Abstract
The loss of crop productivity due to soil salinity is an increasing threat to agriculture worldwide. Calcium (Ca2+) plays vital roles in salt-responsive signaling through the perception of various calcium-binding proteins, such as Ca2+-dependent lipid-binding proteins (CaLBs). Here, CaLBs from barley (Hordeum vulgare), a moderate salt-tolerant crop, and other green plants were selected for evolutionary and bioinformatics analysis. The emergence of the CaLB1 and C2 domains could be traced back to green algae, such as the chlorophyte alga Uronema belka (Uronemataceae). The physiological roles of HvCaLB1 in the salt-tolerant barley accession CM72 were investigated through gene silencing induced by barley stripe mosaic virus. Knockdown of HvCaLB1 significantly and differentially impaired the performance of plant growth, photosynthetic, and chlorophyll fluorescence parameters under the treatments of 200 and 400 mM NaCl. Moreover, the knockdown of HvCaLB1 disrupted the homeostasis of essential elements, particularly in the significant decrease of root potassium (K+) and Ca2+ contents in HvCaLB1 silencing plants compared to the control plants in response to salt stress. Significantly increased accumulation of reactive oxygen species (ROS), reduced cytosolic Ca2+ levels, as well as the decreased expression of HvHVP10 (Vacuolar H+-pyrophosphatase 10) and HvCaM1 (Calmodulin 1), were observed in the roots of the HvCaLB1-silencing plants subjected to 400 mM NaCl treatment compared to those of control plants. Taken together, CaLBs represent an ancient group of Ca2+-binding domain-containing proteins, and HvCaLB1 regulates NaCl-induced ion, ROS homeostasis, and gene expression in barley roots, demonstrating the potential application of CaLBs for crop improvement with increased tolerance to salt stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salinity is one of the major abiotic stresses that adversely affect plant productivity and quality globally (Ponce et al. 2021; Qadir et al. 2014). Soil salinization is aggravated by climate change, poor irrigation practices, and environmental deterioration (Liu et al. 2020; Park et al. 2016; Ziska et al. 2012). Soil salinization causes osmotic stress, ionic stress, and oxidative stress in plants, leading to the disruption of sodium (Na+) and potassium homeostasis, water deficiency, and nutritional imbalance (Shumilina et al. 2019; Tanveer and Shah 2017; Chen et al. 2023). Plants have developed various strategies to alleviate salinity stress by reducing absorption, increasing efflux, and enhancing compartmentalization of salt ions (Solis et al. 2020; Shabala et al. 2020; Zhang et al. 2021). It was revealed that both osmotic and ionic stresses are sensed by membrane-bound sensors that deliver the stress signals to secondary messengers such as Ca2+ and ROS. For example, plasma membrane-localized glycosyl inositol phosphorylceramide sphingolipids (GIPCs) could bind Na+ to gate Ca2+ influx channels, leading to the depolarization of the cell-surface potential, Ca2+ spikes and waves, the activation of Na+/H+ antiporter, and regulation of growth (Munns and Tester 2008; Munns et al. 2020; Ponce et al. 2021; Tyerman et al. 2019; Jiang et al. 2019).
The exposure of plants to salt stress triggers an immediate defined rise in free cytosolic Ca2+ concentration [Ca2+]i spatiotemporally (Steinhorst and Kudla 2019). The Ca2+ signals were transmitted and decoded by an elaborate toolkit of Ca2+-binding proteins (Liu et al. 2017a; Tong et al. 2021). Many Ca2+ components have been identified and characterized in recent years, including cytosolic phospholipase A2 (cPLA2), phospholipase C (PLC), calmodulin, calmodulin-like proteins (CMLs), Ca2+-dependent protein kinases (CDPKs), and CDPK-related kinases (CRKs), and calcineurin B-like proteins (CBLs) and CBL-interacting protein kinases (CIPKs) (Kudla et al. 2018; Tong et al. 2021). The C2 domain, consisting of approximately 130 amino acid residues, was originally identified from cPLA2 and CDPKs (Fu et al. 2019). It has gained increasing attention in decoding Ca2+ signals in recent years (Bondada et al. 2021; Fu et al. 2019). The single C2 domain-containing Ca2+-dependent lipid-binding proteins (CaLBs), which can bind phospholipid membranes in the presence of Ca2+, have been implicated in response to salt stress and other abiotic stresses (de Silva et al. 2011; Xiao et al. 2022). Usually, conformational transitions of functional domains and the activation of membrane–protein interactions, essential for stress signaling cascades mediated by CaLBs, are dependent on the integration of the C2 domain into membrane in the presence of Ca2+ (Fu et al. 2019). The expression level of CaSRC2, encoding a C2 domain-containing protein, was notably up-regulated under abiotic stresses, including salt stress in pepper (Capsicum annuum) (Kim et al. 2008). Ectopic expression of rice (Oryza sativa) small C2 domain protein 1 (OsSMCP1), which encodes a small C2 protein, enhanced salinity tolerance in the transgenic Arabidopsis plants (Yokotani et al. 2009). Similarly, a C2 domains-containing protein, OsC2DP, was explicitly required for salt tolerance by regulating the expression of salt-related genes, particularly salt overly sensitive 1 (OsSOS1) and Na+/H+antiporter 4 (OsNHX4), and it’s also required for maintaining ionic homeostasis in rice (Fu et al. 2019).
Increased [Ca2+]i and ROS production in plant cells were considered as two early responses to salt stress (Xu et al. 2022; Miller et al. 2010; Kandhol et al. 2022). ROS initially interact with polyamines to alter intracellular Ca2+ homeostasis, which synergistically bind to calcium sensor proteins and phosphorylated NADPH [(reduced) nicotinamide adenine dinucleotide phosphate], oxidases leading to elevated production of ROS (Castro et al. 2021; Drerup et al. 2013; Zepeda-Jazo et al. 2011; Wang et al. 2017). Take Arabidopsis as an example, the intercellular Ca2+ wave requires ROS produced by AtRbohD (Arabidopsis respiratory burst oxidase homologues D), while Ca2+ release is partially dependent on the vacuolar channel two-pore cation channel 1 (AtTPC1) (Evans et al. 2016; Shabala et al. 2020). Thus, ROS and calcium signals are interconnected to achieve directional cell-to‐cell systemic signaling in plants under salt stress (Ravi et al. 2023).
As the world’s fourth largest cereal crop, barley is an ideal cereal crop due to its relatively higher resistance to abiotic stress, diploidy, easy genetic transformation, wide adaptability, and economic uses (Zeng et al. 2015; Shen et al. 2020; Feng et al. 2020; Qiu et al. 2022). Barley shows relatively higher salt tolerance than other cereals and is one of the pioneer crops for alleviating soil salinization (Munns and Tester 2008). Many salt-responsive proteins have been identified and functionally characterized in barley during the last decades (Zelm et al. 2020). A vacuolar H+-pyrophosphatase encoded by HvHVP10 could enhance the sequestration of Na+ into root vacuoles, which might synergize with the vacuolar Na+/H+ antiporters (NHX) HvNHX1 and HvNHX4 (Fu et al. 2022). Two barley high-affinity K+ transporters, HvHKT1;1 and HvHKT1;5, could modulate salt tolerance by regulating tissue and cell ion homeostasis. Their expression is regulated by a Ca2+ sensor, HvCaM1 (Han et al. 2018; Huang et al. 2020; Shen et al. 2020). Additionally, a dramatic increase in the transcription of HvSOS1, HvSOS2, and HvSOS3, which was the core competent of the Salt-Overly-Sensitive (SOS) signaling transduction pathway, was observed in the salt-tolerant barley accessions (Jadidi et al. 2022; Yousefirad et al. 2018). Early research has also shown that Ca2+ mitigated the increased Na+ content and the decreased K+ content in the roots of barley exposed to salt (Witzel et al. 2009). However, the regulatory mechanisms underlying NaCl-induced Ca2+ signaling and the involvement of CaLBs in salt response in barley were largely unexplored.
In this study, we hypothesized that CaLB1s are evolutionarily conserved in green plants, and HvCaLB1 is important for salinity tolerance in barley through the regulation of Ca2+ homeostasis. Therefore, we functionally characterized the role of HvCaLB1, a single C2 domain-containing small protein, in salt tolerance through molecular evolutionary and physiological analyses.
Results
HvCaLB1 is evolutionarily unique from its homologous gene due to its significant sequence variability
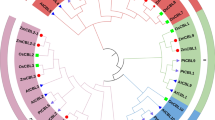
Genome-wide retrieval and analysis of the CaLBs using the highly conserved C2 domain identified 62, 53, and 31 members in Hordeum vulgare (barley), Oryza sativa (rice), and Arabidopsis thaliana, respectively. The phylogenetic tree (Fig. 1A) indicates that a minority of CaLB proteins exist as paralogues (20 pairs) and the majority (30 pairs) as orthologues. Thus, it can be hypothesized that the CaLBs may have experienced several gene duplication events during evolution, and gene duplication activity is maintained after the origin of barley. Intriguingly, AT4G00467.1 and Os02g0665100.1, identified as putative orthologs, were also presented in the phylogenetic relationship, implicating that CaLB genes originated before the differentiation of monocotyledon and dicotyledon species (Fig. 1A).
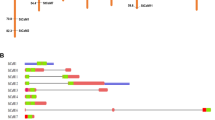
Bioinformatics analysis of CaLBgene family. (A) Phylogenetic relationships of CaLB gene family in barley (red), rice (brown) and Arabidopsis (blue). The unrooted phylogenetic tree was generated based on the amino acid sequences aligning maximum likelihood (ML) method using IQtree. The bootstrap values of 1000 replicates were calculated at each node. (B) Distribution of HvCaLB genes in chromosomes. The location of CaLB gene family members based on barley cultivar ‘Morex’ genome database (Hordeum vulgare. IBSC_v2). The gene pairs highlighted in purple font are located within 100 kb physical distance on chromosomes. (C) Micro-collinearity analysis of HvCaLB1 among 10 representative relative species. The arrows represent genes, the gray straight lines represent chromosomes, and the connecting lines represent homologous relationships, where homologous genes are divided into four types: RBH (Reciprocal Best Hit), SBH (Single-side Best Hit), singleton, and 1-to-many (all putative homologous genes). (D) Evolutionary and structural analysis of the HvCaLB1 homologous genes in related species of barley. Line, intron; bule block, CDS; yellow block, UTR. The red modification highlighted the HvCaLB1.
The centromere and pericentromere regions have been demonstrated as hotspots for gene generation and initiation in plants. Genes within centromere region exhibit an evolutionary trend of escaping outward (Tong et al. 2020; Zhang et al. 2018b). HvCaLBs tend to be located in the noncentromeric regions, and even in the telomeric region (Fig. 1B). Thus, it was proposed that HvCaLBs have undergone a relatively longer evolutionary journey and possess more stable functional properties compared to those in the centromere and pericentromere regions. Alternatively, the gene pairs HORVU5Hr1G052010/HORVU5Hr1G052030, HORVU6Hr1G091350/HORVU6Hr1G091440, HORVU3Hr1G086220/HORVU3Hr1G086230/HORVU3Hr1G086240 were detected within a 100 kb physical distance on chromosomes. The Smith-Waterman algorithm (https://www.ebi.ac.uk/Tools/psa/emboss_water/) was used to determine the similarity of the sequences of the gene pairs, which were found to be 100%, 100%, and 92.9%, respectively (Mcgregor et al. 2017). Thus, the aforementioned gene pairs are considered tandem repeat genes, which may play crucial roles in barley evolution and whole-genome duplication events (Schaper and Anisimova 2015; Marchant et al. 2022).
