Abstract
The herbicide glyphosate inhibits the key enzyme 5-enolpyruvate shikimate-3-phosphate synthase (EPSPS) in the aromatic amino acid synthesis pathway of plants. This study aims to explore the Y-EPSPS gene derived from maize through codon optimization and validate its glyphosate resistance in Arabidopsis Thaliana. Germination rates of seeds under different glyphosate concentrations revealed that seeds overexpressing the Y-EPSPS gene exhibited higher germination rates compared to wild-type seeds. DAB and NBT staining methods were used to measure ROS levels in Arabidopsis plants under 0.8 mM glyphosate stress, showing that plants overexpressing Y-EPSPS had lower ROS levels compared to wild-type plants. Soluble sugar and malondialdehyde (MDA) content were higher in Y-EPSPS overexpressing plants, whereas MDA content was lower, indicating a potential stress response to glyphosate. Chlorophyll content and FV/FW ratio were higher in plants overexpressing Y-EPSPS compared to wild-type plants, suggesting reduced susceptibility to glyphosate. Enzyme activity and gene expression analysis further demonstrated significant increases in POD, SOD, and CAT enzyme activities in Y-EPSPS overexpressing plants compared to wild-type, while SD enzyme activity decreased significantly. Expression levels of ROS detoxification-related genes (AtCAT3 and AtSOD1) and stress defense-related genes (AtLTP3, AtSOS1, and DQSD) were also elevated to varying degrees in Y-EPSPS overexpressing plants compared to wild-type plants. These results indicate that the optimized Y-EPSPS gene confers certain resistance to glyphosate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aromatic amino acids (AAA), including tryptophan, tyrosine, and phenylalanine, serve as precursors for numerous natural products (Parthasarathy et al. 2018) and play crucial roles in plant growth and development (Less et al. 2010). They are involved in various physiological processes such as auxin biosynthesis, enhancing drought resistance, promoting lignin synthesis, participating in signal transduction, and responding to biotic and abiotic stresses (Lynch et al. 2020; Pandey et al. 2017).
Shikimic acid serves as a precursor in the synthesis of AAA, also known as the shikimate pathway. Shikimic acid is synthesized through a series of enzyme-catalyzed reactions involving phosphoenolpyruvate (PEP) and erythros-4-phosphate, including 3-deoxyarabinoheptulose-7-phosphate, 5-enolpyruvate shikimate-3-phosphate synthase (EPSPS), dehydroguanic acid synthase, dehydroguanic acid dehydrase and shikimate dehydrogenase (Maeda et al. 2012; Tzin and Galli 2010; Wu et al. 2022; Suh et al. 1993). Shikimic acid is then used in enzyme-catalyzed reactions involving shikimate kinase, EPSPS enzyme, cladonic acid synthase, AAA, and other enzymes to produce secondary metabolites.
EPSPS enzyme is a key enzyme in the shikimate pathway and the sole target of the herbicide glyphosate (Okumu et al. 2019). Glyphosate is the most widely used broad-spectrum herbicide worldwide, making research on glyphosate and EPSPS enzyme in plants highly significant (Duke 2011). Based on the inherent sensitivity and catalytic efficiency of EPSPS enzyme towards glyphosate, EPSPS enzymes can be classified into two major classes. Class I EPSPS enzymes are found in all plants and certain bacterial species such as Escherichia coli, Klebsiella pneumoniae, and Salmonella typhimurium, which are relatively sensitive to glyphosate inhibition. Class II EPSPS enzymes are only present in bacteria such as Staphylococcus aureus and Agrobacterium tumefaciens. Currently, the most widely used EPSPS gene is derived from Bacillus subtilis and has been extensively studied and applied in plants (Funke et al. 2006; Herrmann and Weaver 1999; Dill et al. 2008; Chhapekar et al. 2015).
The mechanism of action of glyphosate involves mimicking the carbon state of PEP. Glyphosate competitively binds reversibly with EPSPS enzyme to form a stable but non-covalent ternary complex EPSPS-S3P-glyphosate, leading to the loss of EPSPS enzyme activity. This diversion causes a substantial flow of carbon sources towards S3P, resulting in rapid accumulation of shikimic acid within tissues. Consequently, the synthesis of essential aromatic amino acids (AAA) necessary for protein biosynthesis is severely hindered, ultimately inhibiting plant growth (Duke and Powles 2009; Wiersma et al. 2015).
Frequent use of glyphosate can lead to the development of glyphosate resistance in weeds, with amplification of the EPSPS gene being the primary mechanism conferring glyphosate resistance in Amaranthus populations (Mahajan et al. 2018). Chao Ouyang discovered a naturally evolved TIPS-EPSPS mutation that has been shown to confer high resistance to glyphosate in goosegrass, with resistance levels 180 times that of the wild type (WT) (Ouyang et al. 2021). Overexpression of the EPSPS gene and site-specific mutations can enhance plant resistance to glyphosate to varying degrees. When the EPSPS gene is overexpressed, EPSPS enzyme synthesis increases, and recommended field doses of glyphosate do not kill the plants, thereby enhancing glyphosate resistance (Vázquez-García et al. 2020; Chandi et al. 2012; Alcántara-de la Cruz et al. 2016).
