Abstract
NIN-LIKE PROTEIN (NLP) is a conserved plant-specific transcription factor family and has been shown in several plant species to be a key player in regulating nitrogen (N) response. However, little is known about NLP gene family and their characteristics in maize (Zea mays L.). Here we report the identification and characterization of maize NLPs (ZmNLPs), and illustrate the family structure, phylogenetic properties, expression profiles, genetic differentiation between heterotic groups and N response. A total of 9 ZmNLPs from the maize genome were identified, belonging to two subgroups according to conserved domains and gene structure. Their expression profiles were different across tissues and almost all ZmNLPs constitutively expressed in eight different tissues at various developmental stages. Three ZmNLPs (ZmNLP3, 5 and 9) implementing the F ST higher than 0.25, differentiated very greatly between the Iowa Stiff Stalk Synthetic (SS) and Non-Stiff Stalk (NSS) heterotic groups. Quantitative real-time PCR (qPCR) results showed the expression levels of four ZmNLPs (ZmNLP4, 5, 6 and 8) were up-regulated over twofold in response to N treatment, ZmNLP4 and ZmNLP5 showed the largest up-regulation of greater than fivefold at 0.5 h after treatment which was even higher than the benchmark N-responsive gene (ZmNRT2.2) at the same time point, suggesting that they can be involved in the primary nitrogen response. As the first effort aimed to identify and characterize NLP transcription factor gene family in maize, our study also indicates ZmNLPs may have significant roles in maize N response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
NIN-LIKE PROTEIN (NLP), a plant-specific TF family, plays an essential role in nitrate signaling and assimilation. The NIN (for nodule inception) proteins were first identified as a regulator controlling development of symbiotic root nodules in the legume plant Lotus japonicus (Schauser et al. 1999; Suzuki et al. 2013) but were later found widely existing among plant species, including those that do not fix gaseous nitrogen (Schauser et al. 2005). NLPs carry two major conserved domains, a RWP-RK domain for DNA binding, and a PB1 domain that is involved in protein–protein interaction (Chardin et al. 2014; Sumimoto et al. 2007).
The NLP family proteins differ greatly in size as observed from three in Medicago and Brachypodium, six in rice and nine in Arabidopsis (Chardin et al. 2014). Functional studies in non-legume plants were mostly conducted in model organism Arabidopsis, where genetic mutants and transgenic plants demonstrated NLPs in Arabidopsis (AtNLPs) play a central role in orchestrating primary N response by binding to the nitrate-responsive cis-elements (Castaings et al. 2009; Konishi and Yanagisawa 2011, 2013). AtNLP7 was shown to bind with key N pathway genes including ANR1, LBD37/38, NRT1.1, NRT2, and NIA1, and thus able to moderate N assimilation and metabolism by either transcription activation or suppression of the downstream genes (Jian et al. 2015; Marchive et al. 2013). AtNLP8 was demonstrated as a master regulator of nitrate-promoted seed germination, could directly bind to the promoter of the abscisic acid (ABA) catabolic enzyme gene (CYP707A2) and reduce ABA levels in a nitrate-dependent manner (Yan et al. 2016). AtNLP6 and AtNLP7 were found interacting with another key transcriptional regulator teosinte branched1/cycloidea/proliferating cell factor1-20 (TCP20) under continuous nitrate and N-starvation conditions, and these interacting regulators played an important role in controlling the expression of nitrate-responsive genes and the G2/M cell-cycle marker gene (Guan et al. 2017). Furthermore, the nitrate-CPK (Ca2+-sensor protein kinases)-NLP signaling was identified, nitrate triggers Ca2+–CPK signaling and nitrate-coupled CPK signaling phosphorylates NLPs, and this signaling was crucial in nutrient-growth networks (Liu et al. 2017).
Maize (Zea mays L.) is one of the most important crops cultivated worldwide and a significant model plant to study important agronomic traits such as heterosis (Romay et al. 2013) and nitrogen use efficiency (Simons et al. 2014). Iowa Stiff Stalk Synthetic (SS) and Non-Stiff Stalk (NSS) are two major heterotic groups which constitute genetically distinct breeding pools providing superior hybrid performance (van Heerwaarden et al. 2012). Furthermore, maize has one of the highest N responses to supplemental fertilizer, resulting significant amounts of N fertilizers applied annually to maintain high yields (Bi et al. 2014; Humbert et al. 2013). Therefore, discovering the molecular mechanisms of heterosis and N response carries great significance to achieving productive and sustainable maize production.
In this study, we conducted a genome-wide survey of candidate NLPs in maize, and their expression profiles, genetic differentiation between heterotic groups and N response were analyzed. Three ZmNLPs (ZmNLP3, ZmNLP5 and ZmNLP9) differentiated very greatly between the SS and NSS heterotic groups. Four ZmNLPs (ZmNLP4, ZmNLP5, ZmNLP6 and ZmNLP8) were showed to be N responsive genes. Accordingly, we conclude the ZmNLPs as N-responsive putative transcription factors in maize.
