Abstract
Anastrepha striata is widely distributed across the Americas and is a pest of economically important crops, especially crops of the Myrtaceae family. Insect population structures can be influenced by the presence of physical barriers or characteristics associated with habitat differences. This study evaluated the effect of the Western Andes on the population structure of A. striata. Individuals were collected from Psidium guajava fruits from three natural regions of southwestern Colombia (Pacific Coast, mountainous region and the inter-Andean valley of the Cauca River). Based on a 1318 bp concatenated of the genes Cytochrome Oxidase subunit I (COI) and NADH dehydrogenase subunit 6 (ND6), 14 haplotypes with few changes among them (between 1 and 3) were found. There was only one dominant haplotype in all three regions. No genetic structure associated with the three eco-geographical regions of the study was found. Moreover, the Western Andes are not an effective barrier for the genetic isolation of the populations from the Pacific Coast compared with the inter-Andean valley populations. This genetic homogeneity could be partially due to anthropogenic intervention, which acts as a dispersal agent of infested fruits. Another hypothesis to explain the lack of structure would be the relatively recent arrival of A. striata to the region, as indicated by an analysis of the demographic history, which reveals a process of population expansion. This study represents the first attempt to understand the population genetics of A. striata in Colombia and could contribute to the integral management of this pest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Anastrepha (Schiner 1868) of the Tephritidae family, is considered the most economically important genus in Latin America (Castañeda et al. 2010). Anastrepha striata (Schiner 1868), known as the American guava fruit fly, is a quarantine pest according to the USDA-APHIS-PPQ (United States Department of Agriculture, Animal and Plant Health Inspection Service, Plant Protection and Quarantine program) and other regulatory agencies (Norrbom 2003) and is one of five quarantine species in Colombia: A. fraterculus (Wiedemann), A. grandis (Macquart), A. obliqua (Macquart), A. serpentina (Wiedemann) and A. striata (Schinner) (Castañeda et al. 2010). A. striata is a multivoltine and oligophagous species (Insuasty et al. 2007) that infests approximately 37 plant species (Norrbom et al. 1999) including economically important fruits, such as mango (Mangifera indica) and coffee (Coffea arabica). However, they have a clear preference for the genus Psidium (Myrtaceae) and Psidium guajava is its main host (Aluja et al. 2000). A. striata is widely distributed in the Americas, from the southern United States (Texas) to Brazil (Hernandez-Ortiz and Aluja 1993) and has a wide altitudinal range (0-2600 masl) (Arévalo et al. 1997; Martínez and Serna 2005). Along with A. fraterculus and A. obliqua, it has been reported to be one of the most abundant species in Colombia (Nuñez 2010). According to the Corporación Colombiana de Investigación Agropecuaria, CORPOICA, A. striata is a phytosanitary problem in Colombia, causing up to 90% of fruit losses and reaching infestation levels of up to 210 larvae/kg fruit (Insuasty et al. 2007).
Despite its wide distribution and economic importance, studies of A. striata have been restricted to aspects of its biology, behavior, population dynamics and distribution (Aluja et al. 1993; Ramírez et al. 1996; Cruz-López et al. 2015). In Colombia, knowledge of the species is based on studies of its population dynamics in the department of Santander (Olarte 1980) and the diversity of its parasitoids, hosts and distribution in departments such as Tolima, Valle del Cauca and Cundinamarca (Carrejo and González 1994; Martínez and Serna 2005; Ruiz-Hurtado et al. 2013). However, there is no knowledge about its genetic diversity and population structure, which has been demonstrated to be a prerequisite for planning pest control strategies (Roderick and Navajas 2003) because of the information it provides about the insects population dynamics (Aketarawong et al. 2011). This information is important when specialized techniques for control, such as semiochemicals, including sexual or aggregation pheromones (Agelopoulos et al. 1999), or the production and dispersion of sterile insects (SIT) (Karsten et al. 2013) are required.
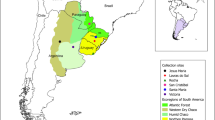
Insect population structures can be influenced by the presence of physical barriers or characteristics associated with habitat differences (Avise 1994). For example, the Andes have played an important role in the diversification of neotropical organisms by their isolation on either side of the mountains or by creating a mosaic of montane habitats and inter-Andean valleys where differentiation processes can occur (Brower 1994; Lynch 1997; Graham et al. 2001; Elias et al. 2009). Southwestern Colombia is a very important fruit-producing region, in which several soils and climates can be found (Orozco 2003). The Andes directly influence these soils and climates by allowing the formation of three well differentiated natural sub-regions: the mountain region, formed by the Western and Central Andes, is mainly characterized by the presence of Andean forests and high humidity. The Pacific Coast, delimited by the Pacific Ocean and the foothills of the West Andes, contains Andean and sub-Andean zones, rainy areas and warm and humid mangroves. The inter-Andean valley of the Cauca River, delimited by the West and Central Andes, is characterized by agricultural zones that have almost entirely replaced the dry and highly dry tropical forests (Salazar et al. 2002). In Colombia, several studies have demonstrated the importance of habitat differences and the physical barrier on the genetic flow of insect species (Díaz et al. 2013; Velasco-Cuervo et al. 2016). The population structure and genetic diversity of insects may also be strongly influenced by the invasion pathways that led to their initial establishment. For example, Karsten et al. (2015) found genetic structuring of Ceratitis capitata between the native region (Africa) and the introduced region (Australia, Greece, Guatemala and Madeira). They also found that the genetic diversity of populations was associated to the initial colonization pathways of Europe from Africa and a secondary colonization of Australia from Europe.