Although EPSPS enzyme is the sole key enzyme in the shikimate pathway (Ge et al. 2012), so far, the Class II EPSPS genes sourced from bacteria have been most widely studied and utilized in plants, while research on Class I EPSPS genes from plants remains insufficiently explored. Further investigation is needed to determine whether overexpressing endogenous EPSPS genes or optimizing them via codon usage can enhance plant resistance to glyphosate. Glyphosate reduces photosynthetic efficiency, increases chlorophyll degradation, inhibits chlorophyll function, carotenoid synthesis, ferredoxin enzyme activity, auxin transduction, and increases auxin oxidation (Ozturk et al. 2008; Vivancos et al. 2011). This study measured changes in chlorophyll content, soluble sugars, malondialdehyde (MDA), relative conductivity, and ROS levels before and after glyphosate treatment in overexpressing and wild-type plants to assess glyphosate’s impact on photosynthesis and plant stress resistance. The study explores whether codon optimization of maize’s endogenous EPSPS gene can enhance glyphosate resistance, optimizing the gene’s impact on the shikimate pathway, and the effects of glyphosate treatment on plants. This research lays a theoretical foundation for breeding glyphosate-resistant maize.
Materials and methods
Optimization of EPSPS gene and construction of overexpression vector
Using the online software GeneScript (https://www.genscript.com.cn/codon-opt.html), the codons of the endogenous EPSPS gene in maize were optimized to construct an expression vector.
Plant materials and growth conditions
Materials included WT Arabidopsis thaliana seeds and T2 seeds overexpressing the Y-EPSPS gene. Seeds were sterilized in 75% ethanol for 1 min, followed by 1% sodium hypochlorite for 10 min, and then rinsed 5–6 times with sterile water. Seeds were placed on 1/2 MS solid medium without glyphosate or with glyphosate, and incubated at 4 °C in the dark for 3 days before being transferred to continuous light for germination. T2 Arabidopsis seeds were germinated in vermiculite under conditions of 220 mmolm-2s-1 light intensity, a 16 h photoperiod, 8 h dark period, temperature maintained at 22–24 °C, relative humidity at 65%, and nutrient solution sprayed every 15 days.
Detection of overexpression positive plants
T2 generation Arabidopsis thaliana overexpressing the Y-EPSPS gene was screened with a 1/1000 concentration of glyphosate, sprayed every 2 days. Surviving seedlings were transplanted into soil (nutrient soil:vermiculite = 8:3). Arabidopsis young leaves were ground with a pestle in 1.5 mL centrifuge tubes containing 200 μL buffer, followed by incubation of Bar test strips in the tubes for 5 min, and subsequent PCR detection. After covering the entire plastic tray with leaves from overexpressing plants, samples were collected for genomic extraction, and PCR detection of the screened marker gene Bar was conducted (primer sequences in Supplementary Material Table 1). Positive plants were utilized for subsequent physiological and biochemical indicator assays.
Glyphosate resistance of overexpressed positive seeds and plants
Wild-type Arabidopsis seeds and seeds overexpressing the Y-EPSPS gene were separately placed on 1/2 MS solid medium without glyphosate or supplemented with 0.5 and 0.8 mM/L glyphosate, and their germination rates were assessed. Arabidopsis plants overexpressing the Y-EPSPS gene were cultivated in nutrient soil. When these plants reached 4 weeks of age, they were subjected to 0.8 mM/L glyphosate stress. One week later, agronomic traits of the plants were statistically analyzed and photographed.
Expression analysis of Y-EPSPS gene under glyphosate stress
Four-week-old T2 generation Arabidopsis overexpressing the Y-EPSPS gene were subjected to 0.8 mM/L glyphosate stress. Leaf samples were collected 3 days post-treatment, rapidly frozen in liquid nitrogen, and RNA was extracted for fluorescence quantitative PCR. The top two positive plants with the highest expression levels of the Y-EPSPS gene were selected for subsequent experiments.
Detection of H2O2 and O2 − content
DAB staining buffer was prepared (0.02 g DAB dissolved in 50 mL water) and placed in 50 mL centrifuge tubes. The fifth leaf of 4-week-old Arabidopsis thaliana was immersed in DAB staining solution and vacuum-infiltrated for 30 min, followed by overnight incubation at room temperature. Chlorophyll was removed in 95% ethanol at 80 °C, and photographs of the staining results were taken (Wang et al. 2022).
NBT staining buffer was prepared (0.5 g NBT, 500 µL sodium azide solution, 500 µL 1 M sodium phosphate buffer, diluted to 50 mL with deionized water) and placed in 50 mL centrifuge tubes. The fifth leaf of 4-week-old Arabidopsis thaliana was soaked in NBT staining buffer for 60 min, washed in 80 °C water containing 95% ethanol until chlorophyll was completely removed, and then photographed for staining results (Geerts and Roels 1981).
Measurement of physiological and biochemical indicators
Hydrogen peroxide, superoxide anion, soluble sugars, malondialdehyde, chlorophyll content, and activities of peroxidase (POD), superoxide dismutase (SOD), catalase (CAT), and shikimate dehydrogenase (SD) were measured. Test kits were purchased from boxbio (http://www.boxbiological.com/) and experiments were conducted according to the manufacturer’s instructions.
Fluorescence quantitative PCR
Total RNA was extracted from maize leaves using the Trizol method. cDNA was synthesized from total RNA using M-MLV reverse transcriptase (Takara). qRT-PCR was performed using SYBR PreMix Ex Taq (Takara) with amplification conditions of: 95 °C for 10 min pre-denaturation, followed by 40 cycles of 95 °C denaturation for 10 s, 58 °C annealing for 20 s, and 72 °C extension for 15 s, with a melt curve analysis. Data were analyzed using the 2−ΔΔt method with AtActin1 gene as an internal reference, and each sample was biologically replicated three times. Primers used in this study are listed in Supplementary Material Table 1.
