Abstract
For the first time, nuclear genome size and ploidy of five Nitraria species from 49 populations were examined by flow cytometry. All populations were also analyzed for the chromosome number. We identified significant differences in the 2C nuclear DNA content among the analyzed species, and this parameter correlated with their ploidy. Diploid (2n = 2x = 24) species N. sibirica and N. tangutorum were found to have smaller genome size (1.24–1.34 and 1.57–1.65 pg) as compared to tetraploid (2n = 4x = 48) species N. komarovii (2.23–2.32 pg), N. pamirica (3.10–3.30 pg), and N. schoberi (2.93–3.39 pg). Intra-population genome size variation was found in examined species, varying from 1.01 to 1.08-fold. Nitraria sibirica has lower inter-population variation of the 2C (1.08-fold) as compared to N. schoberi (1.16-fold). All the Nitraria species are mixoploids. It turned out that an increase in the equatorial axis of Nitraria pollen is associated with an increase in 2C and 1Cx. An exception is N. komarovii, with its intermediate 2C DNA content and the smallest pollen grains. In general, our data confirm Bobrov's previous views on the system of the genus Nitraria, which distinguished ser. Sibiricae and ser. Schoberianae in sect. Nitraria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The nucleus of plant cells contains the material carrying genetic information. Therefore, studying the nuclear DNA content and ploidy is of fundamental importance for answering complex biological questions. Genomes of organisms at the same level of organization are known to vary substantially in the DNA content. The genome size variation is characterized by a difference in the number of chromosomes, nuclear DNA content, and various repetitive DNA sequences (Sedelnikova 2015).
Research has demonstrated the correlations of genome size with breeding systems and species genesis (Albach and Greilhuber 2004; Weiss-Schneeweiss et al. 2005). Intraspecific variation of genome size has been found among plant specimens from geographically separated populations (Jakob et al. 2004; Schmuths et al. 2004; Smarda and Bures 2006), and the nuclear DNA content correlates with environmental factors (Kalendar et al. 2000; Knight and Ackerly 2002) and plant phenotypic traits (Knight et al. 2005; Murray et al. 2005; Beaulieu et al. 2007). The amount of nuclear DNA can influence the phenotype through regulatory processes in the genome and via simple physical effects of the DNA material at the cellular level (Moeller 2018). These effects are known to result in changes in cell cycle duration, pollen maturation timing, and pollen grain size (Bennett 1972, 1987; Leitch and Bennett 2007; Beaulieu et al. 2008; Lomax et al. 2009), guard and epidermal cell size (Snodgrass et al. 2017), and seed size (Beaulieu et al. 2007). Plant ploidy is reported to be related to pollen grain size (Sousa et al. 2013; Srisuwan et al. 2019). An analysis of genome size in terms of phylogenetic relationships among individual taxa revealed that evolutionary trajectories of genome size and pollen size are probably unrelated (Moeller 2018).
Plants of the genus Nitraria L. are halophytes and are usually confined to intrazonal communities. The considerable isolation of numerous populations of Nitraria species from each other makes this genus a unique model plant for research on processes of genetic differentiation.
In this regard, despite the small number of taxa (10–12 species), no obvious patterns in the genus Nitraria (Bobrov 1965; Khalkuziev 1990; Pan et al. 1999; Banaev et al. 2023) and no clear idea of species genesis have been published so far. The issues about the center and time of origin of the genus Nitraria and pathways of species dispersal also remain debatable. The presence of its representatives in Australia (N. billardierei DC.) and the genus’s biology led Komarov (1908) to believe that Nitraria had originated on saline sea coasts of Gondwana as part of tropical flora before the formation of the Asian and Australian deserts. Later on, almost all researchers believed in phylogenetic antiquity of the genus (Il’in 1944, 1958). At the same time, Korovin (1935) and Bobrov (1946) assumed that the center of origin of the genus Nitraria could be deserts of Central Asia, where at present, there is a center of its diversity. Popov (1927) suggested that a new habitat of the genus Nitraria had formed in Central Asia on the basis of data about N. schoberi L. and N. retusa (Forsk.) Aschers, already existing on the African continent in the Cretaceous period. Pan et al. (1999) assumed an African-Mediterranean origin of the genus in accordance with the distribution of diploid Nitraria species. Nonetheless, the same species has been reported to belong to a diploid or a tetraploid group (Pan et al. 2003; Temirbayeva and Zhang 2015; Marhold et al. 2020). The latest fossil pollen evidences suggest a new evolutionary history of Nitraria (Woutersen et al. 2023). Previous molecular genetic research of some Nitraria species from 31 populations in Siberia, the Republics of Kazakhstan and Tajikistan showed a clear separation of a diploid (ser. Sibiricae Bobrov) and a tetraploid (ser. Schoberianae Bobrov) species (Banaev et al. 2023). Biogeographical analysis suggest that the Central Asian species N. sphaerocarpa Maxim. is the oldest species (Paleocene), and the dispersal started from Central Asia to Africa (since the Oligocene) and to Siberia (5.95 Mya) (Late Miocene) and Australia (2.61 Mya) (Late Pliocene) (Zhang et al. 2015). However, as revealed by integration of fossil pollen morphology and molecular data, the split between N. sphaerocarpa and the other Nitraria types did not happen before the Miocene and modern species of Nitraria originate during the Late Miocene (Woutersen et al. 2023).
The nuclear DNA content, estimated by flow cytometry, is an essential genome feature together with the chromosome number (Doležel and Bartoš 2005). Karyotype analysis is an important method for revising species classification and studying phylogenetic relationships (Hong 2021). Flow cytometry can be considered a useful method for understanding taxonomic relationships (Bourge et al. 2018).