It is important to evaluate if the populations of A. striata in southwestern Colombia have a population structure that could be influenced by the Andes or the geographical area it is in. This information will enable us to develop more efficient pest control strategies. For example, to know if management is aimed at a wider or finer scale depending on the level of connectivity of populations.
Mitochondrial genetic markers have proven to be informative for population genetic studies due to their strictly maternal inheritance, lack of recombination, relatively high mutation rate (10-fold faster than single copy nuclear DNA) (Brown et al. 1979) and the availability of very efficient PCR primers (Shi et al. 2005; Xie et al. 2006; Hu et al. 2008). Among these markers, the gene Cytochrome Oxidase subunit I (COI) shows a high degree of polymorphism that makes it appropriate for intraspecific genetic analyses (Mun et al. 2003) and has been used in population genetics studies of different fruit fly species (Hu et al. 2008; Prabhakar et al. 2012, 2013; Meeyen et al. 2014). Although less frequently used, the NADH dehydrogenase subunit 6 (ND6) gene, has similar characteristics to COI (Simon et al. 1994) and a high degree of polymorphism. Therefore, both genes are useful for the evaluation of intraspecific diversity and the population structure of A. striata.
In this study, we used the COI and ND6 gene sequences to assess whether A. striata has a population structure associated with the Western and Central Andes in southwestern Colombia. Additionally, an analysis of the demographic history of A. striata was performed to improve the knowledge about demographic processes that could be shaping the population structure and genetic diversity patterns.
Materials and methods
Sample collection
Fruits from guava (Psidium guajava), mango (Maguifera indica), coffee (Coffea arabica), plum (Spondia purpurea) and carambola (Averrhoa carambolo) were collected in nine locations in southwestern Colombia. The locations were chosen by assuming the West Andes are a physical barrier and to represent each of the different natural regions–the Pacific Coast, the inter-Andean valley of the Cauca River and the mountain region (Table 1; Fig. 1). The specimens were obtained from either fallen fruits or from the tree in non-crop zones (roadside, home garden, small farms, etc.). In an insect-breeding room, the fruits were placed in emergence chambers built from plastic containers containing a sterile mixture of sand and sawdust, with the corresponding fruits on top. The chambers were kept at room temperature until the emergence of adults, which were then preserved in 96% ethanol and stored at −20 °C until further molecular analysis. The specimens were identified using Steyskal’s (1977) and Caraballo’s (2001) taxonomic keys. The following diagnostic features of A. striata were considered: presence of a dark brown-almost black U-shaped mark on the scutum, no connection between the V and S bands in the wing, a yellow to brown subscutellum with lateral dark bands and an aculeus with a wide, blunt and non-serrated apex. For each individual used in this study, a voucher was stored with the date, collector’s name and geographical coordinates in the Museum of Entomology at the Universidad del Valle.
DNA extraction, amplification and sequencing
DNA extraction from the head and legs of adult flies was performed with a DNeasy® Blood and Tissue Kit (QIAGEN, CA, USA) according to the manufacturer’s protocol for insects. A ~ 710 bp region of the COI gene was amplified using the primers LCO1490 (5′-GGTCAACAAATCATAAAGATATTGG-3′) and HCO2198 (5′-TAAACTTCAG GGTGACCAAAAAATCA-3′) (Folmer et al. 1994). The amplification cocktail was modified from Ruiz et al. (2010), for a total volume of 25 μL per reaction and contained 1X PCR buffer, 0.05 mM dNTPs, 2 mM MgCl2, 1 U Taq polymerase, 0.25 μM of each primer and 30 ng of DNA. The following thermal profile was used ,modified from Folmer et al. (1994): initial denaturation at 92 °C for five minutes, 35 cycles of 94 °C for 30 s, annealing at 52 °C for one minute, extension at 72 °C for one minute and a final extension at 72 °C for five minutes. A ~ 725 bp fragment of the ND6 gene was amplified with the primers TT-J-9886 (5′-TAAAAACATTGGTCTTGTAA-3′) (Barr et al. 2006) and ND6r (5′-TTATGATCCAAAATTTCATCA-3′) (Ruiz-Arce et al. 2012) using the protocol proposed by Ruiz-Arce et al. (2012). Amplification of the ND6 and COI genes was confirmed using a 1.5% agarose gel with 0.5X TBE buffer. Finally, the PCR products were sequenced by a specialized service (MACROGEN, INC., MD, USA).