Statistical analysis
All results in this study were repeated at least three times. Experimental quantitative data were statistically analyzed using SPSS 19.0 software (SPSS Inc.Chicago, IL, USA), and differences in treatment outcomes were confirmed using t tests. Significance levels were denoted as *p < 0.05, **p < 0.01 and ***p < 0.001.
Results
Codon optimization results and expression cassette frame in overexpression vectors
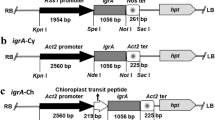
We optimized the codons of the EPSPS gene using online software and observed changes in codon preferences favoring maize compared to the original sequence (Fig. 1). The expression cassette is illustrated in Fig. 2.
Results of overexpression in transgenic plants
Following PCR validation and strip tests (Fig. 3A, C), we successfully obtained 15 positive overexpression transgenic plants. Fluorescence quantitative PCR indicated that plants OE4 and OE13 exhibited the highest gene expression levels (Fig. 3B). Consequently, we selected these strains for propagation and subsequent experiments.
Results of overexpressed positive plants. M: DNA Marker DL 2000; P: positive control; N: water; 1–15: Overexpressed positive plants. A PCR test results. B qPCR test results. C Test result of bar gene test strip. P: Positive control; N: Negative control; 1–15: Overexpressed positive plants. The student’s test was used, with asterisks indicating statistically significant differences (*p < 0.05; **p < 0.01). The data is shown as the mean ± SD from three independent experiments
Glyphosate resistance in Y-EPSPS overexpressing plants
EPSPS enzyme is the sole target of glyphosate action (Ge et al. 2014), reacting to glyphosate. Y-EPSPS overexpressing and wild-type (WT) plants were separately inoculated on 1/2MS solid media containing different concentrations of glyphosate, and seed germination was observed after 8 days. We found that under glyphosate-free conditions, germination in overexpressing plants was similar to WT plants and remained unaffected. However, in media containing 0.5 mM glyphosate, both overexpressing and WT plants showed reduced germination, with higher germination rates observed in overexpressing plants: WT 54.02%, E-OE4 74.44%, E-OE13 80% (Fig. 4A, C). In media with 0.8 mM glyphosate, significant inhibition of germination was observed in all strains: WT 34.48%, E-OE4 52.22%, E-OE13 43.48% (Fig. 4A, C). After spraying 0.8 mM glyphosate solution on overexpressing and WT plants, overexpressing plants remained viable while WT plants perished (Fig. 4B). These results demonstrate that the Y-EPSPS gene can reduce sensitivity of Arabidopsis to glyphosate, enhancing its resistance.
Results of positive seed germination experiment and glyphosate resistance. A Germination results of Arabidopsis overexpressed positive seeds with different concentrations of glyphosate. B Resistance results of overexpressed Arabidopsis positive plants to 0.8 mM glyphosate spray. C Seed germination of Arabidopsis thaliana under different concentrations of glyphosate. The student’s test was used, with asterisks indicating statistically significant differences (*p < 0.05; **p < 0.01)
Glyphosate treatment increases H2O2 and O2 − levels
Reactive oxygen species (ROS) are crucial signaling molecules in response to various stress processes. Changes in hydrogen peroxide and superoxide anion levels play a vital role in plant stress (Sedigheh et al. 2011). Therefore, assessing ROS levels in Y-EPSPS overexpressing and WT plants is crucial. The sixth leaf of control and glyphosate-treated Arabidopsis plants was isolated and immersed in NBT and DAB staining solutions to reveal hydrogen peroxide and superoxide anion levels. Results showed no significant difference between overexpressing plants and WT under normal conditions. However, under glyphosate treatment, overexpressing plants exhibited lighter leaf color compared to WT (Fig. 5A, B). Quantitative results of H2O2 and O2− were consistent with chemical staining results (Fig. 5C, D). These findings indicate that Y-EPSPS overexpression reduces ROS accumulation in Arabidopsis leaves, thereby conferring glyphosate resistance.
H2O2 and O2− content detection results. A, B NBT, DAB staining. Overexpressed and positive plants at 4 weeks of age were stained. C, D The amount of H2O2 and O2−accumulated in plants. The student’s test was used, with asterisks indicating statistically significant differences (*p < 0.05; **p < 0.01). The data is shown as the mean ± SD from three independent experiments
Impact of Y-EPSPS gene on chlorophyll content and Fv/Fm
To further understand the role of the Y-EPSPS gene in glyphosate resistance, two Y-EPSPS overexpressing Arabidopsis plants were genetically transformed, and their chlorophyll content and Fv/Fm were measured. Before glyphosate treatment, chlorophyll content and Fv/Fm levels were similar between overexpressing and WT plants. However, after glyphosate treatment, overexpressing plants showed significantly higher chlorophyll content and Fv/Fm compared to WT plants (Fig. 6). This suggests that overexpressing the Y-EPSPS gene reduces sensitivity of plants to glyphosate, mitigating its toxic effects.
Y-EPSPS gene enhances soluble sugar and MDA content
We measured the levels of soluble sugars and MDA in both Y-EPSPS overexpressing plants and wild-type plants before and after glyphosate treatment. Initially, there was nearly no difference in soluble sugar content between overexpressing and wild-type plants prior to glyphosate treatment. However, post-treatment, the soluble sugar content in overexpressing plants significantly exceeded that of WT plants. Similarly, before glyphosate treatment, there was minimal difference in MDA content between overexpressing and WT plants. Yet, after glyphosate treatment, MDA levels in overexpressing plants notably decreased compared to WT plants (Fig. 7), indicating enhanced stress resistance and antioxidant capability, thereby reducing glyphosate-induced damage and conferring resistance to glyphosate.
