Abstract
Fluorescence in situ hybridization was used to investigate the physical location of oligo-Am1 and (TTG)6 trinucleotide repeats in the metaphase chromosomes of seven diploid species (AA or CC genomes), seven tetraploid species (AABB or AACC genomes), and two hexaploid species (AACCDD genomes) belonging to the genus Avena. The oligo-Am1 probe produced signals that were particularly enriched on almost whole C-genome chromosomes, whereas the (TTG)6 probe was located in the pericentromeric (M), and, occasionally, their telomeric (T) chromosome regions, but showed low matching to C genome. All the species possessed (TTG)6 loci in M regions, and the CC, AABB, and AACCDD species also possessed (TTG)6 loci in T regions. The (TTG)6 signal number is constant in both the AA and CC species but slightly differs in signal intensity, whereas the (TTG)6 signal pattern shows wide diversity in the AABB, AACC and AACCDD species. The probe hybridization results provide key information that can be used in the physical assignment of genome sequences to chromosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Avena contains approximately 30 species, which are widespread throughout Europe, Asia and northwest Africa (Baum 1977). This genus can be divided into diploids, tetraploids, and hexaploids with a base chromosome number of seven. The diploid species have AA or CC genomes, while the tetraploids have AABB or AACC genomes, and the hexaploids have AACCDD genomes. There are five variants of the A genome (Ac, Al, Ad, As, and Ap), whereas the C genome exhibits two different variants (Cp and Cv) (Loskutov and Rines, 2011). The A genome differs distinctly from the C genome but is closely related to both the B and D genomes (Chen and Armstrong 1995). Probable A-genome and C-genome ancestors are relative clear, but the origins of the B and D genomes remain an open question.
The As genome exhibits the highest cytogenetic and molecular similarity to those of the hexaploids (Jellen et al. 1994; Yan et al. 2016a). The Ac genome might be the closest diploid genome to the ancestral D genome donor (Craig et al. 1974). Both Cp and Cv have been proposed as contributors to the C genome in hexaploids (Yan et al. 2016a). Among the four species with the AB genomes, Avena agadiriana Baum et Fedak is cytogenetically distinct from the other three species (Badaeva et al. 2010a). In addition, A. agadiriana shares certain similarities with species possessing the Ac genome (Hayasaki et al. 2001), the AC genomes (Alicchio et al. 1995) and the ACD genomes (Badaeva et al. 2010b) and crosses easily with hexaploids, which might suggest that A. agadiriana participated in the evolution of hexaploids (Thomas 1989). Among the three species with the AC genomes, Avena insularis Ladiz. is morphologically similar to Avena sterilis L. (Ladizinsky1998). Avena magna Gdgr. is thought to have derived from Avena canariensis Baum Raj. et Samp. and Avena ventricosa Bal. ex Coss. (Rajhathy and Thomas 1974).
Information on differences in chromosome structure among closely related species has been recorded based on cytogenetic approaches, such as C-banding (Chaffin et al. 2016; Jellen and Bill 1996), genomic in situ hybridization (GISH) (Chen and Armstrong 1994), and fluorescence in situ hybridization (FISH) (Luo et al. 2014). The cytogenetic positions of specific repetitive sequences, such as AS-1 (Fabijanski et al. 1990), pAvKB30 (Katsiotis et al. 1996), pAs120a (Irigoyen et al. 2002), 5S and 45S rDNA (Badaeva et al. 2010b), A336 (Luo et al. 2015), CCS1 Avena-700 (Tomás et al. 2016), (AC)10 (Fominaya et al. 2017), and (ACT)6 (Luo et al. 2018) have been useful in revealing the structural evolution of the chromosomes of Avena species. The detection of the above repetitive sequences, ranging from 18 to 700 bp, via FISH allowed the identification of certain homologous regions. However, these reported findings are insufficient to clarify the complex homology associations within Avena. Although the hexaploid oat seed transcriptome was annotated by Gutierrez-Gonzalez et al. (2013) and a genotyping-by-sequencing (GBS) marker was used for genomic discovery in cultivated oat by Huang et al. (2014), none of the genome sequences of Avena species have been published to date. Thus, the chromosome distributions of DNA sequences are needed and might provide further clues with respect to this topic.
