Abstract
Wide ranges of microorganisms found in rhizhosphere are able to regulate plant growth and development through production of phytohormones. However, little is known about the effects of these rhizobacteria on phytohormone cytokinin signal transduction in plants. We here found that Bacillus sp. LZR216 can promote growth and alter the root system architecture of Arabidopsis. To study the roles of cytokinin in this process, we used a series of cytokinin signaling mutants. We found that LZR216 inoculation exhibited little effects on the growth promotion and root development of the triple mutant ahp2,3,5. However, LZR216 promoted the growth and altered the root system architecture of single and double mutants of the cytokinin receptor. Furthermore, LZR216 treatment decreased the ARR5-GUS expression in the shoot apical meristem, but increased the expression in root tips. Transcript analysis indicated that LZR216 down-regulates the expressions of AHK3/AHK4, AHP1, and ARR4/5/7/10/12/15 in shoots, but up-regulates the expressions of AHK3/AHK4, AHP1/AHP3, and ARR4/5/7/10/12/15 in roots. Collectively, LZR216 plays different roles in cytokinin signaling between roots and shoots in Arabidopsis, and an intact cytokinin-signaling pathway is necessary for LZR216-promoted plant growth and root system architecture alteration.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As sessile, multicellular organisms, plants rely on developmental and metabolic changes for growth. Roots are plant organs that typically lie below the soil surface, where they grow and respond to a variety of environmental cues. Plant root systems display considerable plasticity in the morphology in response to variable environments. In contrast to primary roots, which are formed during embryogenesis, lateral and adventitious roots are formed postembryonically. Root growth depends on two basic developmental processes: cell division in the root apical meristem and cell elongation that leaves the root meristem (Beemster and Baskin 1998).
Recently, progresses have been made in improving our understanding on the molecular mechanism of cytokinin signaling. The Arabidopsis cytokinin receptor kinases [Arabidopsis HISTIDINE KINASE 2 (AHK2), AHK3, AHK4/CYTOKININ RESPONSE 1 (CRE1)/WOODENLEG (WOL)] display a cyclase/histidine-kinase-associated extracellular (CHASE) domain at N-termini of the proteins. The CHASE domain is the cytokinin-binding region (Anantharaman and Aravind 2001; Mougel and Zhulin 2001). The expression of AHK::GUS fusion genes suggests that AHK genes are expressed ubiquitously, but at low levels (Nishimura et al. 2004). The Arabidopsis genome has five AHP genes and one pseudo AHP gene. The five AHPs contain a highly conserved His residue, which is required for the transfer of phosphate. However, in AHP6, the His residue is replaced by Asn, which makes AHP6 unable to transfer phosphoryl groups to downstream response-regulator genes (ARRs) (Mähönen et al. 2006). Arabidopsis has 23 known ARR genes, which are classified into two major groups based on their sequence similarities, domain structure, and transcriptional responses to cytokinins (D’Agostino et al. 2000). The expression level of type-A ARRs is rapidly elevated in response to exogenous cytokinins. Thus, these ARRs are considered to be primary response genes (D’Agostino et al. 2000; Kiba et al. 2002; Rashotte et al. 2003). The expression of type-B ARRs is not altered by cytokinin. ARR genes have high level of functional redundancy in each of these families (Thomas et al. 1997; Mason et al. 2004; Tajima et al. 2004).
Root development is regulated by a complex network of interactions between different hormones. Cytokinins promote and maintain plant cell division in cultures and are also involved in various differentiation processes including shoot growth, primary root growth, leaf senescence, and seed germination (Riefler et al. 2006). Several cytokinin signal transduction and biosynthesis genes are differentially expressed in root tissues (Higuchi et al. 2004; Mason et al. 2004; To et al. 2004), suggesting that cytokinins play important and complex roles during root development. Auxin plays as important roles as cytokinins in many aspects of plant growth and development (Coenen and Lomax 1997; Rashotte et al. 2005). In fact, auxin and cytokinins interact at different levels. For example, cytokinin regulation of auxin biosynthesis in Arabidopsis involves a homeostatic feedback loop, which is regulated via auxin and cytokinin signal transduction (Jones et al. 2010). In addition, cytokinin regulates root meristem activity via modulation of the polar auxin transport (Růžička et al. 2009). Therefore, auxin and cytokinin play important roles in the regulation of root development.