Available Nitraria-related cytological information that is important for understanding Nitrariaceae evolution is very scarce (Tarnavshi 1948; Reese 1958; Pan et al. 2003; Banaev et al. 2018a,b; Voronkova et al. 2018; Marhold et al. 2020, 2021, 2022). The Plant DNA C-values Database (https://cvalues.science.kew.org/; accessed on 20 June 2023) does not contain information on genome sizes of Nitraria species. Determination of genetic variability of wild plants helps to conserve and use them (Khaleghi and Khadivi 2023).
The purpose of this study was to analyze intra- and inter-population variations of the 2C DNA content and chromosome number and their correlations with pollen grain size of Nitraria species.
Materials and methods
Plant materials
Seeds of five species of the genus Nitraria (N. sibirica Pall., N. schoberi, N. pamirica L.I. Vassiljeva, N. komarovii Iljin & Lava ex Bobrov, and N. tangutorum Bobrov) were collected in 49 natural populations in Russia, Tajikistan, and Kazakhstan from 2009 to 2021 (Table 1, Fig. 1). Twenty-four populations of N. sibirica, 22 populations of N. schoberi, and one population of each of three species (N. pamirica, N. komarovii, and N. tangutorum) were investigated. The vouchers are stored in herbarium NSK (the Dendrology Laboratory of the CSBG SB RAS, Novosibirsk, Russia) and available in the digital herbarium of the CSBG SB RAS, NSK (http://herb.csbg.nsc.ru:8081).
Fresh leaves of Pisum sativum L. ‘Ctirad’ (2C = 9.09 pg) and Raphanus sativus L. ‘Saxa’32 (2C = 1.11 pg) (Doležel et al. 1998) grown from seeds obtained from the Centre of Plant Structural and Functional Genomic at the Institute of Experimental Botany of the Academy of Sciences, AS CR (Olomouc-Holice, Czech Republic) (Doležel et al. 1992) were used as an internal standard.
Flow Cytometry (FCM)
All FCM procedures were performed in the Central Siberian Botanical Garden SB RAS (Novosibirsk, Russia). The analysis was performed on a Cy Flow Space instrument (Sysmex Partec, Norderstedt, Germany) with a 532 nm laser source. The DNA content of plants was determined by FCM with staining of isolated nuclei with propidium iodide (PI). The seeds of Nitraria were analyzed following a previously developed methodology (Banaev et al. 2018b). At least 10 plants were randomly selected in each population of Nitraria species for genome size variation analysis.
Nitraria plant embryos extracted from the seed were ground up using a razor blade in plastic Petri dishes together with an appropriate amount of an internal standard (P. sativum or R. sativus) in 500 µL of chilled extraction buffer (Nuclei Extraction Buffer) (Sysmex Partec, Norderstedt, Germany) according to the manufacturer’s protocol. The plant tissue samples were incubated at room temperature for 2 min. After the extraction of the nuclei, the samples were passed through a 50 µm Celltrics Partec disposable filter (Sysmex Partec, Norderstedt, Germany), followed by the addition of 2 mL of the Staining Solution (for staining) consisting of Staining Buffer (Sysmex Partec, Norderstedt, Germany), PI (50 µg/mL), and RNase A (50 µg/mL). The staining was performed at room temperature in a dark place for 15 min. The prepared nuclei samples were stored in a refrigerator for no more than 4 h.
Next, 15,000 FCM events were collected [the required number is 5,000 to 20,000 (Galbraith et al. 1998; Doležel and Bartoš 2005)] three times per sample; the coefficient of variation of the mean was less than 5%. A relative nuclear DNA content was calculated based on a linear relation between fluorescence signals from stained nuclei of tested specimens and the internal standard (Doležel et al. 2007) and was expressed as an index.
The chromosome number (CHN)
Seeds were stratified on moist filter paper for 1 month and germinated at 27–28 °C. For fixation, roots 0.5–2.5 cm long were selected. After that, 30 individual plant specimens from each population of Nitraria were subjected to the determination of chromosome numbers (2n). Cytological procedures were performed on root meristem. Actively growing seedlings were kept for 3 h at room temperature in a 0.2% colchicine solution and fixed in an ethanol: acetic acid solution (3:1). Seeds were fixed between 10:00 and 11:00 AM (UTC + 7). The preparations were stained with acetohematoxylin according to Smirnov (1968). Chromosome examination and photodocumentation of metaphase plates were carried out under an Axioscope 40 microscope equipped with an AxioCam MRc 5 color digital camera and AxioVision v.4.8 software (Carl Zeiss Ltd., Göttingen, Germany) and under an Axioscope A1 microscope with an Axiocam 506 color digital camera and ZEN 2012 (blue edition) software (Carl Zeiss Ltd., Göttingen, Germany).
The most common number is taken as the value of the chromosome number. The following notation is used for chromosome numbers: modal number and numbers determined.
Morphometric results on pollen grains
For a comparative analysis of the DNA content and pollen characteristics, pollen morphometric data were borrowed from a previously published article (Tomoshevich et al. 2022). The following traits were characterized: polar axis (P, µm), equatorial axis (E, µm), and the P/E ratio.
