Abstract
β-Ketoacyl CoA synthetase (KCS) is a key rate-limiting enzyme for the synthesis of very-long-chain fatty acids (VLCFAs) in plants that determines the synthesis rate and carbon chain length of VLCFAs, ultimately affecting the stress resistance of epidermal wax. In this study, a genome-wide characterization of the KCS family was conducted using high-quality barley (Hordeum vulgare L.) genome sequences. In total, 33 KCS genes were identified and were unevenly distributed in the distal centromere region of each chromosome. These genes were divided 12 subgroups based on sequence alignment and phylogenetic analysis, with members of the same subgroup possessing similar genes and motif structures. Expression analysis showed that the KCS genes demonstrate diverse tissue expression patterns in barley and also have overlapping functions. The barley KCS gene family also exhibited different response characteristics under drought stress. These results provide insights into the evolutionary processes and potential functions of the KCS gene family, offering a theoretical reference for studies on the physiological and biochemical regulation of KCS genes during barley growth and development, as well as for the genetic breeding analysis of wheat crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Very-long-chain fatty acids (VLCFAs) can be modified, derived, esterified, or polymerized to form bioactive products, such as the wax components of the plant epidermis, which play crucial roles in plant development and in the resistance to diverse stress (Haslam and Kunst 2013; Xue et al. 2017). β-Ketoacyl-CoA synthase (KCS), which is embedded in fatty acid elongase (FAE), is related to the synthesis of VLCFAs as an important rate-limiting enzyme (Wettstein-Knowles 1982). It has strict carbon chain-length specificity in substrate and product, influences the composition and content of plant epidermal wax, and modifies plant stress resistance (Vogg et al. 2004).
The first gene encoding a KCS enzyme was identified and isolated from mutant Arabidopsis thaliana plants deficient in VLCFAs (James et al. 1995). The mutant line showed significantly decreased production of C20:0, C20:1, and C22:1 in the seeds, and thus the gene was named fatty acid elongation 1 (FAE1) (James et al. 1995). Subsequently, 25 KCS homologous genes in A. thaliana were identified and functionally characterized through mutation analysis and heterologous expression technology in yeast (Joubès et al. 2008; Dunn et al. 2004). Based on amino acid sequence homology and protein conformation, they were classified into five gene subfamilies: FAE1, FDH, CER6, KCS1, and ELO (Dunn et al. 2004), and most are related to the biosynthesis of wax components in the plant epidermis. Since then, KCS homologous genes have been identified in barley (Li et al. 2018), Simmondsia chinensis (Lassner and Metz 1996), Tropaeolum majus (Mietkiewska et al. 2004), and even early land plants such as Marchantia polymorpha (Kajikawa et al. 2003a, b).
The KCS enzyme, as an endogenous cellular enzyme, consists of a 3-oxoacyl-[acyl-carrier protein (ACP)] synthase III C domain and a Type III polyketide synthase-like protein domain. The former has the core of a conserved domain that catalyzes the extension of the fatty acid chain and also possesses some regions and residues with specific differences. Through a series of biochemical steps, including transfer, decarboxylation, and fixation (Haslam and Kunst 2013), the specific selection of the substrate carbon chain length by each member of KCS and the catalytic synthesis of the corresponding product can be achieved (Lassner and Metz 1996; Millar et al. 1999). According to homology modeling, sequence-structure homology search, and comparative analysis of the structure-function of the KCS protein, all of the members of this family have a similar four-layer αa–βb–αc–αd folding structure (Joubès et al. 2008). The first fold, αa, consists of either one or two alpha helices that for most members of the KCS family are well positioned to serve as a membrane anchor, while the other three-layer βb–αc–αd fold constitutes the soluble core of the KCS enzyme. After comparing and evaluating the size and shape of the substrate-binding domain in the soluble core, it was shown that a certain correlation exists between the size of the domain and the length of the acyl matrix. A site-specific study revealed that Cys223, His302, His387, His391, and His420 are all active sites of KCS condensing enzyme, and these mutations will lead to changes in substrate specificity and even a deficiency in enzyme activity (Blacklock and Jaworski 2002; Ghanevati and Jaworski 2001, 2002).
The KCS gene family determines the content and composition of epidermal wax, actively participates in physiological and biochemical reactions at all stages of plant growth and development, and also shapes plant stress resistance. The fatty acid elongation gene (FAE1/KCS18) in Arabidopsis is related to the synthesis of VLCFAs in the seeds (Kunst et al. 1992) and is a critical erucic acid biosynthesis gene, the deletion mutant of which caused the content of long-chain fatty acids in the seeds to drop precipitously (Kunst et al. 1992). When the soil is short of water or the air humidity is too low, the content of C26–C30 wax alcohols and aldehydes in the Arabidopsis kcs1 mutant become unusually low, with the mutants showing a decline in drought resistance and low humidity tolerance at the seedling stage (Todd et al. 1999). Pm40, the homologous gene of KCS1 in wheat, also participates in the signal transduction of resistance to powdery mildew by regulating photosynthesis and also enhances resistance to powdery mildew (Weidenbach et al. 2015). KCS2/DAISY and KCS20 exhibited similar functions in the biosynthesis of epidermis wax and root suberin, but they demonstrated different regulation and expression under osmotic stress (Lee and Suh 2013). KCS9 is involved in the elongation of C22 to C24 fatty acids, which are essential precursors for the biosynthesis of cuticular waxes, aliphatic suberins, and membrane lipids including sphingolipids and phospholipids (Kim et al. 2013). When the expression of the CER6 gene in the CER6 subfamily was inhibited, the wax in the plant pollen, stem, and pod became critically deficient, possibly causing conditioned male sterility (Millar et al. 1999; Fiebig et al. 2000). Nevertheless, the accumulation of CER6 transcripts increased during osmotic stress and the application of exogenous abscisic acid (Wu 2012). The organs, such as the flowers and young leaves, are the main expression sites of the FDH gene, the mutation of which triggers the post-genital fusion of the leaves and flowers (Pruitt et al. 2000). In addition, genes such as KCS1, KCS2, and FDH could be positively regulated by MYB30 transcription factors under disease stress (Raffaele et al. 2008). However, the specific mechanism still needs further study.
With the large-scale completion of plant genome sequencing and the wide application of bioinformatics technology, 58, 21, 22, and 26 KCS genes have been identified and isolated in G. hirsutum (Xiao et al. 2016), Brassica napus (Wu 2012), pepper (Yi et al. 2020), and even the microalgae Monoraphidium neglectum (Liu et al. 2019), respectively. Barley is one of the most important global food crops that exhibits high adaptability, a wide distribution, and strong stress tolerance (Zhang et al. 2013; Zhu et al. 2020)). In recent years, LEUCINE-RICH REPEAT (CNL) (Andersen et al. 2016), MITOGEN-ACTIVATED PROTEIN KINASE (MAPK) (Cui et al. 2019), NUCLEOTIDE-BINDING SITE (NBS) (Habachi-Houimli et al. 2018), SMALL HEAT SHOCK PROTEIN (HSP20) (Li and Liu 2019), NON-SPECIFIC LIPID TRANSFER PROTEINS (nsLTPs) (Zhang et al. 2019), SQUAMOSA PROMOTER BINDING PROTEIN (SBP) (Tong et al. 2019), and other gene families have been successively identified and analyzed at the genome-wide level in barley. However, there have been no relevant systematic investigations into the barley KCS gene family. In this study, 33 KCS genes in barley were identified by genome-wide analysis technology. The gene structure, physical and chemical characteristics, chromosome location, phylogenetic relationships, and tissue expression patterns were explored, providing a theoretical foundation for the downstream functional analysis of the barley KCS gene. The analysis and quantification of the expression trends of KCS candidate genes under drought stress can also provide an experimental basis for stress resistance breeding and other genetic research in barley.