Oligonucleotides have been widely used as FISH probes for the cytogenetics of many genera (Pedersen et al. 1996; Zhao et al. 2016; Du et al. 2017) but seldomly for oats, except for Fominaya et al. (2017) and Luo et al. (2018). Previously, Yan et al. (2016a) reported the ancestral genomes of Avena species and identified the D-genome chromosomes of hexaploid oat using a GBS marker; these authors obtained approximately 587,510 bp sequences and identified regions of 4D-genome chromosomes showing the strongest differential matching (low matching to the A and C genomes and high matching to the CD genomes). TTG was the trinucleotide that was repeated most often in the above sequence. The oligo-Am1 was an oligonucleotide containing a 51-bp fragment and was developed from pAm1 repeats, a 464-bp fragment described by Fominaya et al. (1995), which was exclusive to whole C-genome chromosomes. The objectives of this study were to locate (TTG)6 trinucleotide repeats and oligo-Am1 in the chromosomes of sixteen Avena species and to explore whether their D-genome origin might allow (TTG)6 repeats to be used as a genome-specific probe in FISH experiments.
Methods
Plant materials and chromosome preparation
Five diploid species with the AA genomes (A. longiglumis, A. nuda, A. brevis, A. canariensis, and A. strigosa), two diploid species with the CC genomes (A. eriantha and A. ventricosa), four tetraploid species with the AABB genomes (A. barbata, A. vaviloviana, A. abyssinica and A. agadiriana), three tetraploid species with the AACC genomes (A. magna, A. murphyi, and A. insularis), and two hexaploid species with the AACCDD genomes (A. sativa and A. fatua) were obtained from the oat germplasm collections of the American National Plant Germplasm System (Pullman, WA, USA) and Plant Gene Resources of Canada (Saskatoon, SK, Canada).
The locations, accession numbers and genomic constitutions of the sixteen Avena species are given in Table 1. All accessions were karyotyped with probes labelled with oligo-Am1 and (TTG)6 DNA. The metaphase chromosomes of all species were prepared from root tips taken from random young seedlings. The excised root tips were pretreated with nitrous oxide for 4 h and then stored in 70% ethanol (Kato et al. 2004). Approximately 1 mm of the root-tip meristem was cut and treated with cellulose and pectinase (2:1); the suspension was then dropped onto slides (Komuro et al. 2013). After air drying, the slides were examined using an Olympus CX21 microscope (Olympus, Japan) and stored at − 20 °C prior to use.
Probe DNA preparation
Two probes were used for the in situ hybridization analyses. The oligo-Am1 probe was an oligonucleotide containing a 51-bp fragment with the sequence 5′-GATCCATGTGTGGTTTGTGGAAAGAACACACATGCAATGACTCTAGTGGTT-3′ and was developed from a 464-bp fragment described by Fominaya et al. (1995). The (TTG)6 probe consisted of the trinucleotide repeat, corresponding to an 18-bp fragment from 4D-genome chromosomes, was first tested in Avena. The two probes were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). The synthetic oligo-Am1 was 5′ end-labelled with 6-carboxytetramethylrhodamine (TAMRM), whereas the (TTG)6 was 5′ end-labelled with 6-carboxyfluorescein (FAM).
FISH analysis
Probe hybridization and signal detection were carried out according to the protocol published by Hao et al. (2013). The preparations were counterstained with 4,6-diamidino-2-phenylindole (DAPI; Vector Laboratories, Inc., Burlingame, VT, USA). The slides were then examined, and selected metaphase plates were photographed with an Olympus BX-63 microscope attached to a Photometric SenSys Olympus DP70 CCD camera (Olympus, Japan). The raw images were processed with Photoshop ver. 7.1 (Adobe Systems Incorporated, San Jose, CA, USA), only using functions that affected the entire image equally. Three to five metaphases were used to FISH analysis for each species.