Materials and methods
Database search for NLP proteins
Raw Hidden Markov Model (HMM) data for the conserved RWP-RK and PB1 domain (RWP-RK. hmm, PF02042; PB1.hmm, PF00564) were downloaded from Pfam database (Finn et al. 2014). HMM search from the HMMER3 package was used to search the maize genome database MaizeGDB (AGPv3; http://www.maizegdb.org/). Proteins contained RWP-RK and PB1 domains with E values < 1E-5 were collected and verified by NCBI Conserved Domains Database (CDD) (http://www.ncbi.nlm.nih.gov/cdd), Plant TFDB Plant Transcription Factor Database (Plant TFDB) (http://planttfdb.cbi.pku.edu.cn/) database. Redundant sequences were removed on the basis of their chromosome locations and sequence similarity. Sequence information of the nine Arabidopsis, six rice (Oryza sativa japonica) and five sorghum NLP proteins was obtained from Plant TFDB, and GRASSIUS (http://grassius.org/tfomecollection.html) databases.
Multiple sequence alignments and phylogenetic analysis
Multiple Sequence Alignments of the amino acid sequences of NLP proteins were performed using Clustal X (version 1.83) (Thompson et al. 1997). The distribution of amino acid residues for the conserved RWP-RK and PB1 domains of ZmNLPs were created using WebLogo [http://weblogo.berkeley.edu/examples.html (Crook s et al. 2004)]. Based on the Multiple Sequence Alignments, the phylogenetic tree was constructed with the MEGA6.0 program [http://www.megasoftware.net/ (Tamura et al. 2007)] using the Neighbor-joining method; and bootstrap tests were carried out with 1000 replicates.
Gene structure and conserved motifs identification
The Gene Structure Display Server program (GSDS 2.0, Hu et al. 2015) was used to illustrate the exon–intron structure for individual ZmNLP genes by comparing the coding sequences with their corresponding genomic DNA sequences from MaizeGDB (http://www.maizegdb.org/). The deduced amino acid sequences of the 29 NLP proteins from maize, Arabidopsis, rice and sorghum were analyzed by the online MEME tool version 4.12.0 (http://meme-suite.org/tools/meme, Bailey et al. 2009). To identify conserved motifs in these sequences, criterion of the maximum number of motifs was set to 15, together with a minimum width of 6 and a maximum width of 50 amino acids, while other factors were set at default values.
Expression analysis of maize NLP genes
The transcriptome profiling data (RNA-seq data) from maize inbred-line B73 during 18 different developmental stages were downloaded from the NCBI SRA database (http://www.ncbi.nlm.nih.gov/sra/; accession number SRR404131-SRR404132, SRR404139-SRR404150, SRR404152-SRR404156, SRR404158-SRR404164, SRR404171-SRR404200, SRR404202-SRR404203). The analysis of RNA-seq data was performed according to the previous study (Trapnell et al. 2013; Wang et al. 2017). FPKM (fragments per kilobase of exon per million fragments mapped) was calculated with Cufflinks (v2.1.1) (Trapnell et al. 2010) representing the expression level. Heat maps were created using MultiExperiment Viewer version 4.9 (MeV4.9, http://www.tm4.org/mev.html) program.
Plant materials and growth conditions
Maize inbred-line B73 was used in this study. Tissues (seeds, roots, stems, leafs) used for expression pattern validation were obtained from at least three healthy B73 plants at the stage of 20 days after pollination, which were cultivated in the field at Jiangsu Academy of Agricultural Sciences(JAAS), Nanjing, China. Nitrate treatment experiments were performed on plants according to previous study (Krouk et al. 2010). Briefly, Seeds were germinated on filter paper soaked in water for 48 h and then transferred into pots 2.5-L volume containing sand. Plants were grown in a greenhouse under a 16-h light/8-h dark photoperiod at a temperature of 20–35 °C (JAAS, Nanjing, China). After grown for 20 days, plants were treated with 15 mM KNO3, the control plants were mock-treated by the same concentration of KCl. Roots were collected at 0 h (h, before treatment) and 0.5, 1, 1.5, 2 h after treatment. Three independent replicates were collected for each time point (treatment and control) and frozen in liquid nitrogen. Samples were then stored at −80 °C for the following RNA isolation.
