Abstract
Common bean is one of the most important legume crops worldwide. Response to selection and success of hybridisation in common bean primarily depends on the nature and magnitude of genetic diversity present in the germplasm used. Germplasm comprising 4274 accessions originating from 58 countries were characterized for 22 phenotypic traits for two years. Genetic diversity for traits such as leaf length (4.5–20.7 cm), leaf width (3.4–17.5 cm), pod length (PL) (3.5–23.5 cm), no of pods/plant (4.2–59.6), seeds/pod (2.1–9.6) and 100-seed weight (SWT) (3.5–96.3 g) was observed in the accessions. Based on multivariate analysis, the entire collection was grouped into 10 genetically diverse clusters irrespective of the origin or place of collection of accessions. First three components obtained through principal component analysis explained 80.44 % of the total variance and it was contributed mainly by PL, pod width (PW), seed length (SL), seed width (SW), pods/plant and SWT. Correlation coefficient of seed weight was positively significant with leaf length, PL, PW, SL and SW while it was negatively correlated with days to flowering, pods/plant and seeds/pod. Regression analysis showed highest direct effect of SW on seed weight followed by SL, and PL. For bean anthracnose, >600 accessions showed resistance under field conditions, however when subjected to screening under artificial conditions against four most prevalent races (03, 515, 598 and 529) of Colletotrichum lindemutianum, we identified 16 accessions which have complete resistance and good agronomic superiority. These accessions may serve as useful genetic material to plant breeders for breeding bean varieties for anthracnose resistance and high yield.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common bean (Phaseolus vulgaris L.) is the most widely cultivated and consumed legume of the genus Phaseolus throughout the world (FAO 2012). It is a non-centric crop (Harlan 1971) having two major eco-geographical gene pools-Mesoamerica and the Andes with multiple domestications (Kwak and Gepts 2009). Characterization of both gene pools based on morphology (Singh et al. 1991b; Madakbas and Ergin 2011; Meza et al. 2013; Stoilova et al. 2013), protein types (Gepts et al. 1986), allozymes (Koenig and Gepts 1989), and molecular markers (Rossi et al. 2009; Kwak and Gepts 2009; Khaidizar et al. 2012; La Fuente et al. 2013) has given an overall indication of occurrence of at least two independent domestication events in the two different hemispheres (Debouck and Tohme 1989; Bitocchi et al. 2012). Two large gene pools that pre-existed domestications have large genetic variability in the wild and cultivated common beans (Rosales-Serna et al. 2005; Chacón et al. 2005; Bitocchi et al. 2013).
Little is known about the dissemination of common bean in India especially in the Himalayan region where major genetic diversity exists (Joshi and Rana 1995; Sofi et al. 2014). However, there are strong indications that many Portuguese, English, Dutch, French who travelled to the Indian subcontinent for trading in early part of the 16th century via Red and Arabian Sea; and Chinese through Hindustan Silk Route must have brought nicely colored and easily transportable beans with them (Simmonds 1976; Joshi and Thomas 1987; Joshi and Mehra 1993; Anononymus 2014). The genetic diversity present in India resembles European and Chinese beans, which depicts the combination of both Mesoamerican and Andean cultivated gene pools (Gepts et al. 1986, Debouck and Smartt 1995; Beebe et al. 2001; Logozzo et al. 2007; Angioi et al. 2010; Akbulut et al. 2013). Wide geographic diffusion among different environments and divergent selection criteria for agronomic and organoleptic qualities adopted by the farmers have further enhanced the range of genetic diversity not only in India but different parts of the world (Rana et al. 2010; Kumar et al. 2009; Singh 1989; Santalla et al. 2002; Meza et al. 2013). Presently, principal cultivated beans in India are dark and light red kidney, white kidney and cranberry beans with seed weight >40 g from Andean gene pool and medium and small-seeded (<40 g) with wide range of colours including white, cream, pinto, navy, yellow, black from Mesoamerican gene pool (Fig. 1).
In India, dry beans are grown in 9,100,000 ha with production of 3,630,000 tons and yield 399.0 kg/ha as compared to World’s area 29,234,228 ha with production 23,139,004 tons and yield 792 kg/ha (FAOSTAT 2013). Here, it is popularly called as rajmash and mainly produced under rainfed conditions in the traditional production system that include rotation with vegetables and intercropping of climbing beans with grain amaranth and maize during rainy season in the mountains while as sole crop of bush types during winters in the Indian plains (Santalla et al. 2004; Sharma and Rana 2005; Tiwari et al. 2005; Sharma et al. 2006; Rana et al. 2012). Common bean is an important source of minerals, i.e., iron and zinc, and many vitamins (Limongelli et al. 1996; Kaur et al. 2009; Singh Narpinder et al. 2010; Khetan et al. 2015), therefore, considered an economically, nutritionally, and socially important crop (Broughton et al.2003; Pujola´ et al. 2007; Mishra et al. 2010; Singh et al. 2012; Parmar et al 2014). In local gastronomy, it is associated with several cultural and culinary traditions (most famous kidney bean and rice based recipe in India is called rajmah-chawal), usually based on the use of local seed production of landraces worldwide (Masi et al. 1999; Polegri and Negri 2010; Negri 2012, Rana et al. 2012; Mercati et al. 2013). The landraces grown in the high mountain regions are highly appreciated and fetch up 2–3 times higher market price than those grown in the plains (Paudel et al. 2003; Rai et al. 2010, Rana et al. 2011; Bist and Rana 2012; Bisht and Rana 2012). Most famous landraces grown in the north-west part of India are known as ‘Chamba rajmah’ ‘Barot’ ‘Kinnauree’ ‘Auli’ ‘Munsiyari’ ‘Harshil’ ‘Bhaderwah’ and ‘Kashmiri’. These landraces have been named on the basis of geographical region in which they are grown. High priority has been accorded to protect these landraces, which are differentiated by morphological, agronomic and nutritional traits and highly adapted to local micro-climatic conditions of their limited geographical areas. Largely, farmers in the traditional production systems grow as many as 10–12 landraces together (Fig. 2), but economic considerations and agronomic developments have led to changed emphasis on single-component varieties over multi component mixtures. As a result, displacement of traditional genetic diversity is now visible not only in the high cropping intensive areas but in the traditional kidney bean growing areas also (Rana et al. 2000; Sofi et al. 2014).
The genetic diversity, which has been collected from different parts of India and introduced from abroad, has been conserved in the national gene bank under long term storage at New Delhi and medium term storage (MTS) at Shimla. However, the value of germplasm conserved in the germplasm bank depends on the information generated through its characterisation and evaluation for different traits (Gepts 2006; Gonçalves et al. 2009; Blair et al. 2010). In this study, we undertook large-scale phenotypic characterization of full range of genetic diversity to select genetically diverse accessions from 4274 accessions conserved in the MTS. Several authors have emphasised that phenotypic and phenological information based on descriptors continues to be the first step for the assessment, description and classification of large germplasm collections to enhance their use in research (Tar’an et al. 2005; Bhullar et al. 2009; Freitas et al. 2010; Szilagyi et al. 2011; Raggi et al. 2013). This paper will give plant breeders an insight of the genetic diversity present in the common bean germplasm in India, which may facilitate their indents to gene bank for germplasm of their own choice.
