Abstract
Thaumatin-like protein (TLP) is one of pathogenesis-related protein family, which plays an important role in plant defense system. While its origin and expansion is unknown in plants. Here, we investigated the evolution of this gene family by performing a genome-wide identification and comparison in Arabidopsis, Oryza, Populus, Zea, Physcomitrella and Chlamydomonas. We found a birth process of this gene family in evolution. Tandem and segmental duplication played dominant roles for their expansion. In addition, TLPs were under purifying selection, and most members were responding to some biotic or abiotic stresses. Functional network analysis exhibited some resistance genes working together with TLPs. Our findings shed some light on this family gene evolution in plants, which might provide a base for further functional investigation of this family.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As sessile organisms, plants do not have an immune system like animals. Therefore, when exposed to biotic or abiotic stresses, they have evolved some adaptive ways to deal with these environmental stresses (Jacobs et al. 1999), including hypersensitive response (HR), lignification of cell wall, and synthesizing a variety of proteins or compounds, such as, antioxidants (like reactive oxygen species, ROS), anti-microbial compounds (like phytoalexins), late embryogenesis-abundant (LEA) proteins, proline, sugars and pathogenesis related (PR) proteins. Based on amino acid composition, serological and biochemical properties, PR proteins have been classified into 17 different families, including β-1,3-glucanases (PR-2), chitinases (PR-3, 4, 8 and 11), thaumatin-like proteins (TLPs) or osmotin (PR-5), proteinase-inhibitor (PR-6), endoproteinase (PR-7), peroxidase (PR-9), defensins (PR-12), thionins (PR-13), lipid-transfer proteins (PR-14), etc. (van Loon et al. 2006).
Plant TLP belongs to the PR-5 family. Historically, it has been called TLP/PR5 or osmotin/osmotin-like protein (OLP). Here, the nomenclature TLP is used to represent this gene family. Most TLPs have 16-cysteine residues, which might form eight disulfide linkages. This structure can stabilize protein, and then let it to resist to pH, proteases and heat-induced denaturation (Ghosh and Chakrabarti 2008). Some smaller TLPs (only containing ten conserved cysteine residues) have also been identified in conifers and monocots (Fierens et al. 2009; Liu et al. 2010a; Petre et al. 2011).
TLPs are involved in plant defense system against various biotic and abiotic stresses (Petre et al. 2011). Over-expression of TLPs can induce stress resistance in different transgenic plants (Liu et al. 1994; Rajam et al. 2007; Datta et al. 1999; Wang et al. 2010; Munis et al. 2010; Subramanyam et al. 2012; Acharya et al. 2013). TLPs can inhibit hyphal growth or spore germination by a membrane permeabilizing mechanism (Abad et al. 1996) or by degradation of cell walls (Osmond et al. 2001; Zareie et al. 2002). In addition to antibiotic activities, TLPs have also been involved in other physiological and developmental roles, including antifreeze activities (Yu and Griffith 1999), abiotic stress tolerance (Subramanyam et al. 2012; Zhu et al. 1995; Zhang and Shih 2007), floral organ formation and fruit ripening (Neale et al. 1990; Salzman et al. 1998), seed germination (Seo et al. 2008), senescence (Sakamoto et al. 2006), and glucanase activity (Osmond et al. 2001; Grenier et al. 1999).
The recent availability of genome sequences of some models plant species provides an opportunity to study the evolution of TLP gene family. Considering the important roles associated with antibiotic activities and developmental and physiological functions, and the number of the TLP genes varied greatly among plant species, it’s of considerable interest to us to investigate how the TLP genes have evolved in Plantae. Here, our results suggest that the TLP gene family has an expansion process in number, and that tandem and segmental duplications play dominant roles in it. Our studies also reveal different expression profiles of the TLP genes in rice and functional network features of the TLP protein in Arabidopsis.
Materials and methods
TLP sequences retrieval and identification in six plant species
We used the Arabidopsis TLP sequences (Liu et al. 2010b) as queries to perform BLAST searches against EnsemblPlants database (http://plants.ensembl.org/index.html) to identify potential members of the TLP gene family in these species. Next, the Conserved Domain Database (CDD) (Marchler-Bauer et al. 2013) was used to confirm whether the returned sequences from such searches encode TLP domain. Predotar program (Small et al. 2004) was used to perform the protein subcellular location.
Phylogenetic analyses of the TLP gene family
We used MUSCLE 3.52 (Edgar 2004) to perform multiple sequence alignments of full-length protein sequences, and used MEGA v5 (Tamura et al. 2011) to carry out phylogenetic analyses of the TLP proteins based on amino acid sequences with the neighbor joining (NJ) method. NJ analyses were done using p-distance methods, pairwise deletion of gaps, and default assumptions. Support for each node was tested with 1000 bootstrap replicates.