Materials and methods
Screening and identification of KCS genes
Twenty-one identified KCS genes were downloaded from the Arabidopsis genome database TAIR (https://www.arabidopsis.org/index.jsp) (Poole 2005). In the light of the conservation of KCS domain evolution, the related Hidden Markov Model profiles of gene family (Accession No.: PF08392 and PF08541) were obtained from the Pfam database (http://pfam.xfam.org/) (Bateman et al. 2004; Finn et al. 2015). HMMER (https://www.ebi.ac.uk/Tools/hmmer/) was used to retrieve online barley protein candidate sequences (E-value < 10− 10) containing KCS conserved domains (FAE1_CUT1_RppA and ACP_syn_III_C). The candidate sequences after removing too short and redundancy were submitted into the NCBI conserved Domains (https://www.ncbi.nlm.nih.gov/structure/cdd/wrpsb.cgi) (Marchler-Bauer et al. 2015) and SMART (http://smart.embl-heidelberg.de/) (Letunic et al. 2012) that access to the revalidation whether they contain KCS domain. Next, the incomplete reading frame, short and redundant sequences were removed manually, after which the KCS gene and protein sequences in barley were finally obtained.
The theoretical isoelectric point and relative molecular mass were predicted by ExPASY (Compute pI/Mw tool) online on the basis of KCS protein sequences. Subcellular localization prediction of the KCS proteins was implemented using Softberry (http://linux1.softberry.com/) and WoLF PSORT (https://wolfpsort.hgc.jp/) (Horton et al. 2006).
Phylogenetic analysis of genes
The acquired full-length KCS protein sequence was aligned using the software MEGA7.0 with the default parameters of ClusterW (Hall 2013), wherein Neighbor-Joining (NJ) method was performed to construct the phylogenetic tree of KCS protein in barley and Arabidopsis with bootstrap test of 1000 times, after which iTOL (https://itol.embl.de/login.cgi) online tool was used visual rendering (Letunic and Bork 2007). Furthermore, the conserved domains was manually selected and intercepted using BioEdit (http://www.mbio.ncsu.edu/bioedit/bioedit.html) (Hall 1999).
Ka/Ks value calculation, structural analysis, and chromosome localization
The substitution rates of non-synonymous (Ka) and synonymous (Ks) were calculated by software DnaSP6 (Rozas et al. 2017). Ks value could be used as molecular clock to reckon the time since gene replication event (Peterson* and Masel† 2009). Ka/Ks rate has been used to determine the type of gene selection during evolution. Ka/Ks = 1, Ka/Ks > 1, and Ka/Ks < 1 represent natural, positive, and purifying selections, respectively. The conserved motifs of barley KCS proteins were performed a thorough investigation with MEME (http://meme-suite.org/tools/meme) (Bailey et al. 2009), the parameters were set as follows: the number of occurrences of the same motif in a sequence is 0 or 1, an optimum motif width of 10 to 300 amino acid residues, and maximum of 5 motifs. The intron/exon distribution pattern diagram of barley KCS genes was generated by the Gene Structure Display Server (GSDS) program with the sequence file and annotation information file (Hu et al. 2014). On account of the location of KCS gene in barley genome, Map Inspect mapping software was carried out to map and depict the physical location of KCS genes to each chromosome.
Expression profile analysis and function identification
To explore the specific expression patterns of KCS gene in barley in different tissues and developmental stages, The corresponding RNA-seq data of 14 developmental stages were downloaded from IPK (http://webblast.ipk-gatersleben.de/barrey’ibsc/index.php) and The James Hutton Institute (https://ics.hutton.ac.uk/morexgenes/index.html) websites, respectively. The gene expression values are represented by fragments per kilobase of exon per million fragments mapped (FPKM). The developmental stages of expression analysis listed hereafter: embryo tissue (EMB); isolate etiolated leaves (ETI), root tissue (ROO); shoots from seedings (10 cm shoot stage) (LEA); epidermal strips (EPI); developing inflorescence tissue (INF); rachis (RAC); third-stem internodes (NOD); isolated lodicules dissected from inflorescences (LOD); lemma dissected from inflorescence (LEM); palea dissected from inflorescence (PAL); grains throughout development (caryopsis, 5 and 15 days post-anthesis) (CAR5, CAR15); and senescent leaves (SEN). These data were submitted to software MeV for hierarchical clustering analysis and expression profile drawing through checking and sorting.
Thirty-three KCS genes in barley were employed for the enrichment analysis of metabolic pathways online combining Blast2go software (Conesa et al. 2005) and KEGG database (https://www.kegg.jp/kegg/pathway.html).
Plant materials and drought treatments, total RNA extraction, and quantitative real-time expression analysis
Seedlings of barley ZJU3, which had been planted in an experimental field, were selected and transplanted into culture pots in the greenhouse. Barley ZJU3 has the advantages of early maturity, lodging resistance, high yield and resistance to yellow mosaic disease. All of the experimental seedlings were cultured under day/night temperatures of 24 °C/20 °C with 16 h of light and 8 h of darkness. After 2 weeks of growth recovery, barley seedings were divided into groups and were selected based on good and consistent growth. The experimental groups were treated with drought for 15 d, whereas the control groups were simultaneously irrigated with distilled water. Six top-second leaves were removed from each pot for subsequent experimental analysis upon completion of the treatment.
An Axygen total RNA kit was used to extract total RNA from the barley leaves by centrifugation. Random primers N6 and oligo (dT)18 primer in the ratio of 1:1 were mixed as reverse transcription primers to improve the performance. Following reverse transcription, the reaction solution was stored at − 20 °C for the short term. For the design of fluorescent quantitative primers, 10 pairs of barley KCS gene primers were designed using the software Primer Premier 5.0 (Lalitha 2000), wherein HvActin was designed with the HORVU1Hr1G002840 gene as the internal reference (Table 1). For simplification, the selected KCS family genes were named from HvqRT1 to HvqRT10. The specificity of primers to their target genes was evaluated on the website EnsemblPlants (http://plants.ensembl.org/hordeum_vulgare/info/index). All of the primers were synthesized by Shanghai Qingke Biotechnology Co., Ltd. The total volume of the quantitative real-time (qRT)-PCR reaction system was 10 µL, which included 5 µL of Hieff® qPCR SYBR Green Master Mix, 1 µL of diluted cDNA template, 1 µL of upstream and downstream primers (10 mmol L− 1), and 3 µL of RNase free ddH2O. The qRT-PCR procedure was 95 °C for 5 min, followed by 40 cycles at 94 °C for 10 s, 60 °C for 20 s, and an elongation step of 20 s at 72 °C. In the elongation step, fluorescence signals were collected, and three biological replicates and three technical replicates were tested for each reaction. The relative expression level of the target gene was calculated using the 2−ΔΔCT method, and single factor analysis of variance was performed in SPSS version 20.0 (IBM Corp, Armonk, NY, USA).
