Abstract
Two subspecies of common reed (Phragmites australis; Poaceae) exist in northern North America: the native P. australis subsp. americanus and the introduced P. australis subsp. australis. There are numerous native populations in Wisconsin, in addition to the more recently established populations of the introduced subspecies. We studied populations of both P. australis subspecies across Wisconsin in order to characterize the genetic diversity of both subspecies and to investigate whether any instances of hybridization could be ascertained in Wisconsin. Using eight microsatellite markers, we found minimal overlap in the alleles that could be recovered from native and introduced plants, and we found no evidence to suggest hybridization between subspecies, even in localities where native and introduced plants grow in close proximity. Overall, we found greater genetic variation in plants of the introduced subspecies relative to the native subspecies, and we observed some geographic patterns of allelic diversity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasive plants disrupt natural ecology in many ways, including by outcompeting native plants and altering trophic relationships (Pyšek et al. 2012; Gallardo et al. 2016). Among the most widespread and ecologically impactful plant invaders in North America is the common reed, Phragmites australis (Cav.) Trin. ex Steud. Two subspecies co-occur in northern North America: the native P. australis subsp. americanus Saltonst., Peterson & Soreng and the introduced P. australis subsp. australis (Saltonstall et al. 2004). The introduced subspecies has been implicated as a noxious weed in Canada and the USA, and in the USA the expansion of introduced plants has contributed to the decline of the native subspecies (Marks et al. 1994; Saltonstall 2002, 2011; Saltonstall and Stevenson 2007; Park and Blossey 2008; Catling and Mitrow 2011; Kettenring et al. 2011; Mozdzer and Megonigal 2012). Phragmites populations must be identified correctly, because the management objectives are entirely different for each subspecies (i.e., conserving native populations versus controlling or eradicating introduced populations; Marks et al. 1994; Kiviat 2013; Hazelton et al. 2014). Although morphological characters are fairly reliable for distinguishing the two Phragmites subspecies (Saltonstall et al. 2004), molecular tools can provide additional confidence in identification (Saltonstall 2002, 2003a, b). Microsatellite markers are arguably the most effective molecular method for characterizing Phragmites populations and have been used to investigate genetic diversity at a variety of geographic scales (e.g., Saltonstall 2003b; Kettenring and Mock 2012; Lambertini et al. 2012; Albert et al. 2015; McCormick et al. 2016).
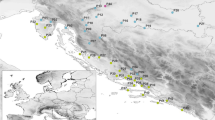
In Wisconsin (USA), Phragmites populations occur most commonly near Lake Michigan, where the density of the introduced subspecies also is greatest (Fig. 1; Granberg and Woods 2017). The native subspecies, in contrast, grows rather sparsely in the majority of Wisconsin counties. Molecular methods for studying Phragmites have not been applied widely in Wisconsin, and these would be able to confirm morphological identifications, as well as provide information about genetic diversity. Microsatellite markers, which are variable among individuals and populations of both subspecies (Saltonstall 2003a, b; Meyerson et al. 2009; Albert et al. 2015; McCormick et al. 2016), offer the potential to estimate the overall genetic diversity of each subspecies and to evaluate geographic patterns of isolation or spread.
Sampling localities and genetic cluster groups for Phragmites plants in Wisconsin. Gray background dots indicate reported Phragmites localities (subspecies unspecified). Sampling sites for this study are indicated with site identifiers (see Supplementary Table 1), and circle size is proportional to the number of genetically distinct individuals that were sampled in each locality. Each locality has a pie chart showing the proportion of individuals that were assigned to each of six genetic cluster groups. Cluster groups ‘A’ and ‘E’ correspond to the native subspecies, and groups ‘B’, ‘C’, ‘D’, and ‘F’ represent the introduced subspecies
Although the two Phragmites subspecies are closely related and wind-pollinated (Ishii and Kadono 2002; Lambert et al. 2007; McCormick et al. 2010), there are relatively few examples of hybridization between them. Hybrids have been induced under controlled conditions (Meyerson et al. 2009) and detected in a small number of natural populations (Paul et al. 2010; Saltonstall et al. 2014, 2016; Wu et al. 2015). Phragmites hybrids exhibit intermediate morphological traits, thereby confounding their identification using morphological characters (Saltonstall et al. 2014). Moreover, hybrids cannot be verified using molecular data from the uniparentally inherited chloroplast and instead must be confirmed using nuclear DNA data such as microsatellites (Saltonstall et al. 2014). In other plant invasions, hybridization has been a driving force enabling the spread of non-native taxa and their displacement of native, congeneric taxa (Thompson 1991; Ellstrand and Schierenbeck 2000; Moody and Les 2007). Genetically, hybridization offers the potential to produce novel gene combinations that might increase competitive ability or broaden environmental tolerances (Ellstrand and Schierenbeck 2000; Saltonstall et al. 2014). Hybridization between the two Wisconsin Phragmites subspecies could accelerate the rate of invasion, and a molecular survey of populations using microsatellite markers would be able to detect hybrids, if they exist.