Semi-quantitative RT-PCR analysis
Total RNA was isolated from collected samples using the TRIzol method as described by the manufacturer. The residual DNA was removed by RNase-free DNase I (Takara, Dalian, China) treatment, and 500 ng of total RNA from each sample was reverse transcribed to cDNA using prime Script™ RT Reagent kit (Takara, Dalian, China). A fourfold dilution of cDNA was used for semi-quantitative RT-PCR analysis. UPF1 was used as an internal control (Lin et al. 2014). Primer pairs for all genes (Table 1) were designed by Primer 3 (http://bioinfo.ut.ee/primer3-0.4.0/), then tested by NCBI Primer BLAST and confirmed by sequencing resulting PCR products. Three biological replicates and 35 cycles for each reaction were performed for semi-quantitative RT-PCR analysis, PCR products were examined by 2% agarose gel electrophoresis.
Genetic differentiation analysis
The DNA-seq data from 289 diverse maize inbred lines were downloaded from the NCBI SRA database (http://www.ncbi.nlm.nih.gov/sra/; accession number SRA049859, SRA051245 and PRJNA260788) (Romay et al. 2013; van Heerwaarden et al. 2012). Raw dates were trimmed to remove low-quality nucleotides via SolexaQA (Cox et al. 2010) with the Phred-Score ≥ 20 longer than 20 bp, SNP sites were analysis using Burrows-Wheeler Alignment tool (BWA, https://sourceforge.net/projects/bio-bwa/files/) and SAMtool (https://sourceforge.net/projects/samtools/files/). FST vaules between two heterotic groups were calculated using vcftools program package (http://samtools.github.io/hts-specs/VCFv4.2.pdf), window-size was set to 5000 and window-step was set to 1000. ZmNLPs involved SNPs were screened in gene internal region.
Quantitative real-time PCR (qPCR)
Total RNA was isolated from KNO3 and KCl treated roots following the cDNA synthesis methods mentioned above. qPCR was performed with three replicates in 96-well plates using a Bio-Rad CFX96 system based on the SYBR Green PCR assay. Standard reactions comprised 5 μl 2 × SYBR® Premix Ex Taq™ II (TaKaRa, Dalian, China), 1 μl diluted cDNA template and 10 μM each primer with a total volume of 10 μl. Each reaction was subjected to the following conditions: an initial step of 3 min at 95 °C, followed by 40 cycles at 95 °C for 10 s, 58 °C for 20 s and 72 °C for 20 s. Each sample had three biological replicates to ensure the accuracy of results. The expression of the ZmNLPs and ZmNRT2.2 were calculated by the 2−ΔΔCt method (Livak and Schmittgen 2001), samples were treated with 15 mM KNO3 (as the experimental treatment) or KCl (as the control treatment). Transcription levels of each tested genes were normalized by UPF1 gene (GRMZM2G163444), such as ΔCt ZmNLP1(KNO3) = Ct ZmNLP1(KNO3) − Ct UPF1 (KNO3), ΔCt ZmNLP1(KCl) = Ct ZmNLP1(KCl) − Ct UPF1 (KCl), ΔΔCt ZmNLP1 = ΔCt ZmNLP1(KNO3) − ΔCt ZmNLP1(KCl). One-Way ANOVA analysis was used to test treatment effects. When effects were significantly different, post hoc multiple comparisons (LSD, p < 0.05) was examined. All data analysis was conducted using DPS v6.55.
Results
Genome-wide identification and conserved domains of maize NLPs
To identify the NLP family in maize, the Pfam HMM profiles (RWP-RK. hmm, PF02042; PB1.hmm, PF00564) were used to perform a HMMER search against the maize ref_v3 genome. Twelve protein sequences that contain conserved RWP-RK and PB1 domains were identified from the genome and further confirmed using NCBI CDD and Plant TFDB. Duplicate entries based on chromosome location and sequence similarity were then consolidated. Finally, nine putative NLP transcription factor-encoding genes were identified in maize genome, named ZmNLP1−ZmNLP9 (Table 2). Except chromosomes 4 and 9, all ten maize chromosomes contain NLPs, with two members on chromosome 2, and one on each of chromosomes 1, 3, 5, 6, 7, 8 and 10. Genomic length of these genes spans from 3.3 to 9.2 kb with an average of 5.9 kb. Encoded proteins consist of 786 to 1089 amino acids (aa) with an average of 912 aa.
All NLP proteins contain conserved RWP-RK and PB1 domains. To investigate the features of the homologous domain sequences, we performed Multiple Sequence Alignment (MSA) using the RWP-RK and PB1 domain amino acid sequences from the nine ZmNLPs (Fig. 1). The predicted DNA-binding RWP-RK domain of maize NLPs showed to be highly conserved with 31 out of 52 amino acids being 100% conserved across family (Fig. 1a; Supplementary Table 1), whereas the protein–protein interaction PB1 domain exhibited more sequence variation, with only 15 out of 84 amino acids being fully conserved in all ZmNLPs (Fig. 1b; Supplementary Table 1).
