Abstract
Glycosphingolipids, including gangliosides, are representative lipid raft markers that perform a variety of physiological roles in cell membranes. However, studies aimed at revealing their dynamic behavior in living cells are rare, mostly due to a lack of suitable fluorescent probes. Recently, the ganglio-series, lacto-series, and globo-series glycosphingolipid probes, which mimic the behavior of the parental molecules in terms of partitioning to the raft fraction, were developed by conjugating hydrophilic dyes to the terminal glycans of glycosphingolipids using state-of-art entirely chemical-based synthetic techniques. High-speed, single-molecule observation of these fluorescent probes revealed that gangliosides were scarcely trapped in small domains (100 nm in diameter) for more than 5 ms in steady-state cells, suggesting that rafts including gangliosides were always moving and very small. Furthermore, dual-color, single-molecule observations clearly showed that homodimers and clusters of GPI-anchored proteins were stabilized by transiently recruiting sphingolipids, including gangliosides, to form homodimer rafts and the cluster rafts, respectively. In this review, we briefly summarize recent studies, the development of a variety of glycosphingolipid probes as well as the identification of the raft structures including gangliosides in living cells by single-molecule imaging.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Previous studies on the dynamic behavior of glycosphingolipids in cell membranes
Glycosphingolipids perform a variety of physiological roles and are involved in many pathological processes in cell membranes, despite being much less abundant than phospholipids and cholesterol [1,2,3,4,5,6,7]. Gangliosides are a family of glycosphingolipids containing one or more N-acetylneuraminic acid (sialic acid) molecules in the carbohydrate chain. Over a hundred different gangliosides exist, and these molecules can be classified into several series (hemato-, ganglio-, globo-, isoglobo-, lacto-, and neolacto-) based on their carbohydrate structure. Gangliosides specifically associate with membrane receptors, such as EGF receptor [8,9,10,11], insulin receptor [12], and AMPA receptor [13], to regulate their activity. Gangliosides also play critical roles in the adhesion between cells [2, 14, 15] and in the invasion of microbial toxins [16, 17], viruses [18], and bacteria [19] into cells. Furthermore, gangliosides are important for promoting the molecular assembly of amyloid β in cell membranes [20,21,22,23]. In many cases, sialic acid from gangliosides is known to be involved in association and dissociation of molecular assembly [24,25,26]. As gangliosides are composed of carbohydrates and ceramide containing long saturated fatty acids (usually from C16:0 to C24:0), gangliosides are representative raft markers in cell plasma membranes (PMs) [27,28,29].
Although glycosphingolipids (including gangliosides) have key roles in important cellular functions, our knowledge of their spatial distributions, interactions with membrane receptors, clustering, and dynamic behavior in living cells remains very limited. Ganglioside-binding proteins, such as cholera toxin subunit B (CTXB), have been used to detect the location of GM1, but CTXB crosslinks five GM1 molecules, which can change their distribution [30,31,32]. Multivalent proteins such as toxins, lectins, and antibodies cannot be used for observation procedures. Even after chemical fixation with 4% paraformaldehyde and 0.3% glutaraldehyde, lipids in cell PMs continue to move [33, 34]. As the glycosphingolipids are moving, staining with multivalent antibodies would induce cluster formation and change their distribution in cell PMs. Therefore, the observation of fluorescent analogs of glycosphingolipids in living cell PMs appears to be the best way to perform detailed investigations of their spatial distribution, clustering, and dynamics. To address these issues, many ganglioside probes conjugated with fluorescent compounds have been synthesized. Examples include GM1 and GM2 analogs conjugated with 7-nitrobenz-2-oxa-1,3-diazol (NBD) in the alkyl chain [35], a GM1 analog with ATTO647N in the sugar chain or alkyl chain [36, 37], and a GM1 analog with Alexa 568 in the carbohydrate chain [38]. However, it has been found these ganglioside probes did not behave in the same way as their parental molecules in terms of partitioning into the liquid ordered (Lo) phase in giant unilamellar vesicles (GUVs) and into Lo-like phase in giant plasma membrane vesicles (GPMVs) [39, 40] and in terms of their binding affinity to cholera toxin subunit B (CTXB) [38]. Therefore, the development of true ganglioside probes that mimic the behavior of the parental molecules is anticipated.
Development of raft-associated glycosphingolipid probes
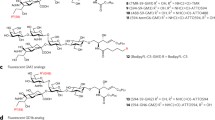
Komura et al. [40, 41] synthesized GM3 probes tagged with fluorescent dyes at the C9 position of sialic acid or at the C6 position of galactose. Interestingly, GM3 labeled with tetramethylrhodamine (TMR) at the C6 position of galactose (TMR-G6-GM3) was almost completely soluble in 1% cold Triton X-100, whereas GM3 with TMR at the C9 position (TMR-S9-GM3) was not. This result suggests that the fluorescent labeling of GM3 at the C9 position of sialic acid may cause a less detrimental reduction in the raft affinity of the probe than labeling at the C6 position of galactose. They subsequently examined the raft affinity of GM3 probes labeled with several different fluorescent dyes (fluorescein [Fl], ATTO488, TMR, ATTO594, ATTO647N) at the C9 position. ATTO647N-S9-GM3 completely partitioned into the liquid-disordered (Ld)-like domains in GPMVs, whereas TMR-S9-GM3 partitioned into both the Lo-like and Ld-like domains. Furthermore, Fl-S9-GM3, ATTO488-S9-GM3, ATTO594-S9-GM3 (Fig. 1a) mainly partitioned into the Lo-like domains [40]. As the hydrophilicity of the dyes follows the order Fl \(\cong\) ATTO488 \(\cong\) ATTO594 > TMR > ATTO647N, these results indicate that the hydrophilic dyes should be conjugated at the C9 position of sialic acid of GM3 to retain the raft affinity [42]. For simplicity, Fl-S9-GM3, ATTO488-S9-GM3, and ATTO594-S9-GM3 are referred to as Fl-GM3, 488-GM3, and 594-GM3 (Fig. 1a), respectively.
a (top) Chemical structure of the GM3 analog conjugated with ATTO594 at the C9 position of sialic acid. (bottom) Chemical structures of ATTO594 (left) and ATTO488 (right). b Schematic representation of glycosphingolipid probes that partition into the detergent-resistant membrane (DRM) fraction and liquid ordered (Lo)-like phase of giant plasma membrane vesicles (GPMVs). Here, the glycosphingolipids are classified into three major series: Ganglio-series (bule), Globo-series (green), and Lacto-series (magenta). A fluorescent dye, ATTO594, was conjugated with terminal glycans such as sialic acid, galactose, or GalNAc. The only the exception is Globo-H, in which galactose next to fucose was conjugated with ATTO594. ATTO594 can be replaced by ATTO488
Using the same strategy, Komura et al. [40, 41] synthesized 594-GM1 and 594-GM2, which mainly partitioned into the cold Triton X-100-insoluble fraction and Lo-like domains in GPMVs. Furthermore, GM1 conjugated with ATTO594 at the C6 position of another terminal glycan, galactose (called 594-termG6-GM1) showed high raft affinity, and so GM2 was conjugated with ATTO594 at the C6 position of terminal N-acetylgalactosamine (GalNAc) (called 594-GN6-GM2) (Fig. 1b). The GD1b analog tagged with ATTO594 at the C6 position of terminal galactose (594-termG6-GD1b) also exhibited high raft affinity. These results explicitly indicate that the hydrophilic dyes should be conjugated at the terminal sugar groups of gangliosides to retain raft affinity (Fig. 1b).