Y-EPSPS gene alters ROS-related enzyme activities
The activity of reactive oxygen species ROS-related enzymes is an important indicator of the plant’s ability to scavenge ROS. Therefore, we assessed the activity of ROS-related enzymes. The results showed that before glyphosate treatment, the levels of POD, SOD, and CAT activities were similar between overexpressing plants and wild-type (WT) plants. After glyphosate treatment, however, the enzyme activities of POD, SOD, and CAT in overexpressing plants were significantly higher than those in WT plants (Fig. 8). This enables the breakdown of higher and more oxidative molecules produced in plants, reducing glyphosate’s toxic effects on plants.
Y-EPSPS gene reduces shikimate dehydrogenase activity
We measured the activity of shikimate dehydrogenase (SD), a bifunctional enzyme essential for shikimate acid synthesis. Before glyphosate treatment, SD activity showed little difference between overexpressing and wild-type plants. However, post-treatment, SD activity decreased in both overexpressing and wild-type plants. Yet, SD activity in overexpressing plants was notably higher than in WT plants, reaching twice the level observed in WT plants (Fig. 9). This suggests that glyphosate reduces EPSPS enzyme activity, leading to excessive accumulation of shikimate acid in plants, thereby inhibiting SD enzyme activity.
Impact of Y-EPSPS gene on ROS detoxification genes, stress response genes, and shikimate synthesis enzyme gene expression under glyphosate stress
Under glyphosate stress, overexpression of the Y-EPSPS gene partially reduces H2O2 and O2− levels in plants. These findings indicate that Y-EPSPS is involved in ROS regulation and participates in plant growth and development. In this study, we analyzed the expression of several ROS detoxification genes (AtCAT3: Gene ID: 838651 and AtSOD1: Gene ID: 837405), stress defense-related genes (AtLTP3: Gene ID: 836051 and AtSOS1: Gene ID: 814729), and the DQSD gene (Gene ID: 819809) (primer sequences in Supplementary Table 1) to determine the role of Y-EPSPS in glyphosate resistance. Results showed that gene expression levels increased post glyphosate treatment (Fig. 10), in response to glyphosate stress, thereby reducing glyphosate toxicity.
Effects of Y-EPSPS gene on the expression of active oxygen-related genes, stress response genes and shikimic acid pathway genes. Expression analysis of ROS (A, B), stress response (C, D) and shikimic acid pathway related genes (E) in Col-0, transgenic lines OE4 and OE13 under control and glyphosate stress conditions. The student’s test was used, with asterisks indicating statistically significant differences (*p < 0.05; **p < 0.01: ***p < 0.001). The data is shown as the mean ± SD from three independent experiments
Discussion
To produce high-quality glyphosate-resistant maize, we utilized the floral dip method to transfer the optimized Y-EPSPS gene into Arabidopsis thaliana (Ali et al. 2022; Zhang et al. 2006), resulting in 15 overexpressing plants. We assessed changes in chlorophyll, soluble sugars, MDA content, relative conductivity, and ROS levels in both overexpressing and wild-type plants before and after glyphosate treatment. Additionally, changes in enzyme activities such as POD, SOD, CAT, and SD were analyzed. Expression patterns of genes related to chlorophyll synthesis, ROS scavenging, and shikimate pathway were also examined.
Codon optimization involves altering gene sequences to enhance the expression of recombinant proteins. The degeneracy of the genetic code allows multiple codon sequences to encode the same protein. Codon usage bias prioritizes the use of specific synonymous codons (Gillen et al. 2021; Fu et al. 2020; López et al. 2020). Therefore, codon optimization can increase gene expression by altering the gene sequence without changing the amino acid sequence, thereby boosting protein content within cells. Traditionally, optimal codons are thought to enhance translation efficiency due to faster ribosomal movement and release along mRNA (Andersson and Kurland 1990; Ehrenberg and Kurland 1984; Sørensen and Pedersen 1991), potentially leading to faster growth rates through more efficient ribosome utilization. Another view suggests that optimal codon usage improves translation accuracy (Akashi 1994).
This study employed codon optimization to make the EPSPS gene sequence more favorable for maize-specific codon usage, enhancing EPSPS expression in plants. This increased the content of EPSP synthase expressed in Arabidopsis plants and verified its activity through indoor glyphosate tolerance experiments, aiming to acquire a novel EPSPS gene that enhances glyphosate tolerance.
SD is a key enzyme catalyzing the fourth step of the shikimate biosynthetic pathway (Yuan et al. 2022). In plants, SDH forms a DQD/SDH complex through its N-terminal to enhance metabolic cycling efficiency, and its activity analysis is crucial for shikimate pathway analysis (Akhlaghi et al. 2018). SD enzyme catalyzes shikimate to 3-dehydrogenated shikimic acid, while EPSPS catalyzes shikimate to 5-enol-shikimic acid, eventually synthesizing the aromatic amino acid precursor chorismate (Choi Si-Sun et al. 2019). Glyphosate competitively binds to EPSPS reversibly, forming a stable but non-covalent ternary complex EPSPS-S3P-glyphosate, leading to loss of EPSPS enzyme activity. This diverts substantial carbon sources towards shikimic acid, resulting in its rapid accumulation within tissues and severely hindering the synthesis of aromatic amino acids essential for protein biosynthesis, ultimately inhibiting plant growth (Duke and Powles 2009; Wiersma et al. 2015). Glyphosate spraying inhibits SD activity in plants. It is speculated that glyphosate forms a ternary complex with EPSPS, blocking the shikimate pathway and causing accumulation of shikimic acid, which in turn downregulates SD activity due to feedback regulation by shikimic acid, leading to decreased enzyme activity.