Statistical analysis
These procedures were carried out in Microsoft Excel 7.0 and STATISTICA 6.0 (correlation analysis, LSD test, Newman–Keuls test, ANOVA, at p ≤ 0.05) (StatSoft Inc., Tulsa, OK, USA). The images and data obtained by FCM were analyzed in the CyFlow® Space software (Sysmex Partec, Norderstedt, Germany). The findings are presented as mean values with standard error, standard deviation, and a coefficient of variation (mean ± SE; mean ± SD; CV, %). To calculate the 1C value, 2C was divided by two, and to calculate 1Cx, 2C was divided by ploidy.
Results
Genome size and ploidy
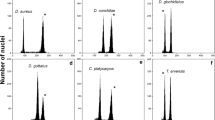
FCM revealed three peaks in N. sibirica, N. komarovii, N. tangutorum, and N. pamirica and four peaks in N. schoberi (Fig. 2). The first peaks in N. schoberi and N. pamirica proved to be internal standard peaks G1 and G2 (R. sativus). The next two peaks represented the G1 and G2 peaks of Nitraria (Fig. 2c, d). In N. sibirica, N. komarovii, and N. tangutorum, the first peaks were peaks G1 and G2 of the Nitraria plant itself, followed by the internal standard (P. sativum) (Fig. 2a, b, e). Figure 2f shows the histograms of N. sibirica, N. komarovii, and N. schoberi without the internal standard and illustrates the differences in genome size among these species.
According to the FCM results, the species in question can be classified into diploids (2n = 2x = 24) and tetraploids (2n = 4x = 48). Statistical analysis revealed four isolated groups based on 2C genome size (Fig. 3). The lowest DNA content was found in N. sibirica (1.24–1.34 pg), and the highest in N. pamirica (3.10–3.30 pg) and N. schoberi (2.93–3.39 pg).
The lowest intra-population variation of the DNA content is characteristic of N. sibirica (1.01–1.02-fold), and the highest variation is characteristic of N. schoberi (1.03–1.08-fold). In N. komarovii, N. tangutorum, and N. pamirica, the intra-population variation of genome size proved to be 1.04-, 1.05-, and 1.06-fold, respectively (Table 2). Additionally, N. sibirica has lower inter-population variation of the DNA content (maximum variation 1.08-fold) as compared to N. schoberi (maximum variation 1.16-fold). A DNA content (2C) analysis of variance (Newman–Keuls test, p ≤ 0.05) subdivided N. schoberi populations into two groups: 2.93–3.10 and 3.17–3.38 pg. No significant differences were found among N. sibirica populations.
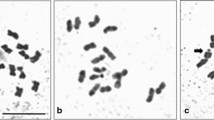
We noticed that in N. pamirica, N. komarovii, and N. schoberi, the modal number of chromosomes is 2n = 48, whereas in N. sibirica and N. tangutorum, it is 2n = 24, confirming the ploidy shown by FCM (Table 2, Fig. 2).
The characterization of the karyotype of Nitraria species helped us to determine the most frequent numbers of chromosomes (Table 2, Fig. 4). For instance, in N. sibirica, cells with 2n = 3x = 36, 2n = 4x = 48, 2n = 5x = 60 were found; in N. tangutorum, 2n≈26; in N. schoberi, 2n = 2x = 24, 2n = 5x = 60, 2n = 6x = 72, 2n = 8x = 96 (Fig. 5); in N. pamirica, 2n = 2x = 24. Cells with 2n≈40 and 2n≈80 were sometimes observed in N. schoberi and N. sibirica.
Different sets of chromosomes in Nitraria species were noted among different plants within populations and among cells of a single plant. The analysis of the nuclear DNA of N. sibirica revealed the presence of endopolyploid nuclei (up to 16C) in plants from different populations (Fig. 6).
Correlation of the nuclear DNA content with pollen grain size
The correlation analysis of datasets of pollen size and DNA contents uncovered a positive correlation between the pollen E and the 2C value (r = 0.52) and a negative correlation between the P/E ratio and 2C (r = –0.54) (Tables 3 and 4), indicating that E goes up with the increasing DNA content along with unchanged or decreasing pollen grain size. Monoploid genome size (1Cx) positively correlated with E even more strongly (r = 0.64).
Discussion
Genome size and ploidy
Interspecies variation of genome size is a well-known fact (Bennet et al. 2000; Doležel et al. 2007), but genome size within a species is thought to be stable (Greilhuber et al. 2005; Lomonosova et al. 2020). Lysak et al. (2000) documented a 1.06-fold variation of genome size in European populations of Sesleria albicans. Similarly, lack of a significant variation, i.e., a 1.06-fold difference, was proved in Pinus nigra (Bogunic et al. 2007). In Trifolium repens and T. fragiferum, only a slight intraspecific variation of genome size was recorded, 1.05- and 1.03-fold, respectively (Lukjanová and Řepková 2021). In a paper about Allium cepa cultivars from different parts of the world, some authors reported exceptional intraspecific stability of genome size (Bennet et al. 2000).
Our results indicate intra-population stability of the nuclear DNA content in all five examined species, ranging from 1.01- to 1.08-fold. A similar magnitude of variation was observed within a single plant of each species of genus Nitraria (Banaev et al. 2018a). This result can be explained by the finding that seeds collected from one bush may have a male gametophyte from another plant (Voronkova et al. 2018).
