Abstract
Hypertrophic scar (HS) is a serious skin fibrotic disease characterized by the excessive proliferation of fibroblasts and often considered as a kind of benign skin tumor. microRNA-155 (miR-155) is usually served as a promising marker in antitumor therapy. In view of the similarities of hypertrophic scar and tumor, it is predicted that miR-155 may be a novel therapeutic target in clinical trials. Here we found the expression levels of miR-155 was gradually down regulated and HIF-1α was upregulated in HS tissue and HS derived fibroblasts (HFs). And cell proliferation was inhibited when miR-155 was overexpressed or HIF-1α was silenced. Moreover, overexpression of miR-155 in HFs could reduce the expression of collagens in vitro and inhibit the collagen fibers arrangement in vivo, whereas miR-155 knockdown gave opposite results. Furthermore, we found that miR-155 directly targeted the HIF-1α, which could also independently inhibit the expression of collagens in vitro and obviously improved the appearance and architecture of the rabbit ear scar in vivo when it was silencing. Finally, we found that PI3K/AKT pathway was enrolled in these processes. Together, our results indicated that miR-155 was a critical regulator in the formation and development of hypertrophic scar and might be a potential molecular target for hypertrophic scar therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hypertrophic scar (HS) is a serious skin fibrotic disease characterized by the excessive proliferation of fibroblasts and deposition of extracellular matrix proteins, mainly including collagen I (Col I), collagen III (Col III) (Armour et al. 2007). About 40–70% of surgical damages and over 91% of burn injuries result in HS and present a major and rising health problem to society (Sideek et al. 2016). HS is usually considered as a benign skin tumor and microRNAs (miRNAs) are hot research targets for antitumor therapy (Huffaker et al. 2012; Zonari et al. 2013). Given that, our study aimed to determine whether microRNAs could exert significant roles in HS therapy.
miRNAs are a class of highly-conserved small RNAs, which usually bind to the 3′-untranslated region of target mRNAs and are capable of regulating post-transcriptional gene by blocking translation or degradation of target mRNAs (Lim et al. 2005; Ruvkun 2001). In general, miRNAs have great effects on the formation and development of many disease, usually serving as therapeutic targets (Fattore et al. 2017; Lin et al. 2016; Mu et al. 2017). Furthermore, recent studies have discovered that miRNAs have close relationships with the formation of HS and might be the key molecules in fibrosis pathogenesis (Li et al. 2017; Wang et al. 2017; Zhou et al. 2017; Zhu et al. 2017). Among the reported miRNAs, microRNA-155 (miR-155) is a well-known oncogenic miRNA and correlates with the pathological degrees of lung fibrosis and liver fibrosis (Christmann et al. 2016; Csak et al. 2015). Moreover, emerging evidences have suggested miR-155−/− mice developed exacerbated lung fibrosis with increasement of Col I and Col III mRNA expression and proteins deposition (Kurowska-Stolarska et al. 2016). However, the concrete mechanisms of miR-155 in HS was reported rarely. Therefore, we explored the roles of miR-155 and elucidated its underlying molecular mechanisms in HS.
Materials and methods
Ethics statement
The protocol for studies using human samples was reviewed and approved by Institutional Ethical Committee of the Fourth Military Medical University (FMMU). Written informed consent was obtained for all patient samples. Animal experiments were conducted in accordance with the Guide for the Care and Use of Laboratory Animals of the FMMU, and all experimental protocols used in this study were approved by the Animal Care Committee of the FMMU.
Tissue samples and cell culture
All HS and normal skin tissues were obtained from 16 patients (Table 1) in Xijing Hospital (Xi’an, China). Before the experiment, all patients were informed about the purpose and procedures of the study and voluntarily agreed to provide tissue. Written consent was obtained from all participants, and all protocols were approved by the Ethics Committee of Xijing Hospital, which is affiliated with the FMMU. All tissues were minced and incubated in a solution of collagenase type I (0.1 mg/ml; Sigma, St. Louis, MO, USA) at 37 °C for 2.5 h to isolate fibroblasts. Fibroblasts were then pelleted and grown in Dulbecco’s Modified Eagle Medium (Gibco,Grand Island, NY, USA) supplemented with 10% fetal calf serum (Gibco), 100 U/ml penicillin, and 100 U/ml streptomycin. Cells were incubated at 37 °C in a 5% (v/v) CO2-humidified atmosphere. All experiments were performed with passage 3–5 cells.