Estimation of the maximum number of gained and lost TLPs
We first divided the phylogeny into different clades to determine the expansion degrees of the gene family in these species (Cao et al. 2015). Nodes were labeled as V: Viridiplantae; Em: Embryophyte; A: Angiosperm; M: Monocots; and E: Eudicots. Notung v2.6 (Chen et al. 2000) was used to infer gene duplication and loss events by reconciling the gene tree with the species tree.
Chromosomal location of the TLP genes and genomic duplication
Annotation information on TAIR (http://www.arabidopsis.org), Rice Genome Annotation Project (http://rice.plantbiology.msu.edu/index.shtml), Populus genome browser (http://www.phytozome.net/poplar) and MaizeSequence (http://www.maizesequence.org) was used to determine the chromosomal locations and intron–extron structures of the TLP genes. SyMAP v3.4 (Soderlund et al. 2011) was used to depict the paralogous regions of the putative ancestral constituents of the genomes. In this study, two patterns of gene expansion (tandem duplication and segmental duplication) were focused on. Tandem duplicated genes were defined as adjacent homologous genes on a single chromosome, separated by no more than one nonhomologous spacer gene (Hanada et al. 2008). Moreover, some tandem duplicated genes were further confirmed in the plant tandem duplicated genes database (PTGBase) (Yu et al. 2015). Segmental duplications of each TLP gene within the family in poplar, rice, maize, and Arabidopsis genomes were searched in the SyMAP v3.4 (Soderlund et al. 2011).
Microarray-based expression analysis
Plant Expression Database (PLEXdb, http://www.plexdb.org/index.php) (Dash et al. 2012) was used for the expression analyses of rice TLP genes. In this study, six experiments (OS8, OS10, OS25, OS85, OS65, and OS92) were selected. And the genesis (v 1.7.6) program (Sturn et al. 2002) was used to normalize the expression data.
Positive selection assessment
We first used the Selecton Server (http://selecton.tau.ac.il/) (Stern et al. 2007) to calculate site-specific selection. Four evolutionary models (M8, M8a, M7 and M5) were used in this study. Each of the models uses different biological assumptions to test different hypotheses. In addition, we also used FEL (fixed-effects likelihood), SLAC (single likelihood ancestor counting), and REL (random-effects likelihood) methods with default settings embedded in the Datamonkey web interface (Delport et al. 2010) to further identify selection in individual codons. Finally, PARRIS was also used to test for the signatures of selection.
Co-expressed network assembly
We used ANAP (Wang et al. 2012), a co-expressed network designed to convert and then integrate 11 Arabidopsis data sets, to analyze TLP co-expressed network in Arabidopsis. Arabidopsis TLP genes were mapped to their corresponding proteins in the network database. Seventeen TLPs were not present in the assembled network database. Resulting interactions were used to build the seven members interaction network.
Results and discussion
Identification of the TLP genes in Arabidopsis, rice, poplar, maize, Physcomitrella and Chlamydomonas genomes
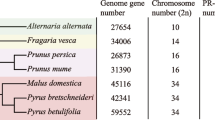
We identified 24 TLP genes in Arabidopsis, 44 in rice, 49 in poplar, 49 in maize, 6 in Physcomitrella and only 1 in Chlamydomonas (Table 1). Compared with other four genomes as described above, Physcomitrella and Chlamydomonas species encoded a much smaller number of TLP genes. This suggested that the expansion of TLP family mainly occurred after the divergence of the Embryophyte leading to vascular plants. The number of TLP genes in rice, poplar and maize is very similar, which are about twice as many TLP genes than that in Arabidopsis.
Phylogenetic analyses and comparison of TLP proteins
A phylogenetic analysis of the predicted TLP protein sequences was performed based on the NJ method. As displayed in Fig. 1, tree branches are colored by species, TLP subclass, the number of introns, and predicted targeting to organelles. TLP genes can be classed into six groups based on their phylogenetic relationships (Fig. 1b). We found that most of the rice TLP proteins belong to the Group II and III. 14 of 15 TLP members come from poplar in Group IV, suggesting that species-specific expansion has happened in these groups (Fig. 1a, b).