Results and analysis
Identification and physicochemical properties of the KCS genes
The candidate sequences related to KCS in barley were retrieved using the Pfam database and HMMER search engine, with the protein sequence of the KCS conserved domain used as the key word. The redundant sequences were then eliminated, the KCS domain was detected again using the SMART and NCBI Conserved Domains websites, and incomplete reading frames and short sequences were manually removed. Ultimately, 33 candidate genes containing the KCS domain were identified in barley. The results are shown in Table S1. The molecular weights and isoelectric points of the KCS family proteins were 44304.8–66238.79 Da and 6.95–10.2, respectively, with a relatively small gap but no obvious regularity. Similarly, the length of these proteins was 398–600 amino acid residues which spanned a relatively small. Subcellular localization results showed that the 33 barley KCS proteins were all located on the cell membrane, indicating that barley KCS proteins possessed highly conserved membrane protein catalytic functions.
Intrachromosomal distribution of KCS genes
The distribution of gene families on the chromosomes was closely related to the participation of the chromosomes in the expression process of gene family members as well as the relevance and importance in plant growth and development processes. Map Inspect software was used to depict the location map of KCS genes on the chromosome in barley (Fig. 1). It can be seen from the figure that the KCS genes were unevenly distributed on all seven of the barley chromosomes, among which only two genes were contained in the 2H and 3H chromosomes, and the 1H, 4H, 5H, and 6H chromosomes contained four, three, five, and five genes, respectively. The 7H chromosome comprised the most genes in terms of quantity, namely, 11, accounting for 33% of KCS gene family members. 1H and 7H both existed the closely coexistence regions, whereas there was no evidence that tandem repeat genes occurred in these regions. The centromere and pericentromere regions, which are hot spots for new gene generation and initiation in plants, experienced dramatic sequence changes and structural remodeling processes, and the genes in the centromere region have an evolutionary tendency of escaping outward (Zhang et al. 2018). In this study, all of the KCS genes were located in the far centromere region, suggesting that the evolutionary process of the barley KCS genes was long and its function was comparatively stable. In addition, HORVU0Hr1G038430 could not be anchored to the chromosome (this gene was located on chrUn) because of the incomplete sequencing and assembly of the barley genome.
Distribution of KCS genes in barley chromosomes. The location of KCS gene family members in the recently release of barley cultivar ‘Morex’ genome database (Hordeum vulgare. IBSC_v2). Data analysis showed that all the KCS genes were distributed far from pericentromeric region. The red dashed box highlighted the gene cluster. (Color figure online)
Phylogenetic analysis of KCS genes
ClustalW alignment of 54 identified KCS protein sequences in Arabidopsis and barley was conducted. The results are shown in Figure S1. It was found that the KCS genes of barley and Arabidopsis had two sequences with highly similar alignments and were approximately 260 and 80 amino acid residues in length, respectively. These were considered as two conserved domains of the KCS gene family, namely, FAE1_CUT1_RppA and ACP_syn_III_C.
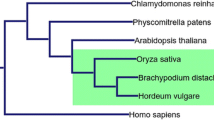
To better understand the evolutionary relationships of the barley KCS gene, 54 KCS protein sequences of Arabidopsis and barley were selected and imported into the software MEGA7.0 to construct an unrooted phylogenetic tree. According to the method about grouping 8 subgroups of Arabidopsis KCS gene family (Joubès et al. 2008; Lessire et al. 1999), the candidate genes could be further categorized into 12 subgroups based on high bootstrap repetitions (Fig. 2). Phylogenetic tree analysis showed that the HORVV-I, HORVV-II, HORVV-III, and HORVV-IV subgroups only consisted of barley KCS proteins; the FDH-like (η), FDH (θ), and FAE-like (β) subgroups only contained Arabidopsis KCS proteins; and both Arabidopsis and barley KCS proteins existed in the remainder of the subgroups. This suggested that the KCS gene originated before the differentiation of monocotyledons and dicotyledons. In addition, based on the proximity of the branches of the evolutionary tree, three pairs of orthologous proteins were identified, including A0A287GND2_HORVV/KCS4_ARATH, F2CSE8_HORVV/KCS1_ARATH, and A0A287 × 585_HORVV/KCS11_ARATH, all of which had a similarity of about 80%. Simultaneously, there were 16 pairs of intraspecific paralogous proteins in the HORVV-I, HORVV-II, and HORVV-IV subgroups of barley, each of which contained 1, 2, and 5 pairs of barley paralogous proteins, respectively. In addition, there was one pair of Arabidopsis paralogous proteins existing separately in the FDH-like (θ), FDH-like (η), CER6 (γ), Fae-like (β), Fae-like (α), FDH-like (ε), KCS1-like (δ), and KCS1-like (ζ) subgroups. In general, 70.37% of KCS proteins exist as homologous proteins within species, and the functional differentiation degree of Arabidopsis KCS protein subgroups is relatively high. In contrast, five of the 17 barley KCS proteins in the HORVV-IV barley subgroups were paralogous proteins, which indicated that the KCS gene still actively duplicated many times after barley speciation. Consequently, the function of the barley KCS protein may have a greater degree of overlap, and the HORVV-IV subgroup is probably particular to Gramineae and even barley (Tong et al. 2019; Guo et al. 2016).
Phylogenetic relationships of KCS gene family in barley (HORVV) and Arabidopsis (ARATH). The unrooted phylogenetic tree was generated based on the amino acid sequences aligning by the neighbor joining (NJ) method using MEGA 7.0. The bootstrap values of 1000 replicates were calculated at each node. All KCS members were classified into 12 groups which differentiate by different colors. (Color figure online)
The Ka/Ks value is an important parameter that is used to evaluate the evolution of coding sequences and determine the type of selection pressure after duplication (Xiong et al. 2019). The Ka/Ks value of the barley KCS gene family was mostly greater than 1, and the average value was 1.3061 (Table S3), indicating that positive selection was the main type of selection. The approximate average differentiation time of barley KCS family genes, which was deduced by calculating the Ks value, was 42.14 million years ago. The differentiation time of A0A287GLF8_HORVV/F2EI13_HORVV in the HORVV-IV subgroup was the shortest at only 5.92 million years ago (Conery and Lynch 2001). Combined with the above, it was inferred that the gene duplication events of the KCS gene in barley were actively distributed throughout the entire barley evolutionary process.