Phragmites plants have been studied extensively in North America, and many eastern USA populations of the native subspecies have effectively been replaced by adventive populations of the introduced subspecies (Saltonstall 2002). Wisconsin offers a rare opportunity to study an ongoing Phragmites invasion, where the introduced populations are relatively few and geographically restricted, whereas the native populations are fairly numerous and widespread. In order to understand more fully the extent of the Phragmites invasion and the current status of the native subspecies in Wisconsin, we sampled populations across the state and analyzed them using microsatellite markers. We set out to ascertain the genetic diversity within and among populations of both subspecies, and to determine whether there was evidence for interspecific hybridization.
Methods
Phragmites samples were obtained from 73 localities in Wisconsin and adjacent states (Fig. 1; Supplementary Table 1). In larger stands, samples were collected from multiple individuals, and these were separated from one another by at least 10 m to reduce the chance of gathering multiple samples from the same genet. The number of samples taken at each locality was roughly proportional to the number of individuals in the stand, up to a maximum of 21 samples. Voucher specimens were deposited in the UW-Whitewater herbarium (UWW). Plants were preliminarily identified on the basis of morphological characters (Saltonstall et al. 2004), and identifications were later confirmed using molecular data from DNA sequencing and microsatellite fragment analysis (see below). Fresh leaf material was kept in a freezer (− 20 °C) or in liquid preservative (Rogstad 1992) until DNA extraction. DNA was extracted using a previously reported protocol (Doyle and Doyle 1987), adjusted to extract from approximately 1 cm2 of leaf blade tissue using an initial buffer volume of 600 µl, and modified to use pure chloroform (without isoamyl alcohol) at the extraction step and ethanol in place of isopropanol at the DNA precipitation step.
Plastid DNA sequences were obtained for the trnT-trnL spacer and trnL intron, which have been used previously to identify Phragmites subspecies, using the ‘a’, ‘b’, ‘c’, and ‘d’ primers (Taberlet et al. 1991), following methods reported previously (Tippery et al. 2018). We sequenced plastid DNA for a subset of individuals from both subspecies, to compare against morphological identifications and microsatellite results. For the microsatellite regions, we initially attempted to amplify the ten markers that were reported by Saltonstall (2003a, b, using the same primers reported in that study. When necessary, we substituted primers that were developed in a subsequent study (Meyerson et al. 2009). Forward primers were modified by appending an oligonucleotide ‘tail’ that served to incorporate fluorescently labeled oligonucleotides using the M13 (FAM fluorophore) or M13A (HEX fluorophore) sequence (Culley et al. 2013). After initial primer screening, we were able to use eight primer sets to collect data (see “Results”).
Polymerase chain reaction (PCR) was conducted separately with each primer set on each sample. Each 10 µl reaction contained 10 ng DNA, 1 µl buffer, 0.15 mM dNTP, 64 nM tailed forward primer, 200 nM fluorescent tail oligonucleotide, 200 nM reverse primer, and 0.25 units Titanium® Taq (Takara Bio Inc., Kusatsu, Shiga, Japan) or AmpliTaq Gold™ polymerase (Thermo Fisher Scientific). PCR reactions were carried out using a Bio-Rad S1000 Thermal Cycler using the following program: 95 °C for 3 min, followed by 40 cycles of 95 °C for 30 s, 55 °C/58° 60 °C (depending on the primer used) for 30 s, 68 °C for 30 s (microsatellites) or 60 s (plastid DNA), with a final extension phase at 68 °C for 3 min. Plastid DNA amplicons were cleaned using the ExoSAP-IT enzyme mix (Thermo Fisher Scientific), then Sanger sequenced (Sanger et al. 1977) using the BigDye® v3.1 reaction mix, with reactions run on an ABI 3730xl DNA Analyzer (Applied Biosystems).