We detected no obvious patterns regarding levels of variation of the DNA content across populations. In particular, among N. schoberi plants from the Eastern Pamir, the variation is 1.07-fold in the Pyandzh1 population and 1.01-fold in the Pyandzh2 population located 20 km away. Among N. sibirica plants, the highest variation of genome size (1.02-fold) was found in populations of Altai Krai (Uglovskoe), Altai Republic (Kosh-Agach), Tuva (Turan), and Kazakhstan (Koktal), located at a substantial distance from each other. Correlations between ecological differentiation and genome size have been reported at both interspecific and intraspecific levels (Knight et al. 2005; Knight and Ackerly 2002), for example, in plant species of genera Larrea (Poggio et al. 1989), Cardiospermum (Urdampilleta et al. 2012), Berberis (Bottini et al. 2000), Coffea (Razafinarivo et al. 2012), and Psidium (Tuler et al. 2019). For instance, it was shown that the species of Larrea, Bulnesia, and Pintoa that inhabit the most arid environments are the ones possessing the highest DNA content (Poggio et al. 1989). In other research, intraspecific DNA content variation has correlated with a geographic environment and ploidy in Festuca pallens (Smarda and Bures 2006) and Miscanthus sp. (Sheng et al. 2016). Our results suggest that among N. schoberi populations, the DNA content is lower in plants growing in the Balkhash-Alakol basin and along the coasts of large water bodies, the Black Sea and Caspian Sea.
Our analysis of DNA content variation showed higher stability of genome size in N. sibirica than in N. schoberi. This finding is consistent with available data on increased variation of genome size in polyploids (Tuna et al. 2017). As stated above, N. sibirica is diploid (2n = 2x = 24), and N. schoberi is tetraploid (2n = 4x = 48).
Our results show that monoploid genome size of Nitraria varies 1.40-fold (0.57–0.80 pg), with the smallest value in N. komarovii. Genome downsizing in the process of polyploidization may increase a plant’s environmental adaptive fitness and facilitate competition with their diploid species. For example, it was demonstrated that altered 1Cx values reflect plasticity of the polyploid genome in various Miscanthus species (Sheng et al. 2016). Furthermore, a decrease in the size of a monoploid genome indicates that the species in question is evolutionarily young (Šmarda et al. 2008). Our finding that the monoploid genome of N. komarovii is the smallest supports the point of view of Bobrov (1946), according to whom N. komarovii is the youngest species associated with the recent history of the Caspian Basin.
It is documented in the literature that in Nitraria species, most frequent chromosome numbers are divisible by the principal number x = 12 or nondivisible by it (Zakharyeva and Astanova 1968; Pan et al. 2002, 2003). For N. schoberi from mountain deserts of Central Asia, 2n = 24 was reported (Reese 1958), and for N. schoberi from southern Romania, it is 2n = 66 (Tarnavshi 1948). The data on chromosome sets for most plants are now readily available and collected in publicly available resources such as the Chromosome Counts Database (CCDB; http://ccdb.tau.ac.il/ (accessed on 20 August 2023), where information (2n) on six species of the genus Nitraria is available: N. sibirica (24, 30, and 60), N. schoberi (24, 34, 42, 48, 60, ⁓66, 72, and 96), N. pamirica (24 and 48), N. komarovii (24), N. tangutorum (24), and N. retusa (18 and 24). In the Index to Plant Chromosome Numbers (IPCN, http://legacy.tropicos.org/Project/IPCN; accessed on 20 August 2023), data (2n) are given for N. sibirica (24 and 60) and N. retusa (24).
Earlier, various cytotypes in the genus Nitraria have been documented; for instance, in N. pamirica, the typical chromosome number proved to be 2n = 48, 2n≈4x, and 2C = 3.15 pg, but only a few specimens showed 2n = 24, 2n≈2x, 2C = 1.50 pg; in N. schoberi the typical number is 2n = 48, 2n≈4x, 2C = 2.98 pg, but some specimens have 2n≈8x≈96, 2C = 5.75 (Marhold et al. 2020). Furthermore, different chromosome numbers have been detected among cells of a single plant, thus pointing to mixoploidy. The latter is a phenomenon characteristic of many woody plant species (Butorina 1989; Butorina and Gavrilov 2001), in particular, it is typical for representatives of families with small chromosomes, including Nitraria (Muratova et al. 2011, 2013). Proportions (%) of cells having different ploidy levels is one of the factors of plant adaptation to new or extreme habitat conditions, e.g., drought, strong light intensity, and high salinity (Cookson et al. 2006; Kunakh 2011; Gegas et al. 2014; Scholes and Paige 2015). According to Sedelnikova (2015), mixoploidy and aneuploidy in Pinaceae family species are often seen in extreme intrazonal bog and mountain ecotypes, and the highest level of mixoploidy for conifers is found in populations of Larix sibirica at the northern border of the species range. The level of endoreduplication may be species-specific or may differ between populations or even between individuals of the same species (Barow and Meister 2003). When researching the family Chenopodiaceae, Skaptsov et al. (2017) reported that endopolyploidy is usually observed in diploid species of the genera Chenopodium, Dysphania, Oxybasis, and Suaeda and not found in polyploid specimens of Suaeda.
The correlation of the nuclear DNA content with pollen grain size
The positive correlation between pollen grain size and the nuclear DNA content is probably the most easily explained because pollen contains only the components necessary for the initiation and maintenance of pollen tube growth and carries a haploid genome. A direct correlation between the nuclear DNA content and pollen size has been found in many plant species and groups (Bennett 1987; Bennett et al. 2005; Sinjushin 2021). Of interest are correlations between the DNA content, ploidy, and pollen grain size. For example, among 17 species of the genus Lippia, a positive association was detected between the chromosome number and pollen grain size as well as between ring length and ring width (Sousa et al. 2013). In Ipomoea batatas and I. trifida, both genome size and ploidy correlate with pollen size (Srisuwan et al. 2019). For species of the genus Plantago, pollen diameter was shown to correlate with 2C but not necessarily with ploidy (Wong and Murray 2012).