NJ distant tree of all TLPs in Arabidopsis, rice, poplar, maize, Physcomitrella and Chlamydomonas. Terminal markers are colored to indicated: a species (Arabidopsis bright green; rice pink; poplar blue; maize black; Physcomitrella red; Chlamydomonas yellow); b subclass (Group I orange; Group II turquoise; Group III crimson; Group IV yellow; Group V deep yellow; Group VI bright green); c number of introns (genes with no intron gray; one intron yellow; two introns turquoise; three introns pink; four introns blue; more than four introns red); and d predicated organelle targeting (ER green; mitochondrial red; plastid blue; elsewhere gray), some no-Met proteins are discarding and are not shown in here
Figure 1c displays the proportions of TLP genes with no intron, one intron, two introns, three introns, four introns and more than four introns in each species. The large majority of the TLP genes contain one or two introns in two eudicots (Arabidopsis and poplar). But, most of TLP genes in two monocots (rice and maize) are intronless. In addition, we also found that two lower plants (Physcomitrella and Chlamydomonas) have more introns. For example, EDP07492 and PP1S412_14V6.1 genes possess five introns, respectively. As we know, introns are important component of eukaryotic genes, and their loss or gain affect the complexity of genetic structure (Koonin 2006). Our results indicated that intron loss/gain events have occurred during the expansion and evolution of TLP paralogs.
Next, we also used the program Predotar (Small et al. 2004) to predict the organelles targeting of the family proteins. As a result, most of the TLP proteins were predicted to be targeted to the endoplasmic reticulum (ER) (Fig. 1d). Proteins targeted in the ER are usually experienced some protein processing, such as glycosylation, disulfide bond formation, folding and so on. Finally, these modified proteins are transported to their destinations when the signal peptides are removed (Trobetta 2003). Moreover, over 80 % of TLP proteins possess the signal peptide identified by the SignalP 4.0 server (Petersen et al. 2011).
Contrasting changes in the numbers of TLP genes
To better understand how TLP genes have evolved in these species, we estimated the number of TLP genes in the most recent common ancestor (MRCA) of Viridiplantae. There were about five ancestral TLP genes in the MRCA of Viridiplantae (V5) by reconciling the gene trees with the species phylogeny. Furthermore, we only identified one orthologous gene in the C. reinhardtii, implying that four of these five ancestral TLP genes have been lost when chlamydomonas is appearing (Fig. 2). The number of TLPs remained relatively stable before Angiosperm. Only after the emergence of Angiosperm species did TLPs once more expand significantly. It suggested that there were about 23 ancestral TLP genes in the MRCA of the green flowering plants. Interestingly, when compared with the MRCA of eudicots and monocots, it appeared that the expansion was uneven before their divergence. The MRCA of monocots has increased in size as much as two times (23/43), while the MRCA of eudicots has a similar number of TLP genes with that of Angiosperm (23/22). The expansion was also unbalanced between plant species since the divergence of eudicots and monocots (Cao and Li 2015). For example, poplar increased over two times (22/49) in size, while Arabidopsis only added two TLP genes (Fig. 2). When compared with the number of ancestral genes, it appeared that, except chlamydomonas, the TLP family had expanded in all the tested species. For instance, there are 24, 49, 44 and 49 genes in Arabidopsis, poplar, rice and maize, respectively; while the estimated number of genes in the MACA of eudicots and monocots is 23. Therefore, Arabidopsis, poplar, rice and maize have netted 1, 26, 21 and 26 genes, respectively, since their splits. Obviously, the numbers of genes gained in the poplar, rice and maize lineages are much greater than that in the Arabidopsis lineage.
Gene gain and loss of TLPs in the evolution of plants. The names of internal nodes are abbreviated (V Viridiplantae, Em Embryophyte, A Angiosperm, M Monocots, E Eudicots). The numbers of common ancestors at the five internal nodes (V, Em, A, M and E) are shown in the quadrates. Numbers after the plus signs the numbers of gene gain events, whereas numbers after the minus signs gene loss events
Chromosomal location and duplication events of the TLP genes
Next, we also investigate the phylogenetic relationship and chromosomal location of each TLP gene. The results indicated that the TLP genes unevenly distributed among different chromosomes of these genomes (Fig. S1), and that the generation of 7 (29.2 % of 24) Arabidopsis, 25 (56.8 % of 44) rice, 28 (57.1 % of 49) poplar and 20 (40.8 % of 49) maize TLP genes could be explained by tandem duplication (Fig. S1). The largest TLP gene clusters are located on chromosome 1 of poplar genome and contain 13 tandem arrayed members, i.e. POPTR_0001s22810.1, POPTR_0001s22830.1, POPTR_0001s22850.1, POPTR_0001s22860.1, POPTR_0001s22870.1, POPTR_0001s22880.1, POPTR_0001s22890.1, POPTR_0001s22900.1, POPTR_0001s22910.1, POPTR_0001s22920.1, POPTR_0001s22930.1, POPTR_0001s22950.1 and POPTR_0001s22960.1 (Fig. S2). Moreover, these genes form a single clade, suggesting that they may come from the recent tandem duplications (Fig. S2). In addition to tandem duplication, segmental duplications also played an important role in the expansion of the TLP family gene. At least 3, 2, 7, and 4 pairs of paraloguos genes come from segmental duplication in Arabidopsis, rice, poplar and maize, respectively (Fig. S1). Within the identified duplication events, some pairs are retained as duplicates, whereas others lost them. It is likely that dynamic changes have occurred following segmental duplication. Therefore, tandem duplication and segmental duplication are the major factor that contributed to the expansion of this gene family.
