Abstract
Ancient species (Triticum monococcum and Triticum dicoccum) of wheat represent a valuable genetic resource for breeding and genetic research. In this study, 81 selected genotypes and 7 commercial cultivars were evaluated with 11 simple sequence repeat (SSR) molecular markers. A total of 93 SSR alleles were detected, giving an average of 8.45 alleles per locus. Consequently, a total of 88 genotypes were assessed for their mean expected heterozygosity (He = 0.486), observed heterozygosity (Ho = 0.121), polymorphism information content (PIC = 0.68), and Shannon’s information index (I = 0.918). The clustering analysis separated the genotypes into five subclusters based on the genetic similarity coefficient. Analysis of molecular variance (AMOVA) was performed to evaluate five different clusters. The result of AMOVA was defined as genetic deviation from expectation for clusters (Fis = 0.877, Fit = 0.903, and Fst = 0.211). In Nei’s pairwise genetic identity, the highest and lowest were observed between P2-P5 populations (0.39) and P1-P2 populations (0.838). The wide variety of wheat lines can be used as a genetic resource in designing a wheat breeding program to develop new cultivars adapted to different geographic and climatic conditions and can also contribute to breeding programs around the world.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The wheat and closely related species are included in the Poaceae (Gramineae) family, which is the genus Triticum (Matsuoka 2011). Ancient wheats, including einkorn (T. monoccocum L.) and emmer wheat (T. dicoccum L.), were discovered for the first time in the Cayonu excavation dating back to 6500–7000 B.C. in Türkiye (Harlan 1998). These species are characterized by hard husks remaining with the grain after threshing.
As with all living species, the evolutionary continuity of plant species is dependent on their ability to adapt to changing environmental conditions. Understanding the genetic diversity of the analyzed plant species is crucial for sustainable agriculture (Sevindik and Efe 2021). It can be possible to develop cultivars that are tolerant and resistant to harsh environmental conditions such as drought with biotechnological applications (Munaweera 2022). The genetic diversity of wheat species is also used in today’s wheat breeding studies (Zhang et al. 2010; Gurcan et al. 2017). Successful breeding programs are closely related to the extent and diversity of genetic material. The genetic diversity of wheat serves as the foundation for a breeding program designed to increase wheat yield. Wheat breeding through hybridization necessitates the selection of multiple genotypes regardless of whether the crop is a pure or hybrid variety (Zeb et al. 2009; Kumar et al. 2016a, b). Genetic characteristics are considered selection criteria in wheat breeding. Molecular markers are used in many areas, such as the preparation of detailed physical and genetic chromosome maps in plant organisms, selection of the desired characteristics in plants, increasing the success of classical breeding studies, determining the characteristics of gene sources in plants, genetic studies, determination of transgenic plants (Gupta et al. 2002; Collard and Mackill 2008; Sönmezoglu and Terzi 2018). Understanding the genetic diversity contained in a germplasm collection will have a substantial effect on wheat breeding. Characterization of the germplasm is regarded as a precondition for breeding since it provides new knowledge that may be used for future breeding actions (Demirel 2020). Molecular markers may give opportunities to discover exact genetic diversity across various farmed and wild wheat species that have varying amounts of ploidy (Gurcan et al. 2021). These opportunities can be found in different wheat species.
The molecular markers are useful for the assessment of genetic diversity such as DAMD (Directed Amplification of Minisatellites DNA), RAPD (Randomly Amplified Polymorphic DNA), ISSR (Inter Simple Sequence Repeats), SSR (Simple Sequence Repeats), IPBS (Inter Primer Binding Site) (Gurcan et al. 2017; Pinar et al. 2019; Demirel 2020; Yildiz et al. 2021a; Karakaya et al. 2023). Molecular markers provide various advantages, including greater information acquisition, independence from the environment, and stability (Bulunuz Palaz et al. 2023). Simple sequence repeats (SSRs) are extensively utilized in wheat due to their high level of polymorphism, uniform dispersion, and codominant inheritance in the wheat genome (Salem et al. 2015; Uzun et al. 2022). In higher organisms, there are randomly repeated DNA regions whose functions are not yet known but that are thought to have regulatory roles (Rafalski and Tingey 1993). These repeats are called microsatellite and minisatellite according to the number of nucleotides. Microsatellites are referred to as SSR (simple sequence repeats) or STR (short tandem repeats) (Liu 1997). The most common among microsatellites are dinucleotide repeats (AT)n consisting of repeats of 1–6 bp. Repeats such as (AAG)n and (AAT)n are very common in plants (Ellegren et al. 1997). SSRs are more plenty, ubiquitous in presence, inherently hypervariable, and have a high polymorphic information content (PIC) (Gupta et al. 2010).