According to Amer and Amany (2014), there is a strong correlation between polyploidy and pollen morphological variation in Atriplex halimus. By contrast, Knight et al. (2010) found no association between pollen size and genome size after examining phylogenetic history of 464 plant species.
We did not see enlargement of the pollen grain with increasing ploidy or increasing genome size in the examined species of Nitraria. Our results indicate a direct correlation between the 2C DNA content, chromosome number, and E of pollen grains in species N. pamirica, N. schoberi, N. sibirica, and N. tangutorum. N. komarovii is an exception because it is a tetraploid but is characterized by the smallest pollen E. The P/E ratio, which is a parameter of pollen grain shape, is lower in tetraploids N. pamirica, N. schoberi, and N. komarovii than in diploids N. sibirica and N. tangutorum.
Conclusions
Our work shows that in species of the genus Nitraria, genome size, ploidy, and pollen grain size are interrelated. We can rank the species under study by genome size and ploidy as follows: diploid (2n = 2x = 24): N. sibirica (1.30 pg) and N. tangutorum (1.59 pg); tetraploid (2n = 2x = 48): N. komarovii (2.28 pg), N. pamirica (3.15 pg), N. schoberi (3.19 pg). By pollen E and 2C (E; 2C), the ranking is as follows: N. sibirica (20.97 μm; 1.30 pg), N. tangutorum (21.14 μm; 1.59 pg) < N. schoberi (24.83 μm; 3.19 pg), N. pamirica (26.50 μm; 3.15 pg). An exception is N. komarovii (16.42 μm; 2.28 pg), having an intermediate DNA content and the smallest pollen width. A comparison of pollen E and 1Cx (E; 1Cx) results in the following ranking: N. komarovii (16.42; 0.57) < N. sibirica (20.97; 0.65) < N. tangutorum (21.14; 0.78) < N. schoberi (24.83; 0.8) < N. pamirica (26.50; 0.79). An increase in E clearly correlates with 1Cx. Our findings indicate the correctness of Bobrov’s (1946) views on relationships within the genus Nitraria, who distinguished two series in sect. Nitraria: ser. Sibiricae (N. tangutorum and N. sibirica) and ser. Schoberianae (N. schoberi and N. komarovii). N. pamirica, which we believe should be placed in the ser. Schoberianae, was not known to E.G. Bobrov because it was described by Vasilieva (1974) later. It is obvious that the five species under study differ in their genesis. As pointed out by Bobrov, the species of the ser. Sibiricae are probably related in origin to ancient deserts of Central Asia, whereas N. schoberi originated in the Aral-Caspian lowlands and spread eastward and southeastward to Central Asia and Western Siberia.
References
Albach DC, Greilhuber JG (2004) Genome size variation and evolution in veronica. Ann Bot 94:897–911. https://doi.org/10.1093/aob/mch219
Amer WM, Amany SA (2014) Infra-specific pollen diversity of Atriplex halimus L. in Egyptian flora. IJRSB 2:36–48
Banaev EV, Tomoshevich MA, Voronkova MS (2018) Flow cytometry analysis of the relative content of nuclear DNA in Nitraria schoberi L. seeds. Botanica Pacifica 7(1):89–92
Banaev EV, Tomoshevich MA, Khozyaykina SA, Erst AA, Erst AS (2023) Integrative taxonomy of Nitraria (Nitrariaceae), description of the new enigmatic species and key to all currently known species. Plants 12:593. https://doi.org/10.3390/plants12030593
Banaev EV, Tomoshevich MA, Ak-Lama TA (2018a) IAPT / IOPB chromosome date 27. In: Marhold K, Breitwieser I (eds) Taxon 67(5):1042, E2. https://doi.org/10.12705/675.24
Barow M, Meister A (2003) Endopolyploidy in seed plants is differently correlated to systematics, organ, life strategy and genome size. Plant Cell Environ 26:571–584
Beaulieu JM, Moles AT, Leitch IJ, Bennett MD, Dickie JB, Knight CA (2007) Correlated evolution of genome size and seed mass. New Phytol 173:422–437. https://doi.org/10.1111/j.1469-8137.2006.01919.x
Beaulieu JM, Leitch IJ, Patel S, Pendharkar A, Knight CA (2008) Genome size is a strong predictor of cell size and stomatal density in angiosperms. New Phytol 179:975–986. https://doi.org/10.1111/j.1469-8137.2008.02528.x
Bennet MD, Bhandol P, Leitch IJ (2000) Nuclear DNA amounts in angiosperms and their modern uses – 807 new estimates. Ann Bot 86:859–909. https://doi.org/10.1006/anbo.2000.1253
Bennett MD (1972) Nuclear DNA content and minimum generation time in herbaceous plants. Proc R SocLond B Biol Sci 181:109–135
Bennett MD (1987) Variation in genomic form in plants and its ecological implications. New Phytol 106(Suppl):S177–S200
Bennett MD, Leitch I, Gregory T (2005) Genome size evolution in plants. In: Ryan G (ed) The Evolution of the Genome books. Elsevier, San Diego, pp 89–162