RT-qPCR
Total RNAs from HS and normal skin tissues and isolated fibroblasts were extracted using RNA-isolation kit (Takara, Japan). The purity of RNA obtained was evaluated by the calculated A260/A280 (1.9–2.0). GAPDH and U6 RNA were used as internal loading controls for mRNA and miRNA, 5′-CTTAATGCTAATCGTGATAGGGGT-3′ or 5′-GTGCTCGCTTCGGCAGCACATAT-3′ for miR-155 or U6 RNA, respectively. 5′-GAGGGCAACAGCAGGTTCACTTA-3′ and 5′-TCAGCACCACCGATGTCCA-3′ for Col I, 5′-CCACGGAAACACTGGTGGAC-3′ and 5′-GCCAGCTGCACATCAAGGAC-3′ for Col III, 5′-GACAATGGCTCTGGGCTCTGTAA-3′ and 5′-TGTGCTTCGTCACCCACGTA-3′ for α-SMA, 5′-AACCCTCGAGGCGTTT CCTAATCTCATT-3′ and 5′-AACCGCGGCCGCAAGCTGGAAGGTTTGTG-3′ for HIF-1α, 5′-GCACCGTCAAGCTGAGAAC-3′ and 5′-TGGTGAAGACGCCAGTGGA-3′ for GAPDH. qPCR reactions were run in triplicate and relative expression levels of miRNA or mRNA were analyzed using the Bio-Rad C1000 Thermal Cycler (Bio-Rad, CA,USA).
Western blot analysis
Fibroblasts were washed with ice-cold phosphate-buffered saline (PBS) and lysed using RIPA buffer supplemented with protease and phosphatase inhibitor mixtures (Heart Biological Technology Co. Ltd., Xi’an, China) on ice. Lysates were separated by centrifugation at 4 °C and 14,000×g for 10 min. Protein concentration was determined by BCA assay (Pierce, USA). 50 mg total protein was subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and transferred to PVDF membranes (Millipore, Bedford, MA, USA). After blocking with 5% non-fat milk, the membranes were incubated with anti-alpha-smooth muscle actin (a-SMA) (1:200, Boster, China) anti-Col I (1:1000, Abcam, UK), anti- Col III (1:1000, Abcam, UK), anti-HIF1α (1:1000, Abcam, UK), anti-P -AKT, anti-AKT (1:1000, Cell signaling technology, USA), goat anti-Actin (1:200, Santa cruz biotechnology, USA) overnight at 4 °C, next day the membranes were incubated with horseradish peroxidase conjugated secondary antibodies (1:3000) 37 °C for 1 h. Then immunoreactive proteins were visualised using ECL western blotting detection reagent (Millipore, Billerica, MA) and detected using MultiImage Light Cabinet Filter Positions (Alpha Innotech, San Leandro, CA).
Cell transfection
Chemically modified sense RNA (miR-155 mimics) or antisense RNA (miR-155 inhibitor) and their cognate control RNAs was synthesized by Qiagen. The sequence was 5′-CTTAATGCTAATCGTGATAGGGGT-3′. HFs were transfected with 80 nM HIF-1α siRNA duplexes sequence (sense, 5′CGAGUGCCUUAUCCAAGAATT-3′) or mock using Lipofectamine 2000 Transfection Reagent (Invitrogen, Carlsbad, CA) and miR-155 scrambled negative control, miR-155 mimics or inhibitor using HiPerFect Transfection Reagent (Qiagen). Briefly, 20 µM of miR-155 mimics or inhibitor was mixed with 12 µl HiPerFect in 100 µl serum-free culture medium for 10 min at room temperature to form transfection complexes. The cells were incubated with the transfection complexes for 48 h.
Flow cytometry analysis
Cell cycle distribution was analyzed by flow cytometry (FACSAria; BD Biosciences). HFs were divided into the following groups: miR-155 scrambled negative control, miR-155 mimics or inhibitor transfected groups. Twenty-four hours after transfection, the cells were harvested, rinsed with PBS, fixed with 95% (v/v) icecold ethanol and resuspended in staining buffer containing FITCAnnexin V and propidium iodide (PI). The mixture was then incubated in the dark at room temperature for 15 min. The DNA contents of the stained nuclei were analyzed, and the number of cells in each cycle phase was calculated.
Cell migration assays
In the present study, the HFs transfected with miR-155 scrambled negative control, miR-155 mimics or inhibitor, were grown to confluence in 12-well plates in DMEM containing 10 µg/ml mitomycin C (Invitrogen) for 1 h to completely inhibit cell proliferation. A straight scratch was made on the HFs using a P200 pipette tip. The cells were then washed with PBS three times, and further cultured in DMEM. After incubating for 0 and 24 h, the gap width of scratch re-population was measured and recorded, and then compared with the initial gap size at 0 h. Using the ImageJ image processing program, the size of the denuded area was determined at each time point from the digital images.