Through this strategy, Konishi et al. developed fluorescent probes for b-series gangliosides of GD3 and GQ1b [43], which are abundant in central nervous tissues and play an important role in nerve processes [44, 45]. The b-series gangliosides are also highly expressed in human gliomas, facilitating the malignant properties of these cells [46, 47]. GD3 and GQ1b were conjugated with the hydrophilic dye, ATTO594, at the C9 position of terminal sialic acid (Fig. 1b). High raft affinity was found for 594-GD3 and 594-GQ1b [43]. Furthermore, Yamaguchi et al. successfully synthesized a GD2 probe by conjugating GD2 at the C6 position of N-acetylgalactosamine (GalNAc) (Fig. 1b) and showed the GD2 probe partitioned into the cold Triton X-100-insoluble fraction [48].
Moreover, Asano et al. developed the fluorescent probes for the globo-series glycosphingolipids, of SSEA-3, SSEA-4 (ganglioside), and Globo-H [49]. The globo-series glycosphingolipids are stage-specific embryonic antigens that are specifically expressed in human-induced pluripotent stem cells [50] and cancer cells [51]. They play important roles in many biological processes, such as cell recognition, cell adhesion, and signal transduction. SSEA-3 and SSEA-4 were conjugated with the hydrophilic dye, ATTO594, at the terminal galactose C3 position and terminal sialic acid C9 position, respectively (Fig. 1b). Meanwhile, Globo-H was labeled with ATTO594 at the C3 position of galactose between the terminal fucose and N-acetylgalactosamine (GalNAc) (Fig. 1b). All the globo-series glycosphingolipids predominantly partitioned into the cold Triton X-100-insoluble fraction and Lo-like phase in GPMVs [49], indicating that they are true raft markers. Although the galactose tagged with ATTO594 is not a terminal glycan in Globo-H (the terminal glycan is fucose), the conjugated ATTO594 is far from the membranes, which may mitigate the detrimental effects to raft affinity.
Furthermore, Takahashi et al. recently synthesized fluorescent probes for lacto-series glycosphingolipids of NeuAcLc4Cer and Lc4Cer [52]. The lacto-series glycosphingolipids are known to be involved in several serious diseases such as lung and digestive system cancers and human gliomas [53]. However, the detailed mechanisms remain unclear. NeuAcLc4Cer and Lc4Cer were conjugated with ATTO594 at the terminal sialic acid and terminal galactose, respectively (Fig. 1b). These lacto-series glycosphingolipid probes mainly partitioned into the cold Triton X-100 insoluble fraction and Lo-like phase in GPMVs [52], indicating that they are true raft markers.
High-speed, single-molecule imaging of ganglioside probes in steady-state cell PMs
Previous studies using stimulated emission depletion microscopy with fluorescence correlation spectroscopy (STED-FCS) showed that the GM1 probes tagged with hydrophobic ATTO647N were temporally confined in small domains (of 20 nm in diameter) for 10–20 ms and for 60%–70% of the time fraction in epithelial Ptk2 cell PMs, whereas ATTO647N-dipalmitoylphosphatidylethanolamine (DPPE), a control lipid probe, was trapped in such a small domain for a much shorter period [36, 54]. Other studies using FCS also demonstrated that the GM1 probe tagged with hydrophobic Bodipy-FL at the alkyl chain was confined in domains of 60–120 nm for 20 ms and for 70% of the time fraction in COS-7 cell PMs, unlike the control lipid probe PC-Bodipy-FL [55, 56]. These studies also showed that the GM1 probe underwent simple Brownian diffusion at 0.5–1.3 µm2/s outside of the temporal confinement area.
However, it has been established that these GM1 probes actually partitioned into Ld phase in GUVs and are therefore not true raft markers, likely because the conjugated dyes are hydrophobic [39, 40]. Therefore, Komura et al. investigated whether the true raft markers of ganglioside probes that they developed were trapped in small domains in steady-state cell PMs [40]. This observation was performed in HBSS without the presence of any ligands and growth factors. Single-fluorescent molecule imaging at high temporal resolution (0.5 ms/frame) revealed that all 594-GM1, 594-GM3, and ATTO594-conjugated dioleoylphosphatodylethanolamine (594-DOPE, non-raft marker) molecules underwent simple Brownian diffusion and were scarcely trapped in small domains (of 100 nm in diameter) for more than 5 ms in all the examined cells (PtK2, T24, NRK, and COS7) at 23 ˚C. Furthermore, Kinoshita et al. reported that single molecules of other representative raft marker probes of sphingomyelin (SM) and distearoylphosphatidylcholine (DSPC), of which choline was conjugated with ATTO594 via a nonaethylene glycol linker (594neg-SM and 594neg-DSPC, respectively), were scarcely trapped in small domains of 100 nm for more than 5 ms in PtK2 and T24 cells at 23 ˚C and 37 ˚C, respectively [57, 58]. These results explicitly indicate that true raft–lipid markers are scarcely confined in tiny domains, but undergo apparent simple Brownian diffusion when observed at 0.5 ms/frame.
Diffusional behavior of gangliosides inferred from the anchored protein picket model
Single-particle tracking (SPT) of 40 nm gold bound to phospholipids, GPI-anchored proteins, or transmembrane proteins at high temporal resolution (20–100 μs/frame) revealed that all phospholipids [59,60,61], GPI-anchored proteins [62], and transmembrane proteins [59, 61, 63] underwent temporally confined diffusion in small domains (30–200 nm), occasionally hopped to adjacent compartments, again being confined in the compartment, and repeated this process, which is called “hop diffusion” (Fig. 2). Phospholipids, GPI-anchored proteins, and transmembrane proteins underwent free simple Brownian diffusion within the compartment, and the microscopic diffusion coefficients in the time window of 100 μs were 5–9 \(\mu\)m2/s [59, 61, 63]. Meanwhile, macroscopic diffusion coefficients in the time window of 100 ms were 0.3–0.5 µm2/s and the ratio of microscopic diffusion coefficients to microscopic diffusion coefficients was more than 10 [59, 61, 63]. Large differences were not found in the trajectories of the membrane molecules on the membrane blebs lacking cortical actin filaments [59, 60]. Rapid-freeze deep-etch electron microscopic tomography revealed the three-dimensional structure of cortical actin filaments in the cytoplasmic membrane surface and showed that the average mesh size made of cortical actin was comparable with that of the compartment size determined by SPT [64]. These results indicate that hop diffusion of membrane molecules was induced by cortical actin filaments. Furthermore, many SPT experiments, combined with Monte Carlo simulation, suggested that transmembrane pickets anchored to the actin filaments can retard the diffusion of phospholipids, GPI-anchored proteins, and transmembrane proteins and induce their compartmentalization into small domains (Fig. 2) [33, 65,66,67,68]. Furthermore, high-speed single-fluorescent molecule imaging also supported this notion [61]. This model is called the “anchored protein picket model”.