Materials and methods
Plant material
The plant material used in this study comprised 4274 accessions, of which 2308 were indigenous collections collected from 18 different states primarily from the Indian Himalaya (1887 accession) and 1966 were exotic collections representing 57 countries of the world (Table 1). Majority of EC accessions represent the countries from Mesoamerican and Andean centres of domestication including Mexico, Colombia and Peru. Data on the germplasm were recorded during two vegetative cycles of 2011 and 2012 in the experimental fields at Regional Station of National Bureau of Plant Genetic Resources (NBPGR 2000) Shimla, Himachal Pradesh. This farm is located at 31°05′53.89″N and 77°09′34.92″E and elevation of 1924 m above sea level. The soil of the experimental farm is sandy loam inclined more towards skeletal nature, dry, shallow and moderately rich in organic matter. Average annual rainfall is 1250 mm, of which around 60–65 % is received during the months from July to September. Mean maximum and minimum temperature ranges between 33.3 and −3.1 °C. The crop was sown in the month of June in each year in 2 rows of 2 m length with 30 cm inter-row spacing for determinate type and 75 cm for indeterminate types and harvested in October–November depending upon the maturity period of accessions. We followed standard cultivation practices recommended for growing common bean in the Indian hill region.
Plant characters and data recording
The data were recorded on 22 traits, of which 11 traits viz. days to flowering (DF), leaflet length (LL), leaflet width (LW), pod length (PL), pod width (PW), pods per plant (PPP), days to maturity (DM), seed length (SL), seed width (SW), seeds per pod (SPP), 100-seed weight (SWT) were measured quantitatively, while another 11 traits viz. early plant vigour, plant growth habit, stem pigmentation, flower colour, leaflet shape, pod colour, pod pubescence, pod shape, seed colour, seed mottling, and bean anthracnose score were assessed qualitatively on different scales. All the traits were measured/assessed based on P. vulgaris descriptors published by IBPGR (1982) on randomly selected five plants of each accession.
For further screening of field resistant lines under artificial conditions against anthracnose, germinated seed dip method of inoculation as described by Champion et al. (1973) was used for evaluation of resistance. Three seeds of each accession were surface sterilized with 0.1 % mercuric chloride followed by three washing with sterilized water were placed on double layers of seed germination paper covered by butter paper to retain the moisture and then kept at 25 ± 1 °C for germination in incubator. After 3 days, seed coats of germinated seeds were removed and dipped in standard spore suspension for 2–3 min. Thereafter seeds were sown in 3 cm deep plastic trays containing sterilized river sand. The trays were kept in growth chamber at 22 + 1 °C with >90 % humidity and 12 h photoperiod for 6 days. The disease reaction was recorded by using 0–5 point scale as described by Drijfhout and Davis (1989), where (0) no disease symptoms, (1) pinpoint lesions, (2) small lesions, (3) large sunken lesions, (4) large deep sunken lesions, (5) graded as susceptible. Plants showing 0, 1 and 2 or either of these were recorded as resistant (r) while those showing 3, 4 and 5 or either of these graded as susceptible (s). Each accession that showed resistance was tested twice to confirm the reaction.
Statistical analysis
We evaluated the germplasm in Augmented Block Design (Federer 1956) with six standard check varieties viz. Jawala, Baspa, Triloki, Kailash, Kentucky Wonder, and Laxmi in 12 blocks. The means were adjusted using online software for augmented block design developed by Rathore et al. 2004. Before undertaking statistical analysis on the basis of adjusted pooled mean values, homogeneity of variance was tested as per Levene (1960). Frequency distribution graphs were obtained for all the traits. The quantitative traits were further analysed for various statistical parameters viz. mean, range, variances, correlations, genetic diversity to find out genetic similarity/dissimilarity and principal component analysis (PCA) using the statistical software SYSTAT-12. Phenotypic and genotypic coefficients of variation (PCV and GCV) for each trait were computed as PCV = √VP/mean × 100, GCV = √VG/mean × 100 as per (Burton 1952) and categorized the range as per Sivasubramanian and Madhavamenon (1978). Broad sense heritability was estimated as h2 (bs) = VG/VP × 100 as per (Lush 1940) and further classified into low, medium and high (Robinson 1996). Expected as EGA = k × VG/VP × √VP as per Johnson et al. 1955. Here the standard value of k is 2.06 assumed at 5 % selection intensity; VG is genotypic variance; and VP is phenotypic variance. Genetic advance was expressed as % of mean as GA (%) = EGA/mean × 100. The significance of variances was tested at the 5 % probability level.
Results
Frequency distribution for qualitative traits
All the traits measured qualitatively showed wide range of variation among 4274 accessions evaluated (Fig. 3). Here in the text we have discussed only those traits and per cent accessions for which frequency of their occurrence was high while detail of all frequencies have been described in Figs. 3, 4. Early plant vigour, which determines the subsequent growth of plant, was good to medium for 98 % accessions. Indeterminate growth habit was observed for 43 % accessions, determinate for 28 and 29 % were intermediate type. The shape of leaflet was ovate for 83 % accessions, round for 14 % and ovate-lanceolate for others. The flower colour was white (39 %) followed by pink (26 %) and lilac (21 %). Green pod colour was predominant with different intensities of dull to shiny green for 87 % accessions. The surface of pods was glabrous for 96 % accessions while pod shape was straight for 69 % accessions. Wide range of seed colours such as white, red, maroon, pink, purple, yellow, black, brown along with varying tonalities was obtained, however, white (31 %) and red (29 %) were predominant. Around 66 % accessions have single seed coat colour while 34 % had mottled seed coats of various colours. We also undertook field screening of accessions against four most prevalent races viz. 03, 515, 529 and 598 of bean anthracnose (Colletotrichum lindemuthianum) causing serious crop loss to bean crop in the Himalayan region (Sharma et al. 2012). Out of 4274 accessions, 1287 were categorised as highly susceptible (scored 4) and, 1618 susceptible (3), 683 moderately resistant (2), 686 resistant (1) and 228 (0) as highly resistant to the disease anthracnose.
Frequency distributions graphs for quantitative traits depicted in the Fig. 4 showed that >70 % accessions fall in the medium group of flowering (46–65 days) and maturity periods (90–130 days). LL and LW revealed wide variation ranging from 4.5–20.7 to 3.4–17.5 cm, respectively. Around 80 % accessions had medium PL (10–16 cm) and PW (0.9–1.2 cm). High variability was recorded for number of pods/plant and it ranged from 10 to25 for 67 % accessions. The SL was 0.5–1.5 cm for 75 % accessions while SW was 0.6–1.0 cm for 74 % accessions. The most common numbers of seeds per pod were 4–8 and occurred in 83 % accessions. The seed weight was found highly variable trait and varied from 3.5 to 96.3 g/100 seed weight.