Results
In situ hybridization with the (TTG)6 and oligo-Am1 probes
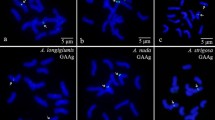
Hybridization of the (TTG)6 probe to the chromosomes of five AA species revealed six green signals located on three A chromosome pairs in pericentromeric regions, whereas two CC species revealed four green signals located on two C chromosome pairs in pericentromeric or telomeric regions (Figs. 1, 4). Six faint signals were located on the A. brevis chromosomes (Fig. 1a); four relative strong signals and two weak signals were located on the A. strigosa chromosomes (Fig. 1b); four strong and two weak signals were observed on the A. canariensis chromosomes (Fig. 1c); two strong and four relative weak signals were observed on the A. longiglumis chromosomes (Fig. 1d); and six relatively strong signals were found on the A. nuda chromosomes (Fig. 1e); two of the weak signals were located in the pericentromeric regions, whereas the other two small signals were located in the telomeric regions of the A. eriantha and A. ventricosa chromosomes (Fig. 1f, g).
FISH using the (TTG)6 trinucleotide repeat probe in the chromosomes of A. brevis (a), A. strigosa (b), A. canariensis (c), A. longiglumis (d), A. nuda (e), A. eriantha (g) and A. ventricosa (h). The chromosomes were probed with the 5′-FAM-labelled (TTG)6 repeat (green) in all images. The (TTG)6 signals are indicated with white arrows. All chromosomes were counterstained with DAPI. Scale bar = 5 μm. (Color figure online)
Hybridization of the (TTG)6 probe to the chromosomes of the tetraploid A. barbata, A. vaviloviana, A. abyssinica, and A. agadiriana revealed eight to fourteen green signals located on four to seven chromosome pairs in their pericentromeric, and, occasionally, telomeric regions (Fig. 2). Eight clear signals (including multi-point signals located on two chromosomes) were located on the A. barbata chromosomes in pericentromeric regions (Fig. 2a). Ten signals (six pericentromeric and four telomeric) were observed on the A. vaviloviana chromosomes (Fig. 2b). Fourteen signals (ten pericentromeric and four telomeric) were observed on the A. abyssinica chromosomes (Fig. 2c). Ten signals (two large, six normal, and two weak signals) were located on the A. agadiriana chromosomes in pericentromeric regions (Fig. 2d).
FISH with the (TTG)6 trinucleotide repeat probe in the chromosomes of A. barbata (a), A. vaviloviana (b), A. abyssinica (c), and A. agadiriana (d). The chromosomes and signals were visualized by the same method as Fig. 1. Scale bar = 5 μm. (Color figure online)
Hybridization of the (TTG)6 probe to chromosomes of three AACC species and two AACCDD species revealed eight to 12 green signals located on 4–6 AC chromosome pairs in pericentromeric regions (Fig. 3a–c), and 16–18 green signals located on 8–9 chromosome pairs in pericentromeric and telomeric regions (Fig. 3d, e). The oligo-Am1 probe produced red signals that were particularly enriched on C-genome chromosomes (Fig. 3). Eight A–C minor translocations were observed on A. magna and A. murphyi chromosomes (Fig. 3a, b), and six A–C minor translocations were observed on A. insularis chromosomes (Fig. 3c). Eight (TTG)6 signals (2 very large, 2 relatively small, and 4 very small) were located on the A chromosomes and four very small (TTG)6 signals were located on the C chromosomes of A. magna (Fig. 3a). Eight (TTG)6 signals (6 relatively large and 2 small) were located on the A chromosomes and two very small (TTG)6 signals were located on the C chromosomes of A. murphyi (Fig. 3b). Eight (TTG)6 signals (2 very large and 6 very small signals) were located on the A chromosomes of A. insularis (Fig. 3c).