Schematic diagram of the anchored protein picket model. (left) Top view from outside the cell. A variety of transmembrane proteins, which are anchored to and aligned along the actin-based membrane skeleton (MSK), form diffusion barriers and compartment boundaries resulting from the hydrodynamic friction-like effects of immobile obstacles. (right) Oblique top view of expanded schematic diagram near the compartment boundaries. Transmembrane proteins, GPI-anchored proteins, and phospholipids occasionally “hop” across the compartment boundaries that are formed by rows of anchored protein pickets
Single-fluorescent molecule imaging at a time resolution of 0.5 ms showed that ganglioside probes exhibited apparent simple Brownian diffusion in PtK2, T24, NRK, and COS7 cell PMs [40]. However, this was a result of the time resolution of the observation. The average compartment size in PtK2, T24, NRK, and COS7 cells was estimated to be 43, 110, 230, and 56 nm, respectively, and the average residency time in each compartment was estimated to be 1.1, 8.9, 13, and 2.8 ms by SPT of 40 nm gold particles bound to phospholipids [60]. Single molecules of ganglioside probes recorded at 0.5 ms resolution resided in the compartments in PtK2, T24, NRK, and COS7 cells for only 2, 17, 26, and 5 frames, respectively. As the microscopic diffusion coefficient inside of the compartment is approximately 9 \(\mu\)m2/s [59], the distance moved in one frame (0.5 ms) can be estimated to be approximately 130 nm. Therefore, ganglioside probes frequently collide with the boundary of the compartments during 0.5 ms, and single molecules of ganglioside probes appear to be localized at the center of the compartments [69]. These results indicate that 0.5 ms/frame is not sufficient time resolution to observe free diffusion in the compartments or hop diffusion beyond many compartments in these cell PMs. As mentioned above, ganglioside should be tagged with hydrophilic dyes and ATTO594 is one of the brightest dyes of the suitable molecules. However, it is very hard to observe single molecules of ATTO594 at a time resolution higher than 0.2 ms/frame, e.g., 50 μs/frame using illumination with higher power lasers because ATTO594 blinks frequently. In the future, if hydrophilic dyes that do not blink when imaged at high-time resolution observation are developed, it would be possible to observe the hop diffusion of ganglioside probes in cell PMs.
Formation of GPI-anchored protein homodimer rafts and cluster rafts by recruiting glycosphingolipids
The behaviors of GPI-anchored proteins, which are representative raft markers, in cell PMs have been investigated because they occupy an important position in the history of raft research. Single-molecule observations of many types of GPI-anchored proteins revealed that they formed transient homodimers with a lifetime of 150–280 ms everywhere in the cell PMs [70,71,72,73,74]. Single-molecule imaging also revealed that homodimer formation was induced by specific ectodomain protein interactions, and was stabilized by cooperative lipid interactions in steady-state cell PMs. Furthermore, CD59, a GPI-anchored protein that is a complement regulatory protein, was shown to form stable homo-oligomers containing up to four CD59 molecules upon stimulation with the natural ligand, membrane attack complex (MAC) consisting of C5b, C6, C7 and C8 [75,76,77,78]. The stimulation of CD59 with MAC is an actual biological event. The stable CD59 homo-oligomers diffused slowly (~ 0.02 µm2/s) and were temporarily immobilized, on average, for 0.6 s and 36% of the time. The immobilization of CD59 clusters was called STALL (Stimulation induced Temporary Arrest of LateraL diffusion). The CD59 clusters recruited raft-associated signaling molecules such as G\(\alpha\)i2 and Lyn at the STALL site, and activated Lyn phosphorylated an as-yet unknown protein, which induced recruitment of PLC\(\gamma\)2 and triggered the intracellular Ca2+ response [75, 76]. The STALL sites were proposed to be a signaling platform for intracellular signaling.
As described above, in steady-state cell PMs, ganglioside probes continuously diffused and exhibit almost no transient trapping in immobile domains. Subsequently, it was investigated whether gangliosides reside in moving rafts. Although homodimers and clusters of GPI-anchored proteins are stabilized by cholesterol, as described above, homodimers were still found to diffuse in PMs [70, 74,75,76]. It is not known if clusters of raft-associated molecules recruit other raft molecules, although it has been shown that simultaneously crosslinked two different raft molecules coalesce with each other [27]. It was also unknown if these secondary non-crosslinked raft elements can be recruited to clustered raft molecules. To address these issues, Komura et al. examined if the ganglioside probes 594-GM1 and 594-GM3 were recruited to diffusing CD59 homodimers tagged with ATTO488 via ACP-tag in CHO-K1 cell PMs by simultaneous two-color, single-molecule imaging [40]. Indeed, these ganglioside probes were recruited to CD59 homodimers for approximately 80 ms, but to CD59 monomers with a lifetime of only approximately 50 ms (Fig. 3). Meanwhile, single molecules of the non-raft unsaturated phospholipid probe, 594-DOPE, were colocalized with fluorescent spots of CD59 homodimers with a lifetime of only approximately 40 ms. Furthermore, Takahashi et al. found that both 594-NeuAcLc4Cer and 594-Lc4Cer were recruited to CD59 homodimers for longer periods (approximately 80 ms) than to CD59 monomers in CHO-K1 cell PMs, whereas 594-DOPE was colocalized with both CD59 homodimers and monomers for very short periods (lifetime of approximately 40 ms) (Fig. 3) [52]. Interestingly, the colocalization lifetimes of lacto-series glycosphingolipids with CD59 homodimers were independent of the presence of sialic acid. Consistent with these results, other representative raftophilic lipid probes, 594neg-SM and 594neg-DSPC, were preferentially recruited to CD59 homodimers rather than the monomers in CHO-K1 cell PMs [57, 58], and the colocalization lifetime of 594neg-SM with CD59 homodimers was comparable with those of 594-GM1, 594-GM3, 594-NeuAcLc4Cer, and 594-Lc4Cer. Therefore, the recruitment of these glycosphingolipid probes to CD59 homodimers is induced by raft–lipid interactions, but not by specific interactions involving glycans (Fig. 3). Furthermore, individual fluorescent spots of 594neg-SM or 594neg-DSPC were colocalized with each other with the lifetimes of approximately 50 ms [57], which is much shorter than the homodimer lifetime of CD59 [70]. These results explicitly indicate that CD59 homodimers serve as primary core molecules to transiently recruit other raftophilic lipids, thereby driving the formation of CD59 homodimer rafts.