Data analysis
The sequences obtained from COI and ND6 were manually edited with Sequencher 4.1.4 (Gene Codes Corporation, Ann Arbor, MI, USA) to obtain final lengths of 658 and 660 bp, respectively. A manual concatenation of ND6 and COI was performed using Sequencher 4.1.4, obtaining a sequence of 1318 bp. Sequence alignments were performed with Clustal X 2.1 (Larkin et al. 2007). The mitochondrial haplotypes and the number of variable sites were identified in the group of sequences using MEGA6 (Tamura et al. 2013). The haplotype and nucleotide diversity were estimated using Arlequin v3.5.1.3 (Excoffier and Lischer 2010). The intraspecific genetic distance was determined using the Kimura 2-parameter (K2P) nucleotide substitution model in MEGA6 (Tamura et al. 2013) because it is the appropriate model for estimating the small intraspecific genetic distances expected for this genomic region (Hebert et al. 2003). To estimate the genealogical relationships between haplotypes, a haplotype network was constructed based on the median-joining network algorithm (MJ) using NETWORK v4.6.1.3 (http://www.fluxus-engineering.com).
The population structure was estimated based on pair-wise FST. The statistical significance of this test was obtained by 1023 permutations. Analysis of molecular variance (AMOVA) was used to test the genetic differentiation among groups of populations from different natural regions. Both population pair-wise FST and AMOVA analysis were done using Arlequin v3.5.1.3 (Excoffier and Lischer 2010). A Mantel test was performed (Mantel 1967) to determine the existing relationship between the genetic distance (Arlequin’s FST) and the geographical distance (km) and thereby evaluate the isolation by distance (IBD) model. This test was performed in IBD v 3.23 (Jensen et al. 2005) using 1000 randomizations.
A mismatch distribution was performed to evaluate the population expansion signals using DNAsp 5.10.1 (Rozas et al. 2010). A population that has recently experienced a population expansion presents a unimodal mismatch distribution (Rogen and Harpending 1992). A sum of squared deviation was used with the Harpending’s index (Harpending 1994) to evaluate the deviation of the observed data from the observations that would be expected under a demographic expansion model. The population expansion time was calculated using the equation τ = 2ut, where u = m T µ (m T is the length of the nucleotide sequence under study, and µ is the mutation rate per nucleotide), and t is the time in generations (Roger and Harpending 1992), assuming a divergence rate of 2.3% per million years for insect mtDNA (Brower 1994) and a mean of six generations per year (Insuasty et al. 2007). To evaluate the population equilibrium, a Fu’s Fs test (Fu 1997) and a Tajima’s D test (Tajima 1989) were performed with DNAsp 5.10.1 (Rozas et al. 2010). Statistically significant negative values are obtained by these tests when a population is expanding.
Results
Of all the fruits processed, P. guajava was the only one that showed A. striata infestation. Eighty-two specimens were processed from guava fruits from the nine locations sampled. In three of the locations sampled (EN, LE and BU), A. fraterculus and A. striata infestations were found together on the same guava tree. However, the A. fraterculus proportion was less than 5% of the A. striata population.
Variation and genealogy of the mitochondrial concatenated sequence
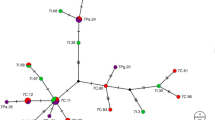
Eighty-two sequences of 1318 bp. each were obtained by concatenation of the COI and ND6 sequences. 14 variable sites were observed (1.06% of the total length), of which seven were unique and seven were parsimony informative. 16 mutational changes, 14 transitions (nine A/G and five T/C) and two transversions (one C/A and one G/T) were observed. 14 haplotypes with intraspecific genetic distances (K2P) between 0 and 0.4% were identified. The sequences for each of these genes were deposited in GenBank under accession numbers KU985248–55 for COI and KU985256–63 for ND6. The haplotype frequencies determined for each location are shown in Fig. 1 and Table 2, where haplotype H1 is the most frequent and is found in all locations, regardless of their differing natural regions. The rest of the haplotypes, except for H12, are private, with a very low frequency (Table 2). The average haplotype diversity was 0.057 and ranged from 0.71 in EN to 0.25 in PR (Table 3). The average nucleotide diversity was 0.0005 and ranged from 0.001 in EN to 0.000 in ZA. The haplotype network (MJ) was structured from a central, most abundant haplotype (H1) and from which the rest of the haplotypes H2 to H14 diverge. All of the haplotypes diverge with a low number of mutational changes; haplotype H11 is the most divergent with three changes (Fig. 2). The second most frequent haplotype was H12, found in six locations (BU, ZA, EN, SE, VE and PO), with no geographical association pattern. The haplotype network showed a star-like shape typical of populations undergoing expansion (Slatkin and Hudson 1991).
Haplotype network of 82 sequences (concatenates of the COI and ND6 genes) from A. striata in three natural regions of southwestern Colombia. Circles represent haplotypes, and their size is relative to the number of individuals sharing the haplotype. The haplotypes are divided by location, which is identified by a texture. The numbers correspond to the position of the nucleotide change. Symbols H1 to H14 represent the different mitochondrial haplotypes
Population genetic structure
The pair-wise FST statistic of nine locations showed that there is no genetic structure of A. striata in the southwestern Colombia, because 97% of the comparisons presented non-significant values below 0.12. The only significant comparison was between CH and BU with an FST value of 0.17 (Table 4). AMOVA analysis by grouping populations according to the natural regions also revealed no significant genetic differentiation among groups (F CT = 0.00006, P = 0.512) (Table 5). Therefore, there is no genetic isolation caused by the West Andes. Mantel test showed no significant relationship between the genetic and geographical distances (r = 0.1397, p = 0.198).