Conserved domains across NLP proteins in maize. The sequence logos of RWP-RK (a) and PB1 (b) are based on full-length alignments of all maize NLP RWP-RK and PB1domians. Multiple alignment analysis of 9 maize NLP proteins was performed with ClustalW. The bit score indicates the information content for each position in the sequence
Phylogenetic relationships and gene structure of maize NLPs
To evaluate the evolutionary structure within the NLP family in maize, a phylogenetic tree was constructed from MSA of all ZmNLP proteins (Fig. 2a; Supplementary Table 1). Based on sequence similarity and phylogenetic tree topology, the maize NLP family was divided into two subgroups (Subgroup 1 and Subgroup 2, or S1 and S2). S1 included five members (ZmNLP1, 2, 3, 7 and 9), and S2 contained four members (ZmNLP4, 5, 6 and 8), with high bootstrap values suggesting a common origin for NLP genes within each subgroup. To understand the structural diversity of NLP genes, we examined the exon–intron structures of ZmNLPs by comparing their cDNA sequences with their genomic sequences (Fig. 2b; Supplementary Table 2). According to the predicted structures, all coding sequences of maize NLPs are disrupted by introns, and consist of four to seven exons varying from 34 to 1674 bp. Notably, the most closely related members in each subgroup were generally conserved in exon length and exon–intron pattern.
Phylogenetic relationship and exon–intron gene structure of maize NLP genes. a Phylogenetic tree of maize NLP proteins by the neighbor-joining method using MEGA6 software with 1000 bootstraps. The two phylogenetic subgroups (S1and S2) are indicated. Scale bar indicates the number of amino acid substitutions per site. b Exon–intron structures of NLP genes. Exons are represented by yellow boxes and introns by black lines. (Color figure online)
Synteny analysis of NLPs in maize, Arabidopsis, rice and sorghum
To better understand molecular evolution of NLP proteins across species, phylogenetic relationships among ZmNLPs and their homologs in Arabidopsis, rice and sorghum were examined. The nine putative ZmNLP proteins were subjected to Multiple Sequence Alignment along with nine Arabidopsis, six rice and five sorghum NLP proteins, and a neighbor-joining tree was constructed (Fig. 3; Supplementary Table 1). In general, the maize NLPs exhibited a close relationship with the ones in the other three species, especially in sorghum. Two putative orthologs (ZmNLP1/SbNLP1, ZmNLP5/SbNLP4) were identified based on the phylogenetic tree with a high degree of homology in the terminal node (99).
Evolutionary relationships of the NLP proteins from maize (ZmNLPs), Arabidopsis (AtNLPs), rice (OsNLPs) and sorghum (SbNLPs). a Phylogenetic tree computed by the MEGA6 software using the neighbor-joining method with 1000 bootstrap. The proteins are clustered into 3 Groups (Group 1–3). Scale bar indicates the number of amino acid substitutions per site. b Motifs identified by MEME online software on the different NLP proteins. The motifs covered describe domains RWP-RK (motifs 4, 1), PB1 (motifs 10, 3). Scale for protein length is indicated at the bottom
In addition, conserved motifs in 29 NLP proteins were predicted by the online MEME tool. Fifteen predicted motifs were identified with sizes varying from 11 to 50 amino acids (Fig. 3; Supplementary Table 3). Almost all of NLP proteins contained the conserved RWP-RK domain (motifs 4, 1, Fig. 3) except OsNLP6. This protein only had the N-terminal part of the domain (motifs 4), the most conserved motif (motif 1) was lost. All NLP proteins contained the PB1 domain (motifs 10, 3, Fig. 3). Additionally, some NLPs were predicted to carry the GAF domain-like in Plant TFDB database, such as ZmNLP5 (start–end: 148–230 aa, motifs 2, Fig. 3) and AtNLP7 (start–end: 214–283 aa, motifs 2, Fig. 3).
Based on the phylogenetic relationships and motif compositions, the NLP proteins from these four higher plants were divided into three major Groups (Group 1, 2 and 3), similar to previous reports in Arabidopsis (Schauser et al. 2005). Group 1–3 contained seven to twelve NLPs. In Group 3, four maize NLP genes (ZmNLP4, ZmNLP5, ZmNLP6 and ZmNLP8) clustered with the AtNLP7, a major player in the primary nitrate response and required for nitrate regulation of N-assimilation in Arabidopsis (Wang et al. 2003; Castaings et al. 2009).