Microsatellite reactions were randomized into positions on 96-well plates to increase our ability to detect cross-contamination. In addition, two out of every 16 PCR reactions used deionized water (negative control) and a Phragmites DNA standard (positive control), respectively, for their templates. A subset of samples from PCR reactions were run on a 1% agarose gel to confirm amplification. The amplified samples were each diluted 1:10 in water, then 1.0 µl of the dilution was combined with 16 µl of Hi-Di™ formamide (Thermo Fisher Scientific) and 0.08 µl of GeneScan™ 400HD ROX™ or 500 LIZ™ (Thermo Fisher Scientific) size standard in preparation for microsatellite analysis. Fragments were analyzed on an ABI 3730xl DNA Analyzer (Applied Biosystems). Fragment sequences for at least one individual of each subspecies were verified against published sequences for each of the microsatellite markers using Sanger sequencing as described above.
Fragment sizes were scored in Geneious® ver. 6.1.8 (Biomatters Ltd.) or the package Fragman ver. 1.0.9 (Covarrubias-Pazaran et al. 2016) in the R statistical environment (R Development Core Team 2019) and exported as allele data for further analysis. Phragmites individuals of both subspecies in northern North America are predominantly tetraploid (Clevering and Lissner 1999; Lambertini et al. 2006), and the presumed tetraploidy of samples in our study prevented effective analyses of heterozygosity and allele transmission (Dufresne et al. 2014). Instead, we evaluated the presence/absence of codominant allelic markers (with presence indicating at least one copy of the allele in question) and recorded the number of alleles (i.e., allelic richness) found in each locality. No more than two alleles were recovered per individual, so the allele data were analyzed like diploid genotypes, as has been done previously for Phragmites (Saltonstall 2003a, b).
Potential clonal genotypes were removed using the clonecorrect function of the poppr package ver. 2.8.6 in R (Kamvar et al. 2014), with individuals stratified by locality (i.e., only one individual of each distinct multilocus genotype was retained for each locality). Next, individuals were binned into six cluster groups using the find.clusters function of the adegenet package ver. 2.1.3 in R (Jombart 2008; Jombart et al. 2010). Clone-corrected genotypes were also evaluated using principal components analysis (PCA) via the dudi.pca function in the R package ade4 ver. 1.7.15 (Thioulouse et al. 1997; Dray and Dufour 2007). Correlations were tested using a linear regression analysis (lm function), and differences were evaluated using a Welch t-test (t.test function; Welch 1947), both in R. When investigating the proportion of presumed clones (i.e., individuals with identical multilocus genotypes to other individuals at the same locality), localities with n = 1 sampled individual were excluded.
Individuals also were clustered using the snmf function in the R package LEA (Frichot and François 2015), which assigns individuals to one or more hypothetical ancestral populations using sparse non-negative matrix factorization, similarly to the method used by the program STRUCTURE (Pritchard et al. 2000; Falush et al. 2003). After examining a range of ancestral population values from 1 to 10, the entropy criterion showed a substantial decrease at K = 2 populations, followed by gradual decreases thereafter. Because we were using the analysis primarily to evaluate the distinctness of the two Phragmites subspecies, we ran the final analysis with K = 2 populations, with ten repetitions, using the repetition with the lowest cross-entropy value.
Results
A total of 202 P. australis subsp. americanus and 346 P. australis subsp. australis individuals were tested, across 29 counties in Wisconsin and adjacent states (Supplementary Table 1; Fig. 1). DNA sequences for the trnT-trnL spacer and trnL intron were obtained for six individuals of the introduced subspecies and eight individuals of the native subspecies (GenBank accession numbers MT316423–MT316436). These were identical, respectively, to previously published sequences for each subspecies (Saltonstall 2003a, b). Initial morphological identifications were consistent, in every case, with DNA sequence and microsatellite data.