Using SSR markers, the purpose of this study was to identify the diversity and population structure among wheat lines that may be employed as parents in breeding and hybridization research.
Materials and methods
Plant materials
The investigation utilized 36 Einkorn (Triticum monococcum L.) wheat lines, 45 Emmer (Triticum dicoccum L.) wheat lines, and seven registered wheat cultivars (Table 1). Plantings of wheat populations that had been gathered from Türkiye’s Kars, Kayseri, and Kastamonu regions were carried out in the Iğdır region. For the purposes of breeding, wheat lines were developed from individual spikes that were chosen from populations of wheat that exhibited a variety of traits. Within the parameters of this research, the wheat lines that were obtained were analyzed to assess the genetic variation. In addition, some Einkorn and Emmer wheat spikes are presented in Suppl. Fig. S1.
DNA extraction and SSR marker analysis
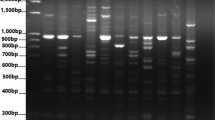
Under the circumstances of a greenhouse, the wheat seeds of each line were planted into individual pots. The newly acquired young leaves were used in an isolation procedure for DNA. The extraction of genomic DNA was accomplished by adhering to the CTAB procedure (Doyle and Doyle 1990) with a few adjustments (Aydin et al. 2018). For determining amplification and polymorphism, the PCR methodology that was established by Celik and Aydin (2023) was used. In this investigation, a total of 11 SSR (Table 2) primer pairs were utilized to determine the level of molecular diversity and population structure (Gurcan et al. 2017; Yildiz et al. 2021b). Amplified PCR (polymerase chain reaction) products were electrophoresed on a 2.5% (w v-1) agarose gel in a Tris-borate-EDTA (TBE) buffer for 120 min; ethidium bromide was used to stain the gel after electrophoresis; and an Imager Gel Doc XR + system (Bio-Rad, United States) was used to observe and photograph the results (Fig. 1). As a molecular size marker, the GeneRuler 100 bp DNA Ladder was utilized (ThermoFischer Scientific Waltham).
Data analysis
For molecular analysis, a genetic similarity and clustering matrix based on the quantity of shared alleles were generated, and the observed heterozygosity (Ho), expected heterozygosity (He), and polymorphism information content (PIC) were computed by the PowerMarker V3.025 software (Liu and Muse 2005). Shannon information index (I) for 11 SSR markers was calculated using the GENALEX V6.5 program (Peakall and Smouse 2006). Population structure of genotypes and the optimal value of ΔK were generated by STRUCTURE software and Structure Harvester (Pritchard et al. 2000; Earl and vonHoldt 2012). For AMOVA and population information (N: sample number, Na: number of different alleles, Ne: number of effective alleles, I: Shannon’s information index, Ho: observed heterozygosity, He: expected heterozygosity, uHe: unbiased expected heterozygosity, and F: fixation index), the GENALEX program was used. In addition, Nei’s genetic identity and distance matrices among subpopulations were created. UPGMA cluster analysis was utilized to assess patterns of diversity among the wheat entries based on the genetic similarity matrix generated by MEGA7 software (Kumar et al. 2016a, b).