Gene expression atlas of ZmNLPs
To investigate the temporal and spatial patterns of NLP gene expression in maize, an expression atlas of ZmNLP from eighteen different tissues were made using the publicly available RNA-seq datasets at NCBI SRA archive (Materials and Methods). ZmNLPs in general exhibited low transcript abundance (FPKM from 0 to 40.7, with an average of 4.6, Fig. 4a; Supplementary Table 4), which is not uncommon for genes encoding transcription factors. Notably, almost all ZmNLPs constitutively expressed in eight different tissues at various developmental stages. Four ZmNLPs (44%) showed highest transcript accumulation level in 24 h after imbibition germinating seed (Fig. 4a, Column 1), in greenhouse grown 6 days-after-sowing (DAS) primary root (Column 2), in the Thirteenth Leaf at vegetative transition (VT) stage (Column 9), the Thirteenth Leaf at reproductive 2 (R2) stage (Column 10). Whereas three ZmNLPs (33%) showed highest transcript accumulation in stem and shoot apical meristem (SAM) at vegetative 3 (V3) stage (Column 3) and leaves at various developmental stages (Columns 4, 5, 6, 8). Especially, ZmNLP2 showed the highest transcript accumulation in all developmental stages tested. Although expression profiles varied for individual ZmNLPs, some conserved patterns were observed. For instance, in Subgroup 1 most ZmNLPs showed higher transcript accumulation in leaf tissues, while in ZmNLP4 and ZmNLP5 from Subgroup 2 expressed with abundance in all leaf tissues (Columns 4, 5, 6, 7, 8, 9, 10).
Expression profiling of NLP genes in maize. a Heat map for ZmNLP gene expression in various maize tissues across development time. The color scale below represents expression values, with higher intensity of red indicating high levels of transcript abundance. Columns 1: 24H_germinating seed; 2: 6DAS_GH_Primary Root; 3: V3_Stem and SAM; 4: V5_Tip of stage-2 Leaf; 5: V9_Eighth Leaf; 6: V9_Eleventh Leaf; 7: V9_Thirteenth Leaf; 8: V9_Immature Leaves; 9: VT_Thirteenth Leaf; 10: R2_Thirteenth Leaf; 11: 10DAP_Whole seed; 12: 12DAP_Whole seed; 13: 14DAP_Whole seed; 14: 16DAP_Whole seed; 15: 12DAP_Endopsperm; 16: 14DAP_Endopsperm; 17: 16DAP_Endosperm; 18:16DAP_Embryo. b Semi-quantitative RT-PCR validation using four representative tissues. Four tissues including seed, root, stem and leaf were assayed. ZmNLPs were compared to the maize reference housekeeping gene UPF1 (UPF1: GRMZM2G163444). (Color figure online)
To validate expression profiles generated from RNA-seq data, we conducted semi-quantitative RT-PCR in four representative tissues (R1_Root, R1_Stem, R1_leaf, and 20DAP_Whole seed, Fig. 4b). All ZmNLP transcripts were detected at all four tissues, with transcripts of ZmNLP4 and ZmNLP8 particularly abundant, in consistence with results from the RNA-seq based expression atlas (Fig. 4b).
Genetic differentiation analysis of ZmNLPs in different heterotic groups
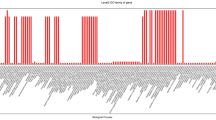
To identify genetic variation of ZmNLPs among different maize germplasm groups, the genetic differentiation analysis was made using the publicly available DNA-seq datasets at NCBI SRA archive (Materials and Methods). The 289 diverse maize inbred lines were classified as members of two major heterotic groups, SS (109 lines) and NSS (180 lines) groups, including 294378 single nucleotide polymorphism (SNP) sites (Romay et al. 2013; van Heerwaarden et al. 2012). Genetic differentiation coefficient (F-statistic, F ST) between two heterotic groups were calculated using vcftools program package. Wright suggested qualitative guidelines for the interpretation of F ST, a range from 0 to 0.05, 0.05 to 0.15 and 0.15 to 0.25 considered to indicate little, moderate and great genetic differentiation respectively, whereas F ST values > 0.25 indicate very great genetic differentiation (Wright 1977). According to chromosome locations of ZmNLP genes, SNPs were screened in gene internal regions. A total of six genes (ZmNLP1, ZmNLP3, ZmNLP5 ZmNLP7, ZmNLP8 and ZmNLP9) in ZmNLP family differentiated significantly between the two groups with F ST values > 0.15, distributing on chromosomes 1, 2, 5, 7, 8 and 10 (Fig. 5), while the rest of ZmNLPs (ZmNLP2, ZmNLP4 and ZmNLP6) showed moderate or little genetic differentiation (Fig. 5). Furthermore, among the six ZmNLPs, the F ST values of ZmNLP3, ZmNLP5 and ZmNLP9 were higher than 0.25 (Fig. 5), indicating these genes differentiate very greatly between the SS and NSS heterotic groups.