Previous studies have found that glyphosate can promote the formation of plant ROS (Jalal et al. 2021). In this study, following glyphosate stress, accumulations of H2O2 and O2− were observed in the leaves of both overexpressing plants and wild-type (WT) plants, though the accumulation in overexpressing plants was lower than in WT plants. Although the accumulation of O2− increased, it was not as significant as the accumulation of the ROS H2O2, which may be due to biochemical differences between these two ROS. Superoxide is a transient molecule, whereas hydrogen peroxide is the most stable ROS, making it easier to accumulate (Shaner et al. 2005). POD, CAT, and SOD constitute the most important protective enzyme systems in plants and are crucial for scavenging reactive oxygen species (ROS) (Shafiq et al. 2021). The activities of POD, CAT, and SOD indirectly reflect the vigor of plants. SOD is responsible for converting O2− to H2O2, while CAT and POD degrade H2O2 to H2O and O2, thereby clearing ROS generated by glyphosate stress and maintaining dynamic balance of ROS in plants (Hu et al. 2018; Jiang et al. 2023; Zhao et al. 2005). This study found that the enzyme activities of POD, SOD, and CAT decreased in plants subjected to glyphosate stress, with overexpressing plants showing higher enzyme activities compared to WT plants, suggesting that the Y-EPSPS gene can mitigate ROS accumulation and oxidative damage caused by glyphosate stress to some extent. After glyphosate stress, both malondialdehyde (MDA) and soluble sugar content increased in both overexpressing and wild-type plants. In this study, the soluble sugar content in overexpressing plants was higher than in wild-type plants, whereas the MDA content was lower (You et al. 2012). MDA, an active metabolic byproduct, is often used as an indicator of oxidative stress. When plant cells undergo oxidative damage, MDA accumulates inside the cells and may react with molecules such as proteins, nucleic acids, and lipids, leading to abnormal cellular functions or even cell death (Mao et al. 2023; Gill and Tuteja 2010; Noctor 2006). Numerous studies have indicated that adverse environments enhance lipid peroxidation in leaf tissues of plants, resulting in increased MDA content and relative conductivity. Lipid peroxidation is a secondary effect of oxidative stress, which has been observed following glyphosate stress in various plants, including soybean, rice (Oryza sativa L.), tobacco (Nicotiana tabacum L.), and peanut (Arachis hypogaea L.) (Li et al. 2019; Vennapusa et al. 2022; Daramola et al. 2023). The content of soluble sugars in plants typically reflects their resistance to stress because sugars serve as both an energy source and antioxidants, aiding plants in coping with adverse conditions (Barco-Antoñanzas et al. 2022). As signaling molecules for nutrition and metabolism, soluble sugars are crucial for maintaining plant structure, metabolism, growth, and cellular osmotic balance (Barco-Antoñanzas et al. 2022; Fernándes-Escalada et al. 2016). Therefore, the content of soluble sugars in plants serves as an important indicator for evaluating their physiological status and stress responses. The increase in soluble sugar content indicates that overexpressing plants exhibit greater tolerance to glyphosate (Li et al. 2020).
After spraying glyphosate, the chlorophyll content of both overexpressing and wild-type plants significantly decreased, while soluble sugars, MDA, relative conductivity, and ROS content markedly increased. The changes in overexpressing plants and wild-type plants were consistent with plant stress responses. Previous research has also indicated that glyphosate spraying reduces leaf area, subsequently lowering photosynthetic rates and aboveground biomass (Hoagland et al. 1980; Ding et al. 2008). Following glyphosate treatment, chlorophyll content in overexpressing plants initially declined but reached levels similar to untreated WT plants by day 30. This suggests glyphosate impacts plants, even those tolerant to it. Additionally, soluble sugars, MDA, relative conductivity, and ROS content in overexpressing plant leaves showed significant changes post-glyphosate treatment, albeit lower than WT plants. Therefore, these overexpressing plants exhibit significant resistance to glyphosate.
In this study, glyphosate treatment affected MDA, soluble sugars, and ROS production. As demonstrated, chlorophyll content decreased in both overexpressing and wild-type Arabidopsis. Glyphosate likely induces oxidative stress, leading to several minor or indirect effects on plant physiology (Gomes et al. 2014). As a result of oxidative stress, glyphosate damages plants, prompting increased production of soluble sugars in response to the damage. Glyphosate disrupts chlorophyll, impacting photosynthetic rates and metabolism. It affects the shikimate pathway, hindering plant growth and development. Ultimately, glyphosate can kill plants. Glyphosate increases ROS activity within plant tissues, and the plant’s ability to scavenge ROS is crucial for enhancing glyphosate tolerance. Overexpressing the Y-EPSPS gene promotes synthesis of aromatic amino acids, enhancing the plant’s ability to clear active substances. Previous studies utilized exogenous EPSPS genes sourced from other plants. This study seeks to identify endogenous EPSPS genes in plants and optimize their codons to enhance glyphosate tolerance. Optimizing endogenous genes can stabilize genetic traits and reduce responses to the biotic environment. To further explore the biological function of the Y-EPSPS gene, future work will involve yeast hybridization, dual luciferase assays, and HPLC detection. A series of experiments will also be conducted on maize to observe phenotypic changes in transgenic maize and measure related physiological and biochemical indicators. This research lays a solid foundation for understanding the herbicide resistance mechanisms in maize, thereby benefiting maize breeding efforts.