Schematic image of the transient recruitment of glycosphingolipids to CD59 homodimers and CD59 clusters, which induce CD59 homodimer rafts and cluster rafts (domains shown in magenta). Glycosphingolipid probes transiently associated with CD59 monomers for short periods, but with CD59 homodimers and liganded CD59 clusters for prolonged periods. The prolonged interaction was dependent on cholesterol, yet independent of the presence of glycan in the glycosphingolipids, which indicated that glycosphingolipid probes were recruited to CD59 homodimers and clusters by raft–lipid interactions
CD59 clusters formed upon stimulation also transiently recruited 594-GM1 and 594-GM3 in T24 cell PMs, and the colocalization lifetime was approximately 100 ms (Fig. 3) [40]. Cholesterol depletion and the replacement of GPI-anchoring chain of CD59 with the non-raft transmembrane domain of the LDL receptor dramatically shortened the colocalization lifetimes and reduced the colocalization frequency. Similar lipid dependency was also observed in the recruitment of other representative raft–lipid probes, 594neg-SM and 594neg-DSPC, to CD59 clusters [57, 58]. These results indicate that the CD59 clusters transiently recruited raft lipids and formed “CD59 cluster rafts” (Fig. 3). As mentioned above, the CD59 clusters recruited raft-associated signaling molecules such as Lyn and G\(\alpha\)i2 into the inner leaflets of the PMs [75]. Therefore, both the outer and inner leaflets of PMs underneath the CD59 clusters may be enriched in raft-lipids. However, it is not known how the signaling molecules in the inner leaflets of PMs are enriched underneath GPI-anchored protein clusters in the outer leaflets; this is described in the following section.
Enrichment mechanisms of signaling molecules underneath domains containing GPI-anchored protein clusters and gangliosides
Through dual-color, single-molecule observation at high temporal resolution (down to 5 ms), Koyama-Honda et al. [79] found that CD59 cluster rafts recruited signaling molecules such as Lyn and H-Ras in the inner leaflets with the colocalization lifetimes of less than 100 ms, and activated these signaling molecules. The recruitment was dependent on cholesterol and the saturated alkyl chains of Lyn and H-Ras. GM1 cluster rafts recruited Lyn and H-Ras as efficiently as CD59 cluster rafts, and deletion mutants of Lyn and H-Ras lacking the protein moieties were still recruited to the cluster rafts, indicating that transbilayer raft phases induced by the cluster rafts in the outer leaflet recruited lipid-anchored signaling molecules by lateral raft–lipid interactions and participated in signal transduction (Fig. 4).
Schematic image of transient recruitment of lipid-anchored signaling molecules to the inner leaflet membrane underneath clusters of CD59 or GM1. Upon crosslinking of CD59 or GM1 in the outer leaflet of cell PMs, transbilayer raft phase is formed, and subsequently, cytoplasmic lipid-anchored signaling molecules such as H-Ras and Lyn are recruited to the transbilayer raft phase in the inner leaflet by lateral raft-lipid interaction
Using the imaging technique of homo-FRET, i.e., FRET between similar fluorophores, Mayor’s group reported that GPI-anchored proteins formed clusters by transbilayer interactions with phosphatidylserine clusters anchored to actin binding proteins [80]. Here, nonspecific transbilayer interdigitation of the fatty acid chains of phosphatidylserine (PS) and GPI-anchored proteins was the driving force inducing GPI-anchored protein clusters. The diameter of the GPI-anchored protein clusters was estimated as 360 nm [81].
Arumugam et al. [82] reported that a GM1 probe conjugated with Alexa 488 at sialic acid via a peptide linker (Alexa488-GM1) partitioned mainly into the Lo phase in GPMVs and bound to CTXB as strongly as endogenous GM1. Homo-FRET observations and fluorescence anisotropy measurements showed that Alexa488-GM1 containing C16:0 formed clusters via transbilayer interactions with PS in the inner leaflets of PMs, whereas Alexa488-GM1 containing C16:1 did not form such clusters. The radii (Rmax) of the GM1 probe clusters was estimated as 115 and 90 nm for C16:0 and C16:1, respectively, by Ripley’s K-function. By super-resolution microscopy, the radii of clusters of CTXB-bound GM1 probe was estimated to be 225 nm, which was much larger than that of the GM1 probe. CD59 was recruited to the CTXB-bound GM1 clusters, but this was not induced by PS in the inner leaflets [82]. These results suggest that the GM1 clusters may recruit CD59 by raft–lipid interactions in the outer leaflets of the PM, which is consistent with the results of Koyama-Honda et al. [79].
Conclusions
A variety of probes for ganglio-, lacto-, globo-series glycosphingolipids, which behave in a similar manner to the parental molecules in terms of raft affinity, have recently developed. High-speed, single-molecule observation in living cell PMs revealed that the ganglioside probes were scarcely trapped within small domains of 100 nm in diameter for more than 5 ms, but instead were transiently recruited to GPI-anchored protein homodimers and clusters for 80–100 ms, which demonstrated the formation of GPI-anchored protein homodimer rafts and cluster rafts. These events may be observed irrespective of cell type. In the future, single-molecule observations of glycosphingolipids conjugated with more photostable and bright dyes at a higher time resolution will facilitate the collection of more detailed molecular interactions in cell PMs. Furthermore, recently developed glycosphingolipid probes can allow us to perform simultaneous multi-color, single-molecule observation of membrane receptors, downstream signaling molecules, and glycosphingolipids, which will help to elucidate the regulatory mechanisms of receptor signaling.