Result and discussion
Molecular characterization
The results from 93 polymorphic bands scored using 11 SSR primers containing divergent linkage groups (LG) were utilized to carry out molecular characterisation and estimate the basics marker characteristics, described in Table 2. The number of alleles found in each marker varied anywhere from 5 to 15, with 8.45 being the mean value. The marker WMC177 provided the greatest number of alleles, which was 15, while the Xgwm135 and Xgwm312 tests produced the fewest number of alleles, each of which was also 5. This finding is much lower than the number that was reported in several earlier research (Teklu et al. 2006; Gurcan et al. 2017), but it is significantly higher than the number that was recorded in a number of earlier studies (Eujay et al. 2002; Salunkhe et al. 2013; Asmamaw et al. 2019). The value of observed heterozygosity (Ho) ranged from 0.037 for marker Xgwm570 to 0.412 for Xgwm135, with a mean of 0.121. The gene diversity (He) of markers ranged from 0.288 to 0.644, with marker Xgwm312 having the lowest score and marker Xgwm427 having the highest score, with the mean He score being 0.486. The Shannon information index (I) of markers ranged from 0.5 to 1.312, with marker Xgwm312 having the lowest score and marker Xgwm427 having the highest score, with the mean I score being 0.918. The average I value determined in this study was lower than the previous study conducted using SSR marker systems by Peng et al. (2009); however, it was higher than the results of Bhandawat et al. (2020). According to the standards established by Vaiman et al. (1994), PIC values were categorized as follows: those with a value of more than 0.5 were considered to have a high value, those with a value ranging from 0.5 to 0.25 were considered to have a medium value, and those with a value of less than 0.25 were considered to have a low value. The PIC values of the loci varied from 0.52 (Xgwm312) all the way up to 0.862 (WMC177), with a mean value of 0.68. The current investigation found that the PIC values for all 11 SSR markers were quite high. In a prior investigation of wheat genotypes using SSR markers, the researchers (Gurcan et al. 2017; Asmamaw et al. 2019) found the same conclusions related to PIC. Previous research has shown that when the PIC is more than 0.5, the marker has the greatest variety, which indicates a significant allelic diversity among germplasms. On the other hand, when PIC is less than 0.25, the marker has the least amount of variety (Kamara et al. 2020; Yildiz et al. 2021b). Considering these findings, which have an average PIC value of 0.68, it may be deduced that there is an adequate level of genetic heterogeneity and variation among the genotypes.
Because of this, the WMC177 marker located in linkage group (LG) 2 A was discovered to be the most polymorphic marker. This means that it has the potential to achieve discriminating ability for more marker-assisted selection studies as well as genetic diversity research in wheat breeding.
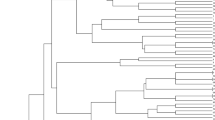
The UPGMA algorithm was used so that the dendrogram could be constructed. In Fig. 2, we presented an optimal tree that had the sum. With the use of the Maximum Composite Likelihood approach, the distances between genotypes were calculated (Tamura et al. 2004). Population genetic structure was also examined among 88 genotypes. The K value of this analysis was determined to be K = 5, implying the existence of five populations (P1, P2, P3, P4, and P5) among the wheat gene pool. The design of populations was visualized in a graph (Fig. 3). The genotypes for which population structure was determined in Fig. 3 were shown with the same colors in the dendrogram (Fig. 2).
The 88 genotypes were divided into two main groups (A and B). Individually from other genotypes, the P4 population’s genotypes were clustered into a singular group (A). The A group consisted exclusively of T. dicoccum L. species. The B group split into two subgroups, C and D. Registered cultivars (except G82) were clustered into the C group in the dendrogram. In addition, they were within P2 in Fig. 3. All the P5 (Fig. 3) genotypes in group D (Fig. 2) were T. monococcum L. species. In general, 11 SSR markers divided different species of wheat into separate clusters. The present results showed similarity with the reports of Gurcan et al. (2017). Therefore, the classification of wheats into various clusters according to their species suggested that there may be evidence of the evolution of diverse progenitors from ancient wheats to cultivated wheats.
Analysis of molecular variance
Analysis of molecular variance (AMOVA) was executed using data from wheat genotypes and STRUCTURE results. AMOVA demonstrated a lower proportion of variety among populations (21%) and within individuals (10%) as compared to variation among individuals within populations (69%) (Table 3). These results agreed with reports from previous studies. They also noted that in their research they employed wheat genotypes that had a large amount of diversity (Ramya et al. 2015; Soriano et al. 2016; Kabbaj et al. 2017). The values of Fit (Individual within the total population), Fis (Individual within the subpopulation) and Fst (Subpopulation within the total population) were 0.903, 0.877, and 0.211 for significance at P < 0.001, respectively. These results were higher than the reported values of Asmamaw et al. (2019). Wright’s F-statistics provide a unified perspective of genetic variation at three hierarchical levels of population structure (Conner and Hartl 2004). As a result, all loci exhibited a high level of inbreeding (0.877), which caused heterozygosity. Fst, a measure of population differentiation due to genetic differentiation, was moderately high (0.211). It could be due to high levels of variation among individuals (Chao et al. 2007).