Bobrov EG (1946) About Asian species of the genus Nitraria L. Sov Bot 14(1):19–30 ((in Russian))
Bobrov EG (1965) On the origin of flora of the deserts of the Old World in conjunction with the review of Nitraria genus. Botanicheskij Zhurnal 50(8):1053–1057 ((in Russian))
Bogunic F, Muratović E, Ballian D, Brown SC (2007) Genome size stability among five subspecies of Pinus nigra Arnold s.l. Environ Exp Bot 59(3):354–360
Bottini MCJ, Greizerstein EJ, Aulicino MB, Poggio L (2000) Relationships among genome size, environmental conditions and geographical distribution in natural populations of nw patagonian species of Berberis L. (Berberidaceae). Ann Bot 86(3):565–573
Bourge M, Brown S, Siljak-Yakovlev S (2018) Flow cytometry as tool in plant sciences, with emphasis on genome size and ploidy level assessment. Gen Appl 2(2):1–12
Butorina AK (1989) Factors of evolution of arboreal karyotypes. Uspekhi Sovremennoy Biologii 108(6):342–357
Butorina AK, Gavrilov IA (2001) Cytogenetic study of some species of the genus Tilia L. Tsitologiya 43(10):934–939
Cookson SJ, Radziejwoski A, Granier C (2006) Cell and leaf size plasticity in Arabidopsis: what is the role of endoreduplication? Plant Cell Environ 29(7):1273–1283
Doležel J, Bartoš J (2005) Plant DNA flow cytometry and estimation of nuclear genome size. Ann Bot 95:99–110
Doležel J, Sgorbati S, Lucretti S (1992) Comparison of three DNA fluorochromes for flow cytometric estimation of nuclear DNA content in plants. Physiol Plant 85:625–631
Doležel J, Greilhuber J, Lucretti S, Meister A, Lysak MA, Nard L, Obermayer R (1998) Plant genome size estimation by flow cytometry: inter-laboratory comparison. Ann Bot 82:17–26
Doležel J, Greilhuber J, Suda J (2007) Estimation of nuclear DNA content in plants using flow cytometry. Nat Protoc 2:2233–2244. https://doi.org/10.1038/nprot.2007.310
Galbraith DW, Lambert GM, Macas J, Doležel J (1998) Analysis of nuclear DNA content and ploidy in higher plants. In: Robinson JP, Darzynkiewicz Z, Dean PN, Dressler LG, Orfao A, Rabinovitch PS, Stewart CC, Tanke HJ, Wheeless LL (eds) Current protocols in cytometry. New York, John Wiley & Sons, 7.6.1–7.6.22.
Gegas VC, Wargent JJ, Pesquet E, Granqvist E, Paul ND, Doonan JH (2014) Endopolyploidy as a potential alternative adaptive strategy for Arabidopsis leaf size variation in response to UV-B. J Exp Bot 65(10):2757–2766
Greilhuber J, Dolezel J, Lysak M, Bennett MD (2005) The origin, evolution and proposed stabilization of the terms ‘genome size’ and ‘C-value’ to describe nuclear DNA contents. Ann Bot 95:255–260
Hong DY (2021) Peonies of the World: Phylogeny and Evolution. Royal Botanic Gardens Kew, Richmond.
Il’in MM (1944) Nitraria and the origin of desert flora. Priroda 5–6:116–118 ((in Russian))
Ilyin MM (1958) Flora of the deserts of Central Asia, its origin and stages of development. Mater History of Flora and Veg USSR 3:129–229 ((in Russian))
Jakob SS, Meister A, Blattner FR (2004) The considerable genome size variation in Hordeum species (Poaceae) is linked to phylogeny, life form, ecology, and speciation rates. Mol Biol Evol 21:860–869
Kalendar R, Tanskanen J, Immonen S, Nevo E, Schulman AH (2000) Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamics in response to sharp microclimatic divergence. Proceed National Acad Sci USA 97:6603–6607
Khaleghi A, Khadivi A (2023) Morphological characterizations of wild nitre-bush (Nitraria schoberi L.) specimens. Genet Resour Crop Evol. https://doi.org/10.1007/s10722-023-01635-3
Khalkuziev P (1990) About family ties of some plant families of desert regions. Publishing house "Fan" of the Uzbekskoy SSR, Tashkent.
Knight CA, Ackerly DD (2002) Variation in nuclear DNA content across environmental gradients: a quantile regression analysis. Ecol Lett 5:66–76. https://doi.org/10.1046/j.1461-0248.2002.00283.x
Knight CA, Molinari N, Petrov DA (2005) The large genome constraint hypothesis: evolution, ecology and phenotype. Ann Bot 95:177–190
Knight CA, Clancy RB, Götzenberger L, Dann L, Beaulieu JM (2010) On the relationship between pollen size and genome size. J Bot. https://doi.org/10.1155/2010/612017
Komarov VL (1908) Introduction to the floras of China and Mongolia. Tr S-Peterb Bot Sada 19:1–179 ((in Russian))
Korovin EP (1935) Essays on the history of vegetation development in Central Asia (Central Kazakhstan). Bull SAGU 20(4). (in Russian)
Kunakh VA (2011) Ontogenetic plasticity of the genome as a basis for plant adaptability. Zhebrakovskiye readings. III. Transformation of genomes, 3–53.