Demographic history
The mismatch distribution of the observed data showed unimodal behavior (Fig. 3), similar to the data simulated from the population expansion model, with a sum of squared deviations of DSS = 0.022 (P = 0.495) and a Harpending’s index of 0.467 (P = 0.683). The negative Fu (−11.834, P = 0.00) and Tajima (−2.002, P = 0.004) values, as the star-like shape haplotype network (Fig. 2), also indicated that the population has undergone expansion. The population expansion time, based on a 2.3% divergence rate (Brower 1994) and assuming six generations per year (Insuaty et al. 2007), was 17,000 years.
Mismatch distribution of 82 sequences (concatenates of the COI and ND6 genes) from A. striata in three natural regions of southwestern Colombia, representing the observed and expected paired differences under the population expansion model. The curve of the observed values shows a unimodal behavior typical of populations undergoing expansion
Discussion
Anastrepha striata was found in all of the locations sampled in greater numbers than A. frateculus; therefore, A. frateculus can be considered a low incidence species in guava fruits, at least in the region studied (5% of individuals collected). The predominance of A. striata was also previously reported by Núñez et al. (2004), who found percentages of 91.72% for A. striata and 8.26% for A. fraterculus in the northeastern region of Colombia.
Hernández-Ortiz et al. (2012), report that in Colombia there is only the Andean morphotype of A. fraterculus, which could explain the low incidence of this species and the predominance of A. striata in guava fruits. Several authors have previously noted that the fraterculus Andean morphotype can infest guava fruits, though it has also frequently found breeding in coffee and Andean berry fruits (Hernández-Ortiz and Morales-Valles 2004, Nuñez et al. 2004; Castañeda et al. 2010). In contrast, the fraterculus Brazilian-1 morphotype is restricted to guava fruits (Selivon et al. 2005). Additionally, A. striata and A. fraterculus proportions can vary depending on the area of study and factors such as the presence of other tephritids and the quality and abundance of alternating hosts. For example, Malavasi and Morgante (1980) found that guava was not very susceptible to A. striata attacks in Brazil, and Swanson and Baranowski (1972) found that A. suspensa was the only species infecting guava in the south Miami (USA) area.
The haplotype and nucleotide diversity of A. striata based on the mitochondrial concatenates data (COI + ND6) was 0.510 and 0.0005, respectively. This report is the first on the genetic diversity of this species in Colombia.
Therefore, there are no reference values for comparisons with other locations in the world. Compared with studies performed on other species of Anastrepha, our results indicate that the diversity of A. striata is low. For example, in A. ludens, Ruiz-Arce et al. (2015) reports haplotype and nucleotide diversities of 0.580 and 0.005, respectively. A recent study in A. obliqua in south western Colombia, with a similar sample size to one used in our study, showed a haplotype and nucleotide diversity of 0.553 and 0.0156, respectively (Aguirre 2016, undergraduate thesis, Universidad del Valle). These differences in genetic diversity between species can be correlated with dietary habits, because A. striata is an oligophagous species and was only found infesting guava fruits. While A. obliqua is a polyphagous species and was found infesting four different fruits in the same region.
Another explanation for the low genetic diversity could be that the haplotypes with low frequency might be present in the location but not detected in the sample due to low sample size. However, in this study, nearly all locations have sample sizes close to 10 individuals (Table 1). According to Luo et al. (2015) in genetic diversity studies with barcoding, sample sizes between 2 and 5 individuals present estimates with a high percentage of error, but values close to 10 generally came close to the correct estimate of diversity.
Despite the existing ecological and geographical differences among the three natural regions, AMOVA revealed no genetic differentiation among A. striata populations. Moreover, populations have the same predominant haplotype in all locations and non-significant FST values in 97% of the comparisons. These results probably indicate that A. striata moves freely across the three natural regions and that there is constant genetic flow. We hypothesized that the West Andes could be a geographical barrier for genetic flow. However, the AMOVA showed no significant genetic differences between the Pacific Coast and in the inner-Andean valley of the Cauca River. This would mean that the West Andes is not acting as an effective genetic barrier. We also found no relationship between genetic and geographical distances, which was expected given that the distance between the sampled localities is much greater than the dispersion capacity that has been reported for species of the genus Anastrepha (240 m of maximum displacement from the release site towards any point, not considering direction; Hernández et al. 2007).
A possible cause for the lack of genetic structure in A. striata is the anthropogenic effect because the transport and commercialization of fruits by humans can facilitate the movement of several insects such as aphids, woodlice, scarabs and larval stages of a variety of insects (Hill 2008). This effect would eliminate the Andes as a geographical barrier and would reduce genetic isolation due to distance or different eco-geographical regions. These dispersion phenomena mediated by humans have been reported in several studies of other fruit fly species (Gasperi et al. 2002; Malacrida et al. 2007; Shi et al. 2014; Karsten et al. 2015; Liebhold et al. 2016).