Of the ten microsatellite primer sets that were attempted, six (PaGT04, PaGT08, PaGT11, PaGT13, PaGT14, and PaGT16) were amplified successfully using the original primer sequences (Saltonstall 2003a, b), and an additional two (PaGT09 and PaGT12) were amplified using modified primers (Meyerson et al. 2009). Nucleotide sequences for each microsatellite marker (i.e., the GT dinucleotide repeat and adjacent sequence) were identical to previously published sequences for each respective subspecies (GenBank accession numbers MT316437–MT316457; Saltonstall 2003b). Two primers (PaGT21 and PaGT22; Saltonstall, 2003b) failed to amplify or failed to produce amplification products that were comparable to those previously reported, and thus were excluded from further analysis. The ‘tailed’ primer method (Culley et al. 2013) successfully produced PCR products that could be detected by fragment analysis, and the fragments amplified using this method were determined to be 16–18 bp longer than the corresponding alleles from prior studies (Saltonstall 2003a, b; Meyerson et al. 2009; Table 1). Meyerson et al. (2009) were determined to be 20 bp longer (PaGT09) and 185 bp longer (PaGT12) than those reported by Saltonstall (2003a, b for the same markers. Missing microsatellite data amounted to 18.5% for P. australis subsp. americanus (range 5.4–42.9% per marker) and 10.0% for P. australis subsp. australis (range 4.3–25.1%).
Alleles for both Phragmites subspecies in Wisconsin were recovered largely at similar frequencies to what Saltonstall (2003a, b) reported previously (Table 1). In most cases the most abundant allele for a subspecies was the same as what had been observed previously. A notable exception was the PaGT12 marker for P. australis subsp. americanus, which included majority alleles 394 and 396 that were not prominent in other studies. Individual marker variation and geographic distributions of allele data are provided as supplementary data. The number of sampled plants per native locality ranged from 1 to 16 (\(\bar{x}\) = 5.46), exhibiting 1–6 (\(\bar{x}\) = 1.78) distinct genotypes (Supplmentary Table 1). In the introduced subspecies, 1–21 (\(\bar{x}\) = 10.18) plants were sampled per locality, and 1–15 (\(\bar{x}\) = 5.21) distinct genotypes were recovered from each locality. Overall, 136 of 202 native individuals (67%) were potentially clones of other individuals collected at the same locality, and for introduced individuals this number was 169 of 346 (49%). Larger stands (i.e., with more individuals sampled) did not have significantly smaller or larger proportions of presumed clones (R2 = 0.044, F(1,64) = 2.976, p = .089). However, the mean proportion of presumed clones per locality was significantly larger for the native subspecies (\(\bar{x}\) = 0.86) than it was for the introduced subspecies (x̄ = 0.58) (t(52.11) = 4.801, p < 10−4).
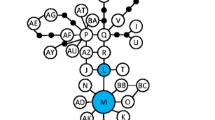
We found the overall genetic diversity to be substantially greater in introduced Phragmites plants than in native plants. Sampled native plants had a mean of 2.0 alleles per marker, whereas this value was 3.5 for introduced plants. In comparison, Saltonstall (2003a, b) reported means of 5.9 alleles per marker for the native subspecies and 6.5 for the introduced subspecies, in a study across the native and introduced ranges of P. australis. Moreover, two microsatellite markers (PaGT04, PaGT08) were entirely homozygous in all native stands studied, and another four markers (PaGT09, PaGT11, PaGT13, PaGT14) were homozygous in over 90% of native individuals (Table 1). Detailed locality information for genotypes was not available from several other studies, so we could not directly compare our data to another study of geographically proximate populations. Individuals of the native subspecies were assigned via find.clusters to cluster groups ‘A’ (n = 123) and ‘E’ (n = 79), whereas individuals of the introduced subspecies were assigned to clusters ‘B’ (n = 95), ‘C’ (n = 104), ‘D’ (n = 81), and ‘F’ (n = 66) (Fig. 1). Each cluster group contained individuals of only one subspecies. The principal components analysis produced two groups, differentiated along the PC1 axis and corresponding to the two subspecies, with no overlap between subspecies groups and no points assigned to an intermediate position (Fig. 2). The snmf analysis cleanly divided individuals into two groups according to their subspecies (Fig. 3). Only 20 individuals (all of the introduced subspecies) were assigned to a group with less than 0.90 ancestry proportion.
