, Yeo GW, Belmonte JCI (2015) Identification of novel long noncoding RNAs underlying vertebrate cardiovascular development. Circulation 131(14):1278–1290. https://doi.org/10.1161/CIRCULATIONAHA.114.013303
Yin A, Feng M, Cheng Z, Zhang Q, Li H, Xu J, Zhang H, Li Y, Qian L (2018) Altered DNA methylation of long noncoding RNA uc.167 inhibits cell differentiation in heart development. Biomed Res Int 2018:4658024. https://doi.org/10.1155/2018/4658024
Gore-Panter SR, Hsu J, Barnard J, Moravec CS, Van Wagoner DR, Chung MK, Smith JD (2016) PANCR, the PITX2 adjacent noncoding RNA, is expressed in human left atria and regulates PITX2c expression. Circ Arrhythm Electrophysiol 9(1):e003197. https://doi.org/10.1161/CIRCEP.115.003197
Matkovich SJ, Edwards JR, Grossenheider TC, de Guzman Strong C, Dorn GW 2nd (2014) Epigenetic coordination of embryonic heart transcription by dynamically regulated long noncoding RNAs. Proc Natl Acad Sci U S A 111(33):12264–12269. https://doi.org/10.1073/pnas.1410622111
Korostowski L, Sedlak N, Engel N (2012) The Kcnq1ot1 long non-coding RNA affects chromatin conformation and expression of Kcnq1, but does not regulate its imprinting in the developing heart. PLoS Genet 8(9):e1002956. https://doi.org/10.1371/journal.pgen.1002956
Han Y, Xu H, Cheng J, Zhang Y, Gao C, Fan T, Peng B, Li B, Liu L, Cheng Z (2016) Downregulation of long non-coding RNA H19 promotes P19CL6 cells proliferation and inhibits apoptosis during late-stage cardiac differentiation via miR-19b-modulated Sox6. Cell Biosci 6:58. https://doi.org/10.1186/s13578-016-0123-5
Zhu JG, Shen YH, Liu HL, Liu M, Shen YQ, Kong XQ, Song GX, Qian LM (2014) Long noncoding RNAs expression profile of the developing mouse heart. J Cell Biochem 115(5):910–918. https://doi.org/10.1002/jcb.24733
Li H, Jiang L, Yu Z, Han S, Liu X, Li M, Zhu C, Qiao L, Huang L (2017) The role of a novel long noncoding RNA TUC40- in cardiomyocyte induction and maturation in P19 cells. Am J Med Sci 354(6):608–616. https://doi.org/10.1016/j.amjms.2017.08.019
Wu R, Xue P, Wan Y, Wang S, Gu M (2018) LncRNA-uc.40 silence promotes P19 embryonic cells differentiation to cardiomyocyte via the PBX1 gene. In Vitro Cell Dev Biol Anim 54(8):600–609. https://doi.org/10.1007/s11626-018-0284-0
Zhang Q, Feng M, Zhang H, Xu J, Zhang L, Wang X, Cheng Z, Qian L (2018) Long noncoding RNA uc.4 inhibits cell differentiation in heart development by altering DNA methylation. J Cell Biochem. https://doi.org/10.1002/jcb.28084
Cheng Z, Zhang Q, Yin A, Feng M, Li H, Liu H, Li Y, Qian L (2018) The long non-coding RNA uc.4 influences cell differentiation through the TGF-beta signaling pathway. Exp Mol Med 50(2):e447. https://doi.org/10.1038/emm.2017.278
Ulitsky I, Shkumatava A, Jan CH, Sive H, Bartel DP (2011) Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147(7):1537–1550. https://doi.org/10.1016/j.cell.2011.11.055
Guo X, Gao L, Wang Y, Chiu DK, Wang T, Deng Y (2016) Advances in long noncoding RNAs: identification, structure prediction and function annotation. Brief Funct Genomics 15(1):38–46. https://doi.org/10.1093/bfgp/elv022
Amaral PP, Leonardi T, Han N, Vire E, Gascoigne DK, Arias-Carrasco R, Buscher M, Pandolfini L, Zhang A, Pluchino S, Maracaja-Coutinho V, Nakaya HI, Hemberg M, Shiekhattar R, Enright AJ, Kouzarides T (2018) Genomic positional conservation identifies topological anchor point RNAs linked to developmental loci. Genome Biol 19(1):32. https://doi.org/10.1186/s13059-018-1405-5
Kirk JM, Kim SO, Inoue K, Smola MJ, Lee DM, Schertzer MD, Wooten JS, Baker AR, Sprague D, Collins DW, Horning CR, Wang S, Chen Q, Weeks KM, Mucha PJ, Calabrese JM (2018) Functional classification of long non-coding RNAs by k-mer content. Nat Genet 50(10):1474–1482. https://doi.org/10.1038/s41588-018-0207-8