Motif composition and structure analysis of KCS genes
Intron/exon structure and intron type and number are typical evolutionary imprints of a gene family (Li et al. 2017). Therefore, further construction of a protein phylogenetic tree and intron/exon structure map of the barley KCS gene family provided an in-depth assessment of the structural characteristics of related genes (Fig. 3). The results indicated that the gene structure similarity of the same branch was high, while that of the different branches was low. Statistical analyses revealed that HORVU7Hr1G023530, HORVU4Hr1G067340, HORVU1Hr1G089710, HORVU7Hr1G084610, HORVU5Hr1G056870, and HORVU2Hr1G122150 contained only one intron, whereas HORVU4Hr1G076940 contained three introns, and the rest of the barley KCS genes did not possess introns, suggesting that the KCS genes of barley were more expressed and more functional compared with other gene families.
Motif distribution and the exon–intron structure of barley KCS family. Different conserved motifs were represented by blocks of five different colors. Motif1, Motif2, Motif3 and Motif4 constituted the FAE1_CUT1_RppA domain of the KCS protein, and ACP_syn_III_C domain included Motif5. Yellow boxes, black lines, and green boxes indicated exons, introns, and non-coding regions, respectively. This graph is depicted by TBtools (Chen et al. 2018). (Color figure online)
Prediction and identification of the motifs contained in the barley KCS protein sequence were made using the MEME analysis tool. The analytical results indicated that the E-values of the five conserved functional motifs were all less than 1e−300 (Motif1: 6.0e−333, Motif2: 7.2e−1160, Motif3: 1.1e−760, Motif4: 3.6e−510, Motif5: 1.9e−2237), indicating that each conserved motif was statistically significant. Motif1, Motif2, Motif3, and Motif4 constituted the FAE1_CUT1_RppA of KCS protein, and Motif5 was the conserved motif of the ACP_syn_III_C domain. In addition, with the exception of HORVU4Hr1G076940, which lacked Motif1, the remaining barley KCS proteins all contained five complete motifs, showing high sequence and functional conservation.
Tissue specific expression of the barley KCS gene
To analyze the expression patterns of the barley KCS gene family, the expression data of barley during different periods as well as tissues and organs (seeds, roots, stems, leaves, flowers and fruits at different developmental stages) were collected from the IPK website to construct the relevant tissue-specific expression profile using a hierarchical clustering model. The results are shown in Fig. 4. HORVU6Hr1G036950 and HORVU1Hr1G089710 were highly expressed in various tissues and fruits at different developmental stages. HORVU4Hr1G076940, HORVU4Hr1G063420, and HORVU4Hr1G067340 indicated similar expression patterns and all clustered with high expression and located on the 4H chromosome, whereas HORVU4Hr1G076940 and HORVU4Hr1G063420 were not expressed in the root tissue and developing inflorescence tissue, respectively. In comparison, HORVU4Hr1G067340 was not expressed in the inflorescence and root tissues. HORVU7Hr1G084610, HORVU7Hr1G023280, and HORVU7Hr1G023530 exhibited pan-tissue expression, but their expression level was relatively low. Great expression fluctuations in the different tissues and periods were observed in the remaining barley KCS genes, which did not express in some tissues or at certain growth stages.
Expression profiles of KCS genes in different tissues and stages of development. Data were downloaded from publicly available databases. Rows represent KCS members, while columns show different developmental stages and tissues. The expression level of KCSs [log2(FPKM + 1)] is shown by the intensity of color, wherein red blocks represent low expression, and green blocks represent high expression: EMB 4-day embryos; ROO roots from seedings (10 cm shoot stage); LEA shoots from seedings (10 cm shoot stage); INF developing inflorescences (1–1.5 cm); NOD developing tillers, 3rd internode (5 DAP); CAR5, CAR15 developing grain (5 DAP, 15 DAP); ETI etiolated seeding, dark cond (10 DAP); EPI epidermal strips (28 DAP); RAC inflorescences, rachis (35 DAP); LEM inflorescences, lemma (42 DAP); LOD inflorescences, lodicule (42 DAP); PAL dissected inflorescences (42 DAP); SEN senescing leaves (56 DAP). (Color figure online)
The expression profile showed that some members of the barley KCS gene family that did not demonstrate clear tissue-specificity played an important role in almost all growth and developmental stages of barley, whereas other members exhibited tissue-specificity to a certain extent. Among them, the HORVU5Hr1G080180 gene was specifically expressed in the developing inflorescence tissue, HORVU7Hr1G042040 showed a relatively high expression level in the embryonic tissue, and HORVU7Hr1G006910 was more highly expressed in the leaf tissue and senescent leaves. Accordingly, combined with studies of gene structure, it was found that the expression of intron-rich genes was biased towards non-tissue specificity.
The results of the Kyoto Encyclopedia of Genes and Genomes (KEGG) online pathway annotation are shown in Fig. 5. Thirty-three barley KCS genes were all enriched in the first step of the condensation reaction of VLCFA synthesis in fatty acid synthesis, indicating the specificity of the biochemical function of barley KCS genes.
Annotation map of KEGG metabolic pathway of KCS gene family in barley using Blast2go software. The green module indicated the step of protein enrichment, and the white module indicated that no protein was enriched in this step. 33 barley KCS proteins were all enriched in the first step of the VLFAs extension biochemical process at the fatty acid extension flow chart, numbered 23.1.199. (Color figure online)
Analysis of the expression pattern of barley KCS genes under drought stress
Combined with the phylogenetic tree and expression characteristics of the barley KCS gene family, 10 KCS genes were screened for the design of specific primers and relative expression analysis of fluorescence quantitative PCR to explore the response characteristics of barley KCS genes to drought stress. The genes were selected from the HORVV-IV, CER6 (γ), Fae-like (α), FDH-like (ε), KCS1-like (δ), and KCS1-like (ζ) subgroups, respectively. The results showed that 10 KCS genes indicated different expression characteristics under drought stress treatment (Fig. 6), and the gene expression patterns of the same subgroup were also distinct. Under drought treatment, the expression level of five KCS genes decreased and other five KCS genes increased. Among them, the relative expression levels of HORVU7Hr1G006910, HORVU5Hr1G087530, HORVU4Hr1G076940, and HORVU4Hr1G063420 decreased by more than one-fold, suggesting that their expression levels were negatively correlated with their functional effects. On the contrary, the relative expression levels of HORVU4Hr1G067340, HORVU6Hr1G004830, HORVU1Hr1G090730, and HORVU6Hr1G07300 increased by more than one-fold, and the activities of these four genes were positively induced by drought stress. Overall, most of the barley KCS genes responded to external drought signals and made positive or negative expression trends to regulate the response to drought, with the regulatory expression trend demonstrating no apparent subgroup characteristics.