FISH with the (TTG)6 trinucleotide repeat and oligo-Am1 probes in the chromosomes of A. magna (a), A. murphyi (b), A. insularis (c), A. sativa (d) and A. fatua (e). The chromosomes and signals were visualized by the same method as Fig. 1, except for the chromosomes were probed with the 5′-TAMRA-labelled oligo-Am1 (red) in all images. Scale bar = 5 μm. (Color figure online)
Twelve A/D-C translocations (including two large translocations) were located on the A. sativa chromosomes (Fig. 3d), and ten A/D-C translocations were located on the A. fatua chromosomes (Fig. 3e). Fourteen (TTG)6 signals were located in pericentromeric regions of the A/D chromosomes of A. sativa and A. fatua, respectively. Two small (TTG)6 signals were located in pericentromeric regions of the C chromosomes of A. sativa and A. fatua, respectively. Two weak (TTG)6 signals were located in telomeric regions of the C chromosomes of A. sativa (Fig. 3d). Four small (TTG)6 signals were located in telomeric regions of the A/D chromosomes of A. fatua, and two of these telomeric signals were located in the A/D-C traditional translocation (Fig. 3e).
Summary of the FISH signal patterns of sixteen Avena species
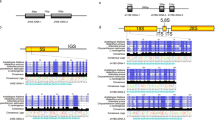
The polymorphisms observed in the FISH signal patterns of the sixteen Avena species based on the FISH signal patterns (signal number, location, intensity) are illustrated in Fig. 4. Chromosomes carrying (TTG)6 and/or oligo-Am1 loci are indicated with AA, CC, AABB, AACC, and AACCDD, representing the genome affiliations of the respective chromosomes. To visually display the associations among the species, the probed chromosomes are presented with the species name.
Genetic associations of Avena species with different genome types based on the FISH signal patterns. Chromosomes carrying (TTG)6 and/or oligo-Am1 loci are indicated with AA, CC, AABB, AACC, and AACCDD, representing the genome affiliations of the respective chromosome. The distributions of the (TTG)6 signal loci in Avena species chromosomes are summarized as two different types: M type, the signal is located in pericentromeric chromosome regions (M), whereas the signal is located in telomeric chromosome regions (T) in T type. (Color figure online)
The (TTG)6 signal number is constant in both the AA and CC species but differs in signal intensity. The (TTG)6 signal pattern is diverse in AABB, AACC and AACCDD species. The AA species possessed six (TTG)6 signal loci but showed slight differences in signal location and intensity. The CC species possessed four (TTG)6 signal loci but slightly differed in signal intensity. The AABB species showed diversity in the (TTG)6 signal number, location, and intensity. Three pericentromeric bands in one pair of chromosome and telomeric signal loci allow distinction between these four AABB species. The AACC species distinguished by oligo-Am1 labelled A–C translocation numbers showed variations in the (TTG)6 signal number, location, and intensity. The (TTG)6 signal number in the C genome was different among these three AACC species, whereas the signal intensity in the A genome greatly differed. The AACCDD species was differentiated by oligo-Am1 labelled A/D-C translocation numbers, and translocation size and showed variations in the (TTG)6 signal number, location, and intensity. Based on the (TTG)6 signal number in the C genome, the telomeric signals differed among these two AACCDD species.
The (TTG)6 signal loci were distributed in pericentromeric chromosome regions (M) or in telomeric chromosome regions (T). Comparison signal patterns from the AA, AABB, AACC, and AACCDD species imply that the AA diploid might have provided M to the AABB, AACC, and AACCDD species. Comparison signal patterns from the CC, AACC, and AACCDD species imply that the CC species might have provided M for the AACC species and MT for the AACCDD species. Comparison signal patterns from all species imply (TTG)6 probe showed low matching to the C genome. The signal number from the AA and AABB species indicated that the B genome possessed (TTG)6 loci, whereas the signal number from the AACC and AACCDD species indicated that the D genome possessed (TTG)6 loci.
Overall, the hybridization patterns of the oligo-Am1 and (TTG)6 probes showed that (1) the oligo-Am1 probe was particularly enriched in C genome, whereas the (TTG)6 probe was located in the pericentromeric, and, occasionally, telomeric chromosome regions, but showed low matching to C genome; (2) loci detected by the (TTG)6 probe were distributed in the AA, CC, AABB, AACC, and AACCDD species and were potentially present in the B and D genomes; (3) all the species possessed (TTG)6 loci in M regions, and the CC, AABB, and AACCDD species also possessed (TTG)6 loci in T regions; (4) the (TTG)6 signal number is constant in both the AA and CC species but slightly differed in signal intensity, whereas the (TTG)6 signal pattern is highly diverse in the AABB, AACC and AACCDD species.