Micro-collinearity analysis of HvCaLB1 among 7 representative grass species, including Hordeum vulgare (two versions), Zea mays, Oryza sativa Japonica, Brachypodium distachyon, Avena strigosa, Aegilops tauschii, and Triticum aestivum, revealed that CaLBs in these species are highly conserved. Favorable inter-genomic collinearity of the representative chromosomes was found among the species based on the distribution of CaLBs, indicating that segmental duplications (SD) and large genomic rearrangements (LGR) did not occur extensively after the origination of the CaLBs (Fig. 1C). Regarding the gene structure, CaLB genes have a similar number and arranged principles of introns and exons, but the coding sequence (CDS) length of HvCaLB1 is relatively longer. Additionally, the evolutionary branch of HvCaLB1 is located at the periphery of the tree and manifested longest, indicating that HvCaLB1 has a relatively distant evolutionary relationship to homologous genes and exhibits the highest sequence variability (Fig. 1D).
CaLB1s showed early evolution from chlorophyte algae
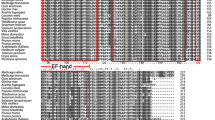
Putative orthologues of CaLB1 have been identified from approximately 82% of the 1,322 plants and algal species presented in the OneKP database (Leebens et al. 2019) (Fig. 2A). Our evolutionary analysis suggests that the homologues of HvCaLB1 were present in chlorophytes and rhodophyta, indicating an earlier evolution predating the emergence of green plants. For instance, CaLB1s in a chlorophyte alga, Pteromonas angulosa (Chlamydomonadales), showed high protein sequence similarity to a streptophyte alga, Coleochaete irregularis (Coleochaetales) (Fig. 2B). Interestingly, two C2 domains were observed in KsCaLB1 derived from the semiterrestrial streptophyte alga Klebsormidium subtile (Fig. 2C). The variations in the amino acid sequences and structural variability of CaLB1s were minimal in the examined basal land plants, including hornworts (Calypogeia fissa), liverworts (Porella navicularis), and mosses (Niphotrichum elongatum) (Fig. 2B, C). However, the similarity of CaLB1s in mosses and lycophytes to angiosperms is lower compared to hornworts, liverworts, and other examined vascular plants (Fig. 2C). The protein sequence similarity of fern CaLB1 to those in seed plants exceeded 50%, and the sequence similarity of CaLB1s between monocots and eudicots further reached more than 60% (Fig. 2B). In addition, the numbers of C2 domains exhibited a sustained increasing trend during the evolution of green plants. The number of C2 domains in streptophyte increased rapidly, which might be attributed to the increased number of C2 domains in the proteins and whole-genome duplication events (Figure S2). These analyses revealed that CaLBs and C2 domains likely evolved early in green algae and play crucial roles in plant adaptation to terrestrial living and potential tolerance to abiotic stresses.
Molecular evolutionary analysis of CaLB1 in plants and algae (A) Phylogenetic trees of CaLB1 protein in representative species of the major lineage of green plants using OneKP and NCBI databases. The tree is constructed based on the ML method. Clades are indicated by different colors. (B) Sequence alignment of the C2 domains among CaLB1 proteins from 12 representative green plant species. (C) Predicted 3D structures of CaLBs in representative plant and algal species. Hv, Hordeum vulgare; Os, Oryza sativa; At, Arabidopsis thaliana; Tc, Taiwania cryptomerioides; Pv, Pteris vittate; Pd, Phylloglossum drummondii; Ne, Niphotrichum elongatum; Pn, Porell navicularis; Cf, Calypogeia fissa; Ks, Klebsormidium subtile; Pa, Pteromonas angulosa; Ci, Coleochaete irregularis
Silencing of HvCaLB1 significantly affects plant growth under saline conditions
Previous quantitative trait loci (QTL) mapping analysis using a barley double haploid (DH) population derived from two genotypes with contrasting salt tolerance, CM72 (salt-tolerant) and Gairdner (salt-sensitive), identified eleven significant QTLs for various traits under saline and control conditions. A locus on chromosome 2 H near the QLT-C.CmGa.2 H marker, within a 17.99 Mb region, was identified as a potential regulator of both ROS and salt stress tolerance in barley (Liu et al. 2017b). Among the candidate genes located in the QTL region, HORVU2Hr1G091320, named as HvCaLB1, was selected for further analysis in this study. Constitutive expression of HvCaLB1 was observed in the roots of barley seedlings with or without 300 mM NaCl treatment, however, the transcription of HvCaLB1 was significantly upregulated by salt in the leaves (Figure S1) (Nefissi Ouertani et al. 2021; Yousefirad et al. 2020).
In this study, the transcripts of HvCaLB1 in the salt-tolerant parent CM72 were suppressed through BSMV-induced gene silencing to elucidate its physiological roles. The expression of HvCaLB1 in the root was slightly higher than that in the leaf under the same treatment (Fig. 3B). Without salt treatment, the expression levels of HvCaLB1 decreased by approximately 84.9% and 82.8% in the leaf and root of HvCaLB1-silencing lines compared to those of EGFP-silencing plants, respectively (Fig. 3B). Furthermore, increased expression levels of HvCaLB1 were observed in both of the roots and leaves when the plants were subjected to salt stress (Fig. 3B).
Plant growth of the EGFP - and HvCaLB1-silencing barley lines. (A) Phenotype after silencing of EGFP and HvCaLB1 in cultivated barley CM72 via BSMV-VIGS. Three concentration gradients were set for the salt treatment experiments, 0, 200, and 400 mM, respectively. Totally, six groups of plants were set up: CK, BSMV:EGFP under 0 mM; C200, BSMV:EGFP under 200 mM; C400, BSMV:EGFP under 400 mM; γ:CK, BSMV: HvCaLB1 under 0 mM; γ200, BSMV: HvCaLB1 under 200 mM; γ400, BSMV: HvCaLB1 under 400 mM. Scale bar = 5 cm. (B) Tissue expression profiles of HvCaLB1 under 3-days salt treatment using real-time PCR analysis. (C) Dry weight measurement of EGFP and HvCaLB1 silencing lines after 12-days salt treatment. The numerical value represents the decreased proportion of the treated group to CK subject to salt. (D) Plant height and root length determination of EGFP and HvCaLB1 silencing lines after 12-days salt treatment. Data are mean ± SD (n = 6). Different small letters indicate significant difference at P < 0.05 by one-way ANOVA test
The EGFP- and HvCaLB1-silencing plants showed no significant difference under the control condition without salt (Fig. 3A, C, D). However, both EGFP- and HvCaLB1-silencing plants displayed significantly retarded growth with the salt treatments for 12 days. The reduction was more remarkable in the plants exposed to 400 mM NaCl than those exposed to 200 mM NaCl (Fig. 3A). Furthermore, the HvCaLB1-downregulated lines displayed more wilted and chlorosis leaves and less dry weight than those of the control when the plants were cultured in the salt treatments (Fig. 3A, C, D). The phenotype was mostly consistent with the dry weight, plant height, and root length of the examined lines (Fig. 3C, D). With the supplement of 200 mM NaCl, the dry weight of leaves and roots of the EGFP-silencing plants decreased by approximately 60% compared to those without salt treatment; the reduction was up to 70% under 400 mM salt. On the other hand, the dry weight of leaves and roots of HvCaLB1-silencing plants decreased by approximately 40% and 90% with the treatments of 200 and 400 mM NaCl, respectively, compared to those without salt stress (Fig. 3C). The plant height and root length of both control and HvCaLB1-silencing plants were reduced by salt treatments. The significantly inhibited root growth attributed to the suppression of HvCaLB1 under salt stress was particularly noticeable in the presence of 400 mM NaCl (Fig. 3D). As a result, silencing of HvCaLB1 significantly impeded plant growth under saline conditions, particularly when the plants were exposed to higher levels of NaCl.
Knockdown of HvCaLB1 reduces photosynthesis parameters under salt stress
Photosynthesis, the critical physiological process in all green plants, is severely affected by various abiotic stresses in all its phases (Muhammad et al. 2021). In Arabidopsis thaliana, AtCaLB specifically bound to the promoter of the thalianol synthase gene (AtTHAS1), a light-induced protein involved in photosynthesis (de Silva et al. 2011), suggesting the potential role of CaLBs in the regulation of photosynthesis. Under the control condition without salt, significant reduction in photosynthetic properties, including the net photosynthetic rate (A), the transpiration rate (Emm), and the stomatal conductance (Gsw and Gtc), were found in the HvCaLB1-silencing barley plants compared to those of EGFP-silencing lines (Fig. 4). The parameter including A, Emm, Gsw, and Gtc of HvCaLB1-silencing plants were not affected by 200 mM salt compared to those under control, however, these parameters were dramatically decreased when the plants were treated with 400 mM salt. By contrast, the parameters of EGFP-silencing plants under 200 mM and 400 mM salt conditions were almost the same and significantly lower than those without salt (Fig. 4). These results indicate that the knockdown of HvCaLB1 suppressed the photosynthesis. The reduction was more pronounced under by 400 mM salt stress compared to 200 mM. However, the photosynthesis in control plants was more sensitive to 200 mM salt stress.
Photosynthetic properties of the EGFP - and HvCaLB1 -silencing barley lines. After the plants completed salt stress, the three main photosynthetic parameters (A) Net Photosynthetic Rate, (B) Transpiration Rate, and (C) Stomatal Conductance of the leaves were measured. Gsw: stomatal conductivity to water vapor; Gtc: stomatal conductivity to CO2. Data are mean ± SD (n = 8). Black dots represent actual data. Different small letters indicate significant difference at P < 0.05 by one-way ANOVA test. Significance (*) means P < 0.05 by independent-sample t test
The chlorophyll content in the leaves of EGFP- and HvCaLB1-silencing plants was estimated using SPAD, and it was decreased with the increasing concentrations of external NaCl. Under control and treatment with 200 mM NaCl for 12 days, the SPAD value showed no difference between EGFP- and HvCaLB1-silencing plants. However, a significant reduction was observed in the HvCaLB1-silencing plants subjected to 400 mM NaCl compared to the control (Fig. 5A). The key chlorophyll fluorescence parameter Fv/Fm decreased in both EGFP- and HvCaLB1-silencing plants when subjected to 400 mM but not 200 mM NaCl for 12 days, compared to the plants without salt treatment (Fig. 5B). The results indicated that 400 mM NaCl significantly reduced the primary light energy conversion efficiency.