Expression of the TLP gene family in rice
Expression profiling is a useful tool for understanding gene function (Durick et al. 1999). To assess the transcriptional characteristics of the TLP genes, we examined some publicly databases in rice. First, we analyzed the spatial- and temporal-specific expression profiles of rice TLP genes in embryo (6 days), endosperm (6 days), root, leaf and seedling. All of the 37 detected transcripts were divergent expressed in different tissues (Fig. S3). Some members of rice TLP gene (such as LOC_Os12g43490, and LOC_Os09g36580) were expressed at the highest levels in the root, implying that they may be involved in the root development. In addition, LOC_Os06g47600 and LOC_Os10g05600 genes were higher expressed in the embryo, suggesting that these TLPs might be associated with early embryonic development of rice. Similar results have also been observed in their homologs in Arabidopsis, which were highly expressed during seed germination (Seo et al. 2008).
Plant growth is affected by some abiotic cues (Han et al. 2015; Jayakannan et al. 2015; Liu et al. 2015). Here, we also examined the expression profiles of rice TLP genes under drought, salt, cold and heat shock stresses. Divergent expression patterns were present among TLP members when exposed to these stress conditions (Fig. S3). Four TLP genes (LOC_Os12g43450, LOC_Os03g46060, LOC_Os07g23470, and LOC_Os03g14050) displayed higher expression levels in these conditions. Interestingly, we also found that, compared with the control, over 70.2 % of rice TLP genes showed higher expression levels under heat shock stress, implying that most TLP genes might be involved in the heat shock response. Infections of some pathogenic bacteria and insect pest are key factors affecting crop quality and yield. Next, we examined some experiments infected by Xanthomonas oryzae pv. oryzae (Xoo) and Blumeria graminis (Bgh) and found that over 67.5 and 75.6 % rice TLP genes exhibited an increase in expression levels under X. oryzae and B. graminis infection, respectively (Fig. S3). In addition, we also examined the expression levels of TLP gene under an insect pest, striped stem borer (SSB) (Fig. S3). The results indicated that expression levels of over 59.4 % TLP genes were increased when attacked by this pest. An increasing number of evidence has suggested that TLPs may function in both biotic and abiotic stress tolerance. Previous studies reported that transgenic plants over-expressing TLP proteins showed enhanced resistance to Alternaria alternate (Velazhahan and Muthukrishnan 2003), Fusarium graminearum (Mackintosh et al. 2007), Verticillium dahliae (Munis et al. 2010), Phaeseoropsis personata (Singh et al. 2013), and so on. Moreover, over-expression of some TLP proteins could confer tolerance during salt, drought and other stresses (Rajam et al. 2007; Munis et al. 2010; Wang et al. 2011; Singh et al. 2013). In addition, several TLPs have been reported to be induced during insect attack (Johnson et al. 2011; Singh et al. 2013). Our study also indicated that most rice TLPs can be induced by these abiotic and biotic stresses, suggesting that they are likely to be required for enhancing resistance to stress.
Different selection regimes in different groups and amino acid sites
K a /K s ratio measures selection pressure on amino acid substitutions. A K a /K s ratio greater than 1 suggests positive selection and a ratio less than 1 suggests purifying selection. The amino acids in a protein sequence are expected to be under different selective pressures and to have distinct K a /K s ratios. To analyze positive or negative selection of specific amino acid sites within the full-length sequences of the TLP proteins in different groups, substitution rate ratios of nonsynonymous (K a ) versus synonymous (K s ) mutations were calculated with the Selecton Server (http://selecton.tau.ac.il) using a Bayesian inference approach (Stern et al. 2007). We performed the tests using four evolution models [M8 (ωs ≥ 1), M8a (ωs = 1), M7 (beta) and M5 (gamma)] implemented in this server. Selection models M8a and M7 do not indicate the presence of positively selected sites, whereas the M8 and M5 models do (Table S1). Moreover, statistical significance of positive selection has been testing for the identified positively selected sites. The results indicated that the K a /K s ratios of the sequences from different TLP groups were significantly different (Table S1). Higher K a /K s values existed in Group IV, indicating a higher evolutionary rate or selective relaxation within members of the Group IV. On the other hand, the K a /K s values in Group I are relatively small, implying a lower evolutionary rate or selective constraint within Group I members. However, despite the differences in K a /K s values, all the estimated K a /K s values are substantially lower than 1, suggesting that the TLP sequences within each group are under purifying selection pressure and that positive selection may have acted only on a few sites during the evolutionary process (Table S1). In addition, we also used SLAC, FEL and REL methods with default settings implemented in the Datamonkey web interface (Delport et al. 2010) to further identify selection in individual codon. The results were shown in Table 2. All the K a /K s ratios were less than 1, indicating that most codons in TLP sequences were under purifying selecting in these six groups. The FEL software detected the largest number of potential positively selected sites for each group. However, SLAC and REL analyses only detected a few. In this study, we used two programs (Selecton and Datamonkey) including seven methods to detect positively selected sites and got similar selection pressures for each group (Tables S1 and 2). Detecting positive selection will help to understand functional residues and functional shift of protein (Loughran et al. 2012; Chen et al. 2014). In this paper, we found that a few sites might undergo positive selection in evolution (Tables S1 and 2); implying that positive selection on these sites might have accelerated functional divergence and then result in the formation of gene subgroups.