Table 4 presents the results of the reporting of genetic parameters of populations obtained by STRUCTURE analysis. The number of different alleles (Na) was most observed in the P3 population with 5.364 and least observed in the P2 population with 2.455. The population with the highest observed number of effective alleles (Ne) was P3, which had 3.246, whereas the population with the lowest recorded number of effective alleles was P4, which had 1.32. The highest value and the lowest value of Shannon’s information index (I) were 1.246 for the P3 population and 0.37 for the P4 population, respectively. The mean of Shannon’s information index was 0.918 for all the populations. The population with the greatest value of observed heterozygosity (Ho) was the P1 population, which had a value of 0.217. The population with the lowest value was the P4 population, which had a value of 0.048. The mean of observed heterozygosity values was 0.121 for all the populations. The highest value and the lowest value of expected heterozygosity (He) were 0.609 for the P3 population and 0.183 for the P4 population, respectively. The mean expected heterozygosity value was 0.486 for all the populations. The highest and lowest values of unbiased expected heterozygosity (uHe) for the P3 and P4 populations, respectively, were 0.631 and 0.19. The mean of unbiased expected heterozygosity values was 0.513 for all the populations. The highest value and the lowest value of fixation index (F) were 0.877 for the P3 population and 0.571 for the P4 population, respectively. The mean unbiased fixation index was 0.713 for all the populations.
In Nei’s pairwise genetic identity, the lowest genetic identity was observed between P2 and P5 populations (0.39). Crossing genotypes from distant clusters may result in wider genetic variability and more heterosis in wheat lines. On the other hand, the highest genetic identity was observed between P1 and P2 populations (0.838). This might be due to the low criteria for wheat lines in this study or the high variance exchange between selecting wheat genotypes. Also, the differences observed in the wheat genotypes could result at least partly from the combined impacts of mutation, genetic drift, selection, and migration (Asmamaw et al. 2019) (Table 5).
In wheat, molecular markers based on simple sequence repeat (SSR) have been the most widely used in recent years due to their wide distribution in the genome, codominance, high polymorphism and reproducibility, and ease of analysis (Gurcan et al. 2017). Several studies have shown that SSR markers are an effective tool for analyzing wheat germplasm collections’ genetic diversity and subpopulation organization (Salem et al. 2015; Soriano et al. 2016; Yildiz et al. 2021b).
Researching wheat’s genetic variety is very important if one wants to comprehend the genetic make-up of wheat (Gurcan et al. 2017). The results of this research may be useful for wheat breeders and anyone who are interested in the improvement of wheat’s genetic resources. As a result, the agricultural gene pool must be appraised according to the degree of genetic diversity (Yildiz et al. 2021b), and the varied genotypes ought to be exploited in a variety of breeding programs.
In hexaploid wheat, just a small portion of the T. monococcum genome is present. Therefore, in breeding tetraploid and hexaploid cultivars, exploiting genetic variation in the T. monococcum genome can be a valuable source for discovering new and extra characteristics (Wang et al. 2014). In this context, knowing the range of variation present in a germplasm collection will majorly affect wheat improvement.
It is critical that the techniques of crop improvement and development make advantage of the native varieties that are available (Pachauri et al. 2017). According to a study by Demirel and Eren (2020), diploid and tetraploid types of wheat are still grown in Türkiye. Few farmers still grow these species in some limited areas, as they adapt better to the local environment than commercial hybrids (Gurcan et al. 2017). The fact that these species are still being cultivated is hopeful for the sustainability of genetic resources.