The Mediterranean fruit fly, Ceratitis capitata, is an example of these insects capacity for dispersion due to human globalization. Karsten et al. (2015), in a study conducted in Africa, Australia, Greece, Guatemala and Madeira suggest that C. capitata had an intercontinental distribution through a migration route from Africa to Europe, Australia and America with a reintroduction to South Africa from Europe. A strong structuring was found between Africa and the invasive localities, which is explained by quarantine export measures. In southwestern Colombia, all of these flies dispersion processes are a direct result of the implementation and modernization of agriculture by humans, largely because this region has been characterized by high agricultural production and possesses a network of main roads, such as the Pan-American Highway and the Panoramic Highway, that allows transportation between the main production and consumption zones of the country. The region also has a main access highway to one of the most important ports in the country, the Buenaventura port (Orozco 2003). Therefore, the high level of commercialization can be proposed as a way to increase the dispersion of larvae of different fruit fly species. This anthropogenic effect on the dispersion of A. striata would directly affect its population structure and genetic diversity.
Another possible cause for the lack of genetic structure of A. striata populations, despite the existence of different geographical barriers, is that the species is undergoing population expansion (Malacrida et al. 2007). This hypothesis is based on the star-like shape of the haplotype network showing the presence of a single common haplotype with a number of rare haplotypes connected to it by a few mutations and the demographic analysis. This pattern suggests that in spite of ecological or geographical isolation of populations, time was too short for mutations to occur and accumulate (Avise 2000), which agrees with the observation that although most haplotypes are not shared among locations, they are still found at very low frequencies. According to Slatkin and Hudson (1991), this type of behavior occurs in cases of recent invasion by a limited number of founders, followed by population growth, which would lead us to think that A. striata is an invasive species, at least in southwestern Colombia.
The demographic history analysis indicates that these recent species expansion signals originated approximately 17,000 years ago, in the last glaciations final epoch. During the Pleistocene glaciations, there was a phase of tropical forest degradation, which led to a cold and dry climate typical of this period (Penny 2001). These conditions could have caused the decrease in A. striata plant hosts. Approximately 18,000 years ago, the expansion of forests began when the climatic conditions became favorable, with a warmer and more humid climate, which corresponds to the onset of the A. striata population expansion. Studies of other insect species in the tropics have also proposed that climate changes in the Pleistocene could have had this effect on those species (Morgan et al. 2011; Meeyen et al. 2014). Therefore, this study supports the idea that this phenomenon was important in the formation of the genetic structure of insect populations.
This study is the first report on the genetic diversity of A. striata in Colombia, from which we conclude that this species is genetically homogeneous in southwestern Colombia; there is no evidence of genetic structure associated with the different eco-geographical regions or the West Andes as a possible geographical barrier. This behavior could be caused by human intervention as a passive dispersal agent of fruit flies, by an invasion followed by population expansion or by a combination of both factors. Therefore, the Andes of southwestern Colombia have not contributed to mitochondrial differentiation between the natural populations of A. striata. This study contributes to the genetic knowledge about this species and can be used for the creation of pest control programs.
References
Agelopoulos N, Birkett MA, Hick AJ, Hooper AM, Pickett JA, Pow EM, Smart LE, Smiley DW, Wadhams LJ, Woodcock CM (1999) Exploiting semiochemicals in insect control. J Pestic Sci 55:225–235
Aketarawong N, Chinvinijkul S, Orankanok W (2011) The utility of microsatellite DNA markers for the evaluation of area-wide integrated pest management using SIT for the fruit fly, Bactrocera dorsalis (Hendel), control programs in Thailand. Genetica 139:129–140