CCK-8 assay
The viability of HFs was detected by CCK-8 Cell Proliferation Assay Kit (Qihai, Shanghai, China) according to the manufacturer’s instructions. Fibroblasts were seeded into 96-well plates at a density of 3 × 103 cells per well. The cells were cultured for12, 24, 48, or 72 h after transfection and then followed by incubation with CCK-8 solution for another 4 h. The amount of newly formed formazan dye was quantitated with a scanning multiwell spectrophotometer at 450 nm. The measured absorbance directly correlates with the number of viable cells.
Luciferase reporter assay
The 3′ untranslated region (UTR) fragments HIF-1α containing the miRNA binding sites were amplified by PCR using the cDNA template obtained from RNA sample of HFs. The wide type 3′ UTRs of HIF-1α as well as mutant 3′ UTRs with nucleotide substitutions in the putative binding sites corresponding to the seed sequence of miR-155 was cloned downstream of the firefly luciferase gene in the pGL3 vector (Promega, Madison, WI). Cells were co-transfected with miR-155 or a control miRNA 0.48 h after transfection, cells were rinsed in PBS and their luciferase activity was measured by a luminometer (Promega), using dual luciferase reporter assay system.
Rabbit ear scar model
The rabbit ear scar model was established as described (12, 14). For short, 10 adult New Zealand white male rabbits (2.0–2.5 kg b.w./each), provided by the Fourth Military Medical University Animal Center, were acclimated and housed under the standard 12 h light: 12 h dark cycle with free access of water and SPF basal diet. Rabbit was first anaesthetized with 1% pentobarbital (1.5 mg/kg b.w.), and then, a dermal punch biopsy (10 × 4 mm) was created down to bare cartilage on the ventral surface of each ear to outline a full thickness wound. Each rabbit thus had eight wounds. For each wound, the epidermis, dermis and perichondrium were completely removed. The wounds were then covered with sterile gauze for 1 day. On postoperative day 28 and afterwards, scars were randomly placed into six groups (six scars to each group): control, saline, mir-155 agomir, mir-155 agomir control, siRNA-HIF1α (siHIF1α) and siRNA-Control groups. They were applied to the scars two times in a week.
Masson staining
After ear wounds healed and scars formed, rabbits were sacrificed and scar tissues in situ were excised for Masson staining using Masson Trichrome Staining Kit (Nanjing Biotech, Nanjing, Jiangsu, China). Paraffin-embedded tissue sections from scars were examined for the expression and arrangement of collagen fibers under a FSX100 microscope (OLYMPUS, Shinjuku-ku, Tokyo, Japan), and images were recorded digitally onto a computer and analyzed with Image-Pro Plus 6.0 system.
Statistical analysis
Each experiment was repeated at least three times, and the data are presented as the means ± SD. Statistical differences between groups were analyzed by the Student’s t test or the Mann–Whitney U test as appropriate using SPSS 13.0. A p value < 0.05 was considered to indicate a statistically significant difference.
Results
miR-155 was down regulated in HS and isolated fibroblasts
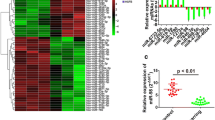
To explore the expression level of miR-155 in HS, we tested the mRNA levels of miR-155 in normal skin (NS), HS and isolated fibroblasts respectively. The results of qRT-PCR analysis revealed that the mRNA levels of miR-155 was significantly decreased by ~ 50% (Fig. 1a, b) in HS and HFs, indicating miR-155 was related with the formation of HS.
Expression of miR-155 in NS and HS and isolated fibroblasts. a, b qRT-PCR results showed the mRNA levels of miR-155 in NS, HS, NFs and HFs. GAPDH served as an internal control. Each data point was normalized against its corresponding GAPDH level. Each bar was a mean ± SD of n = 4. **p < 0.01 compared to control
miR-155 decreased the deposition of collagen in HFs
To investigate the role of miR-155, the protein level of miR-155 was overexpressed or inhibited in HFs by transient transfection of miR-155 mimics or anti-sense oligonucleotides and control oligonucleotides respectively. qRT-PCR detected the efficiency of overexpression or inhibition of miR-155 (Fig. 2a). The major characteristics of fibrosis are excessive abnormal deposition of collagen, mainly including Col I, Col III and a-SMA. Thus, the expression levels of Col I, Col III and α-SMA were detected. The results showed that the expression levels of Col I, Col III were significantly decreased in the group transfected with miR-155 mimics comparing with the cells transfected with control mimics. The experiment of anti-miR-155 gave a reverse results with increasing the protein levels of Col I and Col III comparing with the anti-negative control (NC) oligos. (Fig. 2b–d), while the protein and mRNA levels of α-SMA were relatively unchanged (Fig. 2b, e). These results suggested that miR-155 could decrease the expression of Col I, and Col III and the deposition of collagen. Collectively, it was demonstrated that miR-155 may negatively regulate the expression of these fibrotic makers.