Principal components analysis of clone-corrected data for 176 introduced and 66 native Phragmites australis individuals, derived using genotype data from eight microsatellite loci. PC1 explained 54% of the variation in the data and PC2 explained 11% of the variation in the data. Points are colored according to subspecies, with the introduced subspecies in shades of gray and blue (left side of image), and the native subspecies in shades of orange (right side of image)
Plot of ancestry proportions (i.e., assignment of individuals to one of two hypothetical populations) for all sampled individuals. Individuals are grouped by subspecies, with native individuals at the left-hand side and introduced individuals at the right-hand side. Each individual corresponds to a vertical bar, with colors indicating the proportion of ancestry estimated to be native (orange) or introduced (gray)
Discussion
Introduced Phragmites plants in Wisconsin exhibit higher genetic diversity than plants of the native subspecies, consistent with the higher diversity that was found for this subspecies across North America (Saltonstall 2003a, b). Introduced taxa typically undergo a population bottleneck as they migrate to their adventive ranges, because few individuals are introduced and the starting populations are small with substantial inbreeding (Dlugosch and Parker 2008). However, higher genetic diversity could be achieved in a sexually reproducing taxon that was introduced as a large number of genetically diverse individuals, either through a large initial introduction or repeated introductions from a diverse source population (Roman and Darling 2007; Estoup et al. 2016). Saltonstall (2002) reported that introduced Phragmites plants in northern North America share the same plastid haplotype, also found in plants throughout the P. australis subsp. australis native range. The microsatellite diversity of the introduced subspecies in North America is substantial and suggests that these plants can draw from a broad stock of genetic material as they adapt to novel environments.
In general, a long-established native taxon would be expected to harbor greater variation as a result of accumulating genetic mutations over time and maintaining them through interbreeding among relatively large metapopulations (e.g., Tsai and Manos 2010; Roberts and Hamann 2015). However, Wisconsin habitats are predominantly postglacial, and native Phragmites populations would have recolonized most of the state in relatively recent evolutionary time (Curtis 1959). Recolonizing postglacial populations typically exhibit lower genetic diversity than would be found in longer-established populations, resulting from founder effects in relatively recent time (e.g., Comps et al. 2001; Cheddadi et al. 2006). One point of comparison for Wisconsin may be the Driftless region, an area that was not glaciated during the most recent Ice Age (Clayton et al. 2006). In our study, however, native Phragmites plants from the Driftless region (i.e., localities 45–47 and 50) had comparable genotypes and were assigned to the same cluster groups as plants in other parts of Wisconsin (Fig. 1; Supplementary Table 1).
Phragmites plants are capable of extensive clonal growth using underground rhizomes, and this mechanism of reproduction would be expected to generate large patches of plants that are genetically uniform. We attempted to avoid gathering multiple data points from the same clone by sampling plants that were separated by at least 10 m, however this method could not guarantee that genetically distinct plants (i.e., genets) were sampled. Using the microsatellite marker data, we can estimate the degree to which plants are clonal, by identifying unique genetic signatures, i.e., allelic combinations not found in other plants from the same population. We found a wide range of genetically distinct individuals across the sampled localities, with the introduced subspecies generally having a higher proportion of genetically distinct individuals (Supplementary Table 1). Sampling a larger number of plants in a stand allows for the most thorough estimation of genetic diversity, and we recovered up to 15 genetically distinct individuals in stands of the introduced subspecies. In contrast, all but two stands of the native subspecies had three or fewer distinct genotypes. A comparable study of Phragmites plants in Utah reported several native populations with greater than ten distinct genotypes, and in the introduced subspecies most populations had 15–20 distinct genotypes (Kettenring and Mock 2012). The contrast between the two subspecies also reflects the noted difference in their allelic diversity, because a lower diversity of alleles would limit the number of possible multilocus genotypes.
We detected five alleles (PaGT08: 194, PaGT11: 159, PaGT12: 379, PaGT13: 226, and PaGT14: 202) that were present in both the native and introduced Phragmites subspecies (Table 1), but only two of these were arranged in identical genotypes in both subspecies (PaGT08: 194/194, PaGT13: 226/226). All but two such alleles (PaGT11: 159 and PaGT14: 202) were found previously in both subspecies (Saltonstall 2003a, b; Table 1). All individuals that had less than 0.90 ancestry proportion in the snmf analysis (Fig. 3) also possessed one or more of the alleles and single-locus genotypes that occur in both subspecies. Ambiguous ancestry assignment may reflect intersubspecific hybridization (e.g., Wu et al. 2015); however, in our study the fact that ambiguously assigned individuals all possessed alleles that were not specific to one subspecies decreases the likelihood of this possibility. Moreover, the genotypes of ambiguously assigned individuals at three markers (PaGT09, PaGT14, PaGT16) consisted entirely of alleles that otherwise are found only in the introduced subspecies.