Discussion
The (TTG)6 trinucleotide repeat probes were first tested in Avena species. The 51-bp oligo-Am1 probe identified the C genome in the present study, indicating that it functioned similarly to the 464-bp pAm1 (Sanz et al. 2012). The (TTG)6 sequence observed in 4D-genome chromosomes (Yan et al. 2016a) are a potentially valuable resource for cytogenetic identification of the oat D genome. In the present study, molecular probes coupled with FISH analyses were successfully used to localize a novel probe (TTG)6 originating from four D-genome chromosomes to pericentromeric, and, occasionally, telomeric regions of the chromosomes, showed low matching to C genome of Avena species. Similarly, the A336 FISH probe is a 391-bp fragment from chromosome 18D located in centromeric regions of oat chromosomes (Luo et al. 2015).
The (TTG)6 signal number is constant in both AA and CC species but slightly differs in signal intensity. These results indicate that minor variations occur in species with the AA and CC species, which is in agreement with the results reported by other authors (Loskutov and Rines 2011). The (TTG)6 signal pattern is highly diverse in the AABB, AACC and AACCDD species. The minor differences in the numbers and loci (TTG)6 signals observed in A. barbata, A. vaviloviana and A. abyssinica in the present study are confirmed by C-banding analysis (Badaeva et al. 2010a) and Southern hybridization (Irigoyen et al. 2001). FISH and RAPD analyses showed that A. agadiriana differed from A. abyssinica, A. vaviloviana, and A. barbata (Badaeva et al. 2010a). It was previously shown that A. insularis differs from A. magna and A. murphyi (Yan et al. 2016a), which is mostly consistent with our results. However, A. insularis resembles A. magna, whereas A. murphyi is somewhat distinct from these two species (Cheng et al. 2003). A. fatua is genetically distinct from other hexaploids (including A. sativa) in terms of one unique traditional translocation (Yang et al. 1999), morphological characteristics (Baum 1968) and the differences in the C-banding patterns of chromosome 5C (Badaeva et al. 2011).
The (TTG)6 signal loci are observed in telomeric chromosome regions in the CC and AACCDD species but not in the AACC species, which implies that chromosome construction has changed gradually during polyploidization. This variation is the result of a series of highly active processes, including chromosomal duplication, deletion, inversion and translocation, followed by gene loss and genome rearrangement; the last process often occurs after polyploidy or hybridization (Kellogg and Bennetzen 2004). Most of the Avena polyploids have suffered genome downsizing compared with their diploid ancestors (Yan et al. 2016b). Bennett and Leitch (2012) indicated that the C genome is larger than the A genome. The loss of C-genome sequences might be related to discrepancies in parental genome size in Avena polyploids (Rodrigues et al. 2017). The significantly reduced size, intensity, and number of pTa794 (5S) sites in polyploidy Avena species is evidence of the partial elimination of gene copies at the respective 5S rDNA loci of C-genome chromosomes, and all polyploid Avena species have lost the characteristic “in twos” distribution of 5S rDNA loci on C-genome chromosomes (Badaeva et al. 2010b). Indeed, substantial C genome rRNA sequence elimination has been recorded through molecular analysis (Nikoloudakis and Katsiotis 2008).