Expression profiles of 10 selected barley KCS under drought stress using real-time PCR analysis. The experiments were repeated three times and provided consistent results, and the relevant data were presented as the mean ± SD (n = 3). ZJU3 CK as control group stood for barley with normally watered, and ZJU3 DR as experimental group stood for barley after drought treatment, these groups were represented by different colors (green and orange). (Color figure online)
Discussion
The KCS gene family, which is widely present in terrestrial plants, encodes enzymes that catalyze the rate-limiting condensation reaction and accordingly specifically regulate the elongation of plant fatty acids. They are thus likely to play an essential role in substrate and product specificity. Owing to its important role in the plant epidermal wax synthesis pathway, the KCS gene family has been identified and analyzed at the genome-wide level of various plant species. Xiao et al. (2016) identified 58 KCS genes from G. hirsutum. Some of the gene members respond to exogenous gibberellin A3 (GA) and control fiber cell growth. Wu (2012) cloned 21 BnKCS genes in B. napus, among which the FAE1-like genes could effectively mediate the synthesis of erucic acid, while Yi et al. (2020) identified 22 KCS family genes from the pepper genome. Expression analysis has indicated that the expression of the family members can be activated or inhibited by high temperature, low temperature, and salt (NaCl) treatment. In addition, Monoraphidium, which has a high oil content, also contains 26 KCS genes, and an assessment of its KCS gene family improved the stress resistance of Monoraphidium and the quality characteristics of related microalgae energy (Liu et al. 2019). Therefore, the analysis and identification of the physical and chemical properties, structure, and function of the barley KCS gene family at the whole-genome level have good practical significance and application value, and can provide a theoretical basis for the genetic breeding of further resistance and yield traits in barley.
Through the retrieval, analysis, and organization of the KCS gene family in the genome of barley, a total of 33 barley KCS gene family members were identified. Using the highly conserved KCS domains as keywords, the number of KCS genes was found to be higher than in Arabidopsis (21). This might be related to the large amount of genomic data for barley as well as the high number of repeated sequences. Following sequence alignment and phylogenetic tree construction, the grouping and evolutionary relationships of the barley KCS gene family were determined. The phylogenetic tree categorized the barley KCS gene family into nine subgroups, of which five groups were distributed in A. thaliana. Several KCS genes homologous to A. thaliana (Wang1, WSL4, ONI1, ONI2, etc.) have also been cloned (Wang et al. 2017; Yu et al. 2008), indicating that the basic characteristics of this gene family were formed before the differentiation of monocotyledonous and dicotyledonous plants. In addition, four barley KCS subgroups (HORVV-I ~ IV) contained only a large amount of barley KCS proteins that mostly existed in the form of paralogous proteins within the species. However, there were still three pairs of highly similar orthologous proteins present in Arabidopsis and barley, suggesting that the evolutionary process of the KCS gene family is more complex. There were genes from the vertical lineage evolution during speciation, as well as genes from duplication events and species-specific expansion, which may also be related to the nonuniform replication of ancestral KCS genes (Guo et al. 2016), as has been similarly verified in studies of other plant gene families (Li et al. 2016; Xu et al. 2012; Song et al. 2014). For now, the Ka/Ks values reflect that the KCS genes in barley exhibited positive selection, and their related mutations were also favorable.
The analysis of chromosome location, gene structure, and protein motif revealed that the physical location of barley KCS coding genes was not in the same or adjacent intergenic region; thus, the KCS gene family in barley, like the ARF gene family, mainly expanded through large-scale segmental or whole genome duplications (WGDs) rather than by relying on tandem replication (Guo et al. 2016). In addition, studies on noncoding regions have found that intron length has a strong negative correlation with the expression level of corresponding genes. The intron length of low-level expressed genes was significantly larger than that of high-level expressed genes (Shan 2011), and an increase in intron number will lead to increases in the structural number of nuclear localization signal protein motifs, thus affecting the levels of protein entering the nucleus and the proficiency of DNA binding (Johnston et al. 2005). In this study, most members of the KCS gene family in barley did not contain introns, and with the exception of HORVU4Hr1G076940, which consisted of three introns, there was only one intron in the other genes. Similarly, except for HORVU7Hr1G023530 with an intron length of more than 10 kb, the lengths of the introns in the other gene members were within 3 kb. These conclusions reflect the high expression level and function of the KCS gene from a gene structure perspective.
The phylogenetic tree and KCS gene expression pattern analysis in barley further predicted and revealed the function of the barley KCS genes. The FDH gene in Arabidopsis mutants affected the morphology of the leaves and flowers. Consistent with this, in barley, HORVU4Hr1G076940 within the FDH-like (ε) subgroup was also mainly expressed in the flowers and tender leaves, but not in the root tissues (Joubès et al. 2008; Pruitt et al. 2000). HORVU4Hr1G067340 was not expressed in the inflorescence and root tissues. The function of the KCS5 and KCS6 genes, which were in the same subgroup of CER6 (γ), was also mainly reflected in the synthesis of stem and pollen epidermal wax (Millar et al. 1999; Hooker et al. 2002). However, HORVU4Hr1G063420 showed high expression in the developing inflorescence tissues, whereas the orthologous gene KCS1 of HORVU4Hr1G063420 in Arabidopsis was expressed in all parts of the plant (Lee and Suh 2013). Combined with a study on the KCS gene in Arabidopsis, it was confirmed that the expression pattern and functional characteristics of the KCS gene overlap (Paul et al. 2006; Tresch et al. 2012). This suggests that in addition to barley KCS gene retaining a certain conservative structure and function in its evolution, some sub- and neo-functionalization mutations which were in stable evolutionary screening have also been generated.
By analyzing the expression pattern of the barley KCS genes under drought stress, it was found that the drought signal could induce the expression of barley KCS gene family members at the transcription level, ultimately manifesting as changes in the content and composition of barley epidermal wax. Zhang et al. (2012) reported that barley epidermal wax content and water use efficiency were significantly positively correlated under drought conditions, in that an increase in epidermal wax content could effectively improve barley drought resistance. This conclusion has also been confirmed in other species (Liu et al. 2019, 2020; Su et al. 2020; Jetter and Schäffer 2001). In this study, half of the barley KCS genes could be actively upregulated to promote the synthesis of epidermal waxy component precursor VLCFAs and provide the necessary conditions for increasing the epidermal wax content (Haslam and Kunst 2013). Conversely, an analysis of most relevant mutants to date showed that an increase in epidermal wax permeability is usually accompanied by ultrastructural abnormalities in the stratum corneum, and the wax content was also found to have no correlation with the water limitation capacity of the stratum corneum (Fich et al. 2016; Kissinger et al. 2005; Leide et al. 2007). Furthermore, given that the barley KCS gene has definite and unique substrate specificity, its diverse expression patterns would cause the epidermal wax components, rather than the content, to change dynamically in order to affect the drought resistance of barley plants (Vogg et al. 2004). For example, the C29 ketone could effectively reduce the permeability of the cuticle to increase the drought resistance of plants (Liu et al. 2020). In this study, the barley KCS genes had two positive and negative expression regulation modes under drought stress that could integrate the output of VLCFAs with different chain lengths to change the chemical composition and crystal structure of the epidermal wax and eventually build a more continuous hydrophobic bracket (Fich et al. 2016). In addition, during the evolution of barley KCS genes, there were functional redundancies and overlap caused by sub- and neo-functionalization, which would also make plants maintain redundant functions and generate heavy burden under drought stress (Guo et al. 2016). Therefore, the expression of KCS genes in half of the barley in this study would show a significant decrease, presumably because of the duplication and redundancy of some functions of the KCS genes, making barley more susceptible to drought and other habitat changes. Generally, given the complexity of the evolutionary process of the barley KCS gene family and the regulatory network of the epidermal wax synthesis, the relationship between the barley epidermal wax and the drought resistance of the plant remains a complex issue that needs to be studied more thoroughly under the understanding that the plant epidermal wax content and composition are constantly changing.