Chlorophyll and chlorophyll fluorescence parameters of the EGFP - and HvCaLB1 -silencing barley lines after 12-day treatment. After the plants completed 12-day salt stress, Chlorophyll and chlorophyll fluorescence were measured, including (A) relative content of plant chlorophyll (SPAD value) and (B) chlorophyll fluorescence (Fv/Fm value). Data are mean ± SD (n = 6). Black dots represent actual data. Different small letters indicate significant difference at P < 0.05 by one-way ANOVA test
Silencing of HvCaLB1 significantly disrupts the K and na homeostasis under salt condition
Salt stress always leads to disrupted homeostasis of cations, especially Ca and K in plant tissues (Tanveer and Yousaf 2020). Along with the dramatic increase in Na content in the leaves and roots of the plants subjected to salt stress, significantly reduced K concentrations were observed in the examined tissues of both control and HvCaLB1-silencing lines (Fig. 6A, C). More Na and K were accumulated in the leaves than in the roots when the plants were treated with NaCl (Fig. 6B, D). Compared to the EGFP-silencing plants, the Na content in the leaves of the HvCaLB1-silencing plants decreased in the presence of 400 mM salt, while a reduction in K content in the leaves was observed when the plants were subjected to 200 mM NaCl (Fig. 6A, C). No significant difference in Na content was observed in the roots of the EGFP- and HvCaLB1-silencing plants, but dramatically reduced K content was detected in the roots of HvCaLB1 knockdown lines under 200 mM NaCl (Fig. 6A, C). Thus, the Na/K ratio in the roots of the knockdown lines of HvCaLB1 under 400 mM salt treatment was 14.80, whereas the value was 5.47 in the EGFP-silencing plants (Fig. 6E), indicating severely disrupted ion homeostasis in HvCaLB1-silencing plants.
Mineral elements analysis of the EGFP - and HvCaLB1 -silencing barley plants. Changes in mineral element content in plants after salt stress. (A) Na in leaves and roots, (B) Leaf/root ratio of Na, (C) K in leaves and roots, (D) Leaf/root ratio of K, (E) Na/K ratios in leaves and roots, (F) Ca in leaves and roots, and (G) Leaf/root ratio of Ca. Data are mean ± SD (n = 6). Black dots represent actual data. Different small letters indicate significant difference at P < 0.05 by one-way ANOVA test. Significance (*) represents P < 0.05 and *** represents P < 0.001 by independent-sample t test
The concentrations of phosphorus (P) and magnesium (Mg) in leaves and roots were also determined. Decreased leaf P and increased root P levels were observed in the EGFP-silencing plants when treated salt compared to those without NaCl. No significant difference was detected in the leaves and roots of the HvCaLB1-silencing plants under the three conditions, except for a significant increase of root P under 400 mM NaCl (Figure S3A). The Mg content decreased in the leaves of both control and HvCaLB1-silencing plants under 200 mM NaCl compared to those under control conditions. However, the Mg content increased when the external NaCl concentration was raised to 400 mM (Figure S3B). Furthermore, a significant increase in Mg content was detected in the roots of EGFP-silencing plants in response to NaCl. However, HvCaLB1-silencing plants exhibited lower root Mg levels compared to the control when exposed to 400 mM NaCl (Figure S3B).
Suppression ofHvCaLB1 disrupts the Ca2+ and ROS homeostasis and expression of key genes in roots under salt treatment
HvCaLB1, possessing a single C2 domain containing a cluster of aspartic acid residues that provide negative charge potential for Ca2+ binding, was considered a putative Ca2+-binding protein (Corbin et al. 2007). Here, we found that silencing HvCaLB1 significantly reduced Ca content by 68.03% in the leaves without affecting root Ca content under control conditions (Fig. 6F). NaCl treatments significantly reduced the leaf Ca content in both control and HvCaLB1-silencing plants. Surprisingly, 400 mM NaCl significantly increased root Ca in EGFP-silencing plants by 15.84-fold with no changes to root Ca content in HvCaLB1-silencing plants (Fig. 6F).
The involvement of HvCaLB1 in Ca signaling in the cells of the root tips of the EGFP- and HvCaLB1-silencing plants under 200 mM salt was further validated using a Ca2+ indicator, Fura-2 pentakis (acetoxymethyl) ester (Fura-2 AM), which can be transformed to Fura-2 by esterase once enters into the cell (Chen et al. 2010). Fura-2 could bind with free Ca2+ and produce strong fluorescence under 340 nm excitation light (F340), while the weak signal captured under the 380 nm excitation light (F380) indicates the background. In this way, the F340/F380 ratio represents the intracellular Ca2+ concentration (Chen et al. 2010; Ghosh et al. 2023). Mean F340/F380 ratio was estimated and statistically analyzed for 160 s from a band of pixels equivalent around the eight selected cells of root tips from HvCaBL1- and EGFP-silencing plants to estimate the [Ca2+]i near the plasma membrane, respectively (Fig. 7A, B). Time-course measurements of the F340/F380 ratio demonstrated the real-time changes in apical cells of HvCaLB-1 and EGFP-silencing plants every 10 s. The range of dispersion and kurtosis of [Ca2+]i in the roots of EGFP-silencing plants were higher than those of the HvCaLB1-silencing plants under 200 mM salt treatment. In addition, a significantly reduced [Ca2+]i signal intensity displayed as the ratio of F340/F380 fluorescence was observed in the root tips of HvCaLB1-silencing compared to that of EGFP-silencing plants under 200 mM NaCl (Fig. 7B). Moreover, the significant difference between the two genotypes was observed after 100 s (Figure S4).
Detection and imaging of ROS and Ca2+ signals in roots of the EGFP - and HvCaLB1-silencing plants. (A) Ratiometric Ca2+ imaging in roots was performed using the calcium-sensitive dye Fura-2 AM, eight cells labeled with red curves from fluorescence image pairs collected using a 535-nm interference filter after exciting with 340- and 380-nm light indicated the selected one for [Ca2+]i estimation by using high-speed intracellular ion scale imaging system (B) The mean F340/F380 values in within 2-µm of the periphery of each apical cell at each time point were calculated and statistically presented in a normalized context, and violin plots were plotted based on the actual distribution and dispersion of the values. (C) H2DCF-DA probe was used to detect salt-induced ROS accumulation in roots of seedings with excitation at 488 nm and emission at 505 to 525 nm. (D) Fluorescence emission intensity was estimated and calculated as average grey level value per pixel and corrected for background by ImageJ software. Data are mean ± SD of at least three different experiments and are from three separate root preparations. *, p < 0.05, and ***, p < 0.001 vs. the corresponding control values. vs. the corresponding control values
It’s revealed that Ca2+ interacts with the other intracellular secondary messengers in response to abiotic stresses (Wang et al. 2013; Liu et al. 2021). Therefore, we measured the accumulation of ROS of in the root tips by staining with H2DCF-DA, following the methods described in our previous publications (Cheng et al. 2023; Wang et al. 2017; Zhao et al. 2019; Fan et al. 2023). Compared to the root tips without salt, increased ROS accumulation was observed in plants exposed to NaCl. In addition, a significantly higher level of ROS was clearly shown in the roots of HvCaLB1-silencing lines with the treatment of 400 mM NaCl (Fig. 7D, E).
Additionally, we measured the expression of three key genes (HvHKT1;5, HvHVP10, and HvCaM1) that are known to play important roles in plant salt tolerance (Huang et al. 2020; Shen et al. 2021; Fu et al. 2022). HvHKT1;5, a negative regulator of salt tolerance in barley, was highly upregulated by salinity in both roots and leaves but showed no difference between HvCaLB1 and EGFP-silencing plants (Fig. 8). However, the expression patterns of root HvHVP10 and HvCaM1 showed significant differences between HvCaLB1- and EGFP-silencing plants at 400 mM NaCl (Fig. 8).
Expression of key genes involved in the salt response and Ca2+signalling in shoots and roots of theEGFP- and HvCaLB1-silencing barley plants. Tissue expression profiles of HvHKT1;5, HvP10 and HvCaM1 were counted and plotted under 12-day salt treatment using real-time PCR analysis. Data are mean ± SD (n = 5). Black dots represent actual data. Different small letters indicate significant difference at P < 0.05 by one-way ANOVA test
Discussion
CaLBs represent a group of ancient Ca2+-binding proteins originating from chlorophyte algae
Ca2+-binding proteins are a class of proteins that specifically bind to [Ca2+]i. This binding has been demonstrated in signaling transduction in cells under various stresses (Dominguez et al. 2015; Ketehouli et al. 2022; Liu et al. 2017; Luan and Wang 2021). The C2 domain, a Ca2+-binding motif, is considered a useful model for understanding early evolutionary processes. It is one of the most prevalent lipid-binding domains, expanded and distributed in diverse functional contexts (Zhang and Aravind 2010). The C2 domain is ancient and can be found throughout eukaryotes in a wide variety of proteins (Farah and Sossin 2012). In this study, C2 domain-containing CaLBs were conserved within the grass family through comprehensive analyses, which was consistent with a recent report (Niu et al. 2022). During key evolutionary stages, tandem repeats and positive selection pressure potentially jointly promote the expansion of the CaLB gene family (Hao et al. 2020; Niu et al. 2022; Tiwari and Paliyath 2011).
Our results revealed that CaLBs originated from algae and are abundantly presented in plants across all evolutionary clades. However, the lipid-binding and membrane interaction mode of the C2, as well as the number of C2 domain-contained proteins, were found to be divergent among the examined species (Marchadier et al. 2016; Zhang and Aravind 2010). The quantity of C2 domains appears to increase in algae and rises sharply in bryophytes and ferns. This is accompanied by progressive increases in protein sequences and structural similarities, which may provide the genetic basis for the pivotal transition from aquatic to terrestrial plants, from non-vascular to vascular plants, and from sporophytes to seed plants (Duckett and Pressel 2020; Ishizaki 2017). Our results suggest that C2 domain-containing proteins, such as CaLBs, are likely presented in almost all green plants, despite some divergence in certain species.
Calcium (Ca) was suggested as a “promoter” in the evolution of early life forms (Kazmierczak et al. 2013). The Ca2+-binding motif is widely distributed within classical Ca sensors, such as EF-hand from CaMs, CMLs, CBLs, CIPKs, and CDPKs (Zeng et al. 2017; Mohanta et al. 2019; Tong et al. 2021). Our previous study revealed that CMLs, CaMs, and CIPKs could be traced back to basal lineages of streptophyte algae. CBLs originated from the unicellular species of marine green alga Ostreococcus, while the earliest CDPKs could be found in all green plant lineages and red algae (Tong et al. 2021). Therefore, the origin of CaLBs represents a group of ancient Ca2+ binding proteins that may have emerged in parallel with the appearance of CDPKs in plants (Cui et al. 2023; Hu et al. 2021; Zhang et al. 2022). Numerous reports have demonstrated that streptophyte algae are the sister group of all land plants, and many key protein families for abiotic stress tolerance have originated from streptophyte algae (Zhong et al., 2014; Wang et al. 2020; Zhao et al. 2019; Liu et al. 2021b; Cai et al. 2021; Deng et al. 2021, 2022). However, we found that HvCaLB1 originated from chlorophyte algae instead of from streptophyte algae. One plausible explanation is that CaLB1 is a highly conserved gene that originated from chlorophyte algae, similar to members of protein pumps (e.g., AHAs, VHAs; Chen et al. 2017) and CDPKs (Tong et al. 2021). Another possible reason is that HvCaLB1 was a unique member of the CaLB family that may have evolved earlier from chlorophyte algae than other members of CaLBs, which were later duplicated during the evolution of angiosperms.