Intersubspecific hybridization has been demonstrated several times in Phragmites using principal components analysis, which depicts hybrids as occupying intermediate positions between otherwise differentiated subspecies (Paul et al. 2010; Saltonstall et al. 2014, 2016). There were no individuals in our study group that showed patterns that we would interpret as evidence for interspecific hybridization (Fig. 2). Hybridization has been documented in North American Phragmites using morphological data (Williams et al. 2019) and microsatellite markers (Paul et al. 2010; Saltonstall et al. 2014, 2016; Wu et al. 2015). Additionally, some hybrid plants also exhibit heteroplasmy that manifests as sequence and length variation in plastid loci (Lambertini 2016). We observed no patterns in Wisconsin Phragmites that resembled previously reported genetic data for Phragmites intersubspecific hybrids.
In conclusion, the most substantial results of our study were that (1) we found no evidence for hybridization between native and introduced Phragmites and (2) genetic variation was substantially greater in introduced plants than in native plants. Phragmites plants are capable of both sexual and asexual reproduction, and it will be important for land managers to guard against the potential for hybridization between subspecies. Even without hybridization, the continued sexual reproduction of introduced plants represents a persistent source of genetically variable propagules that may establish as novel aggressive variants of the subspecies. Introduced Phragmites plants have persisted in North America for over 100 years, and they have continued to expand their range over that time (Saltonstall 2002). It will be important to remain vigilant to prevent the establishment of new introduced populations in Wisconsin. To expand upon the results we obtained, it would be worthwhile to investigate genetic patterns in Phragmites populations over a greater geographic area, potentially to uncover evidence for multiple introductions or ecological specialization. In addition, it would be valuable to increase sampling throughout the state, in order to construct a more complete picture of the genetic diversity in native and introduced Phragmites. Collecting more data on the current spread and genetic diversity of Phragmites also could provide a useful benchmark for future studies, in the event that introduced Phragmites plants continue to expand their range.
References
Albert A, Brisson J, Belzile F, Turgeon J, Lavoie C (2015) Strategies for a successful plant invasion: the reproduction of Phragmites australis in north-eastern North America. J Ecol 103:1529–1537
Catling PM, Mitrow G (2011) The recent spread and potential distribution of Phragmites australis subsp. australis in Canada. Can Field 125:95–104
Cheddadi R, Vendramin GG, Litt T, François L, Kageyama M, Lorentz S, Laurent JM, de Beaulieu JL, Sadori L, Jost A, Lunt D (2006) Imprints of glacial refugia in the modern genetic diversity of Pinus sylvestris. Global Ecol Biogeogr 15:271–282
Clayton L, Attig JW, Mickelson DM, Johnson MD, Syverson KM (2006) Glaciation of Wisconsin, 3rd edn. Wisconsin Geological and Natural History Survey, Madison
Clevering OA, Lissner J (1999) Taxonomy, chromosome numbers, clonal diversity and population dynamics of Phragmites australis. Aquat Bot 64:185–208
Comps B, Gömöry D, Letouzey J, Thiébaut B, Petit RJ (2001) Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics 157:389–397
Covarrubias-Pazaran G, Diaz-Garcia L, Schlautman B, Salazar W, Zalapa J (2016) Fragman: an R package for fragment analysis. BMC Genet 17:1–8
Culley TM, Stamper TI, Stokes RL, Brzyski JR, Hardiman NA, Klooster MR, Merritt BJ (2013) An efficient technique for primer development and application that integrates fluorescent labeling and multiplex PCR. Appl Plant Sci 1:1300027
Curtis JT (1959) The vegetation of Wisconsin. The University of Wisconsin Press, Madison
Dlugosch KM, Parker IM (2008) Founding events in species invasions: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17:431–449
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Dray S, Dufour A (2007) The ade4 package: implementing the duality diagram for ecologists. J Stat Softw 22:1–20