This study revealed the origin, replication, expansion, evolution, and possible biochemical functional specialization of the barley KCS gene family using biological information analysis technology. In combination with the real-time measurement of expression under stress, our findings provide a theoretical foundation and reference for the in-depth exploration of biological function in the barley KCS gene family and further genetic breeding analysis of related species.
References
Andersen EJ, Ali S, Reese RN, Yen Y, Neupane S, Nepal MP (2016) Diversity and evolution of disease resistance genes in barley (Hordeum vulgare L.). Evol Bioinform 12:S38085. https://doi.org/10.4137/ebo.s38085
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208. https://doi.org/10.1093/nar/gkp335
Bateman A, Coin L, Durbin R, Finn RD, Hollich V, Griffiths-Jones S, Khanna A, Marshall M, Moxon S, Sonnhammer EL, Studholme DJ, Yeats C, Eddy SR (2004) The Pfam protein families database. Nucleic Acids Res 32:D138–D141. https://doi.org/10.1093/nar/28.1.263
Blacklock BJ, Jaworski JG (2002) Studies into factors contributing to substrate specificity of membrane-bound 3-ketoacyl-CoA synthases. Eur J Biochem 269:4789–4798. https://doi.org/10.1046/j.1432-1033.2002.03176.x
Chen C, Xia R, Chen H, He Y (2018) TBtools, a Toolkit for Biologists integrating various HTS-data handling tools with a user-friendly interface. bioRxiv 289660. https://doi.org/10.1101/289660
Conery JS, Lynch M (2001) Nucleotide substitutions and the evolution of duplicate genes. Pac Symp Biocomput 6:167–178. https://doi.org/10.1142/9789814447362_0018
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676. https://doi.org/10.1093/bioinformatics/bti610
Cui LC, Yang G, Yan J, Pan Y, Nie XJ (2019) Genome-wide identification, expression profiles and regulatory network of MAPK cascade gene family in barley. BMC Genom. https://doi.org/10.1186/s12864-019-6144-9
Dunn TM, Lynch DV, Michaelson LV, Napier JA (2004) A post-genomic approach to understanding sphingolipid metabolism in Arabidopsis thaliana. Ann Bot Lond 93:483–497. https://doi.org/10.1093/aob/mch071
Fich EA, Segerson NA, Rose JKC (2016) The Plant polyester cutin: biosynthesis, structure, and biological roles. Annu Rev Plant Biol. https://doi.org/10.1146/annurev-arplant-043015-111929
Fiebig A, Mayfield JA, Miley NL, Chau S, Fischer RL, Preuss D (2000) Alterations in CER6, a gene identical to CUT1, differentially affect long-chain lipid content on the surface of pollen and stems. Plant Cell 12:2001–2008. https://doi.org/10.2307/3871209
Finn RD, Jody C, William A, Miller BL, Wheeler TJ, Fabian S, Alex B, Eddy SR (2015) HMMER web server: 2015 update. Nucleic Acids Res 1:W30–W38. https://doi.org/10.1093/nar/gkv397
Ghanevati M, Jaworski JG (2001) Active-site residues of a plant membrane-bound fatty acid elongase β-ketoacyl-CoA synthase, FAE1 KCS. Bba-Mol Cell Biol L 1530:77–85. https://doi.org/10.1016/s1388-1981(00)00168-2
Ghanevati M, Jaworski JG (2002) Engineering and mechanistic studies of the Arabidopsis FAE1 β-ketoacyl-CoA synthase, FAE1 KCS. Eur J Biochem 269:3531–3539. https://doi.org/10.1046/j.1432-1033.2002.03039.x
Guo HS, Zhang YM, Sun XQ, Li MM, Hang YY, Xue JY (2016) Evolution of the KCS gene family in plants: the history of gene duplication, sub/neofunctionalization and redundancy. Mol Genet Genom 291:739–752. https://doi.org/10.1007/s00438-015-1142-3
Habachi-Houimli Y, Khalfallah Y, Mezghani-Khemakhem M, Makni H, Makni M, Bouktila D (2018) Genome-wide identification, characterization, and evolutionary analysis of NBS-encoding resistance genes in barley. 3 Biotech 8:453. https://doi.org/10.1007/s13205-018-1478-6
Hall T (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98. https://doi.org/10.1021/bk-1999-0734.ch008
Hall BG (2013) Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol 30:1229–1235. https://doi.org/10.1093/molbev/mst012
Haslam TM, Kunst L (2013) Extending the story of very-long-chain fatty acid elongation. Plant Sci 210C:93–107. https://doi.org/10.1016/j.plantsci.2013.05.008
Hooker TS, Millar AA, Kunst L (2002) Significance of the expression of the CER6 condensing enzyme for cuticular wax production in Arabidopsis. Plant physiol 129:1568–1580. https://doi.org/10.1104/pp.003707
Horton P, Park KJ, Obayashi T, Nakai K (2006) Protein subcellular localization prediction with WoLF PSORT. Proc Asian Pac Bioinform Conf. https://doi.org/10.1142/9781860947292_0007
Hu B, Jin J, Guo AY, Zhang H, Gao G (2014) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296. https://doi.org/10.1093/bioinformatics/btu817
James DW Jr, Lim E, Keller J, Plooy I, Ralston E, Dooner HK (1995) Directed tagging of the Arabidopsis FATTY ACID ELONGATION1 (FAE1) gene with the maize transposon activator. Plant Cell 7:309–319. https://doi.org/10.2307/3869853
Jetter R, Schäffer S (2001) Chemical composition of the Prunus laurocerasus leaf surface. Dynamic changes of the epicuticular wax film during leaf development. Plant Physiol 126:1725–1737. https://doi.org/10.1104/pp.126.4.1725
Johnston CA, Willard FS, Jezyk MR, Fredericks Z, Bodor ET, Jones MB, Blaesius R, Watts VJ, Harden TK, Sondek J, Ramer JK, Siderovski DP (2005) Structure of Galpha(i1) bound to a GDP-selective peptide provides insight into guanine nucleotide exchange. Structure 13:1069–1080. https://doi.org/10.1016/j.str.2005.04.007
Joubès J, Raffaele S, Bourdenx B, Garcia C, Laroche-Traineau J, Moreau P, Domergue F, Lessire R (2008) The VLCFA elongase gene family in Arabidopsis thaliana: phylogenetic analysis, 3D modelling and expression profiling. Plant Mol Biol 67:547–566. https://doi.org/10.1007/s11103-008-9339-z