Aluja M, Jácome M, Birke A, Lozada N, Quintero G (1993) Patterns of Behavior in Wild Anastrepha striata (Diptera: Tephritidae) Flies under Field-Cage Conditions. Ann Entomol Soc Am 86:776–793
Aluja M, Piñero J, López M, Ruíz C, Zúñiga A, Piedra E, Díaz-Fleischer F, Sivinski J (2000) New host plant and distribution records in Mexico for Anastrepha spp., Toxotrypana curvicauda Gerstacker, Rhagoletis zoqui Bush, Rhagoletis sp., and Hexachaeta sp. (Diptera: Tephritidae). Proc Entomol Soc Wash 102:802–815
Arévalo P, Restrepo A, Areiza G (1997) Las moscas de las frutas del género Anastrepha Schiner (Diptera: Tephritidae) en el departamento de Antioquia (Colombia). Aconteceres Entomológicos 271–282
Avise JC (1994) Molecular markers, natural history and evolution. Chapman & Hall, New York
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Barr NB, Copeland RS, Meyer M, Masiga D, Kibogo HG, Billah MK, Osir E, Wharton RA, McPheron BA (2006) Molecular diagnostics of economically important Ceratitis fruit fly species (Diptera: Tephritidae) in Africa using PCR and RFLP analyses. Bull Entomol Res 96:505–521
Brower AVZ (1994) Rapid morphological radiation and convergence among races of the butterfly Heliconius erato, inferred from patterns of mitochondrial DNA evolution. Proc Natl Acad Sci USA 91:6491–6495
Brown WM, George M, Wilson AC (1979) Rapid evolution of animal mitochondrial DNA. PNAS 76:1967–1971
Caraballo J (2001) Diagnosis y clave pictórica para las especies del género Anastrepha Schiner, 1986 (Diptera: Tephritidae) de importancia económica en Venezuela. Entomotrópica 16(3):157–164
Carrejo N, González R (1994) Lista preliminar de las moscas de la fruta del género Anastrepha (Dip.: Tephritidae) en el departamento del Valle del Cauca Cali, Colombia, Boletín del Museo de Entomología de la Universidad del Valle 2(1):85–93
Castañeda M, Osorio A, Canal N, Galeano P (2010) Especies, distribución y hospederos del género Anastrepha Schiner en el departamento del Tolima, Colombia. Agron Colomb 28(2): 265–271
Cruz-López L, Malo E, Rojas J (2015) Sex Pheromone of Anastrepha striata. J Chem Ecol 41:458–464
Díaz AE, Suarez HG, Gallego G, Saldamando CI, Tohme J (2013) Geographic differentiation of Colombian Neulocinodes elegantalis (Lepidoptera: Crambidae) haplotypes: evidence for Solanaceae host plant association and Holdridge life zone for genetic differentiation. Ann Entomol Soc Am 106:586–597
Elias M, Joron M, Willmott K, Silva-Brandão KL, Kaiser V, Arias CF, Gomez Piñerez LM, Uribe S, Brower AV, Freitas AV, Jiggins CD (2009) Out of the Andes: patterns of diversification in clearwing butterflies. Mol Ecol 18:1716–1729
Excoffier L, Lischer H (2010) Arlequín suite ver 3.5: a new series of programs to perform population genetics analyses under linux and windows. Mol Ecol Resour 10(3):564–567
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Marine Biol Biotechnol 3(5):294–299
Fu YX (1997) Statistical test of neutrality of mutations against population growth, hitch hiking and background selection. Genetics 147:915–925
Gasperi G, Bonizzoni M, Gomulski LM, Murelli V, Torti C, Malacrida AR, Guglielmino CR (2002) Genetic differentiation, gene flow and the origin of infestations of the medfly, Ceratitis capitata. Genetica 116:125–135
Graham A, Gregory-Wodzicki KM, Wright KL (2001) Studies in Neotropical botany. XV. A Mio-Pliocene palynoflora from the Eastern Cordillera, Bolivia: implications for the uplift history of the Central Andes. Am J Bot 88:1545–1557
Harpending HC (1994) Signature of ancient population growth in a low-resolution mitochondrial DNA mismatch distribution. Hum Biol 6:591–600
Hebert PDN, Cywinska A, Ball SL, Dewaard JR (2003) Biological identifications through DNA barcodes. Proc Roy Soc Ser B 270:313–321
Hernández E, Orozco D, Flores-Breceda S, Domínguez J (2007) Dispersal and longevity of wild and mass-reared Anastrepha ludens and Anastrepha obliqua (Diptera: Tephritidae). Flo Entomol 90:123–135
Hernández-Ortiz V, Aluja M (1993) Listado de especies del género neotropical Anastrepha (Diptera: Tephritidae) con notas sobre su distribución y plantas hospederas. Folia Entomol 88:89–105
Hernández-Ortiz V, Morales-Valles P (2004) Distribución geográfica y plantas hospederas de Anastrepha fraterculus (Díptera: Tephritidae) en Venezuela. Folia Entomol Mex 43:181–189
Hernández-Ortiz V, Bartolucci F, Morales-Valles P, Frías D, Selivon D (2012) Cryptic species of the Anastrepha fraterculus complex (Diptera: Tephritidae): a multivariate approach for the recognition of South American morphotypes. Ann Entomol Soc Am 105:305–318
Hill DS (2008) Pests of crops in warmer climates and their control. Springer, Dordrecht