Data availability
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
References
Hakomori, S.: Glycosphingolipids in cellular interaction, differentiation, and oncogenesis. Annu Rev Biochem 50(1), 733–764 (1981). https://doi.org/10.1146/annurev.bi.50.070181.003505
Hakomori, S.-I., Handa, K., Iwabuchi, K., Yamamura, S., Prinetti, A.: New insights in glycosphingolipid function: “glycosignaling domain,” a cell surface assembly of glycosphingolipids with signal transducer molecules, involved in cell adhesion coupled with signaling. Glycobiology 8(10), xi-xviii (1998). https://doi.org/10.1093/oxfordjournals.glycob.a018822
Hakomori, S.: Tumor malignancy defined by aberrant glycosylation and sphingo(glyco)lipid metabolism. Cancer Res 56(23), 5309–5318 (1996)
Hakomori, S.: Glycosylation defining cancer malignancy: new wine in an old bottle. Proc Natl Acad Sci USA 99(16), 10231–10233 (2002). https://doi.org/10.1073/pnas.172380699
Mattner, J., DeBord, K.L., Ismail, N., Goff, R.D., Cantu, C., Zhou, D., Saint-Mezard, P., Wang, V., Gao, Y., Yin, N., Hoebe, K., Schneewind, O., Walker, D., Beutler, B., Teyton, L., Savage, P.B., Bendelac, A.: Exogenous and endogenous glycolipid antigens activate NKT cells during microbial infections. Nature 434(7032), 525–529 (2005). https://doi.org/10.1038/nature03408
Zhou, D., Mattner, J., Cantu, C., 3rd., Schrantz, N., Yin, N., Gao, Y., Sagiv, Y., Hudspeth, K., Wu, Y.P., Yamashita, T., Teneberg, S., Wang, D., Proia, R.L., Levery, S.B., Savage, P.B., Teyton, L., Bendelac, A.: Lysosomal glycosphingolipid recognition by NKT cells. Science 306(5702), 1786–1789 (2004). https://doi.org/10.1126/science.1103440
Schnaar, R.L.: Chapter three - the biology of gangliosides. In: Baker, D.C. (ed.) Advances in carbohydrate chemistry and biochemistry, vol. 76, pp. 113–148. Academic Press (2019). https://doi.org/10.1016/bs.accb.2018.09.002
Wang, X.Q., Sun, P., Paller, A.S.: Ganglioside GM3 blocks the activation of epidermal growth factor receptor induced by integrin at specific tyrosine sites. J Biol Chem 278(49), 48770–48778 (2003). https://doi.org/10.1074/jbc.M308818200
Yoon, S.J., Nakayama, K., Hikita, T., Handa, K., Hakomori, S.I.: Epidermal growth factor receptor tyrosine kinase is modulated by GM3 interaction with N-linked GlcNAc termini of the receptor. Proc Natl Acad Sci USA 103(50), 18987–18991 (2006). https://doi.org/10.1073/pnas.0609281103
Kawashima, N., Yoon, S.J., Itoh, K., Nakayama, K.: Tyrosine kinase activity of epidermal growth factor receptor is regulated by GM3 binding through carbohydrate to carbohydrate interactions. J Biol Chem 284(10), 6147–6155 (2009). https://doi.org/10.1074/jbc.M808171200
Guan, F., Handa, K., Hakomori, S.I.: Regulation of epidermal growth factor receptor through interaction of ganglioside GM3 with GlcNAc of N-linked glycan of the receptor: demonstration in ldlD cells. Neurochem Res 36(9), 1645–1653 (2011). https://doi.org/10.1007/s11064-010-0379-9
Kabayama, K., Sato, T., Saito, K., Loberto, N., Prinetti, A., Sonnino, S., Kinjo, M., Igarashi, Y., Inokuchi, J.: Dissociation of the insulin receptor and caveolin-1 complex by ganglioside GM3 in the state of insulin resistance. Proc Natl Acad Sci USA 104(34), 13678–13683 (2007). https://doi.org/10.1073/pnas.0703650104
Prendergast, J., Umanah, G.K., Yoo, S.W., Lagerlöf, O., Motari, M.G., Cole, R.N., Huganir, R.L., Dawson, T.M., Dawson, V.L., Schnaar, R.L.: Ganglioside regulation of AMPA receptor trafficking. J Neurosci 34(39), 13246–13258 (2014). https://doi.org/10.1523/jneurosci.1149-14.2014
Iwabuchi, K., Yamamura, S., Prinetti, A., Handa, K., Hakomori, S.-I.: GM3-enriched microdomain involved in cell adhesion and signal transduction through carbohydrate-carbohydrate interaction in mouse melanoma B16 cells. J Biol Chem 273(15), 9130–9138 (1998). https://doi.org/10.1074/jbc.273.15.9130
Iwabuchi, K., Handa, K., Hakomori, S.-I.: Separation of “Glycosphingolipid signaling domain” from caveolin-containing membrane fraction in mouse melanoma B16 cells and its role in cell adhesion coupled with signaling. J Biol Chem 273(50), 33766–33773 (1998). https://doi.org/10.1074/jbc.273.50.33766
Fishman, P.H.: Role of membrane gangliosides in the binding and action of bacterial toxins. J Membr Biol 69(2), 85–97 (1982). https://doi.org/10.1007/BF01872268
Zuverink, M., Barbieri, J.T.: Chapter eleven - protein toxins that utilize gangliosides as host receptors. In: Schnaar, R.L., Lopez, P.H.H. (eds.) Progress in molecular biology and translational science, vol. 156, pp. 325–354. Academic Press. (2018). https://doi.org/10.1016/bs.pmbts.2017.11.010
Nguyen, L., McCord, K.A., Bui, D.T., Bouwman, K.M., Kitova, E.N., Elaish, M., Kumawat, D., Daskhan, G.C., Tomris, I., Han, L., Chopra, P., Yang, T.-J., Willows, S.D., Mason, A.L., Mahal, L.K., Lowary, T.L., West, L.J., Hsu, S.-T.D., Hobman, T., Tompkins, S.M., Boons, G.-J., de Vries, R.P., Macauley, M.S., Klassen, J.S.: Sialic acid-containing glycolipids mediate binding and viral entry of SARS-CoV-2. Nat Chem Biol 18(1), 81–90 (2022). https://doi.org/10.1038/s41589-021-00924-1
Belotserkovsky, I., Brunner, K., Pinaud, L., Rouvinski, A., Dellarole, M., Baron, B., Dubey, G., Samassa, F., Parsot, C., Sansonetti, P., Phalipon, A.: Glycan-glycan interaction determines shigella tropism toward human T lymphocytes. mBio 9(1) (2018). https://doi.org/10.1128/mBio.02309-17
Yanagisawa, K., Odaka, A., Suzuki, N., Ihara, Y.: GM1 ganglioside-bound amyloid beta-protein (A beta): a possible form of preamyloid in Alzheimer’s disease. Nat Med 1(10), 1062–1066 (1995). https://doi.org/10.1038/nm1095-1062
Matsuzaki, K., Kato, K., Yanagisawa, K.: Ganglioside-mediated assembly of amyloid β-protein: Roles in Alzheimer’s disease. Prog Mol Biol Transl Sci 156, 413–434 (2018). https://doi.org/10.1016/bs.pmbts.2017.10.005