Plant growth-promoting rhizobacteria (PGPR) are natural rhizosphere-inhabiting bacteria. PGPR have positive influences on plant growth and development (Bloemberg and Lugtenberg 2001; Persello-Cartieaux et al. 2003; Kurepin et al. 2015; Islam et al. 2016). Belonging to diverse genera, such as Pseudomonas, Azospirillum and Bacillus, PGPR have been recognized in a wide range of plant species, such as barley, rice, canola, bean, and Arabidopsis (Alstrom 1991; Alexandre et al. 1996; Persello-Cartieaux et al. 2001). The effects of PGPR on plant growth can be exerted by mechanisms including secretion of plant growth-regulating substances such as auxins, cytokinins, and bacterial volatiles (Selvadurai et al. 1991; Lebuhn et al. 1997; Ryu et al. 2003; Arkhipova et al. 2005), and activation of plant defense responses (Pieterse et al. 1996; Schuhegger et al. 2006). Another study showed that the beneficial fungus Trichoderma atroviride regulates the transcription of Arabidopsis WRKY transcription factors, which have been demonstrated to be involved in responses to plant–pathogen interactions (Sáenz-Mata et al. 2014). Bacterial secretion of phytohormones can affect the root architecture by overproducing root hairs and lateral roots and subsequently increase nutrient and water uptake, thus contributing to root growth (Persello-Cartieaux et al. 2003). Furthermore, it was reported that two halotolerant Bacilli sp. isolated from saline habitats of Gujarat alleviate the salt stress in germination of Vigna radiata L. (Patel et al. 2015) and Burkholderia phytofirmans strain PsJN enhances wheat drought tolerance (Naveed et al. 2014).
Cytokinins can be produced by microorganisms (García de Salamone et al. 2001). Thus, plants inoculated with bacterial species capable of producing cytokinins may have increased levels of cytokinins in root tissues. An early report provided important information on the role played by cytokinin receptors in plant growth promotion by rhizobacterium Bacillus megaterium (Ortíz-Castro et al. 2008). Bacillus megaterium alters the root system architecture in inoculated plants, including inhibition in primary root growth, increase in lateral root formation and root hair length. However, in the cre1-12/ahk2-2tk/ahk3-3 triple mutant seedlings, bacterial inoculation fails to stimulate the root growth and development (Ortíz-Castro et al. 2008).
Endophytic Bacillus sp. LZR216 was isolated and identified from Arabidopsis roots, and our previous study showed that Bacillus sp. LZR216 can promote Arabidopsis shoot growth and alter root system architecture (Wang et al. 2015). Although the signaling network between plants and rhizobacteria has been extensively studied over the past 20 years, very few studies on molecular signaling and hormone signaling involved in the interaction of bacteria and plants have been reported until recently (Persello-Cartieaux et al. 2003; Zamioudis et al. 2013). Our work expands the existing knowledge by showing that growth stimulation by Bacillus sp. LZR216 requires an intact cytokinin-signaling pathway in Arabidopsis.
Materials and methods
Materials and growth conditions
Arabidopsis Col-0 was used as wild-type plants. The following mutants were used in this study: ahk2-5, ahk3-7, cre1-2, ahk2-5/ahk3-7, ahk2-5/cre1-2, ahk3-7/cre1-2, ahp1,2,3, ahp2,3,5, arr3,4, arr3,4,5,6, arr3,4,5,6,8,9, arr5,6,8,9, arr8,9, arr10,12. Additionally, ARR5-GUS transgenic lines were also used in this study. Bacillus sp. LZR216 was isolated from surface-sterilized roots of wild-type Arabidopsis Col-0 seedlings (Wang et al. 2015) and was cultured in KB medium, incubated at 28 °C on a shaker at 150 rpm for 24 h (see below for details). Seeds of wild-type and mutants were surface-sterilized and stratified at 4 °C for 2 days before sowing on 1/2 MS agar-solidified medium. To test the effects of LZR216 on plant growth, 5-days-old seedlings were transferred and positioned vertically in a growth chamber under a long photoperiod (16 h light/8 h dark) and 65–70% relative humidity at 22 °C.
Cultivation of LZR216 and induction treatments
Bacillus sp. LZR216 was cultured in KB liquid medium with constant shaking at 150 rpm for 24 h at 28 °C. After 24 h of growth, cells were collected and washed twice in sterile water by centrifugation for 10 min at 7500 rpm, and resuspended in sterile water. The bacterial titer was adjusted to an optical density at 600 nm of 0.5 (5 × 108 colony-forming units mL−1). 20 µL of bacterial suspension (sterile water was used as the control) were spotted on 1/2 MS petri plates. In order to test the effect of LZR216 on promoting Arabidopsis growth, seedlings were grown on horizontal plates for 12 days, and then the shoot fresh weight and total leaf surface area were measured. To examine the effect of LZR216 on the root system architecture, seedlings were grown on vertical plates for 7 days, and then the primary root length was measured and the lateral root number per plant was counted.
GUS histochemical staining
Histochemical assay of the GUS activity was performed as described by Ulmasov et al. (1997) with minor modifications. Five-day-old seedlings were treated with LZR216 for 7 days, then incubated in GUS-staining buffer containing 1 mM X-Gluc, 100 mM sodium phosphate (pH 7.5), 0.5 mM potassium ferricyanide, 0.5 mM potassium ferrocyanide, 10 mM EDTA, and 0.1% Triton X-100. The seedlings were incubated at 37 °C for 12 h. Stained seedlings were cleared in a mixture of chloral hydrate:glycerol:water (8:1:2, w/v/v) and observed with an optical microscope.