Functional network analysis of the TLP genes in Arabidopsis
Genes involved in related biological pathways are usually expressed cooperatively (Eisen et al. 1998). To further investigate which genes are possibly regulated with the TLPs, we assembled a co-expression network (Fig. 3). 7 of 24 TLPs were present in the network, which exhibited 245 physical or functional interactions with 188 genes. Molecular function analysis of these 188 genes showed that genes with ATP or DNA binding, protein binding, kinase activity, hydrolase activity, and transporter activity were overly represented. Among the 188 interactors identified, 102 and 64 genes co-expressed with AT1G18250 and AT4G38660, respectively. Plant resistance is very important to the growth of plant (An et al. 2015). Our co-expressed analysis also revealed that TLP genes might function with some pathogen resistance proteins. C ysteine-rich repeat-like kinases (CRK) is an important group of enzymes involved in pathogen resistance (Chen et al. 2004). Here, three CRKs (AT4G23210, AT4G23150 and AT4G23160) were identified to be co-expressed with the TLP proteins, implying potential interactions between the TLP and CRK genes. In addition, some proteins with transporter activity are also involved in plant resistance. Some of these co-expressed genes included EDS5 (AT4G39030), DND2 (AT5G54250) and TIL1 (AT5G58070). EDS5 encodes an orphan multidrug and toxin extrusion transporter, which is a necessary component of salicylic acid-dependent signaling for disease resistance (Ng et al. 2011). DND2 is a second cyclic nucleotide-gated ion channel gene for hypersensitive response (Jurkowski et al. 2004). TIL1 encodes a temperature-induced lipocalin, which is involved in the thermotolerance (Chi et al. 2009). Plant chitinases are involved in defense responses against pathogen attacks and in tolerance of diverse environmental stresses (Takenaka et al. 2009). A TLP, AT4G11650, was found to be co-expressed with one chitinase, AT3G54420. In addition, another chitinase, CHI (AT2G43570), might be the potential interactors of the TLP, AT1G75040. Thus, whether chitinase could serve as a link to TLP molecular pathways need further experimental confirmation. Although the exact pathways mediated by these genes were still unclear, we speculated that these TLP genes might play critical roles in plant resistance. These observations have led us to hypothesize that TLP could regulate these plant responses through its involvement in different signal pathway in plants. This contributes to the selection of candidate genes for further functional genomics.
Conclusions
This study provided a comparative genomic analysis addressing phylogeny, chromosomal location and duplication, selective pressures, expression profiling, and functional network analysis. Phylogenetic analyses revealed six well-supported groups in the TLP family. The TLP gene family had a birth process only after the emergence of Angiosperm species. Tandem and segmental duplication played a dominant role in the expansion of this gene family. In addition, TLPs were under purifying selection according to estimations of the substitution rates of these genes. Furthermore, comprehensive analysis of the expression profiles provided insights into possible functional divergence among members of the TLP gene family. Functional network analysis was also identified some resistance genes, which might work together with the TLPs. These data may provide valuable information for future functional investigations of this gene family.