Effect of miR-155 on the steady-state levels of Col I, Col III and α-SMA in HFs in vitro. a HFs were transfected with miR-155 mimics, NC mimics, anti-155, or anti-NC at a final concentration of 100 nM. The miR-155 levels in the transfected cells were examined by qRT-PCR analysis. b At 48 h after transfection, western blot analyses and (c, d, e) qRT-PCR were used to evaluate the expression level of Col I, Col III and α-SMA. All values were normalized to either the negative control. Error bars represented means ± SD of n = 4. **p < 0.01 compared to control
miR-155 inhibited HFs proliferation but does not affect HFs migration
For the significantly down-regulation of miR-155 in HS, it was predicted that miR-155 functioned as a HS suppressor. And HFs excessive proliferation are important reasons leading to HS. Therefore, we transfected miR-155 mimics or anti-miR-155 to determine the role of miR-155 in HFs proliferation. The results of CCK-8 experiment exhibited that transfection of HFs with the miR-155 mimics for 0, 12, 24, 48, or 72 h inhibited the viability of cells. However, the anti-miR-155 group gave the opposite results (Fig. 3a). Flow cytometry analysis showed that miR-155 could significantly affect the percentage of cells in the S phase, indicating that miR-155 arrested the cell cycle at the S phase (Fig. 3b). These findings indicated that miR-155 inhibits proliferation of HFs.
Effect of miR-155 on proliferation, migration of HFs in vitro. a Cell proliferation was measured by the CCK-8 assay. b Flow cytometry analysis showing the effect of miR-155 on cell cycle distribution. Cell numbers at G1, G2 and S phases were counted and the percentage was calculated. c Following 1 h of mitomycin C treatment, cells were scratch wounded with a micropipette tip (200 µl). Black dotted lines indicate the wound borders at the beginning of the assay and recorded at 0 and 24 h post-scratching. Scale bar = 100 µM. Relative scratch gap was calculated as the ratio of the remaining scratch gap at given time point and the original gap at 0 h. d Transwell invasion assay was performed in 8 µm transwells inserted in 24-well dishes to detect the migration ability of miR-155 transfected HFs 24 h after plating. The average number of cells traversed across the inner membrance of a transwell was counted. Scale bar = 50 µm. Error bars represented means ± SD of n = 4. *p < 0.05; **p < 0.01 compared to control
To further assess the influence of miR-155 on HFs, we explored its effects on cell migration, another key factor in HS progression and formation. Subsequently, scratch wound healing assays were performed in order to evaluate the effects of miR-155 on HFs migration. The cells were pre-treated with mitomycin C for 1 h to inhibit proliferation prior to the scratch assay. The results demonstrated that the mobility of HFs did not change obviously (Fig. 3c). Furtherly, transwell invasion assay was executed and gave the same results (Fig. 3d). Taken together, these results suggested that miR-155 inhibited HFs proliferation while exerting no marked effect on HFs migration.
HIF1α was a direct target of miR-155 in HFs
The protein expression of HIF1α in NS, HS and isolated fibroblasts were examined (Fig. 4a). The results showed that the protein levels of HIF-1 α in HS and HFs were all higher than that in NS and normal skin derived fibroblasts (NFs), which indicated that HS was a relatively hypoxic tissue. Then, NFs were cultured in 1% oxygen for 0 and 24 h. We found culturing cells in hypoxia significantly increased the expression of HIF-1 α (Fig. 4b). These results suggested that human HS was located in a hypoxia environment.
HIF1α was a direct target of miR-155. a Western blot anlaysis for the protein expression of HIF-1 α in NS, HS, NFs and HFs. b The protein expression of HIF-1 α in dermal fibroblasts cultured under hypoxia conditions (1% O2) for 0 and 24 h. c Alignment of HIF1α 3′-UTR with miR-155 sequences. Bioinformatical analysis was performed to identify the interaction of miR-155 with HIF1α. d HFs were grown and transiently transfected with wide-type or mutated HIF1α 3′-UTR luciferase reporter construct and then with miR-155 mimics for 48 h and then subjected to luciferase assay. e, f Western blot and qRT-PCR analyses of HIF1α expression in HFs that were transfected with the miR-155 mimics, NC mimics, anti-miR-155, or anti-NC. Error bars represented means ± SD of n = 4. **p < 0.01 compared to control
MicroRNAs exert their biological functions mainly through regulating their target genes. To identify the target gene of miR-155 in HFs, the publicly available databases were used and the results discovered that HIF1α owned a potential miR-155 target site in its 3′ untranslated region (UTR). And the complementarity between the target sites and the miR-155 seed region was completely matched (Fig. 4c). As shown in Fig. 4d, luciferase activity was significantly repressed by the miR-155 mimics in the wild-type (WT) 3′ UTR, but did not suppress that of a reporter fused to a mutant (MUT), providing evidence of the direct binding of miR-155 to the 3′-UTR of HIF1α. Furthermore, it was found that overexpression of miR-155 in HFs significantly decreased HIF1α expression at the protein and mRNA levels, whereas inhibition of miR-155 led to an increased expression of HIF1α (Fig. 4e, f). These results strongly indicated that 3′ UTR of HIF1α carried the direct binding seed of miR-155, and miR-155 might target HIF1α and inhibit its expression.