Nitrogen response of NLP genes in maize
Previous studies in Arabidopsis have shown that changes in N status trigger extensive responses in primary and secondary metabolism, physiological and developmental processes, and part of which were due to changes in gene expression (Scheible et al. 2004; Wang et al. 2003). To investigate the early transcriptional response of ZmNLP genes to N in maize, levels of mRNA for ZmNLPs and N-responsive sentinel gene [ZmNRT2.2 (Zamboni et al. 2014)] in response to NO3− treatment were measured (Fig. 6; Supplementary Table 5). The result revealed that, most ZmNLPs demonstrated both up and down regulation with decreased expression at 1 and 2 h after nitrate treatment and increased expression at 0.5 and 1.5 h, except for ZmNLP4 and ZmNLP5 with the expression up-regulated at 0.5 h and then down-regulated at 1–2 h. Four ZmNLPs (ZmNLP4, ZmNLP5, ZmNLP6 and ZmNLP8) have their transcript levels changed significantly after nitrate treatment (up-regulated over twofold in at least one time-point, Fig. 6b) compared to the benchmark N-responsive ZmNRT2.2 (Fig. 6c), while the rest of ZmNLPs showed lesser transcriptional responses (Fig. 6a). Furthermore, ZmNLP4 and ZmNLP5 showed to be very N-responsive by exhibiting the highest up-regulation (> fivefold) at 0.5 h after treatment.
qPCR results for expression patterns of ZmNLPs after nitrate treatment. Levels of mRNA for ZmNLPs and ZmNRT2.2 in maize roots in response to nitrate treatment were detected. 20-day-old plants grown without the presence of nitrate were treated with 15 mM KNO3 or KCl (as a control treatment). Plants were collected at 0 h (h, before treatment) and 0.5, 1, 1.5, 2 h after treatment. Sentinel transcripts were measured in RNA from roots using qPCR and normalized to the maize housekeeping gene UPF1. a, b The groups with different expression patterns in nitrate treatment. c The expression pattern of ZmNRT2.2 (GRMZM2G010251) under nitrate treatment
Discussion
As the first attempt to identify and characterize NLP family in maize, our genome-wide analysis revealed nine maize NLPs. The maize NLP family size is the same as in Arabidopsis (Schauser et al. 2005), but greater than rice with six members (Chardin et al. 2014) and sorghum with five (from this study), implying functional diversification of NLP members. However, the fact that a small-genomed Arabidopsis (135 Mb) has the same number of NLPs as in maize (a genome of 2.5 Gb), suggests the evolutionary conservation of NLPs and their essentiality in maintaining normal plant growth and development.
The conserved nature of NLPs was further supported by structural analysis of protein domains. The RWP-RK domain has 60% amino acids that are 100% conserved within maize NLP family (Fig. 1a). Secondary structure predictions of RWP-RK indicate the presence a basic helix followed by a helix-turn-helix motif and an amphipathic leucine zipper (Schauser et al. 2005), suggesting its involvement in DNA-binding. The PB1 domain at C-terminus contains two α helices, a mixed five-stranded β sheet and an acidic OPCA motif, with a predicted protein-binding ability (Sumimoto et al. 2007). The GAF domain is a ubiquitous motif for signaling and sensory transducing, and has been shown to be associated with gene regulating from bacteria to higher plants (Ho et al. 2000). Even though only 4 NLPs (ZmNLP5, AtNLP3, AtNLP7 and SbNLP4) was predicted to carry the GAF domain-like (Chardin et al. 2014) in Plant TFDB database, a clear sequence homology (motif 2, Fig. 3) can be observed in all Arabidopsis, rice, maize and sorghum NLPs. It is therefore reasonable to assume that the co-existence of a sensing and signaling domain (GAF-like), a DNA-binding domain (RWP-RK) and a protein-binding domain (PB1) underlies the molecular mechanism for NLPs to cross function in various aspects of N response including N status sensing, transcription modulation and signal transduction. The fact that ZmNLPs showed transcript accumulation in almost all tissues examined from root to leaf to developing seeds (Fig. 4), further supported the breadth of involvement of ZmNLPs in maintaining normal plant N metabolism.
In rice, 200 differentially selected regions were identified between the two rice heterotic groups (IndI and IndII), which contained lots of functional genes and loci associated with important agronomic traits (Xie et al. 2015). In these regions, the accumulation of difference loci was detected to be positively correlated with grain yield (Xie et al. 2015). Genes involved in maize heterosis has also been studied such as Zea mays ARGOS1 (ZAR1), transgenic experiments demonstrated that over-expression of ZAR1 improved maize organ growth, grain yield, and drought-stress tolerance (Guo et al. 2014). In our study, three ZmNLPs (ZmNLP3, ZmNLP5 and ZmNLP9) were found differentiated very greatly between the SS and NSS heterotic groups, indicating these genes may contribute to heterosis and have effect on important agronomic traits such as NUE in maize.