CaLBs encode potential ca sensors for salt stress in plants
Ca2+ acts as a crucial second messenger in regulating plant responses to abiotic stresses (Lee and Seo 2021). The conserved functions of the classical Ca2+ sensors in response to salt stress have been revealed in various green plants (Tansley et al. 2022; Shen et al. 2020; Drerup et al. 2013; Dong et al. 2020; Kamachi et al. 2004). The C2 domain of PKCs was considered a Ca sensor, and its Ca2+-dependent binding to phospholipids was thought to be crucial for kinase activation (Farah and Sossin 2012). The C2 domain was also shown to interact with the receptor for activated C-kinase (RACKs), thus contributing to the subcellular localization of PKCs. This indicates their critical roles in sensing the activated signaling pathway in response to external stimuli (Farah and Sossin 2012; Fu et al. 2019; Rupwate and Rajasekharan 2012). Therefore, the widespread presence of the C2 domain-contained CaLBs in green plants was suggested to play pivotal roles in decoding Ca2+ signals.
During the last few decades, the involvement of rice C2 domain-containing proteins, OsSMCP1 and OsC2DP, in response to salt stress has been demonstrated (Fu et al. 2019; Yokotani et al. 2009). OsC2DP was predominantly expressed in the root, and its expression was significantly down-regulated in response to salt stress. Overexpression of OsSMCP1 in Arabidopsis showed enhanced tolerance to high salinity, but the molecular mechanism had not been discovered (Yokotani et al. 2009). OsC2DP was localized in the cytosol but could be translocated to the plasma membrane under salt stress, indicating the potential function of the C2 domain in membrane targeting. Knockout of OsC2DP increased salt sensitivity and Na+ concentration in the shoot through the indirect downregulation of OsSOS1 and OsNHX4 (Fu et al. 2019). Therefore, CaLB is possible to be translocated from the cytosol to the plasma membrane through the C2 domain, a physiological process employing Ca2+ signal pathway in regulation of gene expression in plants (Fu et al. 2019; de Silva et al. 2011).
In this study, the expression level of HvCaBL1 was also highly expressed in the root, and the transcript was induced by salt stress in barley (Fig. 3B). Similar to the loss-of-function of OsC2DP (Fu et al. 2019), knockdown of HvCaBL1 also increased salt sensitivity. This is evident from the decreased dry weight, inhibited root length, reduced chlorophyll content, and chlorophyll fluorescence compared to the EGFP-silencing plants under salt stress. Knockout of OsC2DP and knockdown of HvCaBL1 hardly affect Na+ accumulation in the roots, but significantly reduced Na+ concentration in the leaves. Moreover, the Na/K ratio, one of the most important indicators of salinity tolerance (Munns et al. 2020; Chen et al. 2016), was significantly higher in both leaves and roots of the knockdown lines of HvCaLB1 compared to the EGFP-silencing plants, especially under 400 mM salt. These findings implicate that HvCaLB1 plays a crucial role as a positive regulator in salt tolerance in barley. However, the differential accumulation of Na+ and K+ in the leaves suggests varied regulatory mechanisms of OsC2DP (Fu et al. 2019) and HvCaBL1 on salt tolerance, even though both genes act as positive regulators of salt tolerance. Slightly upregulated expression of HVP10 and HvCaM1 in the roots of HvCaLB1-silencing lines (Fig. 8) implicated the potential linking between the 2 known salt-responsive proteins and HvCaLB1 (Shen et al. 2020; Fu et al. 2022).
[Ca2+]i and ROS are two main second messengers in plant cells in response to a variety of stresses (Cheng et al. 2023; Liu et al. 2021b), including salt stress (Yang and Guo 2018; Zelm et al. 2020). Knockdown of HvCaBL1 significantly reduced Ca contents in the leaves even under control conditions, indicating the potential importance of HvCaBL1 in Ca homeostasis. On the other hand, the Ca content in the roots of HvCaLB1-silencing plants was dramatically decreased compared to that of the EGFP-silencing lines under salt stress, which was consistent with the reduced cytosolic Ca2+ signal intensity in the root tips of HvCaLB1-silencing plants (Fig. 7). Furthermore, silencing HvCaLB1 significantly disrupts the K and Na homeostasis in plants under salinity. The results implicating the role of HvCaLB1 in response to salt stress might rely on the remodeling of ion homeostasis mediated by salt-induced Ca2+ waves, as revealed in previous reports (Choi et al. 2014; Manishankar et al. 2018). In contrast to the decreased [Ca2+]i in the roots, an enhanced ROS signal was observed in the root tip of HvCaLB1-silencing plants. These results demonstrate that HvCaLB1 is involved in Ca and ROS signaling pathways, which are critical for plants in response to salt stress (Ali et al. 2023; Zelm et al. 2020).
Conclusion
The single C2 domain-contained Ca2+-dependent lipid-binding proteins (CaLBs) may have evolved early from chlorophyte algae. Silencing HvCaLB1 can significantly reduce photosynthesis and hinder the morphology of barley under saline conditions, particularly under high salt stress. Taken together, we found that HvCaLB1 represented a group of ancient Ca2+ binding proteins that originated from chlorophyte algae and functioned as potential Ca2+ sensors, positively regulating salt tolerance in plants. The findings provide new insights into the regulation of CaLBs in crop breeding for enhanced salt tolerance.
Materials and methods
Plant materials and growth conditions
A salt-tolerant barley genotype, CM72 (Chen et al. 2016), was used in this study. Barley stripe mosaic virus (BSMV) was employed for the virus-induced gene silencing of CM72. BSMV consists of tripartite short rod-shaped RNA strands: α, β, and γ (He et al. 2015; Feng et al. 2020; Jiang et al. 2023). The cDNA fragments of the enhanced green fluorescent gene (EGFP, 250 bp) and HvCaLB1 (435 bp) were amplified through RT-PCR (γ-CaLB-F: ttttttttttttttagctagcATGGCGCAGGGGACGCT, γ-CaLB-R: gattcttcttccgttgctagcTCAAGATGACTGGTTCCACCCAC, γ-EGFP-F: ttttttttttttttagctagcATGGTGAGCAAGGGCGAGG, γ-EGFP-R: gattcttcttccgttgctagcCTTGTACAGCTCGTCCATGC), and then inserted into the empty RNAγ vectors using the seamless cloning kit (Toroivd, China) to generate BSMV: EGFP and BSMV: HvCaLB1 constructs, respectively. BSMV: HvCaLB1 was used to suppress the expression of HvCaLB1, while BSMV: EGFP was utilized to generate the control plants. In this study. The 10-d-old barely seedlings were cultured in an incubator (22/18°C, day/night) using a modified Hoagland’s solution (Feng et al. 2020) before being inoculated with RNA virus. The plants incubated with BSMV: HvCaLB1 were classified in the γ group, while the seedlings co-cultured with BSMV: EGFP-harboring Agrobacterium were used as the CK group. And then, the plants were treated with 0 mM, 200 mM, and 400 mM NaCl for 12 days. Each treatment was replicated four times, with each replicate consisting of three seedlings.
Genome-wide identification and evolutionary bioinformatics of CaLB1s
The full-length cDNA of HvCaLB1 (HORVU2Hr1G091320) was cloned from the barley cultivar Golden Promise (GP). Candidate orthologs of HvCaLB1 proteins were identified using the 1000 Plant Transcriptome database (OneKP) (https://db.cngb.org/blast/blast/blastp/) (Deng et al. 2021, 2022; Chen et al. 2021; Li et al. 2023; Hu et al. 2020). After multiple alignments by MAFFT software with the default setting (https://mafft.cbrc.jp/alignment/server/), the phylogenetic tree was constructed using the RAxML-HPC2 Workflow in CIPRES Science Gateway V3.3 with 1000 bootstraps (https://www.phylo.org/) (Miller et al. 2010; Jiang et al. 2024). Subsequently, iTOL V5.0 was utilized for visual rendering (https://itol.embl.de/) (Letunic and Bork 2021). Furthermore, the C2 domains were manually selected and analyzed using BioEdit (http://www.mbio.ncsu.edu/bioedit/bioedit.html) and visualized with Jalview (http://www.jalview.org/) (Waterhouse et al. 2009). 3D structures were predictably constructed using the SWISS-MODEL website (https://swissmodel.expasy.org/interactive) (Mather 2014). In addition, the integration of the Hidden Markov Model and Superfamily databases (https://supfam.mrc-lmb.cam.ac.uk/SUPERFAMILY/index.html) enabled the calculation and fitting of the quantity of C2 domains in 35 representative plants and algal species from 11 green plant clades.
In light of the conservation of C2 domain evolution, the corresponding Hidden Markov Model profiles (Accession No. PF00168) were obtained from the Pfam database (http://pfam.xfam.org/) (Finn et al. 2008). Local search for CaLB gene families was conducted in barley, rice, and Arabidopsis using the hmmsearch package with model profiles as keywords (E-value < 10− 5) (Finn et al. 2015). Distribution of HvCaLB genes in chromosomes was required from the Triticeae-Gene Tribe website, whose homology inference strategy also offered microcollinearity, evolutionary, and structural diagrams of the HvCaLB1 orthologs in relatives (http://wheat.cau.edu.cn/TGT/) (Chen et al. 2020). 62 members of barley CaLB genes were identified, and their expression in leaves and roots was obtained through publicly available databases on NCBI (https://www.ncbi.nlm.nih.gov/) and PlantExp (https://biotec.njau.edu.cn/plantExp/).
RNA isolation and quantitative real-time PCR
To confirm the silencing of HvCaLB1 in the incubated seedlings of CM72, leaves and roots from the inoculated seedlings were sampled after three days of exposure to 0, 200, and 400 mM NaCl, respectively. Total RNA was isolated using the TaKaRa MiniBEST Plant RNA Extraction Kit (Takara, Japan), and first-strand cDNA synthesis was carried out with the PrimeScipt™ RT reagent Kit (Takara, Japan). qRT-PCR was performed according to Liu et al. (2014) using the SYBR Green Supermix on a CFX96 system machine (Bio-Rad, USA). The primers used for gene expression analysis of HvCaLB1 were 5’-CGTCATCAAGCTCCTCGACA-3’ and 5’- ACAGTTGGTGGGATGCTTCC-3’, primers for HvHKT1;5 (ABK58096) were 5’-AGTGGAGCAGGGAAGGGAAG-3’ and 5’-ACTTCTTGAGCCTGCCGTAG-3’, primers for HVP10 (HORVU7Hr1G028910) were 5’-AACCCTCTTTGGCGTGGAAA-3’ and 5’-TGTGGCAGTCTGAACCCTTG-3’, primers for HvCaM1 (HORVU0Hr1G001270) were 5’-CATGATCAACGAGGTTGACG-3’ and 5’-GAGATGAAGCCGTTCTGGTC-3’. HvActin was used as an internal standard and the primers were 5’-CGTGTTGGATTCTGGTGATG-3’ and 5’-AGCCACATATGCGAGCTTCT-3’. The relative expression levels were expressed as 2− ρρCt method (Jiang et al. 2023; Chen et al. 2021).