Silencing of HIF1α inhibited the expression of Col I, Col III in HFs and regulated AKT signaling pathway
Since HIF1α was a potential target of miR-155, it was reasoned that ectopic of HIF1α could change the biological phenotypes caused by miR-155 in HFs.
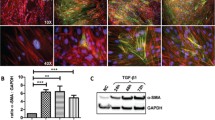
To verify this property, the expression of HIF1α in HFs was silenced by transfection of three different HIF1α specific small interfering RNAs (siRNAs), i.e. HIF1α si-1, HIF1α si-2 and HIF1α si-3, to determine the effects of HIF1α on deposition of collagen (Fig. 5a). The results showed that HIF1α si-2 led to a significant decrease in the expression level of Col I, Col III while the expression levels of α-SMA were relatively unchanged (Fig. 5b), and HIF1α si-2 could also affect the percentage of cells at S phase (Fig. S2), indicating that HIF1α could inhibit the deposition of collagen and HFs proliferation. Moreover, to explore the downstream signaling and underlying mechanism, AKT pathway was investigated after silencing HIF1α or miR-155. The results showed that the phosphorylation levels of AKT were decreased in HIF1α-silenced group and increased in anti-miR-155 group respectively (Fig. 5c, d). Then, AKT inhibitor LY294002 were applied to cultured HFs (Fig. S1a) and the results showed that LY294002 could inhibit the expression level of Col I, Col III and HIF1α and the proliferation of HFs (Fig. S1b, c).
Effect of HIF1α on the expression of Col I, Col III and α-SMA and downstream signaling in HS fibroblasts. a HFs were transfected with 80 nM HIF1α small interfering RNAs (HIF1α si-1, HIF1α si-2 or HIF1α si-3) or siRNA-negative control (Ctr si) for 48 h. Western blot analyzed the silencing efficiency of HIF1α. b The effects of HIF1α si-2 on Col I, Col III and α-SMA protein levels in HFs were analyzed by western blot and normalized to actin. c, d Western blot analyzed the protein levels of P-AKT and AKT following the transfection of negative control (Ctr si), HIF1α si-2, anti-miR-155 and anti-NC. Actin served as a protein loading control. Error bars represented means ± SD of n = 4. *p < 0.05; **p < 0.01 compared to control
These results suggested that the AKT pathways participated in miR-155 induced HIF1α in the formation of HS.
miR-155 agomir or silencing of HIF1α decreased scarring formation in vivo
We next validated whether ectopic expression of miR-155 or HIF1α influences the formation of HS in vivo. To further assess the effect of overexpression miR-155 or silencing of HIF1α on scarring formation in vivo, the rabbit ear scar models were established. On postoperative day 28, cutaneous incision wound models were treated with miR-155 agomir or siHIF1α by subcutaneous injection. Strikingly, 6 weeks later, the wound volume of miR-155 agomir group and siHIF1α group was significant smaller than those in positive control group and control (saline and blank) group (Fig. 6a). The results of Masson trichrome staining showed that miR-155 agomir and siHIF1α remarkably improved the texture of scar tissue, exhibiting significantly lightened collagen staining and more organized arrangement while arranged in a disordered manner, with a bulky appearance in the control group (Fig. 6b). These results proved that overexpression miR-155 and silencing HIF-1a could improve scarring formation in rabbit ear model.
Effect of miR-155 and HIF1α in a rabbit ear scar model in vivo. a Dermal architecture in rabbit ear scar model after miR-155 overexpression or silence of HIF1α, six rabbits in each group (n = 6). 5 nM miR-155 agomir, si HIF1α, positive control or saline were injected into subcutaneous section at 0 and 7th day after scar formation 28 days. b Masson trichrome staining showed the arrangement of collagen, Scale bars = 100 µm
Discussion
HS is a fibrotic disease that is characterized by the over-proliferation and activation of dermal fibroblasts and often considered as a benign skin tumor. Several studies have been published describing the role of miR-155 in tumor progression and served as a tumor suppressor (Chen et al. 2014; Mattiske et al. 2012; Wang and Wu 2012). Given the similarity between HS and tumor, it is reasonable to suppose that miR-155 has close relationship with the formation of HS.