Abbreviations
- DAPI:
-
4,6-Diamidino-2-phenylindole
- FAM:
-
6-Carboxyfluorescein
- TAMRM:
-
6-Carboxytetramethylrhodamine
- FISH:
-
Fluorescence in situ hybridization
- GISH:
-
Genomic in situ hybridization
- GBS:
-
Genotyping-by-sequencing
- M:
-
Pericentromic
- T:
-
Telomeric
References
Alicchio R, Aranci L, Conte L (1995) Restriction fragment length polymorphism based phylogenetic analysis of Avena L. Genome 38:1279–1284. https://doi.org/10.1139/g95-168
Badaeva ED, Shelukhina OY, Diederichsen A, Loskutov IG, Pukhalskiy VA (2010a) Comparative cytogenetic analysis of Avena macrostachya and diploid C-genome Avena species. Genome 53:125–137. https://doi.org/10.1139/g09-089
Badaeva ED, Shelukhina OY, Goryunova SV, Loskutov IG, Pukhalskiy VA (2010b) Phylogenetic relationships of tetraploid AB-genome Avena species evaluated by means of cytogenetic (C-banding and FISH) and RAPD analyses. J Bot 2010:1–13. https://doi.org/10.1155/2010/742307
Badaeva ED, Shelukhina OY, Dedkova OS, Loskutov IG, Pukhalskyi VA (2011) Comparative cytogenetic analysis of hexaploid Avena L. Species. Russ J Genet 47:691–702. https://doi.org/10.1134/s1022795411060068
Baum BR (1968) On some relationships between Avena sativa and A. fatua (Gramineae) as studied from Canadian material. Can J Bot 46:1013–1024. https://doi.org/10.1139/b68-135
Baum BR (1977) Oats wild and cultivated. A monograph of the genus Avena L. (Poaceae). Agriculture Canada. ISBN: 9780660005133
Bennett MD, Leitch IJ (2012) Plant DNA C-values database. Release 6.0. Dec 2012
Chaffin AS, Huang YF, Smith S, Bekele WA, Babiker E, Gnanesh BN, Foresman BJ, Blanchard SG, Jay JJ, Reid RW, Wight CP, Chao S, Oliver R, Islamovic E, Kolb FL, McCartney C, Mitchell Fetch JW, Beattie AD, Bjornstad A, Bonman JM, Langdon T, Howarth CJ, Brouwer CR, Jellen EN, Klos KE, Poland JA, Hsieh TF, Brown R, Jackson E, Schlueter JA, Tinker NA (2016) A consensus map in cultivated hexaploid oat reveals conserved grass synteny with substantial subgenome rearrangement. Plant Genome 9(2):1–21. https://doi.org/10.3835/plantgenome2015.10.0102
Chen Q, Armstrong K (1994) Genomic in situ hybridization in Avena sativa. Genome 37:607–612 PMID:18470104
Chen Q, Armstrong K (1995) Characterization of a library from a single microdissected oat (Avena sativa L.) chromosome. Genome 38:706–714. http://www.ncbi.nlm.nih.gov/pubmed/18470104
Cheng DW, Armstrong KC, Drouin G, McElroy A, Fedak G, Molnar SD (2003) Isolation and identification of Triticeae chromosome 1 receptor-like kinase genes (lrk10) from diploid, tetraploid, and hexaploid species of the genus Avena. Genome 46:119–127. http://www.ncbi.nlm.nih.gov/pubmed/12669804
Craig IL, Murray BE, Rajhathy T (1974) Avena canariensis: morphological and electrophoretic polymorphism and relationship to the A. magna–A. murphyi complex and A. sterilis. Can J Genet Cytol 16:677–689. https://doi.org/10.1139/g74-074
Du P, Zhuang L, Wang Y, Yuan L, Wang Q, Wang D, Dawadondup, Tan L, Shen J, Xu H, Zhao H, Chu C, Qi Z (2017) Development of oligonucleotides and multiplex probes for quick and accurate identification of wheat and Thinopyrum bessarabicum chromosomes. Genome 60:93–103. http://www.ncbi.nlm.nih.gov/pubmed/27936984
Fabijanski S, Fedak G, Armstrong K, Altosaar I (1990) A repeated sequence probe for the c genome in Avena (oats). Theor Appl Genet 79:1–7. https://doi.org/10.1007/BF00223778
Fominaya A, Hueros G, Loarce Y, Ferrer E (1995) Chromosomal distribution of a repeated DNA sequence from C-genome heterochromatin and the identification of a new ribosomal DNA locus in the Avena genus. Genome 38:548–557. http://www.ncbi.nlm.nih.gov/pubmed/7557363