References
Akashi H (1994) Synonymous codon usage in Drosophila melanogaster: natural selection and translational accuracy. Genetics 136(3):927–935. https://doi.org/10.1093/genetics/136.3.927
Akhlaghi M, Ghobadi S, Mohammad Hosseini M, Gholami Z, Mohammadian F (2018) Flavanols are potential anti-obesity agents, a systematic review and meta-analysis of controlled clinical trials. Nutr Metab Cardiovasc Dis 28(7):675–690. https://doi.org/10.1016/j.numecd.2018.04.001
Alcántara-de la Cruz R, Barro F, Domínguez-Valenzuela JA et al (2016) Target and non-target site mechanisms developed by glyphosate-resistant hairy beggarticks (Bidens pilosa L.). Planta 244:365–375. https://doi.org/10.3389/fpls.2016.01492
Ali I, Sher H, Ali A, Hussain S, Ullah Z (2022) Simplified floral dip transformation method of Arabidopsis thaliana. J Microbiol Methods 197:106492. https://doi.org/10.1016/j.mimet.2022.106492
Andersson SG, Kurland CG (1990) Codon preferences in free-living microorganisms. Microbiol Rev 54(2):198–210. https://doi.org/10.1128/mr.54.2.198-210.1990
Barco-Antoñanzas M, Gil-Monreal M, Eceiza MV, Royuela M, Zabalza A (2022) Primary metabolism in an Amaranthus palmeri population with multiple resistance to glyphosate and pyrithiobac herbicides. Plant Sci 318:111212. https://doi.org/10.1016/j.plantsci.2022.111212
Chandi A, Milla-Lewis SR, Giacomini D, Westra P, Preston C, Jordan DL et al (2012) Inheritance of evolved glyphosate resistance in a North Carolina Palmer Amaranth (Amaranthus palmeri) Biotype. Int J Agron 2012:176108. https://doi.org/10.1155/2012/176108
Chhapekar S, Raghavendrarao S, Pavan G, Ramakrishna C, Singh VK, Phanindra ML, Dhandapani G et al (2015) Transgenic rice expressing a codon-modified synthetic CP4-EPSPS confers tolerance to broad-spectrum herbicide, glyphosate. Plant Cell Rep 34:721–731. https://doi.org/10.1007/s00299-014-1732-2
Daramola OS, Kharel P, Iboyi JE, Devkota P (2023) Response of peanut (Arachis hypogaea L.) to sublethal rates of dicamba plus glyphosate at different growth stages. Agron J 115(4):1694–1704. https://doi.org/10.1002/agj2.21372
Dill GM, Cajacob CA, Padgette SR (2008) Glyphosate-resistant crops: adoption, use and future considerations. Pest Manag Sci 64:326–331. https://doi.org/10.1002/ps.1501
Ding X, Cao Y, Huang L, Zhao J, Xu C, Li X, Wang S (2008) Activation of the indole-3-acetic acid-amido synthetase GH3-8 suppresses expansin expression and promotes salicylate- and jasmonate-independent basal immunity in rice. Plant Cell 20(1):228–240. https://doi.org/10.1105/tpc.107.055657
Duke SO (2011) Glyphosate degradation in glyphosate-resistant and susceptible crops and weeds. J Agric Food Chem 59(11):5835–5841. https://doi.org/10.1021/jf102704x
Duke SO, Powles SB (2009) Glyphosate: a once-in-a-century herbicide. Pest Manag Sci 65:319–327. https://doi.org/10.1002/ps.1518
Ehrenberg M, Kurland CG (1984) Costs of accuracy determined by a maximal growth rate constraint. Q Rev Biophys 17(1):45–82. https://doi.org/10.1017/s0033583500005254
Fernandez-Escalada M, Gil-Monreal M, Zabalza A, Royuela M (2016) Characterization of the Amaranthus palmeri physiological response to glyphosate in susceptible and resistant populations. J Agric Food Chem 64:95–106. https://doi.org/10.1021/acs.jafc.5b04916
Fu H, Liang Y, Zhong X et al (2020) Codon optimization with deep learning to enhance protein expression. Sci Rep 10(1):17617. https://doi.org/10.1038/s41598-020-74091-z
Funke T, Han H, Healy-Fried ML, Fischer M, Schönbrunn E (2006) Molecular basis for the herbicide resistance of Roundup Ready crops. Proc Natl Acad Sci 103(35):13010–13015. https://doi.org/10.1073/pnas.0603638103
Ge X, d’Avignon DA, Ackerman JJ (2012) Vacuolar glyphosate-sequestration correlates with glyphosate resistance in ryegrass (Lolium spp.) from Australia, South America, and Europe: a 31P-NMR investigation. J Agric Food Chem 60(5):1243–1250. https://doi.org/10.1021/jf204089c
Gomes MP, Smedbol E, Chalifour A, Hénault-Ethier L, Labrecque M, Lepage L, Lucotte M, & Juneau P (2014) Alteration of plant physiology by glyphosate and its by-product aminomethylphosphonic acid: an overview. J Exp Bot 65:4691–4703. https://doi.org/10.1093/jxb/eru269
Geerts A, Roels F (1981) Quantitation of catalase activity by microspectrophotometry after diaminobenzidine staining. Histochemistry 72(3):357–367. https://doi.org/10.1007/BF00501778
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48(12):909–930. https://doi.org/10.1016/j.plaphy.2010.08.016
Gillen SL, Waldron JA, Bushell M (2021) Codon optimality in cancer. Oncogene 40(45):6309–6320. https://doi.org/10.1038/s41388-021-02022-x
Herrmann KM, Weaver LM (1999) The shikimate pathway. Annu Rev Plant Physiol 50(1):473–503. https://doi.org/10.1146/annurev.arplant.50.1.473