Previous studies have shown N treatment can trigger rapid and extensive transcriptional changes in a wide range of cellular and physiological processes. In this study, qPCR assay of ZmNLPs confirmed the swift transcriptional response (Fig. 6). Notably, the transcription of four ZmNLP genes (ZmNLP4, ZmNLP5, ZmNLP6 and ZmNLP8) responded most prominently to nitrate treatment (Fig. 5b), which happened to be the only four ZmNLPs clustered with AtNLP7 in the same phylogenetic group (Fig. 3a). ZmNLP5 not only has a close phylogenetic relationship with AtNLP7, but even to be the most N-responsive gene in ZmNLP family, and ZmNLP5 also showed significant differentiation in genetic differentiation analysis. Functional experiments and genetic mutant studies in Arabidopsis have demonstrated AtNLP7 as a master regulator in nitrate sensing and signaling (Konishi and Yanagisawa 2013; Marchive et al. 2013). It is then postulated that aforementioned ZmNLP5 may have the similar functions as AtNLP7 in N regulation.
References
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202-8
Bi YM, Meyer A, Downs GS, Shi X, El-Kereamy A, Lukens L, Rothstein SJ (2014) High throughput RNA sequencing of a hybrid maize and its parents shows different mechanisms responsive to nitrogen limitation. BMC Genomics 15:77
Castaings L, Camargo A, Pocholle D, Gaudon V, Texier Y, Boutet-Mercey S, Taconnat L, Renou JP, Daniel-Vedele F, Fernandez E, Meyer C, Krapp A (2009) The nodule inception-like protein 7 modulates nitrate sensing and metabolism in Arabidopsis. Plant J 57(3):426–435
Chardin C, Girin T, Roudier F, Meyer C, Krapp A (2014) The plant RWP-RK transcription factors: key regulators of nitrogen responses and of gametophyte development. J Exp Bot 65(19):5577–5587
Cox MP, Peterson DA, Biggs PJ (2010) SolexaQA: at-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics 11(1):485
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14(6):1188–1190
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42:D222-230
Guan P, Ripoll JJ, Wang R, Vuong L, Bailey-Steinitz LJ, Ye D, Crawford NM (2017) Interacting TCP and NLP transcription factors control plant responses to nitrate availability. Proc Natl Acad Sci USA 114(9):2419–2424
Guo M, Rupe MA, Wei J, Winkler C, Goncalves-Butruille M, Weers BP, Cerwick SF, Dieter JA, Duncan KE, Howard RJ, Hou Z, Loffler CM, Cooper M, Simmons CR (2014) Maize ARGOS1 (ZAR1) transgenic alleles increase hybrid maize yield. J Exp Bot 65(1):249–260
Ho YS, Burden LM, Hurley JH (2000) Structure of the GAF domain, a ubiquitous signaling motif and a new class of cyclic GMP receptor. EMBO J 19(20):5288–5299
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297
Humbert S, Subedi S, Cohn J, Zeng B, Bi YM, Chen X, Zhu T, McNicholas PD, Rothstein SJ (2013) Genome-wide expression profiling of maize in response to individual and combined water and nitrogen stresses. BMC Genomics 14:3
Jian W, Zhang D, Zhu F, Wang S, Zhu T, Pu X, Zheng T, Feng H, Lin H (2015) Nitrate reductase-dependent nitric oxide production is required for regulation alternative oxidase pathway involved in the resistance to Cucumber mosaic virus infection in Arabidopsis. Plant Growth Regul 77(1):99–107
Konishi M, Yanagisawa S (2011) The regulatory region controlling the nitrate-responsive expression of a nitrate reductase gene, NIA1, in Arabidopsis. Plant Cell Physiol 52(5):824–836
Konishi M, Yanagisawa S (2013) Arabidopsis NIN-like transcription factors have a central role in nitrate signalling. Nat Commun 4:1617
Krouk G, Mirowski P, LeCun Y, Shasha DE, Coruzzi GM (2010) Predictive network modeling of the high-resolution dynamic plant transcriptome in response to nitrate. Genome Biol 11(12):R123
Lin F, Jiang L, Liu Y, Lv Y, Dai H, Zhao H (2014) Genome-wide identification of housekeeping genes in maize. Plant Mol Biol 86(4–5):543–554
Liu KH, Niu Y, Konishi M, Wu Y, Du H, Sun Chung H, Li L, Boudsocq M, McCormack M, Maekawa S, Ishida T, Zhang C, Shokat K, Yanagisawa S, Sheen J (2017) Discovery of nitrate-CPK-NLP signalling in central nutrient-growth networks. Nature 545(7654):311–316
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402–408