Quantitative real-time PCR analysis
Total RNA was extracted with Trizol (TaKaRa) from shoots and roots, and then treated with RNase-free DNase (Promega, Madison, WI, USA). First-strand cDNA was synthesized with the PrimeScript II 1st Strand cDNA Synthesis Kit (TaKaRa, Mountain View, CA, USA). Quantitative real-time PCR was performed using the SYBR PrimeScript RT-PCR Kit (Perfect Real Time; TaKaRa). PCR was carried out using a CFX 96 Real-Time system (Bio-Rad, Hercules, CA, USA) with the following standard cycling conditions: 95 °C for 10 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 30 s. Primer sequences used in the study were listed in Supplementary Table 1. The cycle threshold 2[−ΔΔC(T)]-based method was used for relative quantitation of gene expression. Expression levels of genes were normalized to ACTIN2.
Statistical analysis
Each experiment was repeated at least three times. Values were expressed as means ± SE. Statistical analyses were carried out with one way analysis of variance (ANOVA) followed by Duncan’s multiple range test for independent samples or with Student’s t test when two means were compared (SPSS version 17.0). In all cases, the confidence coefficient was set at P < 0.05.
Results
Roles of cytokinin receptors in plant growth stimulation and root system architecture alteration by LZR216
Cytokinins are a class of phytohormones produced by plants and microorganisms (Holland 1997; Arkhipova et al. 2005). The production of cytokinins by PGPR has been well documented and correlated with increased plant growth (Nieto and Frankenberger 1990; García de Salamone et al. 2001; Arkhipova et al. 2005). To investigate whether LZR216-stimulated plant growth involves cytokinin signaling, we compared the responses of wild-type (Col-0) and cytokinin receptor mutants ahk2-5, ahk3-7, cre1-2, ahk2-5/ahk3-7, ahk2-5/cre1-2 and ahk3-7/cre1-2 to LZR216 inoculation. After 12 days of horizontal growth in the presence of LZR216, a significant increase in total leaf surface area and shoot biomass production was observed in seedlings of single or double cytokinin receptor mutants (Fig. 1). After 7 days of vertical growth in the presence of LZR216, the primary root length of all tested mutant seedlings exposed to LZR216 was reduced compared to the control (Fig. 2a), but the lateral number per plant and the lateral root density of seedlings were enhanced, with similar degrees to the control (Fig. 2b, c). Hence, in single and double cytokinin receptor mutants, LZR216 can promote the growth and alter the root system architecture as in wild-type.
Effects of LZR216 inoculation on the growth of cytokinin receptor mutant seedlings. a Photograph of 17-days-old cytokinin receptor mutant seedlings grown on 1/2 MS medium. 5-days-old seedlings were treated with 20 µL 5 × 108 CFU mL−1 suspension in water (20 µL sterilized water as the control) for 12 days and photographed. Bar 1 cm. b The total leaf surface area of 12-days-old horizontally grown seedlings. c Shoot fresh weight of 12-days-old horizontally grown seedlings. Data are means ± SE of three independent experiments (n = 20). Bars with different letters are significantly different at the level of P < 0.05. The experiment was repeated three times with similar results
Effects of LZR216 inoculation on the root system architecture of cytokinin receptor mutant seedlings. a The primary root length of 7-days-old vertically grown seedlings. b The lateral root numbers of 7-days-old vertically grown seedlings. c The lateral root density of 7-days-old vertically grown seedlings. Data are means ± SE of three independent experiments (n = 20). Bars with different letters are significantly different at the level of P < 0.05. The experiment was repeated three times with similar results
Roles of AHP family genes in growth stimulation and root system architecture alteration by LZR216
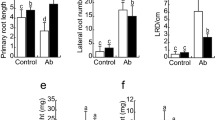
The AHP gene family is important for cytokinin signal transduction. To investigate whether AHPs play some roles during the growth promotion and root system architecture alteration caused by LZR216, we used several AHP family mutants in our subsequent experiments. Results showed that LZR216 significantly increased the total leaf surface area ([−] for mock treatment using H2O, 0.17 ± 0.014, [+] for LZR216 inoculation, 0.99 ± 0.804) and shoot fresh weight ([−] 73.33 ± 1.214, [+] 213.90 ± 34.538) in the triple mutant ahp1,2,3. But no significant increase in total leaf surface area ([−] 0.071 ± 0.0057, [+] 0. 057 ± 0.006) and shoot fresh weight ([−] 25.50 ± 1.389, [+] 26.97 ± 2.603) were observed in the triple mutant ahp2,3,5 (Fig. 3b, c). Similarly, in the triple mutant ahp1,2,3, the primary root length of seedlings exposed to LZR216 ([+] 3.91 ± 0.168) was significantly reduced compared to the control ([−] 6.41 ± 0.071) (Fig. 3d), whereas the lateral number per plant and the lateral root density of seedlings exposed to LZR216 was enhanced compared to the control. However, the root system architecture of the triple mutant ahp2,3,5 (lateral root number per plant, [−] 0, [+] 0) were not altered by LZR216 (Fig. 3e, f). Taken together, LZR216 inoculation failed to stimulate the growth and root development of the triple mutant ahp2,3,5, which may imply that AHP2, AHP3, AHP5 are required for normal growth and development in response to LZR216 inoculation.