Hu J, Zhang JL, Nardi F, Zhang RJ (2008) Population genetic structure of the melon fly, Bactrocera cucurbitae (Diptera: Tephritidae), from China and Southeast Asia. Genetica 134:319–324
Insuasty O, Cuadros J, Monroy R, Bautista J (2007) Manejo Integrado de Moscas de la Fruta de la Guayaba (Anastrepha spp.). Boletín técnico. Publicación CORPOICA. Bogotá
Jensen JL, Bohonak AJ, Kelley ST (2005) Isolation by distance, web service. BMC Genetics 6:13. v.3.23, disponible en: http://ibdws.sdsu.edu/
Karsten M, Vanvuuren BJ, Barnaud A, Terblanche JS (2013) Population genetics of Ceratitis capitata in South Africa: implications for dispersal and pest management. Plos One 8:e54281
Karsten M, Vanvuuren BJ, Addison P, Terblanche JS (2015) Deconstructing intercontinental invasion pathway hypotheses of the Mediterranean fruit fly (Ceratitis capitata) using a Bayesian inference approach: are port interceptions and quarantine protocols successfully preventing new invasions? Divers Distrib 21:813–825
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0″. Bioinformatics 23:2947–2948
Liebhold AM, Yamanaka T, Roques A, Augustin S, Chown SL, Brockerhoff EG, Pyšek P (2016) Global compositional variation among native and non-native regional insect assemblages emphasizes the importance of pathways. Biol invasion 18(4):893–905
Luo A, Lan H, Ling C, Zhang A, Shi L, Ho SYW, Zhu C (2015) A simulation study of sample size for DNA barcoding. Ecol Evol 5(24):5869–5879
Lynch JD, Duellman WE (1997) Frogs of the genus Eleutherodactylus (Leptodactylidae) in western Ecuador: systematics, ecology, and biogeography. Univ. Kansas Spec. Publ. No. 23. Univ. of Kansas Printing Service, Lawrence
Malacrida AR, Gomulski LM, Bonizzoni M, Bertin S, Gasperi G, Guglielmino CR (2007) Globalization and fruitfly invasion and expansion: the medfly paradigm. Genetica 131:1–9
Malavasi A, Morganti JS (1980) Biología da moscas-frutas (DIP: Tephritidae). II. Indices de infestacao em diferentes hospedeiros e localidades. Rev Bras Biol 40(1):17–24
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Martínez A, Serna JS (2005) Identificación y localización geográfica de especies de Anastrepha (Diptera: Tephritidae) en Cundinamarca (Colombia). Agron Colomb 23(1):102–111
Meeyen K, Nanorksopaldawan P, Pramual P (2014) Population structure, population history and DNA barcoding of fruit fly Bactrocera latifrons (Hendel) (Diptera: Tephritidae). Entomol Sci 17:219–230
Morgan K, O’Loughlin SM, Chen B, Linton YM, Thongwat D, Somboon P, Fong MY, Butlin R, Verity R, Prakash A, Htun PT, Hlaing T, Nambanya S, Socheat D, Dinh TH, Walton C (2011) Comparative phylogeography reveals a shared impact of Pleistocene environmental change in shaping genetic diversity within nine Anopheles mosquito species across the Indo–Burma biodiversity hotspot. Mol Ecol 20:4533–4549
Mun J, Bohonak AJ, Roderick GK (2003) Population structure of the pumpkin fruit fly Bactrocera depressa (Tephritidae) in korea and Janpan: Pliocene allopatry or recent invasion? Mol Ecol 12:2941–2951
Norrbom AL (2003) Anastrepha striata (Schiner). The Diptera site. Fruit flies (Diptera:Tephritidae) Databases. Systematic Entomology Laboratory, ARS, USDA. Department of Entomology, NMNH, SI. Disponible en: http://www.sel.barc.usda.gov/diptera/tephriti/Anastrep/striata.htm. Accesed 13 August 2015
Norrbom AL, Carroll LE, Thompson FC, White IM, Freidberg A (1999) Systematic database of names. In: Thompson F. C. (Ed.), Fruit fly expert identification system and systematic information database. Myia (1998) 9, and Diptera Data Dissemination Disk (CD-ROM) (1998) 1:65–251
Núñez L (2010) Contribución al reconocimiento de las moscas de las frutas (Diptera:Tephriridae) en Colombia. Revista del Instituto Colombiano de Agropecuaria 16:173–179
Núñez L, Gómez R, Guarín G, León G (2004) Moscas de las frutas (Díptera: Tephritidae) y parasitoides asociados con Psidium guajava L. y Coffea arabica L. en tres municipios de la Provincia de Vélez (Santander, Colombia) Parte 1: Índices de infestación y daño por moscas de las frutas (Díptera: Tephritidae). Revista Corpoica 51:5–12
Olarte W (1980) Dinámica poblacional del complejo constituido por las moscas de las frutasAnastrepha striataSch. Anastrepha fraterculusWied. En el medio ecológico del sur de Santander. Editorial, Fondo rotatorio de la Imprenta del Departamento de Santander. Universidad industrial de Santander.
Orozco C (2003) Ordenamiento territorial y medio ambiente, Incidencias de las normas jurídicas, Valle del Cauca 1990–1998. Unidad de Artes Gráficas de la Facultad de Humanidades, Universidad del Valle. Cali. CO. pp 153–154.