In this study, it was found that the expression of miR-155 was significantly decreased in HS and HFs comparing with that in the NS and NFs (Fig. 1). In addition, ectopic expression of miR-155 could notably decrease the protein level of Col I and Col III, whereas the protein level of α-SMA, a well-known marker for myofibroblasts, was not changed (Fig. 2). In vivo studies furtherly confirmed above observation, miR-155 agomir treatment significantly improved the appearance of rabbit ear scar and ameliorated collagen arrangement (Fig. 6). Overall, these findings suggested miR-155 may improve the excessive deposition of collagen to reduce HS formation, in accordance with the regulatory role of miR-155 in experimental and idiopathic pulmonary fibrosis (Kurowska-Stolarska et al. 2016).
The aberrant proliferation of fibroblasts is another important character of HS. Thus, based on above findings, we were eager to elucidate whether this characteristic expression pattern of miR-155 was correlated with its biological function in HS. By using CCK-8 and flow cytometry analysis, it was demonstrated that overexpression of miR-155 could inhibit the proliferative activity of HS fibroblasts (Fig. 3a) and increase the percentage of cells in G1 and G2 phases, while decreased that in S phase(Fig. 3b). However, miR-155 had no effects on the migration of HS fibroblast (Fig. 3c, d). Above data, we demonstrated the possible effects of miR-155 on HS were suppression of fibroblast proliferation and had unrelated with cell migration.
miRNAs function as oncogenes or tumor suppressors by regulating their targets on the epigenetic level via decreasing translation of target mRNA or increasing degradation of target mRNA (Breving and Esquela-Kerscher 2010). It has been reported that miR-155 could target PTEN, FoxO3a and SOCS1 (Fu et al. 2017; Qayum et al. 2016; Wu et al. 2016). In this study, we focused on HIF-1α as a potential target of miR-155. Our results exhibited that the expression level of miR-155 was inversely correlated with the expression profile of HIF-1α (Fig. 4), in accordance with a previous report in LPS-induced acute lung injury (Hu et al. 2015). In fact, regulation of HIF-1α by miR-155 was critical in prolonged hypoxia microenvironment (Xie et al. 2015). HIF-1 is an important transcription factor involved in the hypoxic response of cells, and it plays significant roles in tumor development and progression (Lu et al. 2017; Mesarwi et al. 2016). HIF-1α, a subunit of the HIF1, was recently demonstrated to be involved in tissue fibrosis and wound healing (Duscher et al. 2015; Mesarwi et al. 2016). Present researches have shed some light on HIF-1α and HS (Lynam et al. 2015; Zheng et al. 2014). Our results also suggested that HS was located in a local hypoxia environment (Fig. 4b). Moreover, HIF-1α siRNA could down-regulate the expression levels of Col I and Col III in vitro (Fig. 5b) and obviously improved the arrangement of collagen fibres in vivo (Fig. 6). Besides, HIF-1α siRNA suppressing fibroblast proliferation via redistributing cell cycle (Fig. S2). These data demonstrated that the function of miR-155 was related to HIF-1α and miR-155 could modulate the proliferation and secretion of fibroblast though HIF-1α.
There are emerging evidences linking the PI3K/AKT pathway with HIF-1α in tumor cells (Choi et al. 2013; Fang et al. 2005; Joshi et al. 2014). It was reported that inhibition of the PI3K/AKT pathway could block the accumulation of HIF-1α protein and promote the hypoxic degradation of HIF-1α (Joshi et al. 2014). In fact, PI3K/AKT pathway is known to be a major cell survival pathway and responsible for cell survival, energy metabolism and protein synthesis (Castellino et al. 2009; Jiang et al. 2001). Importantly, deactivation of AKT signaling can mediate survival, migration, proliferation of fibroblasts and excessive production of collage, resulting in HS (Shi et al. 2014). Our results also suggest that inhibition of AKT could decrease the protein levels of collagen and HIF-1α and inhibit the proliferation of fibroblasts (Fig. S1). Consistent with these reports, we established here that HIF-1α siRNA could decrease the expression levels of p-AKT, while anti-miR-155 increased the expression levels of p-AKT obviously in HFs (Fig. 5d). It was indicated that miR-155 mediated the proliferation of fibroblasts and excessive production of collage partly attributable to AKT signaling pathway by targeting HIF-1α.
Conclusions
Taken together, our data identified the inhibitory effect of miR-155 on fibrosis in HS. And the AKT signaling pathway was involved in miR-155 modulating processes through HIF-1α (Fig. 7). In summary, the results in our work implied that the function of miR-155 was exerted by regulating HIF-1α/PI3K/AKT pathway. These data highlighted miR-155 was a potential target for HS therapy.