Conclusion
The wheat genotypes used in this study showed a high level of polymorphism. This is promising in terms of sustaining diversity. These results may indicate the potential of the SSR marker technique to discriminate among the wheat genotypes. This is very useful in choosing the parental genotypes used in hybridization and selection breeding programs. The use of these markers in association mapping studies may yield findings related to some characters to be examined in wheat. This investigation was an important study that revealed relationships between ploidy groups of Triticum spp. genotypes using SSR markers. It was observed that the genotypes were grouped clearly according to their ploidy levels. This study can contribute to wheat breeding programs and other breeders worldwide interested in wheat genetic resources.
Data availability
All data needed to conduct this study is provided within the manuscript. Sample spike pictures for einkorn wheat and emmer wheat are included in the supplement.
References
Asmamaw M, Keneni G, Tesfaye K (2019) Genetic diversity of ethiopian durum wheat (Triticum durum Desf.) Landrace collections as reveled by SSR markers. Adv Crop Sci Technol 7:1–9. https://doi.org/10.4172/2329-8863.1000413
Aydin A, Ince AG, Uygur Gocer E, Karaca M (2018) Single cotton seed DNA extraction without the use of enzymes and liquid nitrogen. Fresenius Env Bulletin 27:6722–6726
Bhandawat A, Sharma H, Pundir N, Madhawan A, Roy J (2020) Genome-wide identification and characterization of novel non-coding RNA-derived SSRs in wheat. Mol Biol Rep 47:6111–6125. https://doi.org/10.1007/s11033-020-05687-x
Bulunuz Palaz E, Demirel F, Adali S, Demirel S, Yilmaz A (2023) Genetic relationships of salep orchid species and gene flow among Serapias vomeracea× anacamptis morio hybrids. Plant Biotechnol Report 17:315–327. https://doi.org/10.1007/s11816-022-00782-w
Celik I, Aydin A (2023) SSR mining of black cumin (Nigella sativa L.) transcriptome for molecular marker development. Genet Res Crop Evol 70:629–638. https://doi.org/10.1007/s10722-022-01451-1
Chao S, Zhang W, Dubcovsky J, Sorrells M (2007) Evaluation of genetic diversity and genome-wide linkage disequilibrium among US wheat (Triticum aestivum L.) germplasm representing different market classes. Crop Sci 47:1018–1030. https://doi.org/10.2135/cropsci2006.06.0434
Collard BC, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philosophical Trans Royal Society B Biol Sci 363:557–572. https://doi.org/10.1098/rstb.2007.2170
Demirel F (2020) Genetic diversity of emmer wheats using iPBS markers. Europ J Sci Technolo 1:640–646. https://doi.org/10.31590/ejosat.814537
Demirel F, Eren B (2020) Production projection of einkorn and emmer wheat cultivated in Turkey. J Agri 3:1–5. https://doi.org/10.46876/ja.716882
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:39–40
Earl DA, vonHoldt BM (2012) Structure Harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv Genet Res 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Ellegren H, Moore S, Robinson N, Byrne K, Ward W, Sheldon BC (1997) Microsatellite evolution–a reciprocal study of repeat lengths at homologous loci in cattle and sheep. Mol Biol Evol 14:854–860. https://doi.org/10.1093/oxfordjournals.molbev.a025826
Eujayl I, Sorrells ME, Baum M, Wolters P, Powell W (2002) Isolation of EST-derived microsatellite markers for genotyping the A and B genomes of wheat. Theor Appl Genet 104:399–407. https://doi.org/10.1007/s001220100738
Gupta PK, Varshney RK, Prasad M (2002) Molecular markers: principles and methodology. In: Jain SM, Brar DS, Ahloowalia BS (eds) Molecular techniques in crop improvement. Springer, Dordrecht, pp 9–54
Gupta PK, Langridge P, Mir RR (2010) Marker-assisted wheat breeding: present status and future possibilities. Mol Breed 26:145–161. https://doi.org/10.1007/s11032-009-9359-7