Matsuzaki, K.: Aβ-ganglioside interactions in the pathogenesis of Alzheimer's disease. Biochim Biophys Acta Biomembr 1862(8), 183233 (2020). https://doi.org/10.1016/j.bbamem.2020.183233
Yagi-Utsumi, M., Kato, K.: Conformational variability of amyloid-β and the morphological diversity of its aggregates. Molecules (2022). https://doi.org/10.3390/molecules27154787
Kelm, S., Schauer, R.: Sialic acids in molecular and cellular interactions. Int Rev Cytol 175, 137–240 (1997). https://doi.org/10.1016/s0074-7696(08)62127-0
Schauer, R.: Sialic acids as regulators of molecular and cellular interactions. Curr Opin Struct Biol 19(5), 507–514 (2009). https://doi.org/10.1016/j.sbi.2009.06.003
Schauer, R.: Achievements and challenges of sialic acid research. Glycoconj J 17(7), 485–499 (2000). https://doi.org/10.1023/A:1011062223612
Harder, T., Scheiffele, P., Verkade, P., Simons, K.: Lipid domain structure of the plasma membrane revealed by patching of membrane components. J Cell Biol 141(4), 929–942 (1998). https://doi.org/10.1083/jcb.141.4.929
Sonnino, S., Prinetti, A.: Membrane domains and the “lipid raft” concept. Curr Med Chem 20(1), 4–21 (2013)
Prinetti, A., Chigorno, V., Prioni, S., Loberto, N., Marano, N., Tettamanti, G., Sonnino, S.: Changes in the lipid turnover, composition, and organization, as sphingolipid-enriched membrane domains, in rat cerebellar granule cells developing in vitro. J Biol Chem 276(24), 21136–21145 (2001). https://doi.org/10.1074/jbc.M010666200
Hammond, A.T., Heberle, F.A., Baumgart, T., Holowka, D., Baird, B., Feigenson, G.W.: Crosslinking a lipid raft component triggers liquid ordered-liquid disordered phase separation in model plasma membranes. Proc Natl Acad Sci USA 102(18), 6320–6325 (2005). https://doi.org/10.1073/pnas.0405654102
Lingwood, D., Binnington, B., Rog, T., Vattulainen, I., Grzybek, M., Coskun, U., Lingwood, C.A., Simons, K.: Cholesterol modulates glycolipid conformation and receptor activity. Nat Chem Biol 7(5), 260–262 (2011). https://doi.org/10.1038/nchembio.551
Kabbani, A.M., Raghunathan, K., Lencer, W.I., Kenworthy, A.K., Kelly, C.V.: Structured clustering of the glycosphingolipid GM1 is required for membrane curvature induced by cholera toxin. Proc Natl Acad Sci USA 117(26), 14978–14986 (2020). https://doi.org/10.1073/pnas.2001119117
Kusumi, A., Suzuki, K.: Toward understanding the dynamics of membrane-raft-based molecular interactions. Biochim Biophys Acta 1746(3), 234–251 (2005). https://doi.org/10.1016/j.bbamcr.2005.10.001
Tanaka, K.A., Suzuki, K.G., Shirai, Y.M., Shibutani, S.T., Miyahara, M.S., Tsuboi, H., Yahara, M., Yoshimura, A., Mayor, S., Fujiwara, T.K., Kusumi, A.: Membrane molecules mobile even after chemical fixation. Nat Methods 7(11), 865–866 (2010). https://doi.org/10.1038/nmeth.f.314
Schwarzmann, G., Wendeler, M., Sandhoff, K.: Synthesis of novel NBD-GM1 and NBD-GM2 for the transfer activity of GM2-activator protein by a FRET-based assay system. Glycobiology 15(12), 1302–1311 (2005). https://doi.org/10.1093/glycob/cwj018
Eggeling, C., Ringemann, C., Medda, R., Schwarzmann, G., Sandhoff, K., Polyakova, S., Belov, V.N., Hein, B., von Middendorff, C., Schonle, A., Hell, S.W.: Direct observation of the nanoscale dynamics of membrane lipids in a living cell. Nature 457(7233), 1159–1162 (2009). https://doi.org/10.1038/nature07596
Polyakova, S.M., Belov, V.N., Yan, S.F., Eggeling, C., Ringemann, C., Schwarzmann, G., de Meijere, A., Hell, S.W.: New GM1 ganglioside derivatives for selective single and double labelling of the natural glycosphingolipid skeleton. Eur J Org Chem 2009(30), 5162–5177 (2009). https://doi.org/10.1002/ejoc.200900645
Chinnapen, D.J., Hsieh, W.T., te Welscher, Y.M., Saslowsky, D.E., Kaoutzani, L., Brandsma, E., D’Auria, L., Park, H., Wagner, J.S., Drake, K.R., Kang, M., Benjamin, T., Ullman, M.D., Costello, C.E., Kenworthy, A.K., Baumgart, T., Massol, R.H., Lencer, W.I.: Lipid sorting by ceramide structure from plasma membrane to ER for the cholera toxin receptor ganglioside GM1. Dev Cell 23(3), 573–586 (2012). https://doi.org/10.1016/j.devcel.2012.08.002
Sezgin, E., Levental, I., Grzybek, M., Schwarzmann, G., Mueller, V., Honigmann, A., Belov, V.N., Eggeling, C., Coskun, U., Simons, K., Schwille, P.: Partitioning, diffusion, and ligand binding of raft lipid analogs in model and cellular plasma membranes. Biochim Biophys Acta 1818(7), 1777–1784 (2012). https://doi.org/10.1016/j.bbamem.2012.03.007
Komura, N., Suzuki, K.G., Ando, H., Konishi, M., Koikeda, M., Imamura, A., Chadda, R., Fujiwara, T.K., Tsuboi, H., Sheng, R., Cho, W., Furukawa, K., Furukawa, K., Yamauchi, Y., Ishida, H., Kusumi, A., Kiso, M.: Raft-based interactions of gangliosides with a GPI-anchored receptor. Nat Chem Biol 12(6), 402–410 (2016). https://doi.org/10.1038/nchembio.2059
Komura, N., Suzuki, K.G.N., Ando, H., Konishi, M., Imamura, A., Ishida, H., Kusumi, A., Kiso, M.: Syntheses of fluorescent gangliosides for the studies of raft domains. Methods Enzymol 597, 239–263 (2017). https://doi.org/10.1016/bs.mie.2017.06.004
Suzuki, K.G.N., Ando, H., Komura, N., Fujiwara, T.K., Kiso, M., Kusumi, A.: Development of new ganglioside probes and unraveling of raft domain structure by single-molecule imaging. Biochim Biophys Acta Gen Subj 1861(10), 2494–2506 (2017). https://doi.org/10.1016/j.bbagen.2017.07.012
Konishi, M., Komura, N., Hirose, Y., Suganuma, Y., Tanaka, H.N., Imamura, A., Ishida, H., Suzuki, K.G.N., Ando, H.: Development of fluorescent ganglioside GD3 and GQ1b analogs for elucidation of raft-associated interactions. J Org Chem 85(24), 15998–16013 (2020). https://doi.org/10.1021/acs.joc.0c01493
Cheresh, D.A., Pierschbacher, M.D., Herzig, M.A., Mujoo, K.: Disialogangliosides GD2 and GD3 are involved in the attachment of human melanoma and neuroblastoma cells to extracellular matrix proteins. J Cell Biol 102(3), 688–696 (1986). https://doi.org/10.1083/jcb.102.3.688