Dufresne F, Stift M, Vergilino R, Mable BK (2014) Recent progress and challenges in population genetics of polyploid organisms: an overview of current state-of‐the‐art molecular and statistical tools. Mol Ecol 23:40–69
Ellstrand NC, Schierenbeck KA (2000) Hybridization as a stimulus for the evolution of invasiveness in plants? Proc Natl Acad Sci USA 97:7043–7050
Estoup A, Ravigné V, Hufbauer R, Vitalis R, Gautier M, Facon B (2016) Is there a genetic paradox of biological invasion? Ann Rev Ecol Evol Syst 47:51–72
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Frichot E, François O (2015) LEA: an R package for landscape and ecological association studies. Meth Ecol Evol 6:925–929
Gallardo B, Clavero M, Sánchez MI, Vilà M (2016) Global ecological impacts of invasive species in aquatic ecosystems. Glob Change Biol 22:151–163
Granberg JE, Woods B (2017) Control of non-native Phragmites within the Great Lakes basins: a case study in invasive species strategic planning and implementation. Presented at the 39th annual Wisconsin Lakes Partnership Convention, Stevens Point, Wisconsin, 6 April 2017
Hazelton ELG, Mozdzer TJ, Burdick DM, Kettenring KM, Whigham DF (2014) Phragmites australis management in the United States: 40 years of methods and outcomes. AoB Plants 6:plu001
Ishii J, Kadono Y (2002) Factors influencing seed production of Phragmites australis. Aquat Bot 72:129–141
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281
Kettenring KM, Mock KE (2012) Genetic diversity, reproductive mode, and dispersal differ between the cryptic invader, Phragmites australis, and its native conspecific. Biol Invasions 14:2489–2504
Kettenring KM, McCormick MK, Baron HM, Whigham DF (2011) Mechanisms of Phragmites australis invasion: feedbacks among genetic diversity, nutrients, and sexual reproduction. J Appl Ecol 48:1305–1313
Kiviat E (2013) Ecosystem services of Phragmites in North America with emphasis on habitat functions. AoB Plants 5:plt008
Lambert AM, Casagrande RA (2007) Characteristics of a successful estuarine invader: evidence of self-compatibility in native and non-native lineages of Phragmites australis. Mar Ecol Prog Ser 337:299–301
Lambertini C (2016) Heteroplasmy due to chloroplast paternal leakage: another insight into Phragmites haplotypic diversity in North America. Biol Invasions 18:2443–2455
Lambertini C, Gustafsson MHG, Frydenberg J, Lissner J, Speranza M, Brix H (2006) A phylogeographic study of the cosmopolitan genus Phragmites (Poaceae) based on AFLPs. Plant Syst Evol 258:161–182
Lambertini C, Sorrell BK, Riis T, Olesen B, Brix H (2012) Exploring the borders of European Phragmites within a cosmopolitan genus. AoB Plants pls020
Marks M, Lapin B, Randall J (1994) Phragmites australis (P. communis): threats, management and monitoring. Nat Area J 14:285–294
McCormick MK, Kettenring KM, Baron HM, Whigham DF (2010) Spread of invasive Phragmites australis in estuaries with differing degrees of development: genetic patterns, Allee effects and interpretation. J Ecol 98:1369–1378
McCormick MK, Brooks HE, Whigham DF (2016) Microsatellite analysis to estimate realized dispersal distance in Phragmites australis. Biol Invasions 18:2497–2504
Meyerson LA, Viola DV, Brown RN (2009) Hybridization of invasive Phragmites australis with a native subspecies in North America. Biol Invasions 12:103–111
Moody ML, Les DL (2007) Geographic distribution and genotypic composition of invasive hybrid watermilfoil (Myriophyllum spicatum × M. sibiricum) populations in North America. Biol Invasions 9:559–570
Mozdzer TJ, Megonigal JP (2012) Jack-and-master trait responses to elevated CO2 and N: a comparison of native and introduced Phragmites australis. PLoS ONE 7:e42794
Park MG, Blossey B (2008) Importance of plant traits and herbivory for invasiveness of Phragmites australis (Poaceae). Am J Bot 95:1557–1568
Paul J, Vachon N, Garroway CJ, Freeland JR (2010) Molecular data provide strong evidence of natural hybridization between native and introduced lineages of Phragmites australis in North America. Biol Invasions 12:2967–2973
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pyšek P, Jarošík V, Hulme PE, Pergl J, Hejda M, Schaffner U, Vilà M (2012) A global assessment of invasive plant impacts on resident species, communities and ecosystems: the interaction of impact measures, invading species’ traits and environment. Glob Change Biol 18:1725–1737