Mean, range, variance, coefficient of variance, heritability and genetic advance
Homogeneity of variance was tested using the Levene’s test showed p value 0.93, which was higher than the chosen alpha level (0.05) indicating that the variances of both the years were homogeneous allowing us to do subsequent statistical analysis. The statistical analysis of data on quantitative traits showed wide range of variability among the accessions (Table 2). The mean numbers of days to 50 % flowering were 55, but it ranged from 27 for accession IC370764 to 104 for EC500268 and EC500396. LL and width were highest for EC500242 and EC024955 with an average of 11.5 cm and 8.6 cm, respectively. PL was longest (22.5 cm) for IC328871 and EC271552 and shortest (3.4 cm) for EC531041 with an average value of 12.5 cm. EC500431 has widest pod (2 cm) while it was narrowest (0.4 cm) for IC321182. The average number of pods/plant was 16.3 while range was 4.2–49.6 for IC556535 and EC500299, respectively. PI301808 took longest time of 167 days for maturity as compared EC0944456, which matures in 71 days. IC383388 had maximum SL (2.03 cm) and width (1.2 cm) while IC383008 has maximum number of seeds/pod (10) compared to IC417359, which has minimum (2). The average SWT for all accessions was 29.5 g but it was highest for IC417355 (102.3 g) compared to HUR69 (3.6 g). PCV and GCV were high for pods/plant, SWT, SW, PW, SL and LW and low for rest of the traits. The heritability was found high for all the traits and it was ranging from 71.3 % for DM to 91.3 % for PL. The genetic advance expressed as % of mean was low (<35) for DM, DF, PL, and SPP and high (>35) for rest of the traits.
Correlation and regression
The matrix developed for correlation coefficients (Table 3) showed significant positive correlation of seed weight with LL, PL, PW, SL and SW and negative correlation with DF, seeds/pod and pods/plant. Seeds/pod had positive correlation with DF, leaf width, pods/plant and DM while negative correlation with SL. Other combinations of traits, which have strong positive correlations among themselves were leaf length & leaf width, DF & DM, PL & PW, PL & SL, PW & SW and SL & SW. Least square regression analysis performed for seed weight versus other traits showed that SW had highest direct effect on seed weight followed by PW, SL, PL and leaf length while seeds/pod, pods/plant and leaf width had negative direct effect. The plot of residual versus fitted values showed that residuals of majority of the accessions bounce randomly around 0 line forming horizontal band (data not shown). The variances of the error terms were equal, relationship among accessions was linear and there were only 74 outlier accessions out of 4274.
Genetic diversity and principal component analysis
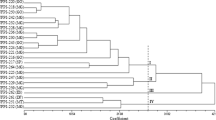
Based on the multivariate analysis performed on 11 quantitatively measured traits, all the 4274 accessions were grouped into 10 clusters and each cluster was found to have varied number of accessions (Table 4). However, it was not feasible to produce a dendrogram due to large number of accessions involved in the analysis. The number of accessions falls in each cluster were highest (562) in cluster 8 (C8) followed by C3 (555), C4 (537), C6 (517), C2 (436), C5 (424), C1 (405), C9 (346), C10 (268) and C7 (224). The mean value of accessions grouped into each cluster (Table 4) showed that accessions in C4 were early in flowering and maturity whereas all accessions of C3 had late maturing types. The accessions with high average number of pods/plant (22.9) were grouped into C8 while seed weight was high (51.7) for accessions grouped in C2, followed by C6 (42.1) and C7 (41.0). High mean (>13 cm) PL and SL (>1.4 cm) were observed for accessions grouped into C2, C4 and C7. Genetic distance measured through multivariate analysis between different clusters was highest in C3 & C4 followed by C3 & C7, C3 & C2, C3 & C8, C6 & C8, C3 & C1, C6 & C7, C1 & C8, C5 & C9, C1 & C4, C4 & C7, C2 & C10 and C7 & C10.
The PCA used to eliminate the redundancy in data set revealed that all the 11 quantitatively measured traits have been loaded on first five components, however, major portion of variance (80.4 %) in bean germplasm is explained by first three components (Fig. 5). The first component (PC1) accounted for 34.5 % of variation, through PL, PW, SL, SW and SWT; PC2 accounted for 27.5 % of variation loaded on leaf length, leaf width, pods/plant and seeds/pod and PC3 contributed 18.4 % variance through DF and DM. PC4 and PC5 shared 8.8 and 7.3 % variance, respectively, and loaded partially on PW, seeds/pod and SW.
Discussion
Common bean ranks high in agriculture worldwide because of its nutritious food components and good market price (Rodin˜o et al. 2006; Rana et al. 2010; Singh et al. 2012; Khetan et al. 2015). The analysis of genetic diversity in large collections conserved in the gene banks is important for deciphering nature and magnitude of variability, genetic relationship between traits for the efficient management and use of germplasm (Stoilova et al. 2005; 2013 Blair et al. 2010; Szilagyi et al. 2011). The simplified phenotypic approach has been found useful for preliminary characterisation and discrimination of accessions to understand the level of genetic diversity (Foschiani et al. 2009; Atilla et al. 2010; Szilagyi et al. 2011). It helps in detecting the occurrence of diversity within population and spelt out precise information on the nature and magnitude of genetic divergence among gene pools for reliable scoring during selection of potential parents for hybridisation (Szilagyi et al. 2011; Mercati et al. 2013; La Fuente et al. 2013).
This study describes one of the first large-scale phenotypic characterisation of the bean collection conserved in the Indian gene bank. The germplasm presented wide range of genetic variability for 22 traits among 4274 accessions. The frequency distribution obtained for 22 traits showed the presence of maximum possible range of variability reported in the Andean and Mesoamerican gene pools of common bean (Singh et al. 1991a; Bitocchi et al. 2012; La Fuente et al. 2013). Among different traits, seed traits have been found most important in common bean and major determinants of commercial acceptability of varieties (Park et al. 2000; Bisht et al. 2014; Rana et al. 2014). Seed traits have also been considered highly heritable traits, therefore important in breeding programmes (Singh et al. 2007; Blair et al. 2010). We found marked variation in seed colour ranging from a single colour viz. white, red, maroon, pink, purple, yellow, black, and brown to mottled seeds with varying tonalities. The preference for seed colour was found varied from place to place in India and it was kidney shaped red, maroon, pink, and yellow beans, which find premium place among bean consumer and producer (Sharma and Rana 2005; Rana et al. 2010). Similar observations on seed colour preference have been recorded in other parts of the world (Trutmann et al. 1996; Wortmann et al. 1998; Fonseca et al. 2007). This suggests that seed colour genes have perhaps been moved between gene pool groups through hybridization, inter-gene pool introgression and extensive selection for different preferences by producers and consumers worldwide. Apart from seed colour, variation found in seed shape and size was significantly wider (3.5–96.3 g/SWT) in the accessions. These results are consistent with findings of different scientists who have reported wide variation in seed shape and size in bean germplasm (Rodin˜o et al. 2003; Rai et al. 2006; Rodin˜o et al. 2006; Cabral et al. 2010; Lioi et al. 2012). The most elaborated work done by Singh (1989) and Singh et al. (1991a, b) have given high taxonomic value for seed size, colour and shape in comparison to the vegetative characteristics of plant in common bean.