Measurement of photosynthetic properties and chlorophyll fluorescence parameters
The photosynthetic properties, including Net photosynthetic rate (A), transpiration rate (Emm), stomatal conductivity to water vapor (Gsw), and total conductivity to CO2 (Gtc) of the third fully expanded leaves after a 12-day treatment, were measured using a portable photosynthesis system (LI-6800, Li-Cor Inc., Lincoln, USA). The measurement conditions were adjusted according to our previous report (Wang et al. 2023), that is flow rate: 500 mol·s− 1, PAR: 1500 mol·m− 2·s− 1, 400 mmol·mol− 1 CO2, and 60% relative humidity. At least eight replicates were performed for each treatment line.
The chlorophyll fluorescence parameters in the leaves of plants from both γ and CK lines were measured on the 12th day treated with salt. The real-time data of photochemical efficiency of PSII in the light (FV/FM) and the relative chlorophyll content were recorded using Chlorophyll fluorometer (OS30p+, Opti-Sciences, USA) (Wang et al. 2016), and chlorophyll meter (SPAD-502, Konica Minolta, Japan) (Wu et al. 2018), respectively.
Determination of mineral elements in leaves and roots
Measurement of mineral elements were conducted according to the method described by Cheng et al. (2023). The barley leaves and roots from hydroponic experiments were washed with deionized water for three times before being harvested and then dried in an oven. After weighing, the dried roots and leaves were digested with 5 ml of concentrated HNO3 and 10 µL H2O2 in a microwave (Multiwave 3000, Anton Paar GmbH, Australia). The digested solution was diluted with double-distilled water, and the content of Na, K, Ca, P, and Mg in the samples was determined simultaneously using an inductively coupled plasma mass spectrometer (ICP-MS, PerkinElmer, USA).
Detection and imaging of ROS and Ca2+ signals in roots
After 12 days of salt exposure, the ROS accumulation in root cells was determined using the fluorescent probe 2’,7’-dichlorodihydrofluorescein diacetate (H2DCF-DA), following the methods described in our previous publications (Cheng et al. 2023; Cai et al. 2021; Chen et al. 2016). Briefly, the roots of the seedlings were stained with 10 µM H2DCF-DA in 10 mM KCl buffered with 5 mM MES [2-(N-morpholino) ethanesulfonic acid] at pH 6.1 adjusted with Ca(OH)2 in the dark for 10 min and then washed three times with 5 mM MES solution at pH 6.1 (Wang et al. 2013). The fluorescence images were detected with excitation at 488 nm and emission at 505 to 525 nm by AZ100 fluorescence stereo microscope (Nikon, Japan). The intensity of ROS fluorescence in the examined roots was estimated using ImageJ (NIH, USA) after normalizing the background (Cai et al. 2021; Zheng et al. 2021; Chen et al. 2016). The experiment was conducted with three biological replicates.
Root staining with ratiometric Ca2+ indicators [Fura-2 pentakis (acetoxymethyl) ester, Fura-2 AM] was performed according to our previous reports (Chen et al. 2010; Wang et al. 2013; Kanchiswamy et al. 2014) with some modifications. Briefly, root tips of salt-free treated seedlings harvested from 22-days post-inoculation were stained with 2 µM Fura-2 AM for 4 hours, which was dissolved in the hydroponic solution employed (pH 6.0). Dye loading was deemed successful based on visual inspections for cytosolic dye distribution and stabilization of the fluorescence ratio signal. After three washes with 20 mM HEPES (N’-a-hydroxythylpiperazine-N’-ethanesulfanic acid) buffer (pH 7.0 adjusted with NaOH), ratiometric Ca2+ imaging of apical cells submerged in HEPES buffer with 200 mM NaCl was performed by high-speed intracellular ion scale imaging system (DG-4, USA). Eight selected independent cells (from eight independent experiments) had their measured [Ca2+]i values determined by fura-2 fluorescence ratio. The specific procedure was referred to the previous methodology (Chen et al. 2010; Wang et al. 2013), [Ca2+]i levels from HvCaLB1- and EGFP-silencing root apical cells (curves plotted in red) were determined from fluorescence ratios in the 2-µm band around the cell periphery. This region is typically subject to Ca2+ entry as well as endomembrane Ca2+ release. Fluorescence image pairs generated from the Ca2+-Fura-2 complex were collected at 10-s intervals using a 535-nm interference filter (± 20 nm bandwidth) after excitation with 340- and 380-nm light (10-nm slit width; Polychrom II; Til Photonics). Measurements were corrected for background using the average of three images collected before dye loading and were analyzed using software ImageJ and MetaFluor (UIC, USA) (Schneider et al. 2012; Ghosh et al. 2023; Wu et al. 2017; Jing et al. 2016; Li et al. 2015; Kanchiswamy et al. 2014; Vafadar et al. 2021). The experiment was conducted with three biological replicates.
Statistical analysis
Data were displayed as means with standard errors based on at least three independent biological replicates. Statistical analysis was conducted using the Statistical Product and Service Solutions software package (IBM, USA). T-test and One-way ANOVA followed by the Tukey’ s Honestly Significant Difference were used to evaluate the significance level of P < 0.05.
Data availability
Data will be made available on request.
References
Ali A, Petrov V, Yun D-J, Gechev T (2023) Revisiting plant salt tolerance: novel components of the SOS pathway. Trends Plant Sci 27:S1360–1385. https://doi.org/10.1016/j.tplants.2023.04.003
Bondada V, Gal J, Mashburn C, Rodgers DW, Larochelle KE, Croall DE, Geddes JW (2021) The C2 domain of calpain 5 contributes to enzyme activation and membrane localization. Biochim Biophys Acta Mol Cell Res 1868(7):119019. https://doi.org/10.1016/j.bbamcr.2021.119019
Cai S, Huang Y, Chen F, Zhang X, Sessa E, Zhao C, Marchant DB, Xue D, Chen G, Dai F, Leebens-Mack JH, Zhang G, Shabala S, Christie JM, Blatt MR, Nevo E, Soltis PS, Soltis DE, Franks PJ, Wu F, Chen ZH (2021) Evolution of rapid blue-light response linked to explosive diversification of ferns in angiosperm forests. New Phytol 230(3):1201–1213. https://doi.org/10.1111/nph.17135
Castro B, Citterico M, Kimura S, Stevens DM, Wrzaczek M, Coaker G (2021) Stress-induced reactive oxygen species compartmentalization, perception and signalling. Nat Plants 7(4):403–412. https://doi.org/10.1038/s41477-021-00887-0
Chen ZH, Chen G, Dai F, Wang Y, Hills A, Ruan YL, Zhang G, Franks PJ, Nevo E, Blatt MR (2017) Molecular evolution of grass stomata. Trends Plant Sci 22(2):124–139. https://doi.org/10.1016/j.tplants.2016.09.005
Chen ZH, Hills A, Lim CK, Blatt MR (2010) Dynamic regulation of guard cell anion channels by cytosolic free Ca2+ concentration and protein phosphorylation. Plant J 61(3):816–825. https://doi.org/10.1111/j.1365-313X.2009.04108.x
Chen ZH, Wang Y, Wang JW, Babla M, Zhao C, Garcia-Mata C, Sani E, Differ C, Mak M, Hills A, Amtmann A, Blatt MR (2016) Nitrate reductase mutation alters potassium nutrition as well as nitric oxide-mediated control of guard cell ion channels in Arabidopsis. New Phytol 209(4):1456–1469. https://doi.org/10.1111/nph.13714
Chen Y, Song W, Xie X, Wang Z, Guan P, Peng H, Jiao Y, Ni Z, Sun Q, Guo W (2020) A collinearity-incorporating homology inference strategy for connecting emerging assemblies in the Triticeae Tribe as a pilot practice in the plant pangenomic era. Mol Plant 13(12):1694–1708. https://doi.org/10.1016/j.molp.2020.09.019
Chen X, Jiang W, Tong T, Chen G, Zeng F, Jang S, Gao W, Li Z, Mak M, Deng F, Chen ZH (2021) Molecular Interaction and Evolution of Jasmonate Signaling with Transport and Detoxification of Heavy Metals and metalloids in plants. Front Plant Sci 12:665842. https://doi.org/10.3389/fpls.2021.665842
Chen X, Zhao C, Yun P, Yu M, Zhou M, Chen ZH, Shabala S (2023) Climate-resilient crops: lessons from xerophytes. Plant J (in Press). https://doi.org/10.1111/tpj.16549
Cheng J, Zhang S, Yi Y, Qin Y, Chen Z-H, Deng F, Zeng F (2023) Hydrogen peroxide reduces root cadmium uptake but facilitates root-to-shoot cadmium translocation in rice through modulating cadmium transporters. Plant Physiol Biochem 200(2):107754. https://doi.org/10.1016/j.plaphy.2023.107754
Choi W-G, Toyota M, Kim S-H, Hilleary R, Gilroy S (2014) Salt stress-induced Ca2 + waves are associated with rapid, long-distance root-to-shoot signaling in plants. Proceedings of the National Academy of Sciences 111(17): 6497–6502. https://doi.org/10.1073/pnas.1319955111
Corbin JA, Evans JH, Landgraf KE, Falke JJ (2007) Mechanism of specific membrane targeting by C2 domains: localized pools of target lipids enhance Ca2+ affinity. Biochemistry 46(14):4322–4336. https://doi.org/10.1021/bi062140c
Cui M, Gupta SK, Bauer P (2023) Role of the plant-specific calcium-binding C2-DOMAIN ABSCISIC ACID-RELATED (CAR) protein family in environmental signaling. Eur J Cell Biol 102(2):151322. https://doi.org/10.1016/j.ejcb.2023.151322