Kajikawa M, Yamaoka S, Yamato KT, Kanamaru H, Ohyama K (2003) Functional analysis of a β-Ketoacyl-CoA synthase gene, MpFAE2, by gene silencing in the liverwort Marchantia polymorpha L. Biosci Biotechnol Biochem 67:605–612. https://doi.org/10.1271/bbb.67.605
Kajikawa M, Yamato KT, Kanamaru H, Sakuradani E, Shimizu S, Fukuzawa H, Sakai Y, Ohyama K (2003) MpFAE3, a β-Ketoacyl-CoA synthase gene in the liverwort Marchantia polymorpha L., is preferentially involved in elongation of palmitic acid to stearic acid. J Agr Chem Soc Jpn 67:1667–1674. https://doi.org/10.1271/bbb.67.1667
Kim J, Jin HJ, Lee SB, Go YS, Kim HJ, Cahoon R, Cahoon EB, Markham JE, Mi CS (2013) Arabidopsis 3-ketoacyl-CoA synthase 9 is involved in the synthesis of tetracosanoic acids as precursors of cuticular waxes, suberins, sphingolipids, and phospholipids. Plant Physiol 162:567–580. https://doi.org/10.1104/pp.112.210450
Kissinger M, Tuvia-Alkalai S, Shalom Y, Fallik E, Elkind Y, Jenks MA, Goodwin MS (2005) Characterization of physiological and biochemical factors associated with postharvest water loss in ripe pepper fruit during storage. J Am Soc Hortic Sci 130:735–741. https://doi.org/10.21273/jashs.130.5.735
Kunst L, Taylor D, Underhill EW (1992) Fatty acid elongation in developing seeds of Arabidopsis thaliana. Plant Physiol Biochem 30:425–434
Lalitha S (2000) Primer premier 5. Biotech Softw Internet Rep 1:270-272
Lassner MW, Metz LJG (1996) A jojoba beta-ketoacyl-CoA synthase cDNA complements the canola fatty acid elongation mutation in transgenic plants. Plant Cell 8:281–292. https://doi.org/10.1105/tpc.8.2.281
Lee SB, Suh MC (2013) Recent advances in cuticular wax biosynthesis and its regulation in Arabidopsis. Mol Plant 6:246–249. https://doi.org/10.1093/mp/sss159
Leide J, Hildebrandt U, Reussing K, Riederer M, Vogg G (2007) The developmental pattern of tomato fruit wax accumulation and its impact on cuticular transpiration barrier properties: effects of a deficiency in a beta-ketoacyl-Coenzyme a synthase (LeCER6). Plant Physiol 144:1667–1679. https://doi.org/10.1104/pp.107.099481
Lessire R, Chevalier S, Lucet-Levannier K, Lellouche JP, Mioskowski C, Cassagne C (1999) Study of the 3-hydroxy eicosanoyl-coenzyme a dehydratase and (E)-2,3 enoyl-coenzyme a reductase involved in acyl-coenzyme a elongation in etiolated leek seedlings. Plant Physiol 119:1009–1016. https://doi.org/10.1104/pp.119.3.1009
Letunic l, Bork P (2007) Interactive tree of life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23:127–128. https://doi.org/10.1093/bioinformatics/btl529
Letunic I, Doerks T, Bork P (2012) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res 40:D302–D305. https://doi.org/10.1093/nar/gkr931
Li J, Liu X (2019) Genome-wide identification and expression profile analysis of the Hsp20 gene family in barley (Hordeum vulgare L.). PeerJ 7:e6832. https://doi.org/10.7717/peerj.6832
Li H, Shi Q, Zhang ZB, Zeng LP, Ma H (2016) Evolution of the LRR-RLK gene family: ancestral copy number and functional divergence of BAM1 and BAM2 in Brassicaceae. J Syst Evol 54:204–218. https://doi.org/10.1111/jse.12206
Li YY, Li J, Liu CN (2017) Genome-wide analysis of SBP-Box transcription factor gene family in Medicago Truncatula. J Nat Sci Hunan Normal Univ 40:24–33. https://doi.org/10.7612/j.issn.1000-2537.2017.06.004
Li C, Haslam TM, Anna K, Schneider LM, Kohei M, Lacey S, Yang H, Ljerka K, Ulrich S, Christiane N (2018) The β-Ketoacyl-CoA Synthase HvKCS1, encoded by Cer-zh, plays a key role in synthesis of barley leaf wax and germination of barley powdery mildew. Plant Cell Physiol 4:806–822. https://doi.org/10.1093/pcp/pcy020
Liu CQ, Wu F, Ren MD, Shi XL (2019) Identification and analysis of the KCS gene family in Microalgae Monoraphidium neglectum. Mol Plant Breed 1–20
Liu XN, Gao S, Liu Y, Cao BL, Chen ZJ, Xu K (2020) Comparative analysis of the chemical composition and water permeability of the cuticular wax barrier in Welsh onion (Allium fistulosum L.). Protoplasma 257:833–840. https://doi.org/10.1007/s00709-019-01470-3
Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Bryant SH (2015) CDD: NCBI’s conserved domain database. Nucleic Acids Res 43:D222–D226. https://doi.org/10.1093/nar/gku1221
Mietkiewska E, Giblin EM, Wang S, Barton DL, Dirpaul J, Brost JM, Katavic V, Taylor DC (2004) Seed-specific heterologous expression of a nasturtium FAE gene in Arabidopsis results in a dramatic increase in the proportion of erucic acid. Plant Physiol 136:2665–2675. https://doi.org/10.1104/pp.104.046839
Millar AA, Clemens S, Zachgo S, Giblin EM, Kunst TL (1999) CUT1, an Arabidopsis gene required for cuticular wax biosynthesis and pollen fertility, encodes a very-long-chain fatty acid condensing enzyme. Plant Cell 11:825–838. https://doi.org/10.2307/3870817
Paul S, Gable K, Beaudoin F, Cahoon E, Jaworski J, Napier JA, Dunn TM (2006) Members of the Arabidopsis FAE1-like 3-ketoacyl-CoA synthase gene family substitute for the Elop proteins of Saccharomyces cerevisiae. J Biol Chem 281:9018–9029. https://doi.org/10.1074/jbc.M507723200
Peterson GI, Masel J (2009) Quantitative prediction of molecular clock and Ka/Ks at short timescales. Mol Biol Evol 26:2595–2603. https://doi.org/10.1093/molbev/msp175
Poole RL (2005) The TAIR Database. Plant Bioinf 406:179–212. https://doi.org/10.1007/978-1-59745-535-0_8
Pruitt RE, Vielle-Calzada JP, Ploense SE, Grossniklaus U, Lolle SJ (2000) FIDDLEHEAD, a gene required to suppress epidermal cell interactions in Arabidopsis, encodes a putative lipid biosynthetic enzyme. Proc Natl Acad Sci USA 97:1311–1316. https://doi.org/10.1073/pnas.97.3.1311
Raffaele S, Vailleau F, Léger A, Joubès J, Miersch O, Huard C, Blée E, Mongrand S, Domergue F, Roby D (2008) A MYB transcription factor regulates very-long-chain fatty acid biosynthesis for activation of the hypersensitive cell death response in Arabidopsis. Plant Cell 20:752–767. https://doi.org/10.1105/tpc.107.054858