Leitch IJ, Bennett MD (2007) Genome size and its uses: the impact of flow cytometry. In: Doležel J, Greilhuber J, Suda J (eds) Flow cytometry with plant cells: analysis of genes, chromosomes and genomes. Wiley-VCH, Weinheim, pp 153–176
Lomax BH, Woodward FI, Leitch IL, Knight CA, Lake JA (2009) Genome size as a predictor of guard cell length in Arabidopsis thaliana is independent of environmental conditions. New Phytol 181:311–314. https://doi.org/10.1111/j.1469-8137.2008.02700.x
Lomonosova MN, Ankova TV, Voronkova MS, Korolyuk EA, Banaev EV, Skaptsov MV (2020) Ploidy level in the representatives of Chenopodiaceae as revealed by genome size and chromosome numbers. Turczaninowia 23(1):24–31
Lukjanová E, Řepková J (2021) Chromosome and genome diversity in the genus Trifolium (Fabaceae). Plants 10:2518. https://doi.org/10.3390/plants10112518
Lysak M, Rostková A, Dixon JM, Dolezel J (2000) Limited genome size variation in Sesleria albicans. Ann Bot 86(2):399–403. https://doi.org/10.1006/anbo.2000.1200
Marhold K, Kučera J, Alexeeva T, Andriyanova E, An’kova TV, Astashenkov AY, Banaev EV, Chepinoga VV, Cheryomushkina VA, Dorogina OV et al (2020) IAPT chromosome data 32. Taxon 69(5):1126–1132. https://doi.org/10.1002/tax.12322
Marhold K, Kučera J, Albach DC, Aleksandrova TG, Banaev EV, Dyubenko TV, Gnutikov AA, Korolyuk EA, Kotseruba VV, Krivenko DA et al (2021) IAPT chromosome data 34/2. Taxon 70(5):1149
Marhold K, Kučera J, Aleksandrova TG, Alexeeva TV, Andriyanova EA, Banaev EV, Bobrov AA, Boltenkov EV, Bondarevich EA, Boyarskikh IG et al (2022) IAPT chromosome data 38/2. Taxon 71(6):1353–1360
Moeller M (2018) Nuclear DNA C-values are correlated with pollen size at tetraploid but not diploid level and linked to phylogenetic descent in Streptocarpus (Gesneriaceae). S Afr J Bot 114:323–344. https://doi.org/10.1016/j.sajb.2017.11.017
Muratova EN, Kvitko OV, Banaev EV, Ts-Ch Z, Wang G (2011) Karyological study of some representatives of Nitraria L. Botan Zhurn 96(1):108–115
Muratova EN, Goryachkina OV, Banaev EV (2013) Karyological study of Siberian species Nitraria L. (Nitrariaceae). Turczaninowia 16(4):50–54
Murray BG, De Lange PJ, Ferguson AR (2005) Nuclear DNA variation, chromosome numbers and polyploidy in the endemic and indigenous grass flora of New Zealand. Ann Bot 96:1293–1305. https://doi.org/10.1093/aob/mci281
Pan XY, Shen GM, Chen P (1999) A preliminary research on taxonomy and systematics genus Nitraria. Acta Bot Yunnan 21(3):287–295
Pan XY, Ca QD, Wei QS, Wang GX (2002) Progress of researches on systematics and biodiversity in the genus Nitraria. Chinese Acad Med J Org 4:1–6
Pan XY, Wei XP, Yu QS, Chen JK, Wang GX (2003) Polyploidy: classification, evolution and applied perspective of the genus Nitraria. Chin Bull Bot 20(5):632–638
Poggio L, Burghardt A, Hunziker J (1989) Nuclear DNA variation in diploid and polyploid taxa of Larrea (Zygophyllaceae). Heredity 63:321–328. https://doi.org/10.1038/hdy.1989.105
Popov MG (1927) The main features of the history of the development of the flora of Central Asia. Bull SAGU 15:239–293 ((in Russian))
Razafinarivo NJ, Rakotomalala JJ, Brown SC, Bourge M, Hamon S, de Kochko A, Poncet V, Dubreuil-Tranchant C, Couturon E, Guyot R, Hamon P (2012) Geographical gradients in the genome size variation of wild coffee trees (Coffea) native to Africa and Indian Ocean islands. Tree Genet 8:1345–1358. https://doi.org/10.1007/s11295-012-0520-9
Reese G (1958) Cyto-systematische Notizen zur Gattung Nitraria (Zygophyllaceae). Flora 146(3):478–487
Schmuths H, Meister A, Horres R, Bachmann K (2004) Genome size variation among accessions of Arabidopsis thaliana. Ann Bot 93:317–321. https://doi.org/10.1093/aob/mch037
Scholes DR, Paige KN (2015) Plasticity in ploidy: a generalized response to stress. Trends Plant Sci 20(3):165–175. https://doi.org/10.1016/j.tplants.2014.11.00
Sedelnikova TS (2015) Variability of the genome size of coniferants under extreme growth conditions. Uspekhi Sovremennoy Biologii 135(5):514–528 ((in Russian))
Sheng J, Hu X, Zeng X, Li Y, Zhou F, Hu Z, Diao Y (2016) Nuclear DNA content in Miscanthus sp and the geographical variation pattern in Miscanthus lutarioriparius. Sci Rep 6(1):1–8
Sinjushin A (2021) The duration of the life cycle is associated with C-value and affects reproductive features in the Fabeae, the tribe with largest genomes in Fabaceae. Flora 285:151954. https://doi.org/10.1016/j.flora.2021.151954
Skaptsov MV, Lomonosova MN, Kutsev MG, Smirnov SV, Shmakov AI (2017) The phenomenon of endopolyploidy in some species of the Chenopodioideae (Amaranthaceae). Bot Lett 164(1):47–53. https://doi.org/10.1080/23818107.2016.1276475
Šmarda P, Bureš P, Horová L, Foggi B, Rossi G (2008) Genome size and GC content evolution of Festuca: ancestral expansion and subsequent reduction. Ann Bot 101(3):421–433. https://doi.org/10.1093/aob/mcm307