de Silva K, Laska B, Brown C, Sederoff HW, Khodakovskaya M (2011) Arabidopsis thaliana calcium-dependent lipid-binding protein (AtCLB): a novel repressor of abiotic stress response. J Exp Bot 62(8):2679–2689. https://doi.org/10.1093/jxb/erq468
Deng F, Zeng F, Chen G, Feng X, Riaz A, Wu X, Gao W, Wu F, Holford P, Chen ZH (2021) Metalloid hazards: from plant molecular evolution to mitigation strategies. J Hazard Mater 409:124495. https://doi.org/10.1016/j.jhazmat.2020.124495
Deng F, Zeng F, Shen Q, Abbas A, Cheng J, Jiang W, Chen G, Shah AN, Holford P, Tanveer M, Zhang D, Chen ZH (2022) Molecular evolution and functional modification of plant miRNAs with CRISPR. Trends Plant Sci 27(6):890–907. https://doi.org/10.1016/j.tplants.2022.01.009
Dominguez DC, Guragain M, Patrauchan M (2015) Calcium binding proteins and calcium signaling in prokaryotes. Cell Calcium 57(3):151–165. https://doi.org/10.1016/j.ceca.2014.12.006
Dong H, Wu C, Luo C, Wei M, Qu S, Wang S (2020) Overexpression of MdCPK1a gene, a calcium dependent protein kinase in apple, increase tobacco cold tolerance via scavenging ROS accumulation. PLoS ONE 15:e0242139. https://doi.org/10.1371/journal.pone.0242139
Drerup MM, Schlucking K, Hashimoto K, Manishankar P, Steinhorst L, Kuchitsu K, Kudla J (2013) The calcineurin B-like calcium sensors CBL1 and CBL9 together with their interacting protein kinase CIPK26 regulate the Arabidopsis NADPH oxidase RBOHF. Mol Plant 6(2):559–569. https://doi.org/10.1093/mp/sst009
Duckett JG, Pressel S (2020) Of mosses and vascular plants. Nat Plants 6(3):184–185. https://doi.org/10.1038/s41477-020-0619-1
Evans MJ, Choi WG, Gilroy S, Morris RJ (2016) A ROS-Assisted calcium Wave Dependent on the AtRBOHD NADPH oxidase and TPC1 Cation Channel propagates the systemic response to salt stress. Plant Physiol 171(3):1771–1784. https://doi.org/10.1104/pp.16.00215
Fan X, Tang H, Chen X, Zeng F, Chen G, Chen ZH, Qin Y, Deng F (2023) Allene oxide synthase 1 contributes to limiting grain arsenic accumulation and seedling detoxification in rice. Stress Biol 3(1):52. https://doi.org/10.1007/s44154-023-00136-8
Farah CA, Sossin WS (2012) The role of C2 domains in PKC signaling. Adv Exp Med Biol 740(16):663–683. https://doi.org/10.1007/978-94-007-2888-2_29
Feng X, Liu W, Qiu CW, Zeng F, Wang Y, Zhang G, Chen ZH, Wu F (2020) HvAKT2 and HvHAK1 confer drought tolerance in barley through enhanced leaf mesophyll H+ homoeostasis. Plant Biotechnol J 18(8):1683–1696. https://doi.org/10.1111/pbi.13332
Finn RD, Tate J, Mistry J, Coggill PC, Sammut SJ, Hotz HR, Ceric G, Forslund K, Eddy SR, Sonnhammer EL (2008) The pfam protein families database. Nucleic Acids Res 32(1):D138. https://doi.org/10.1093/nar/gkaa913
Finn R, Clements J, Arndt W, Miller B (2015) HMMER web server: 2015 update. Nucleic Acids Res 43(2):W30–W38. https://doi.org/10.1093/nar/gkv397
Fu S, Fu L, Zhang X, Huang J, Yang G, Wang Z, Liu YG, Zhang G, Wu D, Xia J (2019) OsC2DP, a novel C2 domain-containing protein is required for Salt Tolerance in Rice. Plant Cell Physiol 60(10):2220–2230. https://doi.org/10.1093/pcp/pcz115
Fu L, Wu D, Zhang X, Xu Y, Kuang L, Cai S, Zhang G, Shen Q (2022) Vacuolar H+-pyrophosphatase HVP10 enhances salt tolerance via promoting Na+ translocation into root vacuoles. Plant Physiol 188(2):1248–1263. https://doi.org/10.1093/plphys/kiab538
Ghosh S, Dahiya M, Kumar A, Bheir M, Pandey GK (2023) Calcium imaging: a technique to monitor calcium dynamics in biological systems. Physiol Mol Biol Plants 29:1777–1811. https://doi.org/10.1007/s12298-023-01405-6
Han Y, Yin S, Huang L, Wu X, Zeng J, Liu X, Qiu L, Munns R, Chen Z-H, Zhang G (2018) A Sodium Transporter HvHKT1;1 confers Salt Tolerance in Barley via regulating tissue and Cell Ion Homeostasis. Plant cell Physiol 59(6):1976–1989. https://doi.org/10.1093/pcp/pcy116
Hao P, Wang H, Ma L, Wu A, Chen P, Cheng S, Wei H, Yu S (2020) Genome-wide identification and characterization of multiple C2 domains and transmembrane region proteins in Gossypium hirsutum. BMC Genomics 21(1):445. https://doi.org/10.1186/s12864-020-06842-1
He X, Zeng J, Cao F, Ahmed IM, Zhang G, Vincze E, Wu F (2015) HvEXPB7, a novel beta-expansin gene revealed by the root hair transcriptome of tibetan wild barley, improves root hair growth under drought stress. J Exp Bot 66(22):7405–7419. https://doi.org/10.1093/jxb/erv436
Hu B, Deng F, Chen G, Chen X, Gao W, Long L, Xia J, Chen ZH (2020) Evolution of Abscisic Acid signaling for stress responses to toxic metals and metalloids. Front Plant Sci 11:909. https://doi.org/10.3389/fpls.2020.00909
Hu Q, Zeng M, Wang M, Huang X, Li J, Feng C, Xuan L, Liu L, Huang G (2021) Family-wide evaluation of multiple C2 domain and transmembrane region protein in Gossypium hirsutum. Front Plant Sci 25(12):767667. https://doi.org/10.3389/fpls.2021.767667
Huang L, Kuang L, Wu L, Shen Q, Han Y, Jiang L, Wu D, Zhang G (2020) The HKT Transporter HvHKT1;5 negatively regulates Salt Tolerance. Plant Physiol 182(1):584–596. https://doi.org/10.1104/pp.19.00882
Ishizaki K (2017) Evolution of land plants: insights from molecular studies on basal lineages. Biosci Biotechnol Biochem 81(1):73–80. https://doi.org/10.1080/09168451.2016.1224641
Jadidi O, Etminan A, Azizi-Nezhad R, Ebrahimi A, Pour-Aboughadareh A (2022) Physiological and molecular responses of Barley genotypes to salinity stress. Genes 13(11):2040. https://doi.org/10.3390/genes13112040
Jiang Z, Zhou X, Tao M, Yuan F, Liu L, Wu F, Wu X et al (2019) Plant cell-surface GIPC sphingolipids sense salt to trigger Ca2+ influx. Nature 572:341–346. https://doi.org/10.1038/s41586-019-1449-z
Jiang W, Tong T, Li W, Huang Z, Chen G, Zeng F, Riaz A, Amoanimaa-Dede H, Pan R, Zhang W, Deng F, Chen ZH (2023) Molecular Evolution of Plant 14-3-3 proteins and function of Hv14-3-3A in Stomatal Regulation and Drought Tolerance. Plant Cell Physiol 63(12):1857–1872. https://doi.org/10.1093/pcp/pcac034
Jiang W, He J, Babla M, Wu T, Tong T, Riaz A, Zeng F, Qin Y, Chen G, Deng F, Chen ZH (2024) Molecular evolution and interaction of 14-3-3 proteins with H+-ATPases in plant abiotic stresses. J Exp Bot 75(3):689–707. https://doi.org/10.1093/jxb/erad414
Jing P, Zou J, Kong L, Hu S, Wang B, Yang J, Xie G (2016) OsCCD1, a novel small calcium-binding protein with one EF-hand motif, positively regulates osmotic and salt tolerance in rice. Plant Sci 247:104–114. https://doi.org/10.1016/j.plantsci.2016.03.011
Kamachi H, Noguchi M, Inoue H (2004) Possible involvement of Ca2+-dependent protein kinases in spore germination of the fern Osmunda Japonica. J Plant Biol 47(3):27–32. https://doi.org/10.1007/BF03030224
Kanchiswamy CN, Malnoy M, Occhipinti A, Maffei ME (2014) Calcium imaging perspectives in plants. Int J Mol Sci 15(3):3842–3859. https://doi.org/10.3390/ijms15033842
Kandhol N, Singh VP, Wang Y, Chen Z-H, Tripathi DK (2022) Ca2+ sensor-mediated ROS homeostasis: defense without yield penalty. Trends Plant Sci 27(3):834–836. https://doi.org/10.1016/j.tplants.2022.05.003
Kazmierczak Jz, Kempe S, Kremer BJCOC (2013) Calcium in the early evolution of Living systems: a Biohistorical Approach. Biology 17(2):1738–1750. https://doi.org/10.1016/j.ejphar.2013.11.013
Ketehouli T, Nguyen Quoc VH, Dong J, Do H, Li X, Wang F (2022) Overview of the roles of calcium sensors in plants’ response to osmotic stress signalling. Funct Plant Biol 49(4):589–599. https://doi.org/10.1071/FP22012
Kim YC, Kim SY, Choi D, Ryu CM, Park JM (2008) Molecular characterization of a pepper C2 domain-containing SRC2 protein implicated in resistance against host and non-host pathogens and abiotic stresses. Planta 227(5):1169–1179. https://doi.org/10.1007/s00425-007-0680-2
Kudla J, Becker D, Grill E, Hedrich R, Hippler M, Kummer U, Parniske M, Romeis T, Schumacher K (2018) Advances and current challenges in calcium signaling. New Phytol 218(2):414–431. https://doi.org/10.1111/nph.14966
Lee H-J, Seo PJ (2021) Ca2+ talyzing initial responses to environmental stresses. Trends Plant Sci 26(3):849–870. https://doi.org/10.1016/j.tplants.2021.02.007
Leebens MJ, Graham SW, Wong GS, Degironimo L, Edger PP, Jordon-Thaden IE, Joya S, Melkonian B, Miles NW, Montero LP, Quigley C (2019) One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574(3):679–685. https://doi.org/10.1038/s41586-019-1693-2
Letunic I, Bork P (2021) Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49(4):W293–W296. https://doi.org/10.1093/nar/gkab301
Li et al (2015) A protocol to measure the cytoplasmic calcium in Arabidopsis Guard Cells, bio-protocol 5. 9e1462. https://doi.org/10.21769/BioProtoc.1462
Li L, Zheng Q, Jiang W, Xiao N, Zeng F, Chen G, Mak M, Chen ZH, Deng F (2023) Molecular regulation and evolution of Cytokinin Signaling in Plant Abiotic stresses. Plant Cell Physiol 63(12):1787–1805. https://doi.org/10.1093/pcp/pcac071
Liu X, Mak M, Babla M, Wang F, Chen G, Veljanoski F, Wang G, Shabala S, Zhou M, Chen ZH (2014) Linking stomatal traits and expression of slow anion channel genes HvSLAH1 and HvSLAC1 with grain yield for increasing salinity tolerance in barley. Front Plant Sci 5(1):634. https://doi.org/10.3389/fpls.2014.00634