Fominaya A, Loarce Y, Montes A, Ferrer E (2017) Chromosomal distribution patterns of the (AC)10 microsatellite and other repetitive sequences, and their use in chromosome rearrangement analysis of species of the genus Avena. Genome 60:216–227. http://www.ncbi.nlm.nih.gov/pubmed/28156137
Gutierrez-Gonzalez JJ, Tu ZJ, Garvin DF (2013) Analysis and annotation of the hexaploid oat seed transcriptome. BMC Genome 14:471. https://doi.org/10.1186/1471-2164-14-471
Hao M, Luo J, Zhang L, Yuan Z, Yang Y, Wu M, Chen W, Zheng Y, Zhang H, Liu D (2013) Production of hexaploid triticale by a synthetic hexaploid wheat-rye hybrid method. Euphytica 193:347–357. https://doi.org/10.1007/s10681-013-0930-2
Hayasaki M, Morikawa T, Leggett JM (2001) Intraspecific variation of 18S-5.8S-26S rDNA sites revealed by FISH and RFLP in wild oat, Avena agadiriana. Genes Genet Syst 76:9–14. http://www.ncbi.nlm.nih.gov/pubmed/11376554
Huang YF, Poland JA, Wight CP, Jackson EW, Tinker NA (2014) Using genotyping-by-sequencing (GBS) for genomic discovery in cultivated oat. PLOS ONE 9:e102448. http://www.ncbi.nlm.nih.gov/pubmed/25047601
Irigoyen ML, Loarce Y, Linares C, Ferrer E, Leggett M, Fominaya A (2001) Discrimination of the closely related a and b genomes in AABB tetraploid species of Avena. Theor Appl Genet 103:1160–1166. https://doi.org/10.1007/s001220100723
Irigoyen ML, Linares C, Ferrer E, Fominaya A (2002) Fluorescence in situ hybridization mapping of Avena sativa L. cv. Sun II and its monosomic lines using cloned repetitive DNA sequences. Genome 45:1230–1237. https://doi.org/10.1139/g02-076
Jellen EN, Bill BS (1996) C-banding variation in the Moroccan oat species Avena agadiriana (2n-2x = 28). Theor Appl Genet 92:726–732. https://doi.org/10.1007/BF00226095
Jellen EN, Gill BS, Cox TS (1994) Genomic in situ hybridization differentiates between a/d- and c-genome chromatin and detects intergenomic translocations in polyploid oat species (genus Avena). Genome 37:613–618. https://doi.org/10.1139/g94-087
Kato A, Lamb JC, Birchler JA (2004) Chromosome painting using repetitive DNA sequences as probes for somatic chromosome identification in maize. Proc Natl Acad Sci USA 101:13554–13559. https://doi.org/10.1073/pnas.0403659101
Katsiotis A, Schmidt T, Heslop-Harrison JS (1996) Chromosomal and genomic organization ofty1-copia-like retrotransposon sequences in the genus Avena. Genome 39:410–417. https://doi.org/10.1139/g96-052
Kellogg EA, Bennetzen JL (2004) The evolution of nuclear genome structure in seed plants. Am J Bot 91:1709–1725. https://doi.org/10.3732/ajb.91.10.1709
Komuro S, Endo R, Shikata K, Kato A (2013) Genomic and chromosomal distribution patterns of various repeated DNA sequences in wheat revealed by a fluorescence in situ hybridization procedure. Genome 56:131–137. https://doi.org/10.1139/gen-2013-0003
Ladizinsky G (1998) A new species of Avena from sicily, possibly the tetraploid progenitor of hexaploid oats. Genet Resour Crop Evol 45:263–269. https://doi.org/10.1023/a:1008657530466
Loskutov IG, Rines HW (2011) Avena. In: Kole C (ed) Wild crop relatives: genomic and breeding resources. Springer, Heidelberg, pp 109–183. https://doi.org/10.1007/978-3-642-14228-4_3
Luo X, Zhang H, Kang H, Fan X, Wang Y, Sha L, Zhou Y (2014) Exploring the origin of the D genome of oat by fluorescence in situ hybridization. Genome 57:469–472. https://doi.org/10.1139/gen-2014-0048