Effects of LZR216 inoculation on the growth of AHP mutant seedlings. a Photograph of 17-days-old AHP mutant seedlings grown on 1/2 MS medium. 5-days-old seedlings were treated with 20 µL 5 × 108 CFU mL−1 suspension in water (20 µL sterilized water as the control) for 12 days and photographed. Bar 1 cm. b Total leaf surface area of 12-days-old horizontally grown seedlings. c Shoot fresh weight of 12-days-old horizontally grown seedlings. d The primary root length of 7-days-old vertically grown seedlings. e The lateral root numbers of 7-days-old vertically grown seedlings. f The lateral root density of 7-days-old vertically grown seedlings. Data are means ± SE of three independent experiments (n = 20). Bars with different letters are significantly different at the level of P < 0.05. The experiment was repeated three times with similar results
Roles of ARR family genes in growth stimulation and root system architecture alteration by LZR216
To investigate the roles of ARR family members in the growth promotion and root system architecture alteration caused by LZR216, we used a series of arr mutants with LZR216 inoculation. Results showed that LZR216 enhances the total leaf surface area and shoot fresh weight in all arr mutants (Fig. 4). Similarly, in all arr mutants, the primary root length of seedlings exposed to LZR216 was reduced compared to the control (Fig. 5a), but the lateral number per plant and lateral root density of seedlings exposed to LZR216 were enhanced compared to the control (Fig. 5b, c).
Effects of LZR216 inoculation on the growth of ARR mutant seedlings. a Photograph of 17-days-old ARR mutant seedlings grown on 1/2 MS medium. 5-days-old seedlings were treated with 20 µL 5 × 108 CFU mL−1 suspension in water (20 µL sterilized water as the control) for 12 days and photographed. Bar 1 cm. b The total leaf surface area of 12-days-old horizontally grown seedlings. c The shoot fresh weight of 12-days-old horizontally grown seedlings. Data are means ± SE of three independent experiments (n = 20). Bars with different letters are significantly different at the level of P < 0.05. The experiment was repeated three times with similar results
Effects of LZR216 inoculation on the root system architecture of ARR mutant seedlings. a The primary root length of 7-days-old vertically grown seedlings. b The lateral root numbers for 7-days-old vertically grown seedlings. c The lateral root density for 7-days-old vertically grown seedlings. Data are means ± SE of three independent experiments (n = 20). Bars with different letters are significantly different at the level of P < 0.05. The experiment was repeated three times with similar results
LZR216 alters the expression of cytokinin signaling genes
The above results indicate that a complete cytokinin-signaling pathway is required for LZR216-promoted growth and root system architecture alteration. As a mechanism in coping with growth promotion and root system architecture alteration, Arabidopsis plants may alter cytokinin-signaling genes in response to LZR216 inoculation. To test this hypothesis, we used the transgene line ARR5::GUS to monitor the expression of cytokinin signaling gene ARR5 following LZR216 treatment. As shown in Fig. 6, LZR216 treatment substantially decreased the ARR5::GUS expression levels in shoot meristem but increased in root tips. LZR216 treatment did not alter the ARR5::GUS levels in the junction between hypocotyls and primary roots. These results, together with the qRT-PCR assays (Fig. 7) revealed that LZR216 down-regulates the expression of ARR4/5/7/10/12/15 in shoots, but up-regulates the expression levels of these ARRs in roots. We also monitored the transcript levels of several cytokinin-signaling genes following LZR216 treatment. As shown in Fig. 7, LZR216 down-regulates the expression of AHK3/AHK4/AHP1 in shoots, but up-regulates the transcript levels of AHK3/AHK4/AHP1/AHP3 in roots. Collectively, our results showed that LZR216 down-regulates the transcript levels of cytokinin-signaling genes in shoots, but up-regulates their expressions in roots.
Quantitative real-time PCR analysis of the relative transcript abundance of gene of cytokinin signaling pathway in WT seedlings. 5-days-old seedlings were treated with 20 µL 5 × 108 CFU mL−1 suspension in water (20 µL sterilized water as the control) for 7 days. All gene transcripts were normalized to ACTIN2. Data are means ± SE of three independent experiments. Asterisks indicate statistically significant differences compared to the control (Student’s t test; P < 0.05). The experiment was repeated three times with similar results
Discussion
Plants and microorganisms have coexisted for millions of years. Plants have complex interactions with microbes that are important for metabolism and immune system of hosts (Brestoff and Artis 2013). PGPR are natural rhizosphere-inhabiting bacteria and generally increase the growth and productivity of plants (Kloepper et al. 1980).