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) A ceRNA hypothesis: the Rosetta stone of a hidden RNA language? Cell 146(3):353–358. https://doi.org/10.1016/j.cell.2011.07.014
Thomson DW, Dinger ME (2016) Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet 17(5):272–283. https://doi.org/10.1038/nrg.2016.20
Denzler R, McGeary SE, Title AC, Agarwal V, Bartel DP, Stoffel M (2016) Impact of microRNA levels, target-site complementarity, and cooperativity on competing endogenous RNA-regulated gene expression. Mol Cell 64(3):565–579. https://doi.org/10.1016/j.molcel.2016.09.027
Cai X, Cullen BR (2007) The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 13(3):313–316. https://doi.org/10.1261/rna.351707
Wang Y, Hu Y, Wu G, Yang Y, Tang Y, Zhang W, Wang K, Liu Y, Wang X, Li T (2017) Long noncoding RNA PCAT-14 induces proliferation and invasion by hepatocellular carcinoma cells by inducing methylation of miR-372. Oncotarget 8(21):34429–34441. https://doi.org/10.18632/oncotarget.16260
DeLaughter DM, Bick AG, Wakimoto H, McKean D, Gorham JM, Kathiriya IS, Hinson JT, Homsy J, Gray J, Pu W, Bruneau BG, Seidman JG, Seidman CE (2016) Single-cell resolution of temporal gene expression during heart development. Dev Cell 39(4):480–490. https://doi.org/10.1016/j.devcel.2016.10.001
See K, Tan WLW, Lim EH, Tiang Z, Lee LT, Li PYQ, Luu TDA, Ackers-Johnson M, Foo RS (2017) Single cardiomyocyte nuclear transcriptomes reveal a lincRNA-regulated de-differentiation and cell cycle stress-response in vivo. Nat Commun 8(1):225. https://doi.org/10.1038/s41467-017-00319-8
Paradis AN, Gay MS, Zhang L (2014) Binucleation of cardiomyocytes: the transition from a proliferative to a terminally differentiated state. Drug Discov Today 19(5):602–609. https://doi.org/10.1016/j.drudis.2013.10.019
Hesse M, Doengi M, Becker A, Kimura K, Voeltz N, Stein V, Fleischmann BK (2018) Midbody positioning and distance between daughter nuclei enable unequivocal identification of cardiomyocyte cell division in mice. Circ Res 123(9):1039–1052. https://doi.org/10.1161/CIRCRESAHA.118.312792
Mohamed TMA, Ang YS, Radzinsky E, Zhou P, Huang Y, Elfenbein A, Foley A, Magnitsky S, Srivastava D (2018) Regulation of cell cycle to stimulate adult cardiomyocyte proliferation and cardiac regeneration. Cell 173(1):104–116 e112. https://doi.org/10.1016/j.cell.2018.02.014
Ye L, D'Agostino G, Loo SJ, Wang CX, Su LP, Tan SH, Tee GZ, Pua CJ, Pena EM, Cheng RB, Chen WC, Abdurrachim D, Lalic J, Tan RS, Lee TH, Zhang J, Cook SA (2018) Early regenerative capacity in the porcine heart. Circulation 138(24):2798–2808. https://doi.org/10.1161/CIRCULATIONAHA.117.031542
Zhu W, Zhang E, Zhao M, Chong Z, Fan C, Tang Y, Hunter JD, Borovjagin AV, Walcott GP, Chen JY, Qin G, Zhang J (2018) Regenerative potential of neonatal porcine hearts. Circulation 138(24):2809–2816. https://doi.org/10.1161/CIRCULATIONAHA.118.034886
Voges HK, Mills RJ, Elliott DA, Parton RG, Porrello ER, Hudson JE (2017) Development of a human cardiac organoid injury model reveals innate regenerative potential. Development 144(6):1118–1127. https://doi.org/10.1242/dev.143966
Lennox KA, Behlke MA (2016) Cellular localization of long non-coding RNAs affects silencing by RNAi more than by antisense oligonucleotides. Nucleic Acids Res 44(2):863–877. https://doi.org/10.1093/nar/gkv1206
Stein CA, Castanotto D (2017) FDA-approved oligonucleotide therapies in 2017. Mol Ther 25(5):1069–1075. https://doi.org/10.1016/j.ymthe.2017.03.023
Collins M, Thrasher A (2015) Gene therapy: progress and predictions. Proc Biol Sci 282(1821):20143003. https://doi.org/10.1098/rspb.2014.3003
Nathwani AC, Reiss UM, Tuddenham EG, Rosales C, Chowdary P, McIntosh J, Della Peruta M, Lheriteau E, Patel N, Raj D, Riddell A, Pie J, Rangarajan S, Bevan D, Recht M, Shen YM, Halka KG, Basner-Tschakarjan E, Mingozzi F, High KA, Allay J, Kay MA, Ng CY, Zhou J, Cancio M, Morton CL, Gray JT, Srivastava D, Nienhuis AW, Davidoff AM (2014) Long-term safety and efficacy of factor IX gene therapy in hemophilia B. N Engl J Med 371(21):1994–2004. https://doi.org/10.1056/NEJMoa1407309