Apart from seed traits, plant growth habit was found varied from bushy to aggressive climbing type. The predominance of one growth habit type is related to ecological adaptation as well as to the cropping system being followed. For instance, in the hill regions beans are generally intercropped with maize and amaranth for which climbing types are more desirable while in the plains beans are grown as sole crop, bush types are preferred. We observed that climbing bean generally have longer life cycle, higher productivity and late maturity while bush type have early maturity, short plant and low productivity. These observations are in agreement with studies conducted by García et al. (1997) who demonstrated that increasing climbing ability increase the time needed for achieving physiological maturity, and plants with indeterminate growth are more productive than plants with determinate growth habit. Piergiovanni and Lucia (2010) described that 90 % of Italian landraces cultivated in Basilicata region have climbing habit. It was found that flowering and maturity were also found to be influenced by environmental parameters viz. temperature and photoperiod length in both bush and climbing types. These results obtained here are in agreement with those obtained in bean germplasm from Iberian Peninsula (Rodin˜o et al. 2003), Nicaragua (Gómez et al. 2004), Mexico (García et al. 1997), Portugal (Coelho Rita et al. 2009) and Andean cultivars (Santalla et al. 2004).
The extent of genetic variation could also be better judged by the estimation of GCV in relation to PCV. Small difference between the two indicates that observed variation and expression of traits is mainly due to genetic factors while larger difference indicate role of environment. However, these parameters of variability alone may not be sufficient for determining of magnitude of heritable variation. High GCV coupled with high heritability and genetic gain is useful for predicting the result of selection based on phenotypic performance. In our germplasm, LL, LW, PL, PW, PPP, SL, SW and SWT showed high level of variance coupled with high heritability and genetic advance. This suggests that these traits may be under the influence of additive gene interactions and use of simple selection methods may bring significant improvement for these traits. Accessions such as IC328541, IC531148, IC326556, EC25506, HUR116, IC328871, IC199249, EC500431, EC500951, EC500299, IC14913 IC383388, NC67302, IC641138, PI1044643, IC417355, NC56079 and IC340902 showed high values for these traits and can be subjected for further testing and selection for yield gains through simple selection based on these traits. On the other hand flowering, maturity and seeds/pod, which have low variance, heritability and genetic advance showed the presence of non-additive gene interactions and needed to be improved by hybridization (Johnson and Gepts 2002; Raffi and Nath 2004; Kumar 2008; Ahmad and Kamaluddin 2011; Rai et al. 2010; Sharma et al. 2012). Accessions viz. IC338701, IC340928, IC328541, EC25501, EC398509, IC383008, IC417353 and IC340964 could be used in the hybridisation for improving on traits showing low heritability and genetic advance.
Correlation coefficient is important in plant breeding because it measures the degree of association (genetic and non-genetic) between two or more traits (Dewey and Lu 1959). In the presence of high correlation between two traits, selection for one trait will cause a change in its mean through additive gene effects of selected individuals and simultaneously cause an indirect change in the mean of the other trait (Dursun 2007; Apostolova et al. 2009; Kumar et al. 2009; Singh et al. 2011; Mudasir et al. 2012). The significant positive correlation of seed weight with LL, PL, PW, SL and SW showed that selection for any of these traits may favour improvement in other traits also, while negative correlation with DF, PPP and SPP may adversely affect the gain. The data on the accessions showed that small leaf and pod size invariably have smaller seeds while large seeded accessions had broader leaf and medium to long and wide pod. The positive association of seed weight with various other yield traits have been reported in bean germplasm (Rai et al. 2006; Blair et al. 2006; Dursun 2007; Bhushan et al. 2008; Apostolova et al. 2009; Krasu and OZ 2011; Kumar et al. 2009; Sofi et al. 2011; Ahmed and Kamaluddin 2013; Sofi et al. 2014). According to Blair et al. (2009), varieties of dry beans that do not meet minimum seed weight criteria can be rejected by the market place while in snap beans varieties with elongated seed shape or smaller seed size are preferred as they are correlated with pod quality. Based on the correlation coefficients, it is possible to construct/select a plant type integrating yield components such as LL, PL, seed weight, seed diameter into one, however, to combine few more traits like pods/plant, seed/pod and early maturity, common bean breeder has to go for extensive crossing followed by selection.
Grouping large number of germplasm accessions into few numbers of homogenous clusters facilitates the selection of diverse parents. It permits precise comparison among all the possible pair of populations and provide an opportunity for bringing together gene constellation yielding desirable progenies. The maximum and minimum difference observed between clusters III & IV and clusters I and VIII, respectively indicating genetic distance and closeness among accessions due to different genetic constitutions. Meaning thereby clusters that have higher mean value and genetically distant could facilitate the selection of accessions for use in the breeding programmes (Sharma et al. 2009; Rana et al. 2013; Meza et al. 2013; Stoilova et al. 2013). The varying levels of similarity and dissimilarity in common bean landraces have been observed in various other studies (Piergiovanni et al. 2000; Franklin et al. 2009; Biswas et al. 2010; Szilagyi et al. 2011; La Fuente et al. 2013). The present collection represents 58 countries of the world but while grouping accessions into various clusters we did not observe any favour to a specific geographical origin. The accessions were grouped into various clusters irrespective of the country/region from where these have been collected/procured. This suggests that due to the free exchange of seed materials among different regions and genetic constellations of character might be associated with particular region in nature, but lose their individuality under human interference (Ramanjulu et al. 1999; Singh et al. 2002; Adesoye and Ojobo 2012). Scientists have discussed genetic drift, selection pressure and environment as other major factors that could cause greater diversity than geographical distance (Sicard et al. 2005; Nehvi et al. 2007; Kwak and Gepts 2009; Angioi et al. 2010).
Similarity indices and pattern of relationships obtained through genetic diversity and PCA are useful to evaluate potential breeding value of germplasm through traits loaded on various components (Keneni et al. 2005). Using PCA, we were able to quantify the degree of divergence between populations to understand the trend of their evolutionary pattern and to assess the relative contribution of different components to the total divergence together with nature of forces operating at intra and inter-cluster levels (Sharma et al. 2009). In the present study, 34.55 % of the total variation was contributed by PL and width, SL and width and seed weight while 27.52 % by leaf length and width, pods/plant and seed/pod indicating that seed and pod traits have contributed highest genetic diversity in common bean germplasm. The results are in agreement with various other studies that reported the maximum contribution of seed weight, pods/plant and seed length towards genetic divergence in common bean (Park et al. 2000; Mirjana 2005; Ceolin et al. 2007; Misra et al. 2010).
Besides, understanding the nature and magnitude of genetic diversity obtained through different traits, it is desirable to indentify the accessions found resistance against most prevalent diseases and pests. Here we simultaneously screened the accession for bean anthracnose, which causes yield losses up to 100 % (Fernandez et al. 2000; Miklas et al. 2006; Sharma et al. 2007). The pathogen possess high degree of pathogenic variability throughout the world including India and >100 races of C. lindemuthianum have been identified worldwide (Rodriguez-Guerra et al. 2003; Gonzalez et al. 2004; Mahuku and Riascos 2004; Sharma et al. 2007). Its management can be done by using certified seed, crop rotation, seeds and foliar treatment with fungicides and genetically resistant varieties. Out of these most of strategies are not followed by marginal and resource poor farmers particularly those living in the mountains and grow bean as major pulse crop. Under such situation use of genetically resistant varieties offer most effective, least expensive and easy to adopt way to the farmer. In the present collection, >600 accessions showed resistance to bean anthracnose under field conditions. Out of >200 accessions indentified as highly resistant in the field 99 had been screened against 4 races viz., 03, 515, 529 and 598 of C. lindemuthianum under laboratory conditions by Sharma et al. (2012). Out of 99 accessions 65 showed resistance to one or other race but we selected 16 accessions viz., IC328537, IC328538, IC448888, IC313194, IC278723, IC339645, IC398530, EC169813, EC398530, EC500226 IC326964, IC370764, EC77006, EC50063, EC50079, and EC530926 which were found resistant against all the races and also showed agronomic superiority. We assumed that being resistant and agronomically superior there accessions could be a good breeding material for imparting anthracnose resistance to bean varieties along with high yield gains.