Hoagland RE (1980) Effects of glyphosate on metabolism of phenolic compounds: VI. Effects of glyphosine and glyphosate metabolites on phenylalanine ammonia-lyase activity, growth, and protein, chlorophyll, and anthocyanin levels in soybean (Glycine max) seedlings. Weed Sci 28(4):393–400. https://doi.org/10.1017/S0043174500055545
Hu Z, Weijian L, Yali F, Huiquan L (2018) Gibberellic acid enhances postharvest toon sprout tolerance to chilling stress by increasing the antioxidant capacity during the short-term cold storage. Sci Hortic 237:184–191. https://doi.org/10.1016/j.scienta.2018.04.018
Jalal A, Oliveira Junior JC, Ribeiro JS, Fernandes GC, Mariano GG, Trindade VDR, Reis ARD (2021) Hormesis in plants: physiological and biochemical responses. Ecotoxicol Environ Saf 207:111225. https://doi.org/10.1016/j.ecoenv.2020.111225
Jiang W, Ye Q, Wu Z, Zhang Q, Wang L, Liu J, Hu X, Guo D, Wang X, Zhang Z, He H, Hu L (2023) Analysis of CAT gene family and functional identification of OsCAT3 in rice. Genes 14(1):138. https://doi.org/10.3390/genes14010138
Less H, Angelovici R, Tzin V, et al (2010) Principal transcriptional regulation and genome-wide system interactions of the Asp-family and aromatic amino acid networks of amino acid metabolism in plants. Amino Acids 39, 1023–1028. https://doi.org/10.1007/s00726-010-0566-7
Less H, Angelovici R, Tzin V et al (2010) Principal transcriptional regulation and genome-wide system interactions of the Asp-family and aromatic amino acid networks of amino acid metabolism in plants. Amino Acids 39(4):1023–1028. https://doi.org/10.1007/s00726-010-0566-7
Li WY, Lu P, Xie H, Li GQ, Wang JX, Guo DY, Liang XY (2019) Effects of glyphosate on soybean metabolism in strains bred for glyphosate-resistance. Physiol Mol Biol Plants 25(2):523–532. https://doi.org/10.1007/s12298-018-0597-1
Li X, Jia J, Zhao P, Guo X, Chen S, Qi D, Cheng L, Liu G (2020) LcMYB4, an unknown function transcription factor gene from sheepgrass, as a positive regulator of chilling and freezing tolerance in transgenic Arabidopsis. BMC Plant Biol 20(1):238. https://doi.org/10.1186/s12870-020-02427-y
López JL, Lozano MJ, Fabre ML, Lagares A (2020) Codon usage optimization in the prokaryotic tree of life: how synonymous codons are differentially selected in sequence domains with different expression levels and degrees of conservation. Mbio 11(4):e00766-20. https://doi.org/10.1128/mBio.00766-20
Lynch JH, Qian Y, Guo L et al (2020) Modulation of auxin formation by the cytosolic phenylalanine biosynthetic pathway. Nat Chem Biol 16:850-856. https://doi.org/10.1038/s41589-020-0519-8
Maeda H, Dudareva N (2012) The shikimate pathway and aromatic amino acid biosynthesis in plants. Annu Rev Plant Biol 63:73–105. https://doi.org/10.1146/annurev-arplant-042811-105439
Mahajan G, Matloob A, Walsh M, & Chauhan BS (2018) Germination Ecology of Two Australian Populations of African turnipweed (Sisymbrium thellungii). Weed Sci 66(6):752–757. https://doi.org/10.1017/wsc.2018.55
Mao H, Zhao W, Yang X, Sheng L, Zhu S (2023) Recruitment and metabolomics between Canna indica and rhizosphere bacteria under Cr stress. Front Microbiol 14:1187982. https://doi.org/10.3389/fmicb.2023.1187982
Noctor G (2006) Metabolic signalling in defence and stress: the central roles of soluble redox couples. Plant Cell Environ 29(3):409–425. https://doi.org/10.1111/j.1365-3040.2005.01476.x
Okumu MN, Vorster BJ, Reinhardt CF (2019) Growth-stage and temperature influence glyphosate resistance in Conyza bonariensis (L.) Cronquist. S Afr J Bot 121:248–256. https://doi.org/10.1016/j.sajb.2018.10.034
Ouyang C, Liu W, Chen S et al (2021) The naturally evolved EPSPS from goosegrass confers high glyphosate resistance to rice. Front Plant Sci 12:756116. https://doi.org/10.3389/fpls.2021.756116
Ozturk L, Yazici A, Eker S, Gokmen O, Romheld V, Cakmak I (2008) Glyphosate inhibition of ferric reductase activity in irondeficient sunflower roots. New Phytol 177(4):899–906. https://doi.org/10.1111/j.1469-8137.2007.02340.x
Pandey P, Irulappan V, Bagavathiannan M, Senthil-Kumar M (2017) Impact of combined abiotic and biotic stresses on plant growth and avenues for crop improvement by exploiting physio-morphological traits. Front Plant Sci 8:537. https://doi.org/10.3389/fpls.2017.00537
Parthasarathy A, Cross PJ, Dobson RCJ, Adams LE, Savka MA, Hudson AO (2018) A three-ring circus: metabolism of the three proteogenic aromatic amino acids and their role in the health of plants and animals. Front Mol Biosci 5:29. https://doi.org/10.3389/fmolb.2018.00029
Sedigheh HG, Mortazavian M, Norouzian D, Atyabi M, Akbarzadeh A, Hasanpoor K, Ghorbani M (2011) Oxidative stress and leaf senescence. BMC Res Notes 4:477. https://doi.org/10.1186/1756-0500-4-477