The first step of cytokinin perception requires the binding of cytokinin to a transmembrane histidine kinase receptor (Inoue et al. 2001). Following autophosphorylation, the phosphate group is then transferred to a conserved Asp residue within the receiver domain of the AHK protein, and subsequently transferred to a member of the ARABIDOPSIS HIS PHOSPHOTRANSFER PROTEIN (AHP) family (Hwang and Sheen 2001; Hutchison et al. 2006). In the nucleus, phosphorylated AHPs transfer the phosphate group to the nuclear-localized ARABIDOPSIS RESPONSE REGULATOR (ARR) proteins (Hutchison et al. 2006). Cytokinins play major roles in many aspects of plant growth and development, including cell division, regulation of root and shoot growth and branching, chloroplast development, and leaf senescence (Mok and Mok 2001). Many literatures show that cytokinin can regulate root development through interacting with auxin (Růžička et al. 2009, Su et al. 2011; Chang et al. 2013). Cytokinin signaling is important for microbes-promoted plant growth (Ortíz-Castro et al. 2008). Biosynthesis of tZ- but not cZ-type cytokinins appears to be required for P. indica-induced Arabidopsis growth (Vadassery et al. 2008). Nod factor signaling by rhizobia in the legume Lotus japonicas requires the cytokinin receptor LHK1 for cell division and initiation of nitrogen-fixing nodule development (Murray et al. 2007; Tirichine et al. 2007). Bacillus megaterium promotes plant growth and alters root system architecture, but the effect is completely abolished in the triple mutant cre1-1/ahk2-2/ahk3-3, indicating the importance of an intact cytokinin-signaling pathway in the response to B. megaterium (Ortíz-Castro et al. 2008). The roots of the double mutant cre1/ahk2 do not respond to P. indica, while other double mutants cre1/ahk3 and ahk2/ahk3 respond to the fungus in a manner similar to the wild-type (Vadassery et al. 2008). In this study, we demonstrated that in single or double mutants of CRE1, AHK2, and AHK3, LZR216 promotes the growth in terms of shoot fresh weight and total leaf surface area compared to the control; and the primary root length and lateral root numbers of all cytokinin receptor mutants are inhibited and increased, respectively. A recent research showed that the Pseudomonas type III effector HopQ1 activates cytokinin signaling and interferes with plant innate immunity (Hann et al. 2014), indicating that cytokinin signaling is involved in plant–microbe interactions. The transcript levels of all three cytokinin receptors are not regulated by P. indica (Vadassery et al. 2008). As a mechanism of plant growth promotion and root system architecture alteration, cytokinin signaling may be altered in response to LZR216 treatment, as our results showed that LZR216 up-regulates the transcript levels of AHK3/AHK4 in roots, but down-regulates the transcript levels of AHK3/AHK4 in shoots.
The five Arabidopsis AHP genes are expressed ubiquitously. Single and various double mutants of AHP genes exhibit no differences in cytokinin responsiveness. The triple mutant ahp1/ahp2/ahp3 displays reduced sensitivity to exogenous cytokinin in root elongation assays and the quintuple mutant ahp1/ahp2/ahp3/ahp4/ahp5 is severely impaired in root development (Hutchison et al. 2006), indicating that AHPs are positive, redundant elements in the cytokinin signal transduction pathway (Ferreira and Kieber 2005). So far, studies on AHP genes in the plant–microbe interaction are limited. Our results showed that AHP2/AHP3/AHP5 play a role in LZR216-promoted growth and alteration of root system architecture. LZR216 can promote the growth and alter root morphology of the triple mutant ahp1,2,3 but not the triple mutant ahp2,3,5, suggesting that AHP5 is more important than AHP1 in this process. The qRT-PCR results showed that LZR216 down-regulates the transcript levels of AHP1 in shoots, but up-regulates the transcript levels of AHP1 and AHP3 in roots. Therefore, LZR216 plays different roles on cytokinin signaling between roots and shoots in Arabidopsis.