Conclusion
Progress in breeding high yielding varieties of common bean has been modest and one of the constraints regarded by many plant breeders is the narrow genetic base available within the breeders’ stock (Cooper et al. 1997; Kelly et al. 1998; Perseguini et al. 2011). Given this constraint, plant breeders need genetically diverse germplasm not only for yield but its contributing traits also. The results obtained based on various genetic parameters analysed on wide range of traits, especially seed and pod traits; it is concluded that determinant genetic effects of the phenotypic expression of these traits are fundamentally additive type. Therefore, a high response to selection should be achievable after several selection cycles. The identification of agronomically superior and anthracnose resistant accessions will be useful in minimising the linkage drag usually breeders come across while transferring disease resistance in already available high yielding but susceptible varieties. This collection represents a valuable genetic patrimony with great potential for the future and as such, germplasm from Himalayan region is an integral part of the secondary diversity of common beans. As a further step, the development of core sets and traits specific sub sets based on this data and subsequent analysis of diversity using more specific molecular markers are recommended to elucidate more information on the overall genetic diversity and particular genes responsible for specific characters of agronomic interest in the present collection.
References
Adesoye AI, Ojobo OA (2012) Genetic diversity assessment of Phaseolus vulgaris L. landraces in Nigeria’s mid-altitude agro ecological zone. Int J Biodivers Conserv 4(13):453–460
Ahmed S, Kamaluddin (2013) Correlation and path analysis for agro morphological traits in rajmash beans underBaramulla- Kashmir region. Afri J Agric Res 8(18):2027–2032
Akbulut B, Karakurt Y, Tonguc M (2013) Molecular characterization of common bean (Phaseolus vulgaris L.) genotypes. Akdeniz Univ J Fac Agric 26(2):105–108
Angioi SA, Rau D, Attene G, Nanni L, Bellucci E, Logozzo G, Negri V, SpagnolettiZeuli PL, Papa R (2010) Beans in Europe: origin and structure of the European landraces of Phaseolus vulgaris L. Theor Appl Genet 121:829–843
Anononymus (2014) Encyclopedia of India-China Cultural Contacts. MaXposure Media Group (I) Pvt. Ltd. 2nd floor Mira Corporate Suites, Ishwar Nagar, Mathura Road, New Delhi, 110065, India
Apostolova E, Dimova D, Svetleva E (2009) Correlation dependence between twenty one traits of some Bulgarian common bean genotypes. Acta Horti 830:191–197
Atilla D, Kamil H, Melek E (2010) Characterization of breeding lines of common bean as revealed by RAPD and relationship with morphological traits. Pak J Bot 4(6):3839–3845
Beebe S, Rengifo J, Gaitan E, Duque MC, Tohme J (2001) Diversity and origin of Andean landraces of common bean. Crop Sci 41:854–962
Bhullar NK, Street K, Mackay M, Yahiaoui N, Keller B (2009) Unlocking wheat genetic resources for the molecular identification of previously un described functional alleles at the Pm3 resistance locus. Proc Nat Acad Sci USA 106:9519–9524
Bhushan KB, Jadli S, Verma O, Goswami AK (2009) Plant characters correlation and path coefficient analysis of seed yield in exotic French bean (Phaseolus vulgaris L.) germplasm. Intern J Agric Sci 4(2):667–669
Bisht IS, Rana JC (2012) Best practices in rajmash. SLEM-CPP News lett 3(1):13–15
Bisht IS, Pandravada SR, Rana JC, Malik SK, Archna Singh, Singh PB, Firoz Ahmad, Bansal KC (2014) Subsistence farming, agro-biodiversity and sustainable agriculture: a case study. Journal of agro-ecology and sustainable food systems 38(8):890–912
Biswas MS, Hassan J, Hossain MM (2010) Assessment of genetic diversity in French bean (Phaseolus vulgaris L.) based on RAPD marker. Afr J Biotechnol 9(32):5073–5077
Bitocchi E, Nanni L, Bellucci E, Rossi M, Giardini A, Zeuli PS, Logozzo G, Stougaard J, Mc Clea P, Atten G, Papa R (2012) Mesoamerican origin of the common bean (Phaseolus vulgaris L.) is revealed by sequence data. Proc Natl. Acad Sci USA 09:E788–E796
Bitocchi E, Bellucci E, Giardin A, Rau D, Rodriguez M (2013) Molecular analysis of the parallel domestication of the common bean (Phaseolus vulgaris) in mesoamerica and the andes. New Phytol 197:300–313
Blair MW, Giraldo MC, Buendia HF, Tovar E, Duque MC, Beebe S (2006) Microsatellite marker diversity in common bean (Phaseolus vulgaris L.). Theor Appl Genet 113:100–109
Blair MW, Lucy MD, Hector FB, Myriam CD (2009) Genetic diversity, seed size associations and population structure of a core collection of common beans (Phaseolus vulgaris L.). Theor Appl Genet 119:955–972
Blair MW, Laura FG, Paul M, Louis B (2010) Genetic diversity, inter-gene pool introgression and nutritional quality of common beans (Phaseolus vulgaris L.) from Central Africa. Theor Appl Genet 121:237–248
Broughton WJ, Hernandez G, Blair M, Beebe S, Gepts P, Vanderleyden J (2003) Beans (Phaseolus spp)–model food legumes. Plant Soil 252:55–128
Burton GW (1952) Quantitative inheritance in grasses. Proc Int Grassl Cong 1:277–283
Cabral PDS, Taís CBS, Leandro SAG, Antonio TAJ, Andreia BPL, Rosana R, de Frederico PM (2010) Quantification of the diversity among common bean accessions using Ward-MLM strategy. Pesq Agropecu Brasília 45(10):1124–1132
Ceolin ACG, Vidigal MCG, Filho PSV, Kvitschal MV, Gonela A, Scapim CA (2007) Genetic divergence of the common bean (Phaseolus vlgaris L.) group carioca using morpho-agronomic traits by multivariate analysis. Hereditas 144:1–9
Chacón SMI, Pickersgill B, Debouck DG (2005) Domestication patterns in common bean (Phaseolus vulgaris L.) and the origin of the Mesoamerican and Andean cultivated races. Theor Appl Genet 110:432–444
Champion MR, Brunet D, Maudit ML, Lami RI (1973) Method of testing resistance of bean varieties to Colletotrichum lindemuthianum (Sac. and Magn) Briosi and Cav. Acad de Agric de Fr 59:951–958
Coelho Rita C, Miguel AF, Joana R, Reis Aida, Maria BPP, Oliveira Eugenia N (2009) Assessing genetic variability in germplasm of Phaseolus vulgaris L. collected in Northern Portugal. Sci Hort 122:333–338
Cooper M, Stucker RE, De Lacy IH, Harch BD (1997) Wheat breeding nurseries, target environments, and indirect selection for grain yield. Crop Sci 37:1168–1176