Gurcan K, Demirel F, Tekin M, Demirel S, Akar T (2017) Molecular and agro-morphological characterization of ancient wheat landraces of turkey. BMC Plant Biol 17:171. https://doi.org/10.1186/s12870-017-1133-0
Harlan JR (1998) The living fields: our agricultural heritage. Cambridge University Press, UK
Hosking LK, Boyd PR, Xu CF, Nissum M, Cantone K, Purvis IJ, Khakhar R, Barnes MR, Liberwirth U, Hagen Mann K, Ehm MG, Riley JH (2002) Linkage disequilibrium mapping identifies a 390 kb region associated with CYP2D6 poor drug metabolising activity. Pharmacogenomics J 2:165–175. https://doi.org/10.1038/sj.tpj.6500096
Kabbaj H, Sall AT, Al-Abdallat A, Geleta M, Amri A, Filali-Maltouf A, Belkadi B, Ortiz R, Bassi FM (2017) Genetic diversity within a global panel of durum wheat (Triticum durum) landraces and modern germplasm reveals the history of alleles exchange. Fronti Plant Sci 8:1277. https://doi.org/10.3389/fpls.2017.01277
Kamara MM, Rehan M, Ibrahim KM, Alsohim AS, Elsharkawy MM, Kheir AM, Hafez EM, El-Esawi MA (2020) Genetic diversity and combining ability of white maize inbred lines under different plant densities. Plants 9:1140. https://doi.org/10.3390/plants9091140
Karakaya O, Yaman M, Balta F, Yilmaz M, Balta MF (2023) Assessment of genetic diversity revealed by morphological traits and ISSR markers in hazelnut germplasm (Corylus avellana L.) from Eastern Black Sea Region, Turkey. Genet Res Crop Evol 70:525–537. https://doi.org/10.1007/s10722-022-01444-0
Kumar S, Stecher G, Tamura K (2016a) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kumar P, Yadava RK, Kumar S, Kumar P (2016b) Molecular diversity analysis in wheat genotypes using SSR markers. Elect J Plant Breed 7:464–468. http://dx.doi.org/10.5958/0975-928X.2016b.00060.0
Liu BH (1997) Statistical genomics: linkage, mapping and QTL Analysis. CRS Press, Boca Raton, pp 10–24
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Matsuoka Y (2011) Evaluation of polyploid Triticum wheats under cultivation: the role of domestication, natural hybridization and allopolyploid specification in their diversification. Plant Cell Physiol 52:750–764. https://doi.org/10.1093/pcp/pcr018
Munaweera TIK, Jayawardana NU, Rajaratnam R, Dissanayake N (2022) Modern plant biotechnology as a strategy in addressing climate change and attaining food security. Agr Food Security 11:1–28. https://doi.org/10.1186/s40066-022-00369-2
Pachauri A, Sarawgi AK, Bhandarkar S, Ojha GC (2017) Genetic variability and association study for yield contributing traits of promising core rice germplasm accessions (Oryza sativa L). Res Crops 18:133–138. https://doi.org/10.5958/2348-7542.2017.00023.7
Peakall ROD, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Peng JH, Bai Y, Haley SD, Lapitan NLV (2009) Microsatellite-based molecular diversity of bread wheat germplasm and association mapping of wheat resistance to the russian wheat aphid. Genetica 135:95–122. https://doi.org/10.1007/s10709-008-9262-x
Pinar H, Uzun A, Unlu M, Yaman M (2019) Genetic diversity in turkish banana (Musa cavendishii) genotypes with DAMD markers. Fresen Env Bulletin 28:459–463
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Rafalski JA, Tingey SV (1993) Genetic diagnostics in plant breeding: RAPDs, microsatellites and machines. Trend Genet 9:275–280. https://doi.org/10.1016/0168-9525(93)90013-8
Ramya P, Jain N, Singh GP, Singh PK, Prabhu KV (2015) Population structure, molecular and physiological characterisation of elite wheat varieties used as parents in drought and heat stress breeding in India. Ind J Genet Pl Breed 75:250–252. https://doi.org/10.5958/0975-6906.2015.00038.3
Salem KF, Röder MS, Börner A (2015) Assessing genetic diversity of egyptian hexaploid wheat (Triticum aestivum L.) using microsatellite markers. Genet Res Crop Evol 62:377–385. https://doi.org/10.1007/s10722-014-0159-5
Salunkhe A, Tamhankar S, Tetali S, Zaharieva M, Bonnett D, Trethowan R, Misra S (2013) Molecular genetic diversity analysis in emmer wheat (Triticum dicoccon Schrank) from India. Genet Res Crop Evol 60:165–174. https://doi.org/10.1007/s10722-012-9823-9