Penny D (2001) A 40,000 year palynological record from northeast Thailand, implications for biogeography and palaeoenvironmental reconstruction. Palaeogeogr Palaeoclimatol Palaeoecol 171:97–128
Prabhakar CS, Mehta PK, Sood P, Singh SK, Sharma P, Sharma PN (2012) Population genetic structure of the melón fly, Bactrocera cucurbitae (Coquillett) (Diptera: Tephritidae) based on mitochondrial cytochrome oxidase (COI) gene sequences. Genetica 140:83–91
Prabhakar CS, Sood P., Metha PK, Sharma PN (2013) Population genetic structure of the pumpkin fruit fly, Bactrocera tau (Walker) (Diptera: Tephritidae) in Himachal Pradesh, India. Biochem Syst Ecol 51:291–296
Ramírez A, Hernández-Ortiz V, Martínez I (1996) Maduración Ovárica en la mosca de la guayaba Anastrepha striata Schiner (Diptera: Tephritidae). Acta Zool Mex nueva serie (69):105–116
Roderick G, Navajas M (2003) Genes in new environments: genetics and evolution in biological control. Nat Rev Genet 4:889–899
Roger AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Rozas J, Librado P, Sánchez-Del Barrio JC, Messeguer X, Rozas R (2010) DnaSP Version 5 Help Contents [Help File]. Disponible en http://www.ub.edu/dnasp/
Ruiz F, Linton YM, Ponsonby DJ, Conn JE, Herrera M, Quiñones ML, Vélez ID, Wilkerson RC (2010) Molecular comparison of topotypic specimens confirms Anopheles (Nyssorhynchus) dunhami Causey (Diptera: Culicidae) in the Colombian Amazon. Mem Inst Oswaldo Cruz 105(7):899–903
Ruiz-Arce R, Barr NB, Owen CL, Thomas DB, McPheron BA (2012) Phylogeography of Anastrepha obliqua inferred with mtDNA sequencing. J Econ Entomol 105:2147–2160
Ruiz-Arce R, Owen CL, Thomas DB, Barr NB, McPheron BA (2015) Phylogeographic structure in Anastrepha ludens (Diptera: Tephritidae) populations inferred with mtDNA sequencing. J Econ Entomol. doi:10.1093/jee/tov082
Ruiz-Hurtado FM, Ramirez-Quimbayo JH, Rojas-Plazas B, Galeano-Olay PE, Canal N (2013) Diversidad de parasitoides (hymenoptera) de moscas frugívoras (díptera: tephritoidea) en dos áreas cafeteras del departamento del Tolima, Colombia. Revista Tumbaga. 2(8):29–53.
Salazar RMI, Gómez N, Vargas WG, Reyes M, Castillo LE, Bolívar W (2002) Bosques secos y muy secos del departamento del Valle del Cauca. Serie de Ecosistemas del Valle del Cauca. Corporación Autónoma Regional del Valle del Cauca CVC.
Selivon D, Perondini ALP, Morgante JS (2005) A genetic-morphological characterization of two cryptic species of the Anastrepha fraterculus complex. (Diptera: Tephritidae). Ann Entomol Soc Am 98:367–381
Shi W, Kerdelhue C, Ye H (2005) Population genetics of the oriental fruit fly, Bactrocera dorsalis (Diptera: Tephritidae), in Yunnan (China) Based on Mitochondrial DNA sequences. Environ Entomol 34:977–983
Shi W, Kerdelhue C, Ye H (2014) Genetic structure and colonization history of the fruit fly Bactrocera tau (Diptera: Tephritidae) in China and Southeast Asia. J Econ Entomol 107(3):1256–1265
Simon C, Frati F, Beckenbach A, Crespi B, Liu H, Flook P (1994) Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann Entomol Soc Am 87:651–701
Slatkin M, Hudson RR (1991) Pairwise comparisons of mitocondrial DNA sequences in stable and exponentially growing populations. Genetics 129:555–562
Steyskal GC (1977) Pictorial key to species of the genus Anastrepha (Diptera: Tephritidae). Entomological Society of Washington press, Washington
Swanson R, Baranowski RW (1972) Host range and infestation by the Caribbean Fruit Fly Anastrepha suspensa (DIP: Tephritidae) In South Florida. Fla State Hortic Soc 85:271–274
Tajima F (1989) Statistical methods for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA 6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30:2725–2729
Velasco-Cuervo SM, Espinosa L, Duque-Gamboa DN, Castillo-Cardenas MF, Hernández LM, Guzmán YC, Manzano MR, Toro-Perea N (2016) Barcoding, population structure, and demographic history of Prodiplosis longifila associated with the Andes. Entomol Exp Appl 158: 217–227
Xie L, Hong XY, Xue XF (2006) Population genetic structure of the twospotted spider mite (Acari: Tetranychidae) from China. Ann Entomol Soc Am 99:959–965
Acknowledgements
This research was funded by the Universidad del Valle through an internal call for proposals for the project “Códigos de Barras de especímenes frescos y de museo de la mosca de la fruta Anastrepha fraterculus y A. obliqua: Contribución al conocimiento de su historia demográfica y estructura poblacional en el Valle del Cauca”. We are grateful to Nelson Rivera of the human genetics research group at the Universidad del Valle for his technical assistance and contributions to the data analysis. We thank Maria del Rosario Castañeda and Mauricio Peñuela for their help improving this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declares that they have no conflict of interest.
Informed consent
Informed consent was obtained from all individual participants included in this study.
Rights and permissions
About this article
Cite this article
Gallo-Franco, J.J., Velasco-Cuervo, S.M., Aguirre-Ramirez, E. et al. Genetic diversity and population structure of Anastrepha striata (Diptera: Tephritidae) in three natural regions of southwestern Colombia using mitochondrial sequences. Genetica 145, 79–89 (2017). https://doi.org/10.1007/s10709-016-9947-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-016-9947-5