Abbreviations
- HS:
-
Hypertrophic scar
- HFs:
-
Hypertrophic scar derived fibroblasts
- NS:
-
Normal skin
- NSFs:
-
Normal skin derived fibroblasts
- HIF1α:
-
Hypoxia inducible factor1α
- Col I:
-
Collagen I
- Col III:
-
Collagen III
- a-SMA:
-
a-smooth muscle actin
References
Armour A, Scott PG, Tredget EE (2007) Cellular and molecular pathology of HTS: basis for treatment. Wound Repair Regen 15(Suppl 1):S6–S17. https://doi.org/10.1111/j.1524-475X.2007.00219.x
Breving K, Esquela-Kerscher A (2010) The complexities of microRNA regulation: mirandering around the rules. Int J Biochem Cell Biol 42:1316–1329. https://doi.org/10.1016/j.biocel.2009.09.016
Castellino RC, Muh CR, Durden DL (2009) PI-3 kinase-PTEN signaling node: an intercept point for the control of angiogenesis. Curr Pharm Des 15:380–388
Chen Z, Ma T, Huang C, Hu T, Li J (2014) The pivotal role of microRNA-155 in the control of cancer. J Cell Physiol 229:545–550. https://doi.org/10.1002/jcp.24492
Choi YH, Jin GY, Li LC, Yan GH (2013) Inhibition of protein kinase C delta attenuates allergic airway inflammation through suppression of PI3K/Akt/mTOR/HIF-1 alpha/VEGF pathway. PLoS ONE 8:e81773. https://doi.org/10.1371/journal.pone.0081773
Christmann RB et al (2016) miR-155 in the progression of lung fibrosis in systemic sclerosis. Arthritis Res Ther 18:155. https://doi.org/10.1186/s13075-016-1054-6
Csak T, Bala S, Lippai D, Kodys K, Catalano D, Iracheta-Vellve A, Szabo G (2015) MicroRNA-155 deficiency attenuates liver steatosis and fibrosis without reducing inflammation in a mouse model of steatohepatitis. PLoS ONE 10:e0129251. https://doi.org/10.1371/journal.pone.0129251
Duscher D et al (2015) Fibroblast-specific deletion of hypoxia inducible factor-1 critically impairs murine cutaneous neovascularization and wound healing. Plast Reconstr Surg 136:1004–1013. https://doi.org/10.1097/PRS.0000000000001699
Fang J, Xia C, Cao Z, Zheng JZ, Reed E, Jiang BH (2005) Apigenin inhibits VEGF and HIF-1 expression via PI3K/AKT/p70S6K1 and HDM2/p53 pathways. FASEB J 19:342–353. https://doi.org/10.1096/fj.04-2175com
Fattore L et al. (2017) MicroRNAs in melanoma development and resistance to target therapy. Oncotarget. https://doi.org/10.18632/oncotarget.14763
Fu X et al (2017) MicroRNA-155-5p promotes hepatocellular carcinoma progression by suppressing PTEN through the PI3K/Akt pathway. Cancer Sci. https://doi.org/10.1111/cas.13177
Hu R, Zhang Y, Yang X, Yan J, Sun Y, Chen Z, Jiang H (2015) Isoflurane attenuates LPS-induced acute lung injury by targeting miR-155-HIF1-alpha. Front Biosci 20:139–156
Huffaker TB et al (2012) Epistasis between microRNAs 155 and 146a during T cell-mediated antitumor immunity. Cell Rep 2:1697–1709. https://doi.org/10.1016/j.celrep.2012.10.025
Jiang BH, Jiang G, Zheng JZ, Lu Z, Hunter T, Vogt PK (2001) Phosphatidylinositol 3-kinase signaling controls levels of hypoxia-inducible factor 1. Cell Growth Differ 12:363–369
Joshi S, Singh AR, Zulcic M, Durden DL (2014) A macrophage-dominant PI3K isoform controls hypoxia-induced HIF1alpha and HIF2alpha stability and tumor growth, angiogenesis, and metastasis. Mol Cancer Res 12:1520–1531. https://doi.org/10.1158/1541-7786.MCR-13-0682
Kurowska-Stolarska M et al (2016) The role of microRNA-155/liver X receptor pathway in experimental and idiopathic pulmonary fibrosis. J Allerg Clin Immunol. https://doi.org/10.1016/j.jaci.2016.09.021
Li Y et al (2017) MicroRNA-192 regulates hypertrophic scar fibrosis by targeting SIP1. J Mol Histol 48:357–366. https://doi.org/10.1007/s10735-017-9734-3
Lim LP et al (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773. https://doi.org/10.1038/nature03315
Lin Q, Ma L, Liu Z, Yang Z, Wang J, Liu J, Jiang G (2016) Targeting microRNAs: a new action mechanism of natural compounds. Oncotarget. https://doi.org/10.18632/oncotarget.14392