Sevindik E, Efe F (2021) Molecular genetic diversity and phylogenetic analyses of Punica granatum L. populations revealed by ISSR markers and chloroplast (cpDNA) trnL-F region. Erwerbs-Obstbau 63:339–345. https://doi.org/10.1007/s10341-021-00581-7
Sönmezoglu ÖA, Terzi B (2018) Characterization of some bread wheat genotypes using molecular markers for drought tolerance. Physiol Mol Biol Plant 24:159–166. https://doi.org/10.1007/s12298-017-0492-1
Soriano JM, Villegas D, Aranzana MJ, García del Moral LF, Royo C (2016) Genetic structure of modern durum wheat cultivars and Mediterranean landraces matches with their agronomic performance. PLoS ONE 11:e0160983. https://doi.org/10.1371/journal.pone.0160983
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceed Nation Acad Sci 101:11030–11035. https://doi.org/10.1073/pnas.0404206101
Teklu Y, Hammer K, Huang XQ, Röder MS (2006) Analysis of microsatellite diversity in Ethiopian tetraploid wheat landraces. Genet Res Crop Evol 53:1115–1126. https://doi.org/10.1007/s10722-005-1146-7
Uzun A, Pinar H, Yaman M, Yigit MA, Cakiroglu Y, Karakaya A, Uysal M, Ozturk G, Yilmaz KU, Gurcan K, Ercisli S (2022) Identification of genetic diversity in wild pear (Pyrus elaeagrifolia Pall.) Genotypes collected from different regions of turkey with SSR marker system. Genetika 54:109–118. https://doi.org/10.2298/GENSR2201109U
Vaiman D, Mercier D, Moazami-Goudarzi K, Eggen A, Ciampolini R, Lépingle A, Velmala R, Kaukinen J, Varvio SL, Martin P, Levéziel H, Guérin G (1994) A set of 99 cattle microsatellites: characterization, synteny mapping, and polymorphism. Mamm Genome 5:288–297. https://doi.org/10.1007/BF00389543
Wang S, Wong D, Forrest K, Allen A, Chao S et al (2014) Characterization of polyploid wheat genomic diversity using a high-density 90 000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796. https://doi.org/10.1111/pbi.12183
Yildiz E, Pinar H, Uzun A, Yaman M, Sumbul A, Ercisli S (2021a) Identification of genetic diversity among Juglans regia L. genotypes using molecular, morphological, and fatty acid data. Genet Res Crop Evol 68:1425–1437. https://doi.org/10.1007/s10722-020-01072-6
Yildiz A, Hajyzadeh M, Ozbek K, Nadeem MA, Hunter D (2021b) Molecular characterisation of the oldest domesticated turkish einkorn wheat landraces with simple sequence repeat (SSR) markers. Biotechnol Biotechnol Equip 35:1291–1300. https://doi.org/10.1080/13102818.2021.1970023
Zeb B, Khan I, Ali S, Bacha S, Mumtaz S, Swati Z (2009) Study on genetic diversity in pakistani wheat varieties using simple sequence repeat (SSR) markers. Afr J Biotech 8:4016–4019
Zhang D, Bai G, Zhu C, Yu J, Carver BF (2010) Genetic diversity, population structure, and linkage disequilibrium in US elite winter wheat. Plant Genome 3:117–127. https://doi.org/10.3835/plantgenome2010.03.0004
Funding
No funding was received to conduct this study.
Author information
Authors and Affiliations
Contributions
SD and FD conceptualized and established the methodology, while SD performed molecular characterization. FD analyzed the statistical data. The manuscript was written by SD and FD.
Corresponding author
Ethics declarations
Confict of interest
The author’s confirm that this article content has no confict of interest.
Consent to participate and Publish
The authors reviewed the manuscript and expressed a desire to publish this study.
Ethics approval
This article does not contain any animal or human participant investigations conducted by the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Demirel, S., Demirel, F. Molecular identification and population structure of emmer and einkorn wheat lines with different ploidy levels using SSR markers. Genet Resour Crop Evol 71, 363–372 (2024). https://doi.org/10.1007/s10722-023-01627-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01627-3