Tsuji, S., Arita, M., Nagai, Y.: GQ1b, a bioactive ganglioside that exhibits novel nerve growth factor (NGF)-like activities in the two neuroblastoma cell lines. J Biochem 94(1), 303–306 (1983). https://doi.org/10.1093/oxfordjournals.jbchem.a134344
Ohkawa, Y., Zhang, P., Momota, H., Kato, A., Hashimoto, N., Ohmi, Y., Bhuiyan, R.H., Farhana, Y., Natsume, A., Wakabayashi, T., Furukawa, K., Furukawa, K.: Lack of GD3 synthase (St8sia1) attenuates malignant properties of gliomas in genetically engineered mouse model. Cancer Sci 112(9), 3756–3768 (2021). https://doi.org/10.1111/cas.15032
Iwasawa, T., Zhang, P., Ohkawa, Y., Momota, H., Wakabayashi, T., Ohmi, Y., Bhuiyan, R.H., Furukawa, K., Furukawa, K.: Enhancement of malignant properties of human glioma cells by ganglioside GD3/GD2. Int J Oncol 52(4), 1255–1266 (2018). https://doi.org/10.3892/ijo.2018.4266
Yamaguchi, E., Komura, N., Tanaka, H.-N., Imamura, A., Ishida, H., Groux-Degroote, S., Mühlenhoff, M., Suzuki, K.G.N., Ando, H.: Fluorescent GD2 analog for single-molecule imaging. Glycoconj J 40(2), 247–257 (2023). https://doi.org/10.1007/s10719-023-10102-1
Asano, S., Pal, R., Tanaka, H.N., Imamura, A., Ishida, H., Suzuki, K.G.N., Ando, H.: Development of fluorescently labeled SSEA-3, SSEA-4, and Globo-H glycosphingolipids for elucidating molecular interactions in the cell membrane. Int J Mol Sci 20(24) (2019). https://doi.org/10.3390/ijms20246187
Takahashi, K., Tanabe, K., Ohnuki, M., Narita, M., Ichisaka, T., Tomoda, K., Yamanaka, S.: Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131(5), 861–872 (2007). https://doi.org/10.1016/j.cell.2007.11.019
Hung, T.C., Lin, C.W., Hsu, T.L., Wu, C.Y., Wong, C.H.: Investigation of SSEA-4 binding protein in breast cancer cells. J Am Chem Soc 135(16), 5934–5937 (2013). https://doi.org/10.1021/ja312210c
Takahashi, M., Komura, N., Yoshida, Y., Yamaguchi, E., Hasegawa, A., Tanaka, H.N., Imamura, A., Ishida, H., Suzuki, K.G.N., Ando, H.: Development of lacto-series ganglioside fluorescent probe using late-stage sialylation and behavior analysis with single-molecule imaging. RSC Chem Biol 3(7), 868–885 (2022). https://doi.org/10.1039/d2cb00083k
Wikstrand, C.J., Longee, D.C., McLendon, R.E., Fuller, G.N., Friedman, H.S., Fredman, P., Svennerholm, L., Bigner, D.D.: Lactotetraose series ganglioside 3’,6’-isoLD1 in tumors of central nervous and other systems in vitro and in vivo. Cancer Res 53(1), 120–126 (1993)
Sahl, S.J., Leutenegger, M., Hilbert, M., Hell, S.W., Eggeling, C.: Fast molecular tracking maps nanoscale dynamics of plasma membrane lipids. Proc Natl Acad Sci USA 107(15), 6829–6834 (2010). https://doi.org/10.1073/pnas.0912894107
Lenne, P.F., Wawrezinieck, L., Conchonaud, F., Wurtz, O., Boned, A., Guo, X.J., Rigneault, H., He, H.T., Marguet, D.: Dynamic molecular confinement in the plasma membrane by microdomains and the cytoskeleton meshwork. Embo J 25, 3245–3256 (2006). https://doi.org/10.1038/sj.emboj.7601214
Wenger, J., Conchonaud, F., Dintinger, J., Wawrezinieck, L., Ebbesen, T.W., Rigneault, H., Marguet, D., Lenne, P.F.: Diffusion analysis within single nanometric apertures reveals the ultrafine cell membrane organization. Biophys J 92(3), 913–919 (2007). https://doi.org/10.1529/biophysj.106.096586
Kinoshita, M., Suzuki, K.G., Matsumori, N., Takada, M., Ano, H., Morigaki, K., Abe, M., Makino, A., Kobayashi, T., Hirosawa, K.M., Fujiwara, T.K., Kusumi, A., Murata, M.: Raft-based sphingomyelin interactions revealed by new fluorescent sphingomyelin analogs. J Cell Biol 216(4), 1183–1204 (2017). https://doi.org/10.1083/jcb.201607086
Kinoshita, M., Suzuki, K.G.N., Murata, M., Matsumori, N.: Evidence of lipid rafts based on the partition and dynamic behavior of sphingomyelins. Chem Phys Lipids 215, 84–95 (2018). https://doi.org/10.1016/j.chemphyslip.2018.07.002
Fujiwara, T., Ritchie, K., Murakoshi, H., Jacobson, K., Kusumi, A.: Phospholipids undergo hop diffusion in compartmentalized cell membrane. J Cell Biol 157(6), 1071–1081 (2002). https://doi.org/10.1083/jcb.200202050
Murase, K., Fujiwara, T., Umemura, Y., Suzuki, K., Iino, R., Yamashita, H., Saito, M., Murakoshi, H., Ritchie, K., Kusumi, A.: Ultrafine membrane compartments for molecular diffusion as revealed by single molecule techniques. Biophys J 86(6), 4075–4093 (2004). https://doi.org/10.1529/biophysj.103.035717
Fujiwara, T.K., Iwasawa, K., Kalay, Z., Tsunoyama, T.A., Watanabe, Y., Umemura, Y.M., Murakoshi, H., Suzuki, K.G., Nemoto, Y.L., Morone, N., Kusumi, A.: Confined diffusion of transmembrane proteins and lipids induced by the same actin meshwork lining the plasma membrane. Mol Biol Cell 27(7), 1101–1119 (2016). https://doi.org/10.1091/mbc.E15-04-0186
Umemura, Y.M., Vrljic, M., Nishimura, S.Y., Fujiwara, T.K., Suzuki, K.G., Kusumi, A.: Both MHC class II and its GPI-anchored form undergo hop diffusion as observed by single-molecule tracking. Biophys J 95(1), 435–450 (2008). https://doi.org/10.1529/biophysj.107.123018
Suzuki, K., Ritchie, K., Kajikawa, E., Fujiwara, T., Kusumi, A.: Rapid hop diffusion of a G-protein-coupled receptor in the plasma membrane as revealed by single-molecule techniques. Biophys J 88(5), 3659–3680 (2005). https://doi.org/10.1529/biophysj.104.048538
Morone, N., Fujiwara, T., Murase, K., Kasai, R.S., Ike, H., Yuasa, S., Usukura, J., Kusumi, A.: Three-dimensional reconstruction of the membrane skeleton at the plasma membrane interface by electron tomography. J Cell Biol 174(6), 851–862 (2006). https://doi.org/10.1083/jcb.200606007
Kusumi, A., Suzuki, K., Koyasako, K.: Mobility and cytoskeletal interactions of cell adhesion receptors. Curr Opin Cell Biol 11(5), 582–590 (1999). https://doi.org/10.1016/s0955-0674(99)00020-4