de La Fuente M, Gonza´lez A, de Ron AM, Ron S (2013) Patterns of genetic diversity in the Andean gene pool of common bean reveal a candidate domestication gene. Mol Breed 31:501–516
Debouck DG, Smartt J (1995) Beans. In: Smartt J, Simmonds NW (eds) Evolution of crop plants. Longman Scientific & Technical, Harlow, pp 287–296
Debouck DG, Tohme J (1989) Implications for bean breeders of studies on the origins of common beans, Phaseolus vulgaris L. In: Beebe S (ed) Current topics in breeding of common bean 47:3–42. Bean Program, Centro Internacional de Agricultura Tropical (CIAT), Cali
Dewey DR, Lu KH (1959) A correlation and path coefficient analysis of components of crested wheat grass seed production. Agron J 51:515–518
Drijfhout E, Davis JHC (1989) Selection of a new set of homogeneously reacting beans (Phaseolus vulgaris L.) differentials to differentiate races of Colletotrichum lindemuthianum. Plant Pathol 38:391–396
Dursun A (2007) Variability, heritability and correlation studies in bean (Phaseolus vulgaris L.) genotypes. World J Agrl Sci 3(1):12–16
FAO–Food and Agriculture Organization of the United Nations (2012) Food and agricultural commodities production. http://faostat.fao.org/site/339/default.aspx
Federer WT (1956) Augmented (or hoonuiaku) designs. Hawaii Plant Record 55:191–208
Fernandez MT, Fernandez M, Casares A, Rodr´ıguez R, Fueyo M (2000) Bean germplasm evaluation for anthracnose resistance and characterization of agronomic traits: a new physiological strain of C. lindemuthianum infecting Phaseolus vulgaris L. in Spain. Euphytica 114:143–149
Fonseca JR, Marques EMG, Vieira EHN, Silva HT (2007) Algumas características do germoplasma de feijão (Phaseolus vulgaris L.) coletado no Espírito Santo. Revista Ceres 54:358–362
Foschiani A, Miceli F, Vischi M (2009) Assessing diversity in common bean (Phaseolus vulgaris L.) accessions at phenotype and molecular level: a preliminary approach. Genet Resour Crop Evol 56:445–453
Franklin CJ, Thomas MMS, Varghese G, Selvaraj N, Darai M (2009) Genetic diversity and conservation of common bean (Phaseolus vulgaris L., Fabaceae) landraces in Nilgiris. Curr Sci 97(2):227–235
Freitas G, Jose F, Gananca T, Humberto N, Elvio N, Graca C, Jan J, Slaski Miguel AA, de Carvalho Pinheiro (2010) Morphological evaluation of common bean diversity on the Island of Madeira. Genet Resour Crop Evol 58:861–874
García EH, Pen˜a-Valdivia CB, Aguirre JRR (1997) Morphological and agronomic traits of a wild population and an improved cultivar of common bean (Phaseolus vulgaris L.). Ann Bot 79(2):207–213
Gepts P (2006) Plant genetic resources conservation and utilization: the accomplishments and future of a societal insurance policy. Crop Sci 46:2278–2292
Gepts P, Osborn TC, Rashka K, Bliss FA (1986) Phaseolin-protein variability in wild forms and landraces of the common bean (Phaseolus vulgaris): evidence for multiple centers of domestication. Econ Bot 40:451–468
Gómez OJ, Blair MW, Frankow-Lindberg BE, Gullberg U (2004) Molecular and phenotypic diversity of common bean landraces from Nicaragua. Crop Sci 44:1412–1418
Gonzalez CM, Guerrn RR, Hernandez-Godinez F, Acosta-GallegosJA Martinez, de la Vega O, Simpson J (2004) Analysis of pathotypes of Colletotrichum lindemuthianum found in central region of Mexico and resistance in elite germplasm of Phaseolus vulgaris. Plant Dis 88:152–156
Harlan JR (1971) Agricultural origins: centers and noncenters. Science 174:468–474
IBPGR (1982) Descriptors of Phaseolus vulgaris. IBPGR (now Bioversity International), Crop Genetic Resources Centre, Rome, p 37
Johnson WC, Gepts P (2002) The role of epistasis in controlling seed yield and other agronomic traits in an Andean × Mesoamerican cross of common bean (Phaseolus vulgaris L.). Euphytica 125:69–79
Johnson HW, Robinson HF, Comstock RE (1955) Estimates of genetic and environmental variability in Soybean. Agron J 47:314–318
Joshi BD, Mehra KL (1993) Adaptability in French bean. Indian J Pl Genet Resour 6(1):73–77
Joshi BD, Rana RS (1995) French bean in India. National Bureau of Plant Genetic Resources Regional station, Shimla
Joshi BD, Thomas TA (1987) Genetic resources in temperate grain legumes. In: Paroda RS, Arora RK, Chandel KPS (eds) Plant genetic resources-Indian perspectives. NBPGR, New Delhi, pp 255–260
Kamaluddin Ahmad S (2011) Variability, correlation and path analysis for seed yield and yield related traits in common beans. Indian J Hort 68(1):56–60
Kaur Seeratpreet, Narpinder S, Navdeep SS, Rana JC (2009) Diversity in properties of seed and flour of kidney bean germplasm. Food Chem 117(2):282–289
Kelly JD, Kolkman JM, Schneider K (1998) Breeding for yield in dry bean (Phaseolous vulgaris L.). Euphytica 102:343–356
Keneni G, Jarso M, Wolabu T, Dino G (2005) Extent and pattern of genetic diversity for morpho-agronomic traits in Ethiopian highland pulse landraces: 1. Field pea (Pisum sativum L.). Genet Res Crop Evol 52:539–549
Khaidizar MI, Haliloglu K, Elkoca E, Aydin M, Kantar F (2012) Genetic diversity of common bean (Phaseolus vulgaris L.) landraces grown in northeast Anatolia of Turkey assessed with simple sequence repeat markers. J Field Crops 17(2):145–150
Khetan S, Singh N, Kaur A, Rana JC (2015) Structural and functional characterization of kidney bean and field pea protein isolates: a comparative study. Food Hydrocoll 43:679–689
Koenig R, Gepts P (1989) Allozyme diversity in wild Phaseolus vulgaris: further evidence for two major centers of genetic diversity. Theor Appl Genet 78:809–817
Krasu A, OZ M (2011) A study on coefficient analysis and association between agronomical characters in dry bean (Phaseolus vulgaris L.). Bulg J Agric Res 16(2):203–211
Kumar K (2008) Variability, heritability and genetic advance in pea (Pisum sativum L.). Int J Plant Sci 3(1):211–212
Kumar A, Singh A, Singh P, Singh SB, Singh V (2009a) Relationship and path analysis for green pod yield and its contributing characters over environments in French bean (Phaseolus vulgaris L.). Legume Res 32(4):270–273
Kumar V, Sharma S, Sharma AK, Sharm S, Bhat KV (2009b) Comparative analysis of diversity based on morpho-agronomic traits and microsatellite markers in common bean. Euphytica 170:249–262
Kwak M, Gepts P (2009) Structure of genetic diversity in the two major gene pools of common bean (Phaseolus vulgaris L. Fabaceae). Theor Appl Genet 118:979–992