Shafiq S, Akram NA, Ashraf M, García-Caparrós P, Ali OM, Latef AAHA (2021) Influence of glycine betaine (natural and synthetic) on growth, metabolism and yield production of drought-stressed maize (Zea mays L.) Plants. Plants 10(11):2540. https://doi.org/10.3390/plants10112540
Shaner DL et al (2005) Glyphosate-resistant crops: history, status and future. Pest Manag Sci 61(3):219–224. https://doi.org/10.1002/ps.1008
Si-Sun C, Seung-Yeul S, Sun-Ok P, Han-Na L, Ji-soo S, Ji-yeon K, Ji-Hoon P, Sangyong K, Joung LS, Gie-Taek C, Eung-Soo K (2019) Cell factory design and culture process optimization for dehydroshikimate biosynthesis in Escherichia coli. Front Bioeng Biotechnol 2019:7–241. https://doi.org/10.3389/fbioe.2019.00241
Sørensen MA, Pedersen S (1991) Absolute in vivo translation rates of individual codons in Escherichia coli. The two glutamic acid codons GAA and GAG are translated with a threefold difference in rate. J Mol Biol 222(2):265–280. https://doi.org/10.1016/0022-2836(91)90211-n
Suh H, Hepburn AG, Kriz AL, Widholm JM (1993) Structure of the amplified 5-enolpyruvylshikimate-3-phosphate synthase gene in glyphosate-resistant carrot cells. Plant Mol Biol 22:195–205. https://doi.org/10.1007/BF00014928
Tzin V, Galili G (2010) New insights into the shikimate and aromatic amino acids biosynthesis pathways in plants. Mol Plant 3(6):956–972. https://doi.org/10.1093/mp/ssq048
Vázquez-García JG, Alcántara-de la Cruz R, Palma-Bautista C, Rojano-Delgado AM, Cruz-Hipólito HE, Torra J, Barro F, De Prado R (2020) Accumulation of target gene mutations confers multiple resistance to ALS, ACCase, and EPSPS inhibitors in lolium species in chile. Front Plant Sci 11:553948. https://doi.org/10.3389/fpls.2020.553948
Vennapusa AR, Agarwal S, Rao Hm H, Aarthy T, Babitha KC, Thulasiram HV, Kulkarni MJ, Melmaiee K, Sudhakar C, Udayakumar M, Vemanna SR (2022) Stacking herbicide detoxification and resistant genes improves glyphosate tolerance and reduces phytotoxicity in tobacco (Nicotiana tabacum L.) and rice (Oryza sativa L.). Plant Physiol Biochem 189:126–138. https://doi.org/10.1016/j.plaphy.2022.08.025
Vivancos PD et al (2011) Perturbations of amino acid metabolism associated with glyphosate-dependent inhibition of shikimic acid metabolism affect cellular redox homeostasis and alter the abundance of proteins involved in photosynthesis and photores-piration. Plant Physiol 157:256–268. https://doi.org/10.1104/pp.111.181024
Wang C, Chen N, Liu J et al (2022) Overexpression of ZmSAG39 in maize accelerates leaf senescence in Arabidopsis thaliana. Plant Growth Regul 98:451–463. https://doi.org/10.1007/s10725-022-00874-1
Wiersma AT, Gaines TA, Preston C, Hamilton JP, Giacomini D, Robin Buell C et al (2015) Gene amplification of 5-enol-pyruvylshikimate-3-phosphate synthase in glyphosate-resistant Kochia scoparia. Planta 241(2):463–474. https://doi.org/10.1007/s00425-014-2197-9
Wu S, Chen W, Lu S, Zhang H, Yin L (2022) Metabolic engineering of shikimic acid biosynthesis pathway for the production of shikimic acid and its branched products in microorganisms: advances and prospects. Molecules 27:4779. https://doi.org/10.3390/molecules27154779
You J, Hu H, Xiong L (2012) An ornithine δ-aminotransferase gene OsOAT confers drought and oxidative stress tolerance in rice. Plant Sci 197:59–69. https://doi.org/10.1016/j.plantsci.2012.09.002
Yuan J, Zhong S, Long Y, Guo J, Yu Y, Liu J (2022) Shikimate kinase plays important roles in anthocyanin synthesis in petunia. Int J Mol Sci 23(24):15964. https://doi.org/10.3390/ijms232415964
Zhang X, Henriques R, Lin SS, Niu QW, Chua NH (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1(2):641–646. https://doi.org/10.1038/nprot.2006.97
Zhao HC, Zhao H, Wang BC, Wang JB (2005) Effect of local stress induction on resistance-related enzymes in cucumber seeding. Colloids Surf B 43(1):37–42. https://doi.org/10.1016/j.colsurfb.2005.01.017
Acknowledgements
We thank SYG and YYM conceived research plans and designed experiments.
Funding
This work was supported by Jilin Province Science and Technology Development Plan Project [20230202003NC].
Author information
Authors and Affiliations
Contributions
SYG and YYM conceived research plans and designed experiments. FHW, ZC conducted experiments. FHW wrote the draft. CLW, XTW analyzed the data. SYL, SYG and YYM reviewed and edited this article and provided helpful comments and discussions. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval and consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, FH., Zhang, C., Wang, CL. et al. Estimating the role of maize Y-EPSPS gene in glyphosate resistance in Arabidopsis transgenic lines. Plant Growth Regul (2024). https://doi.org/10.1007/s10725-024-01188-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10725-024-01188-0