Study of single and multiple loss-of-function type-A arr mutants, including a hextuple arr3,4,5,6,8,9 mutant, has demonstrated that only multiple type-A arr mutants are hypersensitive to cytokinins (To et al. 2004). Some reports demonstrated that loss-of-function mutants of six type-B ARRs (ARR1, ARR2, ARR10, ARR11, ARR12, and ARR18) show a gradient of inhibition in the root elongation and inhibition of lateral root formation (Mason et al. 2005). So far, studies on arr mutants in plant–microbe interaction are scarce. ARR genes have high level of functional redundancy observed in each of these families (Thomas et al. 1997; Mason et al. 2004; Tajima et al. 2004). We used a series of arr mutants to examine the response to LZR216 treatment. Results showed that LZR216 promotes the growth and alters root morphology of arr mutants. The cytokinin-responsive ARR5 gene is highly expressed in the base of the lateral root primordia and in root tips during lateral root development in Arabidopsis (Lohar et al. 2004). The ARR5 mRNA level in P. indica-treated roots is higher than that in control roots, indicating that cytokinin-regulated genes may play a role in the interaction between P. indica and Arabidopsis (Vadassery et al. 2008). Similarly, our results indicated that LZR216 treatment substantially decreased the ARR5::GUS levels in shoot meristem, but increased in root tips. These results, together with the qRT-PCR assays, revealed that LZR216 down-regulates the expression of ARR4/5/7/10/12/15 in shoots, but up-regulates the transcript levels of ARR4/5/7/10/12/15 in roots. Taken together, Arabidopsis differently alters cytokinin signaling in roots and shoots in response to LZR216 treatment.
References
Alexandre G, Jacoud C, Faure D, Bally R (1996) Population dynamics of a motile and a non-motile Azospirillum lipoferum strain during rice root colonization and motility variation in the rhizosphere. FEMS Microbiol Ecol 19:271–278
Alstrom S (1991) Induction of disease resistance in common bean susceptible to halo blight bacterial pathogen after seed bacterization with rhizosphere pseudomonads. J Gen Appl Microbiol 37:495–501
Anantharaman V, Aravind L (2001) The CHASE domain: a predicted ligand-binding module in plant cytokinin receptors and other eukaryotic and bacterial receptors. Trends Biochem Sci 26:579–582
Arkhipova T, Veselov S, Melentiev A, Martynenko E, Kudoyarova G (2005) Ability of bacterium Bacillus subtilis to produce cytokinins and to influence the growth and endogenous hormone content of lettuce plants. Plant Soil 272:201–209
Beemster GT, Baskin TI (1998) Analysis of cell division and elongation underlying the developmental acceleration of root growth in Arabidopsis thaliana. Plant Physiol 116:1515–1526
Bloemberg GV, Lugtenberg BJ (2001) Molecular basis of plant growth promotion and biocontrol by rhizobacteria. Curr Opin Plant Biol 4:343–350
Brestoff JR, Artis D (2013) Commensal bacteria at the interface of host metabolism and the immune system. Nat Immunol 14:676–684
Chang L, Ramireddy E, Schmülling T (2013) Lateral root formation and growth of Arabidopsis is redundantly regulated by cytokinin metabolism and signalling genes. J Exp Bot 64:5021–5032
Coenen C, Lomax TL (1997) Auxin–cytokinin interactions in higher plants: old problems and new tools. Trends Plant Sci 2:351–356
D’Agostino IB, Deruère J, Kieber JJ (2000) Characterization of the response of the Arabidopsis response regulator gene family to cytokinin. Plant Physiol 124:1706–1717
Ferreira FJ, Kieber JJ (2005) Cytokinin signaling. Curr Opin Plant Biol 8:518–525
García de Salamone IE, Hynes RK, Nelson LM (2001) Cytokinin production by plant growth promoting rhizobacteria and selected mutants. Can J Microbiol 47:404–411
Hann DR, Domínguez-Ferreras A, Motyka V, Dobrev PI, Schornack S, Jehle A, Felix G, Chinchilla D, Rathjen JP, Boller T (2014) The Pseudomonas type III effector HopQ1 activates cytokinin signaling and interferes with plant innate immunity. New Phytol 201:585–598
Higuchi M, Pischke MS, Mähönen AP, Miyawaki K, Hashimoto Y, Seki M, Kobayashi M, Shinozaki K, Kato T, Tabata S (2004) In planta functions of the Arabidopsis cytokinin receptor family. Proc Natl Acad Sci USA 101:8821–8826
Holland MA (1997) Occam’s razor applied to hormonology (are cytokinins produced by plants?) Plant Physiol 115:865
Hutchison CE, Li J, Argueso C, Gonzalez M, Lee E, Lewis MW, Maxwell BB, Perdue TD, Schaller GE, Alonso JM (2006) The Arabidopsis histidine phosphotransfer proteins are redundant positive regulators of cytokinin signaling. Plant Cell 18:3073–3087