R Development Core Team (2019) R: A language and environment for statistical computing. http://www.R-project.org
Roberts DR, Hamann A (2015) Glacial refugia and modern genetic diversity of 22 western North American tree species. Proc R Soc B 282:20142903
Rogstad SH (1992) Saturated NaCl-CTAB solution as a means of field preservation of leaves for DNA analyses. Taxon 41:701–708
Roman J, Darling JA (2007) Paradox lost: genetic diversity and the success of aquatic invasions. Trends Ecol Evol 22:454–464
Saltonstall K (2002) Cryptic invasion by a non-native genotype of the common reed, Phragmites australis, into North America. Proc Natl Acad Sci USA 99:2445–2449
Saltonstall K (2003a) A rapid method for identifying the origin of North American Phragmites populations using RFLP analysis. Wetlands 23:1043–1047
Saltonstall K (2003b) Microsatellite variation within and among North American lineages of Phragmites australis. Mol Ecol 12:1689–1702
Saltonstall K (2011) Remnant native Phragmites australis maintains genetic diversity despite multiple threats. Conserv Genet 12:1027–1033
Saltonstall K, Stevenson JC (2007) The effect of nutrients on seedling growth of native and introduced Phragmites australis. Aquat Bot 86:331–336
Saltonstall K, Peterson PM, Soreng RJ (2004) Recognition of Phragmites australis subsp. americanus (Poaceae: Arundinoideae) in North America: evidence from morphological and genetic analyses. Sida 21:683–692
Saltonstall K, Castillo HE, Blossey B (2014) Confirmed field hybridization of native and introduced Phragmites australis (Poaceae) in North America. Am J Bot 101:211–215
Saltonstall K, Lambert AM, Rice N (2016) What happens in Vegas, better stay in Vegas: Phragmites australis hybrids in the Las Vegas Wash. Biol Invasions 18:2463–2474
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad of Sci USA 74:5463–5467
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Thioulouse J, Chessel D, Dolédec S, Olivier J (1997) Ade-4: a multivariate analysis and graphical display software. Stat Comput 7:75–83
Thompson JD (1991) The biology of an invasive plant. Bioscience 41:393–401
Tippery NP, Sears NL, Zentner AB, Sivadas V (2018) Evidence for allopolyploid speciation in Nymphoides (Menyanthaceae). Syst Bot 43:117–129
Tsai YHE, Manos PS (2010) Host density drives the postglacial migration of the tree parasite, Epifagus virginiana. Proc Natl Acad Sci USA 107:17035–17040
Welch BL (1947) The generalization of “Student’s” problem when several different population variances are involved. Biometrika 34:28–35
Williams J, Lambert AM, Long R, Saltonstall K (2019) Does hybrid Phragmites australis differ from native and introduced lineages in reproductive, genetic, and morphological traits? Am J Bot 106:29–41
Wu CA, Murray LA, Heffernan KE (2015) Evidence for natural hybridization between native and introduced lineages of Phragmites australis in the Chesapeake Bay watershed. Am J Bot 102:805–812
Acknowledgements
We are extremely grateful for the individuals who supplied samples: K. Anderson, C. Bohn, C. Curney, S. Davis-Faust, K. Dolata, J. Estrada, D. Feirer, S. Garske, S. Goodwin, M. Hess, S. Johnson, S. Kirby, J. Lepsch, C. MacDonald, R. Parchim, J. Scherer, I. Shackleford, R. Sharka, A. Smith, M. Sundeen, J. Wahls, C. Weber-Starling, A. Wirt, and B. Woods; K. Monson, A. Olson, and A. Vang provided laboratory assistance. We are grateful to the Wisconsin Department of Natural Resources for funding and supporting this study. B. Woods was instrumental in providing us with samples, and S. Schumacher and J. Granberg provided helpful logistical support. Additional support was provided by the University of Wisconsin-Whitewater Undergraduate Research Program and the Department of Biological Sciences, University of Wisconsin-Whitewater.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tippery, N.P., Pesch, J.D., Murphy, B.J. et al. Genetic diversity of native and introduced Phragmites (common reed) in Wisconsin. Genetica 148, 165–172 (2020). https://doi.org/10.1007/s10709-020-00098-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-020-00098-z