Levene H (1960) Robust testes for equality of variances. In: Olkin I (ed) Contributions to probability and statistics. Stanford University Press, Palo Alto, pp 278–292 MR0120709
Limongelli G, Laghetti G, Perrino P, Piergiovanni AR (1996) Variation of seed storage proteins in landraces of common bean (Phaseolus vulgaris L.) from Basilicata, southern Italy. Euphytica 92:393–399
Lioi L, Alberto N, Bruno C, Angela RP (2012) Assessment of genetic variation in common bean (Phaseolus vulgaris L.) from Nebrodi mountains (Sicily, Italy). Genet Resour Crop Evol 59:455–464
Logozzo G, Rosa D, Leonardo M, Roberto P, Helmut K, Pierluigi SZ (2007) Analysis of the contribution of Mesoamerican and Andean gene pools to European common bean (Phaseolus vulgaris L.) germplasm and strategies to establish a core collection. Genet Resour Crop Evol 54:1763–1779
Lush JL (1940) Intrusive collection of regression of offspring on dams as a method of estimating heritability of characters. Proc Am Soc Anim Prod 33:293–301
Madakbas SY, Ergin M (2011) Morphological and phenological characterization of Turkish bean (Phaseolus vulgaris L.) genotypes and their present variation states. Afr J Agric Res 6(28):6155–6166
Mahuku GS, Riascos JJ (2004) Virulence and molecular diversity within Colletotrichum lindemuthianum isolates from Andean and Mesoamerican bean varieties and regions. Eur J Plant Pathol 110:253–263
Masi P, Figliuolo G, Spagnoletti Zeuli PL (1999) Landraces of bean (Phaseolus vulgaris L.) collected in Basilicata Italy. Plant Genet Res 119:51–55
Mercati F, Maria L, Antonio L, Agostino S, Monica B, Maria RA, Francesco S (2013) Genetic diversity and population structure of a common bean (Phaseolus vulgaris L.) collection from Calabria (Italy). Genet Resour Crop Evol 60:839–852
Meza N, Juan CR, Juan PM, Jesu´s MO (2013) Biodiversity of common bean (Phaseolus vulgaris L.) in Honduras, evidenced by morphological characterization. Genet Resour Crop Evol 60:1329–1336
Miklas P, Kelly J, Beebi S, Blair M (2006) Common bean breeding for resistance against biotic and abiotic stresses from classical to MAS breeding. Euphytica 147:105–131
Mirjana V (2005) Principal component analysis of dry bean collection. Bean Improv Corp 48:16–17
Mishra S, Sharma MK, Singh M, Yadav SK (2010) Genetic diversity of French bean (Bush type) genotypes in north-west Himalayas. Indian J Plant Genet Resour 23(3):285–287
Mudasir S, Sofi PA, Khan MNM, Sofi NR, Dar ZA (2012) Research article genetic diversity, variability and character association in local common Bean (Phaseolus vulgaris L.) germplasm of Kashmir. Electron J Plant Breed 3(3):883–891
Narpinder Singh, Kaur Navpreet, Rana Jai Chand, Sharma Shyam Kumar (2010) Diversity in seed and flour properties in field pea (Pisum sativum) germplasm. Food Chem 122(3):518–525
NBPGR (2000) Minimal descriptors (Part–I). Agro biodiversity (PGR)-9. National Bureau of Plant Genetic Resources, New Delhi
Negri V (2012) Policies supportive of on-farm conservation and their impact on custodians farmers in Italy. In: Padulosi S, Bergamini N, Lawrence T (eds) On farm conservation of neglected and underutilized species: status, trends and novel approaches to cope with climate change. Proceedings of an International Conference, Frankfurt, 14–16. Biodiversity International, Rome, p 211
Nehvi FA, Singh G, Manzar A, Allai BA (2007) Evaluation of local land races of French bean under temperate conditions of kashmir valley. J Food Legum 20:41–42
Park SO, Coyne DP, Jung G, Skroch PW, Arnaud-Santana E, Steadman JR, Ariyarathne HM, Nienhuis J (2000) Mapping of QTL for seed size and shape traits in common bean. J Am Soc Hort Sci 125:466–475
Parmar N, Virdi AS, Singh N, Kaur A, Ritika B, Rana JC, Lalit A, Nautiyal CS (2014) Evaluation of physicochemical, textural, mineral and protein characteristics of kidney bean grown at Himalayan region. Food Res Int 66:45–57
Paudel KB, Chaudhary JN, Gautam IP, Upadhaya KP, Khatri B (2003) Performance of bush type French bean in the Hills of Nepal. Proceeding of 2nd SAS-N Convention, 30 July-1 August, Kathmandu
Perseguini J, Chioratto A, Zucchi M, Colombo C, Carbonell S (2011) Genetic diversity in cultivated Carioca common beans based on molecular marker analysis. Genet Mol Biol 34(1):88–102
Piergiovanni AR, Lucia L (2010) Italian common bean landraces: history genetic diversity and seed quality. Diversity 2:837–862
Piergiovanni AR, Cerbino D, Brandi M (2000) The common bean populations from Basilicata (southern Italy). An evaluation of their variation. Genet Resour Crop Evol 47:489–495
Polegri L, Negri V (2010) Molecular markers for promoting agrobiodiversity conservation: a case study from Italy. How cowpea landraces were saved from extinction. Genet Resour Crop Evol 57:867–880
Pujola M, Farreras A, Casan˜ as F (2007) Protein and starch content of raw, soaked and cooked beans (Phaseolus vulgaris L.). Food Chem 102:1034–1041
Raffi SA, Nath UK (2004) Variability, heritability, genetic advance and relationships of yield and yield contributing characters in dry bean (Phaseolus vulgaris L.). J Biol Sci 4:157–159
Raggi L, Barbara T, Valeria N (2013) Italian common bean landraces: diversity and population structure. Genet Resour Crop Evol 60:1515–1530
Rai N, Asati BS, Singh AK, Yadav DS (2006) Genetic variability, character association and path coefficient study in pole type French bean. Indian J Hort 63:188–191
Rai N, Singh PK, Verma A, Yadav PK, Choubey T (2010) Hierarchical analysis for genetic variability in pole type French bean. Indian J Hort 67:150–153
Ramanjulu V, Reddy DR, Babu GP, Gopal K (1999) Identification of source of resistance to bean common mosaic virus in common bean (Phaseolus vulgaris L.). Pl Des Res 19:134–136
Rana JC, Sharma BD, Gautam PL (2000) Agri-diversity Erosion in the north-west Indian Himalayas-some case studies. Indian J Pl Genet Resour 13(3):252–258
Rana JC, Singh A, Sharma Y, Pradheep K, Mendiratta N (2010) Dynamics of plant bio-resources in western Himalayan region of India–watershed based case study. Curr Sci 98(2):192–203
Rana JC, Pradheep K, Chaurasia OP, Sood S, Sharma RM, Singh A, Negi R (2011) Genetic resources of wild edible plants and their uses among tribal communities of cold arid region of India. Genet Resour Crop Evol 59(1):135–149