Hwang I, Sheen J (2001) Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413:383–389
Inoue T, Higuchi M, Hashimoto Y, Seki M, Kobayashi M, Kato T, Tabata S, Shinozaki K, Kakimoto T (2001) Identification of CRE1 as a cytokinin receptor from Arabidopsis. Nature 409:1060–1063
Islam F, Yasmeen T, Arif MS, Ali S, Ali B, Hameed S, Zhou W (2016) Plant growth promoting bacteria confer salt tolerance in Vigna radiata by up-regulating antioxidant defense and biological soil fertility. Plant Growth Regul 80:23–36
Jones B, Gunnerås SA, Petersson SV, Tarkowski P, Graham N, May S, Dolezal K, Sandberg G, Ljung K (2010) Cytokinin regulation of auxin synthesis in Arabidopsis involves a homeostatic feedback loop regulated via auxin and cytokinin signal transduction. Plant Cell 22:2956–2969
Kiba T, Yamada H, Mizuno T (2002) Characterization of the ARR15 and ARR16 response regulators with special reference to the cytokinin signaling pathway mediated by the AHK4 histidine kinase in roots of Arabidopsis thaliana. Plant Cell Physiol 43:1059–1066
Kloepper J, Schroth M, Miller T (1980) Effects of rhizosphere colonization by plant growth-promoting rhizobacteria on potato plant development and yield. Phytopathology 70:1078–1082
Kurepin LV, Park JM, Lazarovits G, Hüner NP (2015) Involvement of plant stress hormones in Burkholderia phytofirmans-induced shoot and root growth promotion. Plant Growth Regul 77:179–187
Lebuhn M, Heulin T, Hartmann A (1997) Production of auxin and other indolic and phenolic compounds by Paenibacillus polymyxa strains isolated from different proximity to plant roots. FEMS Microbiol Ecol 22:325–334
Lohar DP, Schaff JE, Laskey JG, Kieber JJ, Bilyeu KD, Bird DM (2004) Cytokinins play opposite roles in lateral root formation, and nematode and rhizobial symbioses. Plant J 38:203–214
Mähönen AP, Bishopp A, Higuchi M, Nieminen K, Kinoshita K, Törmäkangas K, Ikeda Y, Oka A, Kakimoto T, Helariutta Y (2006) Cytokinin signaling and its inhibitor AHP6 regulate cell fate during vascular development. Science 311:94–98
Mason MG, Li J, Mathews DE, Kieber JJ, Schaller GE (2004) Type-B response regulators display overlapping expression patterns in Arabidopsis. Plant Physiol 135:927–937
Mason MG, Mathews DE, Argyros DA, Maxwell BB, Kieber JJ, Alonso JM, Ecker JR, Schaller GE (2005) Multiple type-B response regulators mediate cytokinin signal transduction in Arabidopsis. Plant Cell 17:3007–3018
Mok DW, Mok MC (2001) Cytokinin metabolism and action. Annu Rev Plant Biol 52:89–118
Mougel C, Zhulin IB (2001) CHASE: an extracellular sensing domain common to transmembrane receptors from prokaryotes, lower eukaryotes and plants. Trends Biochem Sci 26:582–584
Murray JD, Karas BJ, Sato S, Tabata S, Amyot L, Szczyglowski K (2007) A cytokinin perception mutant colonized by rhizobium in the absence of nodule organogenesis. Science 315:101–104
Naveed M, Hussain MB, Zahir ZA, Mitter B, Sessitsch A (2014) Drought stress amelioration in wheat through inoculation with Burkholderia phytofirmans strain PsJN. Plant Growth Regul 73:121–131
Nieto K, Frankenberger W (1990) Microbial production of cytokinins. Soil Biochem 6:191–248
Nishimura C, Ohashi Y, Sato S, Kato T, Tabata S, Ueguchi C (2004) Histidine kinase homologs that act as cytokinin receptors possess overlapping functions in the regulation of shoot and root growth in Arabidopsis. Plant Cell 16:1365–1377
Ortíz-Castro R, Valencia-Cantero E, López-Bucio J (2008) Plant growth promotion by Bacillus megaterium involves cytokinin signaling. Plant Signal Behav 3:263–265
Patel RR, Patel DD, Thakor P, Patel B, Thakkar VR (2015) Alleviation of salt stress in germination of Vigna radiata L. by two halotolerant Bacilli sp. isolated from saline habitats of Gujarat. Plant Growth Regul 76:51–60
Persello-Cartieaux F, David P, Sarrobert C, Thibaud M-C, Achouak W, Robaglia C, Nussaume L (2001) Utilization of mutants to analyze the interaction between Arabidopsis thaliana and its naturally root-associated Pseudomonas. Planta 212:190–198
Persello-Cartieaux F, Nussaume L, Robaglia C (2003) Tales from the underground: molecular. Plant Cell Environ 26:189–199
Pieterse C, Van Wees S, Hoffland E, Van Pelt JA, Van Loon LC (1996) Systemic resistance in Arabidopsis induced by biocontrol bacteria is independent of salicylic acid accumulation and pathogenesis-related gene expression. Plant Cell 8:1225–1237