Smarda P, Bures P (2006) Intraspecific DNA content variability in Festuca pallens on different geographical scales and ploidy levels. Ann Bot 98:665–678. https://doi.org/10.1093/aob/mcl150
Smirnov YuA (1968) Accelerated method for studying somatic chromosomes in fruit trees. Tsitologiya 10:1132–1134 ((in Russian))
Snodgrass SJ, Jareczek J, Wendel JF (2017) An examination of nucleotypic effects in diploid and polyploid cotton. AoB Plants 9(1):plw082
Sousa SM, Pierre PM, Torres GA, Davide LC, Viccini LF (2013) Relationship between pollen morphology and chromosome numbers in Brazilian species of Lippia L. (Verbenaceae). An Acad Bras Cienc 85(1):147–157
Srisuwan S, Sihachakr D, Martín J, Valles J, Ressayre A, Brown SC, Siljak-Yakovlev S (2019) Change in nuclear DNA content and pollen size with polyploidisation in the sweet potato (Ipomoea batatas, Convolvulaceae) complex. Plant Biol (stuttg) 21(2):237–247. https://doi.org/10.1111/plb.12945
Tarnavshi I (1948) Die chromosomenzahlen der Anthophyten-Flora von Rumanien mit einem Ausblick auf das polyploidie-Problem. Buletinul Gradinii Botanice Si Al Muzeului Botanic De La Universitatea Din Cluj 28:1–130
Temirbayeva K, Zhang ML (2015) Molecular phylogenetic and biogeographical analysis of Nitraria based on nuclear and chloroplast DNA sequences. Plant Syst Evol 30:1897–1906. https://doi.org/10.1007/s00606-015-1202-5
Tomoshevich M, Banaev E, Khozyaykina S, Erst A (2022) Pollen morphology of some species from genus Nitraria. Plants 11:2359. https://doi.org/10.3390/plants11182359
Tuler AC, Carrijo TT, Peixoto AL, Garbin ML, da Silva Ferreira MF, Carvalho CR, Spadeto MS, Clarindo WR (2019) Diversification and geographical distribution of Psidium (Myrtaceae) species with distinct ploidy levels. Trees 33:1101–1110. https://doi.org/10.1007/s00468-019-01845-2
Tuna GS, Duyu G, Uzun K, Yücel G, Tuna M (2017) Determination of nuclear DNA content and ploidy of Hypericum perforatum L accessions collected from Western Turkey. Tarim Bilimleri Dergisi-J Agricultural Sci 23:395–403
Urdampilleta JD, Coulleri JP, Ferrucci MS, Forni-Martins ER (2012) Karyotype evolution and phylogenetic analyses in the genus Cardiospermum L. (Paullinieae, Sapindaceae). Plant Biol (Stuttg) 15(5):868–881
Vasil’eva LI (1974) A new species of the genus Nitraria L. from the Pamirs. Nov Sist Vyssh Rast 11:341–344
Voronkova MS, Banaev EV, Tomoshevich MA, Ak-Lama T (2018) Variation of nuclear DNA content in seeds of Nitraria schoberi L. BIO Web of Conferences 11:00046. https://doi.org/10.1051/bioconf/20181100046
Weiss-Schneeweiss H, Greilhuber J, Schneeweiss GM (2005) Genome size evolution in holoparasitic Orobanche (Orobanchaceae) and related genera. Am J Bot 93:148–156
Wong Ch, Murray BG (2012) Variable changes in genome size associated with different polyploid events in Plantago (Plantaginaceae). J Hered 103(5):711–719. https://doi.org/10.1093/jhered/ess049
Woutersen A, Jardine Ph, Silvestro D, Bogotá-A RG, Zhang HX, Meijer N, Bouchal J, Barbolini N, Dupont-Nivet G, Koutsodendris A, Antonelli A, Hoorn C (2023) The evolutionary history of the Central Asian steppe-desert taxon Nitraria (Nitrariaceae) as revealed by integration of fossil pollen morphology and molecular data. Bot J Linn. https://doi.org/10.1093/botlinnean/boac050
Zakharyeva OI, Astanova SB (1968) Chromosomal numbers of some wild species of flowering plants in Central Asia. Doklady Akademii Nauk Tadzhikskoy SSR 11(11):72–75 ((in Russian))
Zhang ML, Temirbayeva K, Sanderson S, Chen X (2015) Young dispersal of xerophil Nitraria lineages in intercontinental disjunctions of the Old World. Sci Rep 5:13840. https://doi.org/10.1038/srep13840
Acknowledgements
During the preparation of this publication, materials of bioresource scientific collections of the CSBG SB RAS “Collections of living plants indoors and outdoors” (unique scientific unit USU_440534) and “Herbarium of higher plants, lichens and fungi (NS, NSK)” (USU-4450537) were used.
Funding
This work was supported by the Ministry of Science and Higher Education of the Russian Federation program of scientific research “Theoretical and applied aspects of studying gene pools of natural plant populations and conservation of plant diversity ‘outside the typical environment’ (ex situ)” (project No. AAAA-A21-121011290027–6).
Author information
Authors and Affiliations
Contributions
EVB: conceptualization, project administration, methodology, resources, and writing–review and editing. MAT: conceptualization, writing–original draft preparation, resources, methodology, investigation, and visualization. AAE: software, visualization, and formal analysis. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Banaev, E.V., Tomoshevich, M.A. & Erst, A.A. The nuclear DNA content, ploidy, and chromosome numbers in some species of Nitraria and associations with pollen characteristics. Genet Resour Crop Evol 71, 2843–2858 (2024). https://doi.org/10.1007/s10722-023-01811-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01811-5