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34:3299–3302. https://doi.org/10.1093/molbev/msx248
Shan YJ (2011) Study on the relationship between intron, gene spacer length and gene expression. Hebei Univ. https://doi.org/10.7666/d.y1634211
Song X, Liu G, Duan W, Liu T, Huang Z, Ren J, Li Y, Hou X (2014) Genome-wide identification, classification and expression analysis of the heat shock transcription factor family in Chinese cabbage. Mol Genet Genom 289:541–551. https://doi.org/10.1007/s00438-014-0833-5
Su R, Chen L, Wang Z, Hu Y (2020) Differential response of cuticular wax and photosynthetic capacity by glaucous and non-glaucous wheat cultivars under mild and severe droughts. Plant Physiol Biochem 147:303–312. https://doi.org/10.1016/j.plaphy.2019.12.036
Todd J, Post-Beittenmiller D, Jaworski JG (1999) KCS1 encodes a fatty acid elongase 3-ketoacyl-CoA synthase affecting wax biosynthesis in Arabidopsis thaliana. Plant J 17:119–130. https://doi.org/10.1046/j.1365-313x.1999.00352.x
Tong T, Fang YX, Zhang ZL, Zheng JJ, Xue DW (2019) Genome-wide identification, phylogenetic and expression analysis of SBP-box gene family in barley (Hordeum vulgare L.). Plant Growth Regul 90:1–13. https://doi.org/10.1007/s10725-019-00559-2
Tresch S, Heilmann M, Christiansen N, Looser R, Grossmann K (2012) Inhibition of saturated very-long-chain fatty acid biosynthesis by mefluidide and perfluidone, selective inhibitors of 3-ketoacyl-CoA synthases. Phytochemistry 76:162–171. https://doi.org/10.1016/j.phytochem.2011.12.023
Vogg G, Fischer S, Leide J, Emmanuel E, Jetter R, Levy A, Riederer M (2004) Tomato fruit cuticular waxes and their effects on transpiration barrier properties: functional characterization of a mutant deficient in a very-long-chain fatty acid beta-ketoacyl-CoA synthase. J Exp Bot 55:1401–1410. https://doi.org/10.1093/jxb/erh149
Wang X, Guan Y, Zhang D, Dong X, Tian L, Qu LQ (2017) A β-Ketoacyl-CoA synthase is involved in rice leaf cuticular wax synthesis and requires a CER2-LIKE protein as a cofactor. Plant Physiol 173:944–955. https://doi.org/10.1104/pp.16.01527
Weidenbach D, Jansen M, Bodewein T, Nagel KA, Schaffrath U (2015) Shoot and root phenotyping of the barley mutant kcs6 (3-ketoacyl-CoA synthase6) depleted in epicuticular waxes under water limitation. Plant Signal Behav 10:1–3. https://doi.org/10.1080/15592324.2014.1003752
Wettstein-Knowles PV (1982) Genes, elongases and epicuticular wax biosynthesis. Plant Physiol Biochem 20:797–809. https://doi.org/10.1007/978-1-4684-5263-1_88
Wu YH (2012) Expression profile and functional characterization of KCS gene family in Brassica napus. The Chinese Academy of Agricultural Sciences, Beijing, China
Xiao GH, Wang K, Huang G, Zhu YX (2016) Genome-scale analysis of the cotton KCS gene family revealed a binary mode of action for gibberellin a regulated fiber growth. J Integr Plant Biol 58:577–589. https://doi.org/10.1111/jipb.12429
Xiong JS, Jiang YT, Wang L, CHENG ZM (2019) Evolution of the R2R3-MYB gene family in six Rosaceae species and expression in woodland strawberry. J Integr Agric 12:2753–2770. https://doi.org/10.1016/s2095-3119(19)62818-2
Xu X, Qiu J, Xu Y, Xu C-W (2012) Molecular evolution and expression analysis of subfamily ABCB transporter genes in rice. Chi J Rice Sci 26:127–136. https://doi.org/10.3969/j.issn.1001-7216.2012.02.001
Xue D, Zhang XQ, Lu XL, Chen G, Chen ZH (2017) Molecular and evolutionary mechanisms of cuticular wax for plant drought tolerance. Front Plant Sci 8:621–633. https://doi.org/10.3389/fpls.2017.00621
Yi T, Zhang ZS, Tang BQ, Xie LL, Zou XX (2020) Identification and expression analysis of the KCS gene family in pepper. Acta Hortic Sin 47:1–12. https://doi.org/10.16420/j.issn.0513-353x.2019-0574
Yu D, Ranathunge K, Huang H, Pei Z, Franke R, Schreiber L, He C (2008) Wax crystal-sparse leaf1 encodes a β-ketoacyl CoA synthase involved in biosynthesis of cuticular waxes on rice leaf. Planta 228:675–685. https://doi.org/10.1007/s00425-008-0770-9
Zhang HL, Qi JC, Wang XJ (2012) Effects of water stress on epicuticular wax content and main physiological parameters of barley. J Triticeae Crops 2:280–283. https://doi.org/10.7606/j.issn.1009-1041.2012.02.016
Zhang XQ, Xue DW, Fei-Bo WU, Zhang GP (2013) Genotypic and environmental variations of arabinoxylan content and endoxylanase activity in barley grains. J Integr Agric 008:1489–1494. https://doi.org/10.1016/s2095-3119(13)60555-9
Zhang Y, Li D, Zhang DB, Zhao XG, Cao XM, Dong LL, Liu JX, Chen KL, Zhang HW, Gao CX, Wang DW (2018) Analysis of the functions of TaGW2 homoeologs in wheat grain weight and protein content traits. Plant J 94:857–866. https://doi.org/10.1111/tpj.13903
Zhang MY, Kim YJ, Zong J, Lin H, Dievart A, Li HJ, Zhang DB, Liang WQ (2019) Genome-wide analysis of the barley non-specific lipid transfer protein gene family. Crop J 007:65–76. https://doi.org/10.1016/j.cj.2018.07.009
Zhu J, Fan Y, Li C, Shabala S, Zhao C, Hong Y, Lv C, Guo B, Xu R, Zhou M (2020) Candidate genes for salinity tolerance in barley revealed by RNA-seq analysis of near-isogenic lines. Plant Growth Regul. https://doi.org/10.1007/s10725-020-00662-9
Acknowledgements
This research was supported by the National Natural Science Foundation of China (31401316) and Hangzhou Scientific and Technological Program (20140432B03). We thank LetPub (www.letpub.com) for its linguistic assistance during the preparation of this manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tong, T., Fang, Yx., Zhang, Z. et al. Genome-wide identification and expression pattern analysis of the KCS gene family in barley. Plant Growth Regul 93, 89–103 (2021). https://doi.org/10.1007/s10725-020-00668-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-020-00668-3