Liu J, Niu Y, Zhang J, Zhou Y, Ma Z, Huang X (2017a) Ca2+ channels and Ca2+ signals involved in abiotic stress responses in plant cells: recent advances. Plant Cell Tiss Org 132(3):413–424. https://doi.org/10.1007/s11240-017-1350-0
Liu X, Fan Y, Mak M, Babla M, Holford P, Wang F, Chen G, Scott G, Wang G, Shabala S, Zhou M, Chen ZH (2017b) QTLs for stomatal and photosynthetic traits related to salinity tolerance in barley. BMC Genomics 18(1):9. https://doi.org/10.1186/s12864-016-3380-0
Liu M, Pan T, Allakhverdiev SI, Yu M, Shabala S (2020) Crop Halophytism: an environmentally sustainable solution for Global Food Security. Trends Plant Sci 25(7):630–634. https://doi.org/10.1016/j.tplants.2020.04.008
Liu W, Feng X, Cao F, Wu D, Zhang G, Vincze E, Wang Y, Chen ZH, Wu F (2021b) An ATP binding cassette transporter HvABCB25 confers aluminum detoxification in wild barley. J Hazard Mater 401(6):123371. https://doi.org/10.1016/j.jhazmat.2020.123371
Luan S, Wang C (2021) Calcium Signaling mechanisms across kingdoms. Annu Rev Cell Dev Biol 37(3):311–340. https://doi.org/10.1146/annurev-cellbio-120219-035210
Manishankar P, Wang N, Köster P, Alatar AA, Kudla J (2018) Calcium signaling during salt stress and in the regulation of ion homeostasis. J Exp Bot 69(17):4215–4226. https://doi.org/10.1093/jxb/ery201
Marchadier E, Oates ME, Fang H, Donoghue PC, Hetherington AM, Gough J (2016) Evolution of the calcium-based Intracellular Signaling System. Genome Biol Evol 8(7):2118–2132. https://doi.org/10.1093/gbe/evw139
Marchant DB, Chen G, Cai S, Chen F, Schafran P, Jenkins J, Shu S, Plott C, Webber J, Lovell JT, He G, Sandor L, Williams M, Rajasekar S, Healey A, Barry K, Zhang Y, Sessa E, Dhakal RR, Wolf PG, Harkess A, Li FW, Rossner C, Becker A, Gramzow L, Xue D, Wu Y, Tong T, Wang Y, Dai F, Hua S, Wang H, Xu S, Xu F, Duan H, Theissen G, McKain MR, Li Z, McKibben MTW, Barker MS, Schmitz RJ, Stevenson DW, Zumajo-Cardona C, Ambrose BA, Leebens-Mack JH, Grimwood J, Schmutz J, Soltis PS, Soltis DE, Chen ZH (2022) Dynamic genome evolution in a model fern. Nat Plants 8(2):1038–1051. https://doi.org/10.1038/s41477-022-01226-7
Mather C (2014) Swiss model. New Sci 222(2968):31–31. https://doi.org/10.1093/nar/gky427
Mcgregor N, Yin V, Tung CC, Van PF, Brumer H (2017) Crystallographic insight into the evolutionary origins of xyloglucan endotransglycosylases and endohydrolases. Plant J 89(5):651. https://doi.org/10.1111/tpj.13421
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees in Proceedings of the Gateway Computing Environments Workshop (GCE). New Orleans, LA pp 1–8. https://doi.org/10.1109/GCE.2010.5676129
Mohanta TK, Yadav D, Khan AL, Hashem A, Abd Allah EF, Al-Harrasi A (2019) Molecular players of EF-hand containing Calcium Signaling event in plants. Int J Mol Sci 20(1):1476. https://doi.org/10.3390/ijms20061476
Muhammad I, Shalmani A, Ali M, Yang QH, Ahmad H, Li FB (2021) Mechanisms regulating the dynamics of Photosynthesis under Abiotic stresses. Front Plant Sci 28(11):615942. https://doi.org/10.3389/fpls.2020.615942
Munns R, Day DA, Fricke W, Watt M., Arsova B, Barkla BJ, Bose J, Byrt CS, Chen Z-H, Foster KJ, Gilliham M, Henderson SW, Jenkins CLD, Kronzucker HJ, Miklavcic SJ, Plett D, Roy SJ, Shabala S, Shelden MC, Soole KL, Taylor NL, Tester M, Wege S, Wegner LH and Tyerman SD (2020) Energy costs of salt tolerance in crop plants. New Phytol 225:1072–1090. https://doi.org/10.1111/nph.15864
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/annurev-arplant-050718-100005
Nefissi Ouertani R, Arasappan D, Abid G, Ben Chikha M, Jardak R, Mahmoudi H, Mejri S, Ghorbel A, Ruhlman TA, Jansen RK (2021) Transcriptomic analysis of salt-stress-responsive genes in Barley roots and leaves. Int J Mol Sci 22(15):8155. https://doi.org/10.3390/ijms22158155
Niu J, Li Z, Zhu J, Wu R, Kong L, Niu T, Li X, Cheng X, Li J, Dai L (2022) Genome-wide identification and characterization of the C2 domain family in Sorghum bicolor L. and expression profiles in response to saline-alkali stress. Physiol Mol Biol Plants 28(13):1695–1711. https://doi.org/10.1007/s12298-022-01222-3
Park HJ, Kim WY, Yun DJ (2016) A new insight of salt stress signaling in plant. Mol Cells 39(3):447–459. https://doi.org/10.14348/molcells.2016.0083
Ponce KS, Meng L, Guo L, Leng Y, Ye G (2021) Advances in sensing, response and regulation mechanism of Salt Tolerance in Rice. Int J Mol Sci 22(15):2254. https://doi.org/10.3390/ijms22052254
Qadir M, Quillérou E, Nangia V, Murtaza G, Singh M, Thomas RJ, Drechsel P, Noble AD (2014) Economics of salt-induced land degradation and restoration. Nat Resour Forum 38(4):282–295. https://doi.org/10.1111/1477-8947.12054
Qiu CW, Ma Y, Liu W, Zhang S, Wang Y, Cai S, Zhang G, Chater CCC, Chen ZH, Wu F (2022) Genome resequencing and transcriptome profiling reveal molecular evidence of tolerance to water deficit in barley. J Adv Res 9(1):008. https://doi.org/10.1016/j.jare.2022.09.008
Ravi B, Foyer CH, Pandey GK (2023) The integration of reactive oxygen species (ROS) and calcium signalling in abiotic stress responses. Plant Cell Environ 46(4):1985–2006. https://doi.org/10.1111/pce.14596
Rupwate SD, Rajasekharan R (2012) C2 domain is responsible for targeting rice phosphoinositide specific phospholipase C. Plant Mol Biol 78(11):247–258. https://doi.org/10.1007/s11103-011-9862-1
Schaper E, Anisimova M (2015) The evolution and function of protein tandem repeats in plants. New Phytol 206(16):397–410. https://doi.org/10.1111/nph.13184
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7):671–675. https://doi.org/10.1038/nmeth.2089
Shabala S, Chen G, Chen ZH, Pottosin I (2020) The energy cost of the tonoplast futile sodium leak. New Phytol 225(7):1105–1110. https://doi.org/10.1111/nph.15758
Shen Q, Fu L, Su T, Ye L, Huang L, Kuang L, Wu L, Wu D, Chen ZH, Zhang G (2020) Calmodulin HvCaM1 negatively regulates Salt Tolerance via Modulation of HvHKT1s and HvCAMTA4. Plant Physiol 183(4):1650–1662. https://doi.org/10.1104/pp.20.00196
Shumilina J, Kusnetsova A, Tsarev A, van Janse HC, Medvedev S, Demidchik V, Van den Ende W, Frolov A (2019) Glycation of Plant Proteins: Regulatory roles and Interplay with Sugar Signalling? Int J Mol Sci 20(9):2366. https://doi.org/10.3390/ijms20092366
Solis CA, Yong MT, Vinarao R, Jena K, Holford P, Shabala L, Zhou M, Shabala S, Chen ZH (2020) Back to the Wild: on a Quest for donors toward Salinity Tolerant Rice. Front Plant Sci 11(2):323. https://doi.org/10.3389/fpls.2020.00323
Steinhorst L, Kudla J (2019) How plants perceive salt. Nature 572(7769):318–320. https://doi.org/10.1038/d41586-019-02289-x
Tansley C, Houghton J, Rose AME, Witek B, Payet RD, Wu T, Miller JB (2022) CIPK-B is essential for salt stress signalling in Marchantia polymorpha. New Phytol 237(5):2210–2223. https://doi.org/10.1111/nph.18633
Tanveer M, Shah AN (2017) An insight into salt stress tolerance mechanisms of Chenopodium album. Environ Sci Pollut Res Int 24(19):16531–16535. https://doi.org/10.1007/s11356-017-9337-2
Tanveer M, Yousaf U (2020) Chap. 23 - Plant single-cell biology and abiotic stress tolerance, in: Tripathi DK, Pratap Singh V, Chauhan DK, Sharma S, Prasad SM, Dubey NK, Ramawat, N (Eds.), Plant Life Under Changing Environment. Academic Press, pp 611–626. https://doi.org/10.1016/B978-0-12-818204-8.00026-6
Tiwari K, Paliyath G (2011) Cloning, expression and functional characterization of the C2 domain from tomato phospholipase Dα. Plant Physiol Biochem 49(1):18–32. https://doi.org/10.1016/j.plaphy.2010.09.015
Tong T, Fang Y-x, Zhang Z, Zheng J, Zhang X, Li J, Niu C, Xue D, Zhang X (2020) Genome-wide identification and expression pattern analysis of the KCS gene family in barley. Plant Growth Regul 93(1):89–103. https://doi.org/10.1007/s10725-020-00668-3
Tong T, Li Q, Jiang W, Chen G, Xue D, Deng F, Zeng F, Chen ZH (2021) Molecular Evolution of Calcium Signaling and Transport in Plant Adaptation to Abiotic Stress. Int J Mol Sci 22(22):12308. https://doi.org/10.3390/ijms222212308
Tyerman SD, Munns R, Fricke W, Arsova B, Barkla BJ, Bose J, Bramley H, Byrt C, Chen Z, Colmer TD, Cuin T, Day DA, Foster KJ, Gilliham M, Henderson SW, Horie T, Jenkins CLD, Kaiser BN, Katsuhara M, Plett D, Miklavcic SJ, Roy SJ, Rubio F, Shabala S, Shelden M, Soole K, Taylor NL, Tester M, Watt M, Wege S, Wegner LH, Wen Z (2019) Energy costs of salinity tolerance in crop plants. New Phytol 221(13):25–29. https://doi.org/10.1111/nph.15555
Vafadar F, Amooaghaie R, Ehsanzadeh P, Ghanati F (2021) Melatonin improves the photosynthesis in Dracocephalum Kotschyi under salinity stress in a Ca2+/CaM-dependent manner. Funct Plant Biol 49:89–101. https://doi.org/10.1071/FP21233
Wang Y, Chen ZH, Zhang B, Hills A, Blatt MR (2013) PYR/PYL/RCAR abscisic acid receptors regulate K+ and cl channels through reactive oxygen species-mediated activation of Ca2+ channels at the plasma membrane of intact Arabidopsis guard cells. Plant Physiol 163(2):566–577. https://doi.org/10.1104/pp.113.219758