Lu Y, Ji N, Wei W, Sun W, Gong X, Wang X (2017) MiR-142 modulates human pancreatic cancer proliferation and invasion by targeting hypoxia-inducible factor 1 (HIF-1alpha) in the tumor microenvironments. Biol Open. https://doi.org/10.1242/bio.021774
Lynam EC, Xie Y, Dawson R, McGovern J, Upton Z, Wang X (2015) Severe hypoxia and malnutrition collectively contribute to scar fibroblast inhibition and cell apoptosis. Wound Repair Regen 23:664–671. https://doi.org/10.1111/wrr.12343
Mattiske S, Suetani RJ, Neilsen PM, Callen DF (2012) The oncogenic role of miR-155 in breast cancer. Cancer Epidemiol Biomark Prev 21:1236–1243. https://doi.org/10.1158/1055-9965.EPI-12-0173
Mesarwi OA, Shin MK, Bevans-Fonti S, Schlesinger C, Shaw J, Polotsky VY (2016) Hepatocyte hypoxia inducible factor-1 mediates the development of liver fibrosis in a mouse model of nonalcoholic fatty liver disease. PLoS ONE 11:e0168572. https://doi.org/10.1371/journal.pone.0168572
Mu P, Akashi T, Lu F, Kishida S, Kadomatsu K (2017) A novel nuclear complex of DRR1, F-actin and COMMD1 involved in NF-kappaB degradation and cell growth suppression in neuroblastoma. Oncogene 36:5745–5756. https://doi.org/10.1038/onc.2017.181
Qayum AA et al (2016) IL-10-induced miR-155 targets SOCS1 to enhance IgE-mediated mast cell function. J Immunol 196:4457–4467. https://doi.org/10.4049/jimmunol.1502240
Ruvkun G (2001) Molecular biology. Glimpses of a tiny RNA world. Science 294:797–799. https://doi.org/10.1126/science.1066315
Shi H et al (2014) Ethanol extract of Portulaca oleracea L. reduced the carbon tetrachloride induced liver injury in mice involving enhancement of NF-kappaB activity. Am J Transl Res 6:746–755
Sideek MA, Teia A, Kopecki Z, Cowin AJ, Gibson MA (2016) Co-localization of LTBP-2 with FGF-2 in fibrotic human keloid and hypertrophic scar. J Mol Histol 47:35–45. https://doi.org/10.1007/s10735-015-9645-0
Wang J, Wu J (2012) Role of miR-155 in breast cancer. Front Biosci 17:2350–2355
Wang B, Yao K, Wise AF, Lau R, Shen HH, Tesch GH, Ricardo SD (2017) MicroRNA-378 reduces mesangial hypertrophy and kidney tubular fibrosis via MAPK signaling. Clin Sci. https://doi.org/10.1042/CS20160571
Wu H et al (2016) MiR-155 is involved in renal ischemia-reperfusion injury via direct targeting of FoxO3a and regulating renal tubular cell pyroptosis. Cell Physiol Biochem 40:1692–1705. https://doi.org/10.1159/000453218
Xie S, Chen H, Li F, Wang S, Guo J (2015) Hypoxia-induced microRNA-155 promotes fibrosis in proximal tubule cells. Mol Med Rep 11:4555–4560. https://doi.org/10.3892/mmr.2015.3327
Zheng J, Song F, Lu SL, Wang XQ (2014) Dynamic hypoxia in scar tissue during human hypertrophic scar progression. Dermatol Surg 40:511–518. https://doi.org/10.1111/dsu.12474
Zhou B, Zhu H, Luo H, Gao S, Dai X, Li Y, Zuo X (2017) MicroRNA-202-3p regulates scleroderma fibrosis by targeting matrix metalloproteinase 1. Biomed Pharmacother 87:412–418. https://doi.org/10.1016/j.biopha.2016.12.080
Zhu J et al (2017) NF-kappaB p65 overexpression promotes bladder cancer cell migration via FBW7-mediated degradation of RhoGDIalpha protein. Neoplasia 19:672–683. https://doi.org/10.1016/j.neo.2017.06.002
Zonari E, Pucci F, Saini M, Mazzieri R, Politi LS, Gentner B, Naldini L (2013) A role for miR-155 in enabling tumor-infiltrating innate immune cells to mount effective antitumor responses in mice. Blood 122:243–252. https://doi.org/10.1182/blood-2012-08-449306
Funding
This work was supported by National Natural Science Foundation of China (81601693, 81372069, 81703924).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, X., Li, J., Yang, X. et al. miR-155 inhibits the formation of hypertrophic scar fibroblasts by targeting HIF-1α via PI3K/AKT pathway. J Mol Hist 49, 377–387 (2018). https://doi.org/10.1007/s10735-018-9778-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10735-018-9778-z