Rashotte AM, Carson SD, To JP, Kieber JJ (2003) Expression profiling of cytokinin action in Arabidopsis. Plant Physiol 132:1998–2011
Rashotte AM, Chae HS, Maxwell BB, Kieber JJ (2005) The interaction of cytokinin with other signals. Physiol Plant 123:184–194
Riefler M, Novak O, Strnad M, Schmülling T (2006) Arabidopsis cytokinin receptor mutants reveal functions in shoot growth, leaf senescence, seed size, germination, root development, and cytokinin metabolism. Plant Cell 18:40–54
Růžička K, Šimášková M, Duclercq J, Petrášek J, Zažímalová E, Simon S, Friml J, Van Montagu MC, Benková E (2009) Cytokinin regulates root meristem activity via modulation of the polar auxin transport. Proc Natl Acad Sci USA 106:4284–4289
Ryu C-M, Farag MA, Hu C-H, Reddy MS, Wei H-X, Paré PW, Kloepper JW (2003) Bacterial volatiles promote growth in Arabidopsis. Proc Natl Acad Sci USA 100:4927–4932
Sáenz-Mata J, Salazar-Badillo FB, Jiménez-Bremont JF (2014) Transcriptional regulation of Arabidopsis thaliana WRKY genes under interaction with beneficial fungus Trichoderma atroviride. Acta Physiol Plant 36:1085–1093
Schuhegger R, Ihring A, Gantner S, Bahnweg G, Knappe C, Vogg G, Hutzler P, Schmid M, Van Breusegem F, Eberl L (2006) Induction of systemic resistance in tomato by N-acyl-l-homoserine lactone-producing rhizosphere bacteria. Plant Cell Environ 29:909–918
Selvadurai EL, Brown AE, Hamilton J (1991) Production of indole-3-acetic acid analogues by strains of Bacillus cereus in relation to their influence on seedling development. Soil Biol Biochem 23:401–403
Su Y-H, Liu Y-B, Zhang X-S (2011) Auxin–cytokinin interaction regulates meristem development. Mol Plant 4:616–625
Tajima Y, Imamura A, Kiba T, Amano Y, Yamashino T, Mizuno T (2004) Comparative studies on the type-B response regulators revealing their distinctive properties in the His-to-Asp phosphorelay signal transduction of Arabidopsis thaliana. Plant Cell Physiol 45:28–39
Thomas T, Hare P, Van Staden J (1997) Phytochrome and cytokinin responses. Plant Growth Regul 23:105–122
Tirichine L, Sandal N, Madsen LH, Radutoiu S, Albrektsen AS, Sato S, Asamizu E, Tabata S, Stougaard J (2007) A gain-of-function mutation in a cytokinin receptor triggers spontaneous root nodule organogenesis. Science 315:104–107
To JP, Haberer G, Ferreira FJ, Deruère J, Mason MG, Schaller GE, Alonso JM, Ecker JR, Kieber JJ (2004) Type-A Arabidopsis response regulators are partially redundant negative regulators of cytokinin signaling. Plant Cell 16:658–671
Ulmasov T, Murfett J, Hagen G, Guilfoyle TJ (1997) Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9:1963–1971
Vadassery J, Ritter C, Venus Y, Camehl I, Varma A, Shahollari B, Novák O, Strnad M, Ludwig-Müller J, Oelmüller R (2008) The role of auxins and cytokinins in the mutualistic interaction between Arabidopsis and Piriformospora indica. Mol Plant Microbe Interact 21:1371–1383
Wang JF, Zhang YQ, Li Y, Wang XM, Nan WB, Hu YF, Zhang H, Zhao CZ, Wang F, Li P, Shi HY, Bi YR (2015) Endophytic microbes Bacillus sp. LZR216-regulated root development is dependent on polar auxin transport in Arabidopsis seedlings. Plant Cell Rep 34:1075–1087
Zamioudis C, Mastranesti P, Dhonukshe P, Blilou I, Pieterse CMJ (2013) Unraveling root developmental programs initiated by beneficial Pseudomonas spp. bacteria. Plant Physiol 162:304–318
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31671595; 31670244), Foundation of Science and Technology Program of Gansu Province (1506RJZA209), The Agricultural Biotechnology Research and Application Development Program of Gansu Province (GNSW-2016-23), The Fundamental Research Funds for the Central Universities (lzujbky-2015-ct01; lzujbky-2016-80), The Foundation of Science and Technology Program of Lanzhou City (2015-3-53), The Project of Qinghai Science & Technology Department (2016-ZJ-Y01), The Open Project of State Key Laboratory of Plateau Ecology and Agriculture, Qinghai University (201-KF-05). We thank Ping Li for her help and advice for data analysis during the preliminary stages of this project, and the two anonymous reviewers for reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, J., Zhang, Y., Jin, J. et al. An intact cytokinin-signaling pathway is required for Bacillus sp. LZR216-promoted plant growth and root system architecture altereation in Arabidopsis thaliana seedlings. Plant Growth Regul 84, 507–518 (2018). https://doi.org/10.1007/s10725-017-0357-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0357-1