References
Abad LR, D’Urzo MP, Liu D, Narasimhan ML, Reuveni M, Zhu JK, Niu X, Singh NK, Hasegawa PM, Bressan RA (1996) Antifungal activity of tobacco osmotin has specificity and involves plasma membrane permeabilization. Plant Sci 118:11–23
Acharya K, Pal AK, Gulati A, Kumar S, Singh AK, Ahuja PS (2013) Overexpression of Camellia sinensis thaumatin-like protein, CsTLP in potato confers enhanced resistance to Macrophomina phaseolina and Phytophthora infestans infection. Mol Biotechnol 54:609–622
An X, Liao Y, Zhang J, Dai L, Zhang N, Wang B, Liu L, Peng D (2015) Overexpression of rice NAC gene SNAC1 in ramie improves drought and salt tolerance. Plant Growth Regul 76:211–223
Cao J, Li X (2015) Identification and phylogenetic analysis of late embryogenesis abundant proteins family in tomato (Solanum lycopersicum). Planta 241:757–772
Cao J, Li X, Lv Y, Ding L (2015) Comparative analysis of the phytocyanin gene family in 10 plant species: a focus on Zea mays. Front Plant Sci 6:515
Chen K, Durand D, Farach-Colton M (2000) NOTUNG: a program for dating gene duplications and optimizing gene family trees. J Comput Biol 7:429–447
Chen K, Fan B, Du L, Chen Z (2004) Activation of hypersensitive cell death by pathogen-induced receptor-like protein kinases from Arabidopsis. Plant Mol Biol 56:271–283
Chen Y, Hao X, Cao J (2014) Small auxin up-regulated RNA (SAUR) gene family in maize: identification, evolution and its phylogenetic comparison with Arabidopsis, rice and sorghum. J Integr Plant Biol 56:133–150
Chi WT, Fung RW, Liu HC, Hsu CC, Charng YY (2009) Temperature-induced lipocalin is required for basal and acquired thermotolerance in Arabidopsis. Plant, Cell Environ 32:917–927
Dash S, Van Hemert J, Hong L, Wise RP, Dickerson JA (2012) PLEXdb: gene expression resources for plants and plant pathogens. Nucleic Acids Res 40:D1194–D1201
Datta K, Velazhahan R, Oliva N, Mew T, Khush GS, Muthukrishnan S, Datta K (1999) Over-expression of cloned rice thaumatin-like-protein (PR-5) gene in transgenic rice plants enhances environmental friendly resistance to Rhizoctonia solani causing sheath blight disease. Theor Appl Genet 98:1138–1145
Delport W, Poon AF, Frost SD, Kosakovsky Pond SL (2010) Datamonkey 2010: a suite of phylogenetic analysis tools for evolutionary biology. Bioinformatics 26:2455–2457
Durick K, Mendlein J, Xanthopoulos KG (1999) Hunting with traps: genome-wide strategies for gene discovery and functional analysis. Genome Res 9:1019–1925
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Fierens E, Gebruers K, Voet ARD, De Maeyer M, Courtin CM, Delcour JA (2009) Biochemical and structural characterization of TLXI, the Triticum aestivum L. thaumatin-like xylanase inhibitor. J Enzyme Inhib Med Chem 24:646–654
Ghosh R, Chakrabarti C (2008) Crystal structure analysis of NP24-I: a thaumatinlike protein. Planta 228:883–890
Grenier J, Potvin C, Trudel J, Asselin A (1999) Some thaumatin-like proteins hydrolyse polymeric beta-1,3-glucans. Plant J 19:473–480
Han Y, Yin S, Huang L (2015) Towards plant salinity tolerance-implications from ion transporters and biochemical regulation. Plant Growth Regul 76:13–23
Hanada K, Zou C, Lehti-Shiu MD, Shinozaki K, Shiu SH (2008) Importance of lineage-specific expansion of plant tandem duplicates in the adaptive response to environmental stimuli. Plant Physiol 148(2):993–1003
Jacobs AK, Dry IB, Robinson SP (1999) Induction of different pathogenesis-related cDNAs in grapevine infected with powdery mildew and treated with ethephon. Plant Pathol 48:325–336
Jayakannan M, Bose J, Babourina O, Rengel Z, Shabala S (2015) Salicylic acid in plant salinity stress signaling and tolerance. Plant Growth Regul 76:25–40
Johnson ET, Dowd PF, Liu ZL, Musser RO (2011) Comparative transcription profiling analyses of maize reveals candidate defensive genes for seedling resistance against corn earworm. Mol Genet Genomics 285:517–525
Jurkowski GI, Smith RK Jr, Yu IC, Ham JH, Sharma SB, Klessig DF, Fengler KA, Bent AF (2004) Arabidopsis DND2, a second cyclic nucleotide-gated ion channel gene for which mutation causes the “defense, no death” phenotype. Mol Plant Microbe Interact 17:511–520
Koonin EV (2006) The origin of introns and their role in eukaryogenesis: A compromise solution to the introns-early versus introns-late debate? Biol Direct 1:22