Luo X, Tinker NA, Zhang H, Wight CP, Kang H, Fan X, Wang Y, Sha L, Zhou Y (2015) Centromeric position and genomic allocation of a repetitive sequence isolated from chromosome 18D of hexaploid oat, Avena sativa L. Genet Resour Crop Evol 62:1–4. https://doi.org/10.1007/s10722-014-0170-x
Luo X, Tinker NA, Zhou Y, Wight PC, Wan W, Chen L, Peng Y (2018) Genomic relationships among sixteen Avena taxa based on (ACT)6 trinucleotide repeat FISH. Genome 61:63–70. https://doi.org/10.1139/gen-2017-0132
Nikoloudakis N, Katsiotis A (2008) The origin of the C-genome and cytoplasm of Avena polyploids. Theor Appl Genet 117:273–281. https://doi.org/10.1007/s00122-008-0772-9
Pedersen C, Rasmussen SK, Linde-Laursen I (1996) Genome and chromosome identification in cultivated barley and related species of the Triticeae (Poaceae) by in situ hybridization with the GAA-satellite sequence. Genome 39:93–104. https://doi.org/10.1139/g96-013
Rajhathy T, Thomas H (1974) Cytogenetics of oats (Avena L.). Genetics Society of Canada, Ottawa. Miscellaneous Publ Genet Soc Can, p 90. https://www.researchgate.net/publication/37875550_Cytogenetics_of_oats_Avena_L
Rodrigues J, Viegas W, Silva M (2017) 45S rDNA external transcribed spacer organization reveals new phylogenetic relationships in Avena genus. PLoS ONE 12:e0176170. https://doi.org/10.1371/journal.pone.0176170
Sanz MJ, Loarce Y, Ferrer E, Fominaya A (2012) Use of tyramide-fluorescence in situ hybridization and chromosome microdissection for ascertaining homology relationships and chromosome linkage group associations in oats. Cytogenet Genome 36:145–156. https://doi.org/10.1159/000335641
Thomas H (1989) New species of Avena. In: Proceedings of the 3rd international oat conference, Lund, Sweden, pp 18–23. https://doi.org/10.1017/s1477200007002447
Tomás D, Rodrigues J, Varela A, Veloso MM, Viegas W, Silva M (2016) Use of repetitive sequences for molecular and cytogenetic characterization of Avena species from Portugal. Int J Mol Sci 7:203. https://doi.org/10.3390/ijms17020203
Yan H, Bekele WA, Wight CP, Peng Y, Langdon T, Latta RG, Fu YB, Diederichsen A, Howarth CJ, Jellen EN, Boyle B, Wei Y, Tinker NA (2016a) High-density marker profiling confirms ancestral genomes of Avena species and identifies D-genome chromosomes of hexaploid oat. Theor Appl Genet 129:2133–2149. http://www.ncbi.nlm.nih.gov/pubmed/27522358
Yan H, Martin SL, Bekele WA, Latta RG, Diederichsen A, Peng Y, Tinker NA (2016b) Genome size variation in the genus Avena. Genome 59:209–220. http://www.ncbi.nlm.nih.gov/pubmed/26881940
Yang Q, Hanson L, Bennett MD, Leitch IJ (1999) Genome structure and evolution in the allohexaploid weed Avena fatua L. (Poaceae). Genome 42:512–518. https://doi.org/10.1139/g98-154
Zhao LB, Ning SZ, Yu JJ, Hao M, Zhang LQ, Yuan ZW, Zheng YL, Liu DC (2016) Cytological identification of an Aegilops variabilis chromosome carrying stripe rust resistance in wheat. Breed Sci 66:522–529. https://doi.org/10.1270/jsbbs.16011
Acknowledgements
This study was supported by the Natural Science Foundation of China (31500993). The authors greatly appreciate the American National Plant Germplasm System (Pullman, WA, USA) and Plant Gene Resources of Canada (Saskatoon, SK, Canada) for providing the material for the investigation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Luo, X., Tinker, N.A., Zhou, Y. et al. Chromosomal distributions of oligo-Am1 and (TTG)6 trinucleotide and their utilization in genome association analysis of sixteen Avena species. Genet Resour Crop Evol 65, 1625–1635 (2018). https://doi.org/10.1007/s10722-018-0639-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-018-0639-0