Wang F, Chen ZH, Liu X, Colmer TD, Zhou M, Shabala S (2016) Tissue-specific root ion profiling reveals essential roles of the CAX and ACA calcium transport systems in response to hypoxia in Arabidopsis. J Exp Bot 67(4):3747–3762. https://doi.org/10.1093/jxb/erw034
Wang F, Chen ZH, Liu X, Colmer TD, Shabala L, Salih A, Zhou M, Shabala S (2017) Revealing the roles of GORK channels and NADPH oxidase in acclimation to hypoxia in Arabidopsis. J Exp Bot 68:3191–3204. https://doi.org/10.1093/jxb/erw378
Wang S, Li L, Li H, Sahu SK, Wang H, Xu Y, Xian W, Song B, Liang H, Cheng S, Chang Y, Song Y, Çebi Z, Wittek S, Reder T, Peterson M, Yang H, Wang J, Melkonian B, Van de Peer Y, Xu X, Wong GK, Melkonian M, Liu H, Liu X (2020) Genomes of early-diverging streptophyte algae shed light on plant terrestrialization. Nat Plants 6(1):95–106. https://doi.org/10.1038/s41477-019-0560-3
Wang Y, Chen G, Zeng F, Han Z, Qiu CW, Zeng M, Yang Z, Xu F, Wu D, Deng F, Xu S, Chater C, Korol A, Shabala S, Wu F, Franks P, Nevo E, Chen ZH (2023) Molecular evidence for adaptive evolution of drought tolerance in wild cereals. New Phytol 237(2):497–514. https://doi.org/10.1111/nph.18560
Waterhouse AM, Procter JB, Martin D, Clamp M, Barton GJ (2009) Jalview version 2: a multiple sequence alignment and analysis Workbench. Bioinformatics 25(9):1189–1191. https://doi.org/10.1093/bioinformatics/btp033
Witzel K, Weidner A, Surabhi GK, Börner A, Mock HP (2009) Salt stress-induced alterations in the root proteome of barley genotypes with contrasting response towards salinity. J Exp Bot 60(12):3545–3557. https://doi.org/10.1093/jxb/erp198
Wu J, Wang Y, Kim SG, Jung KH, Gupta R, Kim J, Park Y, Kang KY, Kim ST (2017) A secreted chitinase-like protein (OsCLP) supports root growth through calcium signaling in Oryza sativa. Physiol Plant 161:273–284. https://doi.org/10.1111/ppl.12579
Wu H, Shabala L, Azzarello E, Huang Y, Pandolfi C, Su N, Wu Q, Cai S, Bazihizina N, Wang L, Zhou M, Mancuso S, Chen Z, Shabala S (2018) Na+ extrusion from the cytosol and tissue-specific Na+ sequestration in roots confer differential salt stress tolerance between durum and bread wheat. J Exp Bot 69(7):3987–4001. https://doi.org/10.1093/jxb/ery194
Wu X, Wang Y, Bian Y, Ren Y, Xu X, Zhou F, Ding H (2022) A critical review on plant annexin: structure, function, and mechanism. Plant Physiol Biochem 190(3):81–89. https://doi.org/10.1016/j.plaphy.2022.08.019
Xiao S, Wu C, Zuo D, Cheng H, Zhang Y, Wang Q, Lv L, Song G (2022) Systematic analysis and comparison of CaLB genes reveal the functions of GhCaLB42 and GhCaLB123 in fiber development and abiotic stress in cotton. Ind Crops Prod 184:115030. https://doi.org/10.1016/j.indcrop.2022.115030
Xu T, Niu J, Jiang Z (2022) Sensing mechanisms: Calcium Signaling mediated abiotic stress in plants. Front Plant Sci 13:925863. https://doi.org/10.3389/fpls.2022.925863
Yang Y, Guo Y (2018) Unraveling salt stress signaling in plants. J Integr Plant Biol 60(4):796–804. https://doi.org/10.1111/jipb.12689
Yokotani N, Ichikawa T, Kondou Y, Maeda S, Iwabuchi M, Mori M, Hirochika H, Matsui M, Oda K (2009) Overexpression of a rice gene encoding a small C2 domain protein OsSMCP1 increases tolerance to abiotic and biotic stresses in transgenic Arabidopsis. Plant Mol Biol 71(4):391–402. https://doi.org/10.1007/s11103-009-9530-x
Yousefirad S, Soltanloo H, Ramezanpour SS, Zaynalinezhad K, Shariati V (2018) Salt oversensitivity derived from mutation breeding improves salinity tolerance in barley via ion homeostasis. Biol Plant 62(3):775–785. https://doi.org/10.1007/s10535-018-0823-2
Yousefirad S, Soltanloo H, Ramezanpour SS, Zaynali Nezhad K, Shariati V (2020) The RNA-seq transcriptomic analysis reveals genes mediating salt tolerance through rapid triggering of ion transporters in a mutant barley. PLoS ONE 15:e0229513. https://doi.org/10.1371/journal.pone.0229513
Zelm E, Zhang Y, Testerink C (2020) Salt tolerance mechanisms of plants. Annu Rev Plant Biol 71(5):403–433. https://doi.org/10.1146/annurev-arplant-050718-100005
Zeng X, Long H, Wang Z, Zhao S, Tang Y, Huang Z, Wang Y, Xu Q, Mao L, Deng G, Yao X, Li X, Bai L, Yuan H, Pan Z, Liu R, Chen X, WangMu Q, Chen M, Yu L, Liang J, DunZhu D, Zheng Y, Yu S, LuoBu Z, Guang X, Li J, Deng C, Hu W, Chen C, TaBa X, Gao L, Lv X, Abu YB, Fang X, Nevo E, Yu M, Wang J, Tashi N (2015) The draft genome of tibetan hulless barley reveals adaptive patterns to the high stressful tibetan Plateau. Proc Natl Acad Sci U S A 112(2):1095–1100. https://doi.org/10.1073/pnas.1423628112
Zeng H, Zhang Y, Zhang X, Pi E, Zhu Y (2017) Analysis of EF-Hand Proteins in soybean genome suggests their potential roles in environmental and nutritional stress signaling. Front Plant Sci 24:8:877. https://doi.org/10.3389/fpls.2017.00877
Zepeda-Jazo I, Velarde-Buendia AM, Enriquez-Figueroa R, Bose J, Shabala S, Muniz-Murguia J, Pottosin II (2011) Polyamines interact with hydroxyl radicals in activating Ca2+ and K+ transport across the root epidermal plasma membranes. Plant Physiol 157(17):2167–2180. https://doi.org/10.1104/pp.111.179671
Zhang D, Aravind L (2010) Identification of novel families and classification of the C2 domain superfamily elucidate the origin and evolution of membrane targeting activities in eukaryotes. Gene 469(6):18–30. https://doi.org/10.1016/j.gene.2010.08.006
Zhang Y, Li D, Zhang D, Zhao X, Cao X, Dong L, Liu J, Chen K, Zhang H, Gao C, Wang D (2018b) Analysis of the functions of TaGW2 homoeologs in wheat grain weight and protein content traits. Plant J 94(9):857–866. https://doi.org/10.1111/tpj.13903
Zhang H, Zhu J, Gong Z, Zhu JK (2021) Abiotic stress responses in plants. Nat Rev Genet 23(2):104–119. https://doi.org/10.1038/s41576-021-00413-0
Zhang H, Zeng Y, Seo J, Kim YJ, Kim ST, Kwon SW (2022) Global identification and characterization of C2 Domain-Containing Proteins Associated with abiotic stress response in Rice (Oryza sativa L). Int J Mol Sci 23(3):2221. https://doi.org/10.3390/ijms23042221
Zhao C, Wang Y, Chan KX, Marchant DB, Franks PJ, Randall D, Tee EE, Chen G, Ramesh S, Phua SY, Zhang B, Hills A, Dai F, Xue D, Gilliham M, Tyerman S, Nevo E, Wu F, Zhang G, Wong GK, Leebens-Mack JH, Melkonian M, Blatt MR, Soltis PS, Soltis DE, Pogson BJ, Chen ZH (2019) Evolution of chloroplast retrograde signaling facilitates green plant adaptation to land. Proc Natl Acad Sci U S A 116(9):5015–5020. https://doi.org/10.1073/pnas
Zheng M, Lin J, Liu X, Chu W, Li J, Gao Y, An K, Song W, Xin M, Yao Y, Peng H, Ni Z, Sun Q, Hu Z (2021) Histone acetyltransferase TaHAG1 acts as a crucial regulator to strengthen salt tolerance of hexaploid wheat. Plant Physiol 186(4):1951–1969. https://doi.org/10.1093/plphys/kiab187
Zhong B, Xi Z, Goremykin VV, Fong R, McLenachan PA, Novis PM, Davis CC, Penny D (2014) Streptophyte algae and the origin of land plants revisited using heterogeneous models with three new algal chloroplast genomes. Mol Biol Evol 31(1):177–183. https://doi.org/10.1093/molbev/mst200
Ziska LH, Bunce JA, Shimono H, Gealy DR, Baker JT, Newton PC, Reynolds MP, Jagadish KS, Zhu C, Howden M, Wilson LT (2012) Food security and climate change: on the potential to adapt global crop production by active selection to rising atmospheric carbon dioxide. Proc Biol Sci 279(1745):4097–4105. https://doi.org/10.1098/rspb.2012.1005
Acknowledgements
We’d like to thank Dr. Yizhou Wang and Mengmeng Rui from Zhejiang University for the facilities, we also thank Dr. Shengguan Cai, Qi Li, Xiachen Lv, Qiufang Shen from Zhejiang University and Dr. Liuhui Kuang from Hunan Agricultural University for the discussion on the draft preparation.
Funding
This research was financially supported by the National Natural Science Foundation of China (32170276, 32272053), Major International (Regional) Joint Research Project from NSFC-ASRT (32061143044), and Yangtze University. This work was also funded by Australian Research Council (FT210100366), Grains Research & Development Corporation (WSU2303-001RTX), and Horticulture Innovation Australia (LP18000) to Zhong-Hua Chen.
Author information
Authors and Affiliations
Contributions
Conceptualization, Zhong-Hua Chen and Fenglin Deng; methodology, Zhong-Hua Chen, Fenglin Deng, Yuan Qin, Fanrong Zeng and Xueyi Zhou; data analysis, Tao Tong and Yanli Zhang; experimental manipulation, Tao Tong, Wei Jiang and Guang Chen; writing—original draft preparation, Tao Tong, Yuan Qin, Fenglin Deng and Zhong-Hua Chen; writing—review and editing, Fenglin Deng, Yuan Qin, Tao Tong and Yanli Zhang; supervision, Fenglin Deng, Zhong-Hua Chen, Dawei Xue and Fanrong Zeng; funding acquisition, Zhong-Hua Chen, Fenglin Deng, Guang Chen and Fanrong Zeng. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tong, T., Zhang, Y., Zhou, X. et al. Molecular and physiological evidence of HvCaLB1 regulates calcium homeostasis and salt tolerance in Barley. Plant Growth Regul (2024). https://doi.org/10.1007/s10725-024-01176-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10725-024-01176-4