Liu D, Raghothama KG, Hasegawa PM, Bressan RA (1994) Osmotin overexpression in potato delays development of disease symptoms. Proc Natl Acad Sci USA 91:1888–1892
Liu JJ, Sturrock R, Ekramoddoullah AK (2010a) The superfamily of thaumatin-like proteins: its origin, evolution, and expression towards biological function. Plant Cell Rep 29:419–436
Liu JJ, Zamani A, Ekramoddoullah AK (2010b) Expression profiling of a complex thaumatin-like protein family in western white pine. Planta 231:637–651
Liu H, Liu J, Zhao MM, Chen JS (2015) Overexpression of ShCHL P in tomato improves seedling growth and increases tolerance to salt, osmotic, and oxidative stresses. Plant Growth Regul 77:211–221
Loughran NB, Hinde S, McCormick-Hill S, Leidal KG, Bloomberg S, Loughran ST, O’Connor B, O’Fágáin C, Nauseef WM, O’Connell MJ (2012) Functional consequence of positive selection revealed through rational mutagenesis of human myeloperoxidase. Mol Biol Evol 29:2039–2046
Mackintosh CA, Lewis J, Radmer LE, Shin S, Heinen SJ, Smith LA, Wyckoff MN, Dill-Macky R, Evans CK, Kravchenko S, Baldridge GD, Zeyen RJ, Muehlbauer GJ (2007) Overexpression of defense response genes in transgenic wheat enhances resistance to Fusarium head blight. Plant Cell Rep 26(4):479–488
Marchler-Bauer A, Zheng C, Chitsaz F, Derbyshire MK, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Lu S, Marchler GH, Song JS, Thanki N, Yamashita RA, Zhang D, Bryant SH (2013) CDD: conserved domains and protein three-dimensional structure. Nucleic Acids Res 41:D348–D352
Munis MF, Tu L, Deng F, Tan J, Xu L, Xu S, Long L, Zhang X (2010) A thaumatin-like protein gene involved in cotton fiber secondary cell wall development enhances resistance against Verticillium dahliae and other stresses in transgenic tobacco. Biochem Biophys Res Commun 393:38–44
Neale AD, Wahleithner JA, Lund M, Bonnett HT, Kelly A, Meeks-Wagner DR, Peacock WJ, Dennis ES (1990) Chitinase, beta-1,3-glucanase, osmotin, and extensin are expressed in tobacco explants during flower formation. Plant Cell 2:673–684
Ng G, Seabolt S, Zhang C, Salimian S, Watkins TA, Lu H (2011) Genetic dissection of salicylic acid-mediated defense signaling networks in Arabidopsis. Genetics 189:851–859
Osmond RI, Hrmova M, Fontaine F, Imberty A, Fincher GB (2001) Binding interactions between barley thaumatin-like proteins an (1, 3)-β-d-glucans. Eur J Biochem 15:4190–4199
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Petre B, Major I, Rouhier N, Duplessis S (2011) Genome-wide analysis of eukaryote thaumatin-like proteins (TLPs) with an emphasis on poplar. BMC Plant Biol 11:33
Rajam MV, Chandola N, Goud PS, Singh D, Kashyap V, Choudhary ML, Sihachakr D (2007) Thaumatin gene confers resistance to fungal pathogens as well as tolerance to abiotic stresses in transgenic tobacco plants. Biol Plant 51:135–141
Sakamoto Y, Watanabe H, Nagai M, Nakade K, Takahashi M, Sato T (2006) Lentinula edodes tlg1 encodes a thaumatin-like protein that is involved in lentinan degradation and fruiting body senescence. Plant Physiol 141:793–801
Salzman RA, Tikhonova I, Bordelon BP, Hasegawa PM, Bressan RA (1998) Coordinate accumulation of antifungal proteins and hexoses constitutes a developmentally controlled defense response during fruit ripening in grape. Plant Physiol 117:465–472
Seo PJ, Lee AK, Xiang F, Park CM (2008) Molecular and functional profiling of Arabidopsis pathogenesis-related genes: insights into their roles in salt response of seed germination. Plant Cell Physiol 49:334–344
Singh NK, Kumar KR, Kumar D, Shukla P, Kirti PB (2013) Characterization of a pathogen induced thaumatin-like protein gene AdTLP from Arachis diogoi, a wild peanut. PLoS ONE 8(12):e83963
Small I, Peeters N, Legeai F, Lurin C (2004) Predotar: a tool for rapidly screening proteomes for N-terminal targeting sequences. Proteomics 4:1581–1590
Soderlund C, Bomhoff M, Nelson WM (2011) SyMAP v3.4: a turnkey synteny system with application to plant genomes. Nucleic Acids Res 39:e68
Stern A, Doron-Faigenboim A, Erez E, Martz E, Bacharach E, Pupko T (2007) Selecton 2007: advanced models for detecting positive and purifying selection using a Bayesian inference approach. Nucleic Acids Res 35:W506–W511
Sturn A, Quackenbush J, Trajanoski Z (2002) Genesis: cluster analysis of microarray data. Bioinformatics 18:207–208