Rana JC, Dutta M, Rathi RS (2012) Plant genetic resources of the Indian Himalayan region–an overview. Indian J Genet 72(2):115–129
Rana JC, Banyal DK, Sharma KD, Sharma MK, Gupta SK (2013) Screening of pea germplasm for resistance to powdery mildew. Euphytica 189(2):271–282
Rana PK, Kumar P, Singhal VK, Rana JC (2014) Uses of local plant biodiversity among the tribal communities of Pangi Valley of district Chamba in cold desert Himalaya, India. World Sci J 2014:753289
Rathore A, Parsad R, Gupta VK (2004) Computer aided construction and analysis of augmented designs. J Indian Soc Agric Stat 57:320–344
Robinson HF (1966) Quantitative genetics in relation to breeding of the centennial of Mendalism. Indian J Genet 26:171–187
Rodin˜o AP, Santalla M, de Ron AM, Singh SP (2003) A core collection of common bean from the Iberian Peninsula. Euphytica 131:165–175
Rodin˜o AP, Santalla M, Gonza´lez AM, de Ron AM, Singh SP (2006) Novel genetic variation in common bean from the Iberian Peninsula. Crop Sci 46:2540–2546
Rodriguez-Guerra R, Ramirez-Rueda MT, Martinez de la Vega O, Simpson J (2003) Variation in genotype, pathotype and anastomosis groups of Colletotrichum lindemuthianum isolates from Mexico. Plant Pathol 52:228–235
Rosales-Serna R, Hernández-Delgad S, González-Paz M, Acosta-Gallegos JA, Mayek-Pére N (2005) Genetic relationshipsand diversity revealed by AFLP markers in Mexican common bean bred cultivars. Crop Sci 45:1951–1957
Rossi M, Bitocchi E, Bellucci E, Nanni L, Rau D (2009) Linkage disequilibrium and population structure in wild and domesticated populations of Phaseolus vulgaris L. Evol Appl 2:504–522
Santalla M, Rodiño AP, De Ron AM (2002) Allozyme evidence supporting southwestern Europe as a secondary center of genetic diversity for common bean. Theor Appl Genet 104:934–944
Santalla M, Carmen MS, Ana B, Monteagudo Antonio M, Ron De (2004) Genetic diversity of Argentinean common bean and its evolution during domestication. Euphytica 135:75–87
Sharma BD, Rana JC (2005) Plant genetic resources of western Himalaya: status and prospects. Bishan Singh Mahaendra Singh, Dehradun 457
Sharma TR, Rana JC, Sharma R, Rathour R, Sharma PN (2006) Genetic diversity analysis on exotic and Indian accessions of common bean using RAPD markers. Indian J Genet 66(4):275–278
Sharma PN, Bilal A, Sharma OP, Pathania A, Sharma P (2007) Pathological and molecular diversity in Colletotrichum lindemuthianum (bean anthracnose) across Himachal Pradesh-A north- western Himalayan state of India. Australas Plant Pathol 36:191–197
Sharma MK, Mishra S, Rana NS (2009) Genetic divergence in French bean (Phaseolus vulgaris L.) pole type cultivars. Legume Res 32(3):220–223
Sharma PN, Banyal K, Rana JC, Nag R, Sharma SK, Pathania A (2012) Screening of common bean germplasm against Colletotrichum lindemuthianum causing bean anthracnose. Indian Phytopathol 65(1):99–102
Singh SP (1989) Patterns of variation in cultivated common bean (Phaseolus vulgaris, Fabaceae). Econ Bot 43:39–57
Singh SP, Gepts P, Debouck DG (1991a) Races of common bean (Phaseolus vulgaris, Fabaceae). Econ Bot 45:379–396
Singh SP, Gutie´rrez JA, Molina A, Urrea C, Gepts P (1991b) Genetic diversity in cultivated common bean. II. Marker based analysis of morphological and agronomic traits. Crop Sci 31:19–23
Singh NK, Tiwari M, Kumar N (2002) Seed protein studies in common bean (Phaseolus vulgaris L.) gremplasm grown at high altitude. Indian J Genet 63:267–268
Singh KP, Minakshi J, Minakshi B (2007) Genetic variability in French bean (Phaseolus vulgaris L.). Res Crops 8:636–637
Singh A, Singh S, Babu JDB (2011) Heritability, character association and path analysis in early segregating populations of field pea. Int J Plant Breed Genet 5(1):86–92
Singh N, Kaur S, Rana JC, Nakaura Y, Inouchi N (2012) Isoamylase debranched fractions and granule size in starches from kidney bean germplasm: distribution and relationship with functional properties. Food Res Int 47:174–181
Sofi PA, Zargar MY, Debouck D, Graner A (2011) Evaluation of common bean (Phaseolus vulgaris L) germplasm under temperate conditions of Kashmir Valley. J Phytol 3(8):47–52
Sofi PA, Rana JC, Bhat NA (2014) Pattern of variation in common bean (Phaseolus vulgaris L) genetic resources of Jammu and Kashmir. J Food Legumes 27(3):197–201
Stoilova T, Pereira G, Tavares de souse MM, Carnide V (2005) Diversity in common bean landraces (Phaseolus vulgaris L.) from Bulgaria and Portugal. J Cent Eur Agric 6(4):443–448
Stoilova T, Pereira G, Tavares-De-Sousa M (2013) Morphological characterization of a small common bean (Phaseolus vulgaris L.) collection under different environments. J Cent Eur Agric 14(3):854–864
Szilagyi L, Taygar S, Ciuca M (2011) Evaluation of genetic diversity in common bean (P. vulgaris L) using RAPD markers and morpho-agronomic traits. Rom Biotechnol Lett 16(1):98–105
Tar’an B, Zhang C, Warkentin T, Tullu A, Vandenberg A (2005) Genetic diversity among varieties and wild species accessions of pea (Pisum sativum L.) based on molecular markers, and morphological and physiological characters. Genome 48:257–272
Tiwari M, Singh NK, Rathore M, Kumar N (2005) RAPD markers in the analysis of genetic diversity among common bean germplasm from Central Himalaya. Genet Resour Crop Evol 52:315–324
Trutmann P, Voss J, Fairhead J (1996) Local knowledge and farmer perceptions of bean diseases in the Central Africa Highlands. Agric Hum Values 13:112–125
Wortmann CS, Kirkby RA, Eledu CA, Allen DJ (1998) Atlas of common bean (Phaseolus vulgaris L.) production in Africa. CIAT Pan-African Bean Research Alliance, Cali
Acknowledgments
We sincerely thank the all the farming communities across the world who have been conserving the germplasm on farm and curators of bean germplasm in different countries for sharing the germplasm of common bean among different researchers. We also acknowledge Director, ICAR-NBPGR for providing facilities to conduct the research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rana, J.C., Sharma, T.R., Tyagi, R.K. et al. Characterisation of 4274 accessions of common bean (Phaseolus vulgaris L.) germplasm conserved in the Indian gene bank for phenological, morphological and agricultural traits. Euphytica 205, 441–457 (2015). https://doi.org/10.1007/s10681-015-1406-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-015-1406-3