Nathwani AC, Tuddenham EG, Rangarajan S, Rosales C, McIntosh J, Linch DC, Chowdary P, Riddell A, Pie AJ, Harrington C, O'Beirne J, Smith K, Pasi J, Glader B, Rustagi P, Ng CY, Kay MA, Zhou J, Spence Y, Morton CL, Allay J, Coleman J, Sleep S, Cunningham JM, Srivastava D, Basner-Tschakarjan E, Mingozzi F, High KA, Gray JT, Reiss UM, Nienhuis AW, Davidoff AM (2011) Adenovirus-associated virus vector-mediated gene transfer in hemophilia B. N Engl J Med 365(25):2357–2365. https://doi.org/10.1056/NEJMoa1108046
Haemmig S, Feinberg MW (2017) Targeting LncRNAs in cardiovascular disease: options and expeditions. Circ Res 120(4):620–623. https://doi.org/10.1161/CIRCRESAHA.116.310152
Ha D, Yang N, Nadithe V (2016) Exosomes as therapeutic drug carriers and delivery vehicles across biological membranes: current perspectives and future challenges. Acta Pharm Sin B 6(4):287–296. https://doi.org/10.1016/j.apsb.2016.02.001
Zangi L, Lui KO, von Gise A, Ma Q, Ebina W, Ptaszek LM, Spater D, Xu H, Tabebordbar M, Gorbatov R, Sena B, Nahrendorf M, Briscoe DM, Li RA, Wagers AJ, Rossi DJ, Pu WT, Chien KR (2013) Modified mRNA directs the fate of heart progenitor cells and induces vascular regeneration after myocardial infarction. Nat Biotechnol 31(10):898–907. https://doi.org/10.1038/nbt.2682
Mahmoud AI, O'Meara CC, Gemberling M, Zhao L, Bryant DM, Zheng R, Gannon JB, Cai L, Choi WY, Egnaczyk GF, Burns CE, Burns CG, MacRae CA, Poss KD, Lee RT (2015) Nerves regulate cardiomyocyte proliferation and heart regeneration. Dev Cell 34(4):387–399. https://doi.org/10.1016/j.devcel.2015.06.017
Aurora AB, Porrello ER, Tan W, Mahmoud AI, Hill JA, Bassel-Duby R, Sadek HA, Olson EN (2014) Macrophages are required for neonatal heart regeneration. J Clin Invest 124(3):1382–1392. https://doi.org/10.1172/JCI72181
Boeckel JN, Perret MF, Glaser SF, Seeger T, Heumuller AW, Chen W, John D, Kokot KE, Katus HA, Haas J, Lackner MK, Kayvanpour E, Grabe N, Dieterich C, von Haehling S, Ebner N, Hunecke S, Leuschner F, Fichtlscherer S, Meder B, Zeiher AM, Dimmeler S, Keller T (2018) Identification and regulation of the long non-coding RNA Heat2 in heart failure. J Mol Cell Cardiol 126:13–22. https://doi.org/10.1016/j.yjmcc.2018.11.004
Weirick T, Militello G, Uchida S (2018) Long non-coding RNAs in endothelial biology. Front Physiol 9:522. https://doi.org/10.3389/fphys.2018.00522
Hou J, Wang L, Wu Q, Zheng G, Long H, Wu H, Zhou C, Guo T, Zhong T, Wang L, Chen X, Wang T (2018) Long noncoding RNA H19 upregulates vascular endothelial growth factor a to enhance mesenchymal stem cells survival and angiogenic capacity by inhibiting miR-199a-5p. Stem Cell Res Ther 9(1):109. https://doi.org/10.1186/s13287-018-0861-x
Ounzain S, Pedrazzini T (2015) The promise of enhancer-associated long noncoding RNAs in cardiac regeneration. Trends Cardiovasc Med 25(7):592–602. https://doi.org/10.1016/j.tcm.2015.01.014
Gomes CPC, Salgado-Somoza A, Creemers EE, Dieterich C, Lustrek M, Devaux Y, Cardiolinc n (2018) Circular RNAs in the cardiovascular system. Noncoding RNA Res 3 (1):1–11. doi:https://doi.org/10.1016/j.ncrna.2018.02.002
Acknowledgments
I would like to express my deepest appreciation to Abdelaziz AI for his guidance and continuous support.
Funding
None.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None to declare.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Afify, A.R.Y. The long non-coding road to endogenous cardiac regeneration. Heart Fail Rev 24, 587–600 (2019). https://doi.org/10.1007/s10741-019-09782-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10741-019-09782-5