Kusumi, A., Koyama-Honda, I., Suzuki, K.: Molecular dynamics and interactions for creation of stimulation-induced stabilized rafts from small unstable steady-state rafts. Traffic 5(4), 213–230 (2004). https://doi.org/10.1111/j.1600-0854.2004.0178.x
Kusumi, A., Shirai, Y.M., Koyama-Honda, I., Suzuki, K.G., Fujiwara, T.K.: Hierarchical organization of the plasma membrane: investigations by single-molecule tracking vs. fluorescence correlation spectroscopy. Febs Lett 584(9), 1814–1823 (2010). https://doi.org/10.1016/j.febslet.2010.02.047
Kusumi, A., Suzuki, K.G., Kasai, R.S., Ritchie, K., Fujiwara, T.K.: Hierarchical mesoscale domain organization of the plasma membrane. Trends Biochem Sci 36(11), 604–615 (2011). https://doi.org/10.1016/j.tibs.2011.08.001
Ritchie, K., Shan, X.Y., Kondo, J., Iwasawa, K., Fujiwara, T., Kusumi, A.: Detection of non-Brownian diffusion in the cell membrane in single molecule tracking. Biophys J 88(3), 2266–2277 (2005). https://doi.org/10.1529/biophysj.104.054106
Suzuki, K.G., Kasai, R.S., Hirosawa, K.M., Nemoto, Y.L., Ishibashi, M., Miwa, Y., Fujiwara, T.K., Kusumi, A.: Transient GPI-anchored protein homodimers are units for raft organization and function. Nat Chem Biol 8(9), 774–783 (2012). https://doi.org/10.1038/nchembio.1028
Suzuki, K.G., Kasai, R.S., Fujiwara, T.K., Kusumi, A.: Single-molecule imaging of receptor-receptor interactions. Methods Cell Biol 117, 373–390 (2013). https://doi.org/10.1016/b978-0-12-408143-7.00020-7
Suzuki, K.G.: New insights into the organization of plasma membrane and its role in signal transduction. Int Rev Cell Mol Biol 317, 67–96 (2015). https://doi.org/10.1016/bs.ircmb.2015.02.004
Suzuki, K.G.: Single-Molecule Imaging of Signal Transduction via GPI-Anchored Receptors. Methods Mol Biol 1376, 229–238 (2016). https://doi.org/10.1007/978-1-4939-3170-5_19
Tiwari, S.S., Shirai, Y.M., Nemoto, Y.L., Kojima, K., Suzuki, K.G.N.: Native prion protein homodimers are destabilized by oligomeric amyloid β 1–42 species as shown by single-molecule imaging. NeuroReport 29(2), 106–111 (2018). https://doi.org/10.1097/wnr.0000000000000916
Suzuki, K.G., Fujiwara, T.K., Sanematsu, F., Iino, R., Edidin, M., Kusumi, A.: GPI-anchored receptor clusters transiently recruit Lyn and G alpha for temporary cluster immobilization and Lyn activation: single-molecule tracking study 1. J Cell Biol 177(4), 717–730 (2007). https://doi.org/10.1083/jcb.200609174
Suzuki, K.G., Fujiwara, T.K., Edidin, M., Kusumi, A.: Dynamic recruitment of phospholipase C gamma at transiently immobilized GPI-anchored receptor clusters induces IP3-Ca2+ signaling: single-molecule tracking study 2. J Cell Biol 177(4), 731–742 (2007). https://doi.org/10.1083/jcb.200609175
Suzuki, K.G.: Lipid rafts generate digital-like signal transduction in cell plasma membranes. Biotechnol J 7(6), 753–761 (2012). https://doi.org/10.1002/biot.201100360
Kusumi, A., Fujiwara, T.K., Chadda, R., Xie, M., Tsunoyama, T.A., Kalay, Z., Kasai, R.S., Suzuki, K.G.: Dynamic organizing principles of the plasma membrane that regulate signal transduction: commemorating the fortieth anniversary of Singer and Nicolson’s fluid-mosaic model. Annu Rev Cell Dev Biol 28, 215–250 (2012). https://doi.org/10.1146/annurev-cellbio-100809-151736
Koyama-Honda, I., Fujiwara, T.K., Kasai, R.S., Suzuki, K.G.N., Kajikawa, E., Tsuboi, H., Tsunoyama, T.A., Kusumi, A.: High-speed single-molecule imaging reveals signal transduction by induced transbilayer raft phases. J Cell Biol 219(12) (2020). https://doi.org/10.1083/jcb.202006125
Raghupathy, R., Anilkumar, A.A., Polley, A., Singh, P.P., Yadav, M., Johnson, C., Suryawanshi, S., Saikam, V., Sawant, S.D., Panda, A., Guo, Z., Vishwakarma, R.A., Rao, M., Mayor, S.: Transbilayer lipid interactions mediate nanoclustering of lipid-anchored proteins. Cell 161(3), 581–594 (2015). https://doi.org/10.1016/j.cell.2015.03.048
Saha, S., Das, A., Patra, C., Anilkumar, A.A., Sil, P., Mayor, S., Rao, M.: Active emulsions in living cell membranes driven by contractile stresses and transbilayer coupling. Proc Natl Acad Sci USA 119(30), e2123056119 (2022). https://doi.org/10.1073/pnas.2123056119
Arumugam, S., Schmieder, S., Pezeshkian, W., Becken, U., Wunder, C., Chinnapen, D., Ipsen, J.H., Kenworthy, A.K., Lencer, W., Mayor, S., Johannes, L.: Ceramide structure dictates glycosphingolipid nanodomain assembly and function. Nat Commun 12(1), 3675 (2021). https://doi.org/10.1038/s41467-021-23961-9
Acknowledgements
This work was supported in part by Japan Society for the Promotion of Science (JSPS) KAKENHI Grant Nos. JP21H02424 (K. G. N. S.), JP20K21387 (K. G. N. S.), JP20K15412 (N. K.), JP18H0392 (H. A.); JSPS Core-to-Core Program Grant No. JPJSCCA20200007 (H.A.); Japan Science and Technology Agency (JST) CREST “Elucidation of Biological Mechanisms of Extracellular Fine Particles and the Control System” Grant No. JPMJCR18H2 (K. G. N. S. and H. A.); JST FOREST Grant No. JPMJFR2004 (N.K.); the Takeda Foundation (K. G. N. S. and H. A.); the Mizutani foundation for Glycoscience (K. G. N. S. and H. A.); SUNBOR Grant from the Suntory Foundation for Life Sciences (N. K.).
Funding
This work was supported by the funding organizations listed in the Acknowledgments
Author information
Authors and Affiliations
Contributions
All authors wrote the paper, and participated in the revisions.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Ethics approval
This work does not include any studies involving humans or animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Suzuki, K.G.N., Komura, N. & Ando, H. Recently developed glycosphingolipid probes and their dynamic behavior in cell plasma membranes as revealed by single-molecule imaging. Glycoconj J 40, 305–314 (2023). https://doi.org/10.1007/s10719-023-10116-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-023-10116-9