Subramanyam K, Arun M, Mariashibu TS, Theboral J, Rajesh M, Singh NK, Manickavasagam M, Ganapathi A (2012) Overexpression of tobacco osmotin (Tbosm) in soybean conferred resistance to salinity stress and fungal infections. Planta 236:1909–1925
Takenaka Y, Nakano S, Tamoi M, Sakuda S, Fukamizo T (2009) Chitinase gene expression in response to environmental stresses in Arabidopsis thaliana: chitinase inhibitor allosamidin enhances stress tolerance. Biosci Biotechnol Biochem 73:1066–1071
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Trobetta ES (2003) The contribution of N-glycans and their processing in the endoplasmic reticulum to glycoprotein biosynthesis. Glycobiology 13:77R–91R
van Loon LC, Rep M, Pieterse CM (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
Velazhahan R, Muthukrishnan S (2003) Transgenic tobacco plants constitutively overexpressing a rice thaumatin-like protein (PR-5) show enhanced resistance to Alternaria alternate. Bio Plant 47:347–354
Wang X, Tang C, Deng L, Cai G, Liu X, Liu B, Han Q, Buchenauer H, Wei G, Han D, Huang L, Kang Z (2010) Characterization of a pathogenesis-related thaumatin-like protein gene TaPR5 from wheat induced by stripe rust fungus. Physiol Plant 139:27–38
Wang Q, Li F, Zhang X, Zhang Y, Hou Y, Zhang S, Wu Z (2011) Purification and characterization of a CkTLP protein from Cynanchum komarovii seeds that confers antifungal activity. PLoS ONE 6:e16930
Wang C, Marshall A, Zhang D, Wilson ZA (2012) ANAP: an integrated knowledge base for Arabidopsis protein interaction network analysis. Plant Physiol 158:1523–1533
Yu XM, Griffith M (1999) Antifreeze proteins in winter rye leaves form oligomeric complexes. Plant Physiol 119:1361–1369
Yu J, Ke T, Tehrim S, Sun F, Liao B, Hua W (2015) PTGBase: an integrated database to study tandem duplicated genes in plants. Database 2015:bav017
Zareie R, Melanson DL, Murphy PJ (2002) Isolation of fungal cell wall degrading proteins from barley (Hordeum vulgare L.) leaves infected with Rhynchosporium secalis. Mol Plant Microbe Interact 15:1031–1039
Zhang Y, Shih DS (2007) Isolation of an osmotin-like protein gene from strawberry and analysis of the response of this gene to abiotic stresses. J Plant Physiol 164:68–77
Zhu BL, Chen THH, Li PH (1995) Expression of 3 osmotin-like protein genes in response to osmotic stress and fungal infection in potato. Plant Mol Biol 28:17–26
Acknowledgments
This project is supported by Grants from the National Science Foundation of China (Nos. 31100923, 31200209), the National Science Foundation of Jiangsu Province (BK2011467), the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD), and Jiangsu University “Youth Backbone Teacher Training Project” (2012–2016).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Likelihood values and parameter estimates for the TLP genes (DOC 51 kb)
Fig. S1
Gene locations and genomic duplication in poplar (A), Arabidopsis (B), rice (C) and maize (D). TLP genes are mapped to the relevant chromosomes. Paralogous regions in the putative ancestral constituents of the three genomes are depicted using SyMAP v3.4 (Soderlund et al. 2011) (PPT 489 kb)
Fig. S2
Origin and evolution of one subgroup of poplar TLP genes by tandem duplication. Letter “D” on the nodes of the phylogenetic tree indicates the positions where tandem duplication has occurred (PPT 62 kb)
Fig. S3
Expression profiles of the rice TLP genes. The data come from the following six experiments. OS8: Expression data from rice embryo, endosperm, root, leaf and seedling. OS10: Expression data for stress treatment (drought, salt and cold) in rice seedling. OS25: Expression data for heat shock in rice seedlings. OS85: Expression data from infected (Xanthomonas oryzae pv. Oryzae (P10) (Xoo) leaf in japonica rice (cv. Y73). OS65: Whole-genome analysis of SSB larvae infestation regulated rice genes. OS92: Rice gene global expression during nonhost interaction with Blumeria graminis f. sp. hordei (Bgh) (PPT 144 kb)
Rights and permissions
About this article
Cite this article
Cao, J., Lv, Y., Hou, Z. et al. Expansion and evolution of thaumatin-like protein (TLP) gene family in six plants. Plant Growth Regul 79, 299–307 (2016). https://doi.org/10.1007/s10725-015-0134-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-015-0134-y