Marchive C, Roudier F, Castaings L, Brehaut V, Blondet E, Colot V, Meyer C, Krapp A (2013) Nuclear retention of the transcription factor NLP7 orchestrates the early response to nitrate in plants. Nat Commun 4:1713
Romay MC, Millard MJ, Glaubitz JC, Peiffer JA, Swarts KL, Casstevens TM, Elshire RJ, Acharya CB, Mitchell SE, Flint-Garcia SA, McMullen MD, Holland JB, Buckler ES, Gardner CA (2013) Comprehensive genotyping of the USA national maize inbred seed bank. Genome Biol 14(6):R55
Schauser L, Roussis A, Stiller J, Stougaard J (1999) A plant regulator controlling development of symbiotic root nodules. Nature 402(6758):191–195
Schauser L, Wieloch W, Stougaard J (2005) Evolution of NIN-like proteins in Arabidopsis, rice, and Lotus japonicus. J Mol Evol 60(2):229–237
Scheible WR, Morcuende R, Czechowski T, Fritz C, Osuna D, Palacios-Rojas N, Schindelasch D, Thimm O, Udvardi MK, Stitt M (2004) Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, and the regulatory infrastructure of Arabidopsis in response to nitrogen. Plant Physiol 136(1):2483–2499
Simons M, Saha R, Guillard L, Clement G, Armengaud P, Canas R, Maranas CD, Lea PJ, Hirel B (2014) Nitrogen-use efficiency in maize (Zea mays L.): from ‘omics’ studies to metabolic modelling. J Exp Bot 65(19):5657–5671
Sumimoto H, Kamakura S, Ito T (2007) Structure and function of the PB1 domain, a protein interaction module conserved in animals, fungi, amoebas, and plants. Sci STKE 2007(401):re6
Suzuki W, Konishi M, Yanagisawa S (2013) The evolutionary events necessary for the emergence of symbiotic nitrogen fixation in legumes may involve a loss of nitrate responsiveness of the NIN transcription factor. Plant Signal Behav 8(10):e25975
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28(5):511–515
Trapnell C, Hendrickson DG, Sauvageau M, Goff L, Rinn JL, Pachter L (2013) Differential analysis of gene regulation at transcript resolution with RNA-sEq. Nat Biotechnol 31(1):46–53
van Heerwaarden J, Hufford MB, Ross-Ibarra J (2012) Historical genomics of North American maize. Proc Natl Acad Sci USA 109(31):12420–12425
Wang R, Okamoto M, Xing X, Crawford NM (2003) Microarray analysis of the nitrate response in Arabidopsis roots and shoots reveals over 1000 rapidly responding genes and new linkages to glucose, trehalose-6-phosphate, iron, and sulfate metabolism. Plant Physiol 132(2):556–567
Wang Y, Liu H, Wang S, Li H (2017) Genome-wide identification and expression analysis of the YUCCA gene family in soybean (Glycine max L.). Plant Growth Regul 81(2):265–275
Wright S (1977) Evolution and the genetics of populations. Variability within and among natural populations, vol 3. University of Chicago Press, Chicago, IL
Xie W, Wang G, Yuan M, Yao W, Lyu K, Zhao H, Yang M, Li P, Zhang X, Yuan J, Wang Q, Liu F, Dong H, Zhang L, Li X, Meng X, Zhang W, Xiong L, He Y, Wang S, Yu S, Xu C, Luo J, Li X, Xiao J, Lian X, Zhang Q (2015) Breeding signatures of rice improvement revealed by a genomic variation map from a large germplasm collection. Proc Natl Acad Sci USA 112(39):E5411-5419
Yan D, Easwaran V, Chau V, Okamoto M, Ierullo M, Kimura M, Endo A, Yano R, Pasha A, Gong Y, Bi YM, Provart N, Guttman D, Krapp A, Rothstein SJ, Nambara E (2016) NIN-like protein 8 is a master regulator of nitrate-promoted seed germination in Arabidopsis. Nat Commun 7:13179
Zamboni A, Astolfi S, Zuchi S, Pii Y, Guardini K, Tononi P, Varanini Z (2014) Nitrate induction triggers different transcriptional changes in a high and a low nitrogen use efficiency maize inbred line. J Integr Plant Biol 56(11):1080–1094
Acknowledgements
We sincerely thank editorial reviewers for valuable comments that helped improve the quality of this manuscript. This work was supported by grant from the National Natural Science Foundation of China (No. 31401394) and Jiangsu Agriculture Science and Technology Innovation Fund [CX(14)2009].
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ge, M., Liu, Y., Jiang, L. et al. Genome-wide analysis of maize NLP transcription factor family revealed the roles in nitrogen response. Plant Growth Regul 84, 95–105 (2018). https://doi.org/10.1007/s10725-017-0324-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0324-x









