Abstract
The suh gene is crucial in Notch pathway and regulates mammalian gonad development. In this study, the sequences of suh1 and suh2 genes in Yellow River carp (Cyprinus carpio) were verified. The partial 5′-flanking regions of suh1 and suh2 were analyzed and several potential transcription factor-binding sites were identified. Phylogenetic, gene structure, and chromosome synteny analyses revealed that carp suh1 and suh2 were orthologs and homologous to vertebrate suh. Investigation of the expression profiles of suh1 and suh2 with qPCR showed that these genes were abundant in the brain and gonad of carp, with suh1 exhibiting sexual dimorphism expression pattern in gonad. To study the relationship between gonad differentiation and Notch signaling, primordial gonads were exposed to DAPT, an inhibitor of Notch signaling, in vitro and in vivo. The results revealed a significant downregulation of suh1 and other Notch genes in vitro. In addition, expression of male-biased genes, such as amh, dmrt1, etc., was downregulated, whereas that of female-biased genes, such as foxl2, gdf9, etc., was upregulated. When the primordial gonads were subjected to long-term DAPT exposure, an increased proportion of ovary and delay in testis development were observed. These results suggest that suh gene may have a conservative function between teleosts and mammals. Furthermore, Notch signaling was found to be involved in gonad differentiation in Yellow River carp, and DAPT was noted to inhibit and enhance the expression of male- and female-biased genes, respectively, and induce the increase in number of females.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Notch gene was first identified by Morgan in 1917 and was named after the wing edge notch observed in fruit fly ( Drosophila melanogaster ) following deletion of this gene. In 1983, Artavanis-Tsakonas first cloned the Notch gene, which encodes a large class of transmembrane receptors (Artavanis-Tsakonas et al., 1983). The Notch signaling pathway is a highly conserved cell signaling system present in most of the vertebrates and invertebrates. In mammals, four Notch family receptors, encoded by notch1, notch2, notch3, and notch4 genes, have been described. The Notch ligands are encoded by Jagged1, Jagged2, Deltalike1, Deltalike3, and Deltalike4 genes. When Notch ligands bind to Notch receptors, the receptors become susceptible to proteolytic cleavage mediated by γ-secretase complex, which is a large protease complex composed of a catalytic subunit (psen-1 or psen-2) and accessory subunits (pen2, aph1, and nicastrin), for releasing the intracellular domain of Notch (NICD) (Struhl and Greenwald 2001; Greenwald 2012; Jarriault et al. 1995). The NICD then translocates into the nucleus, where it interacts with the suppressor of hairless (suh) gene, which is a transcription factor (TF) known as recombination signal binding protein for immunoglobulin kappa J region (RBPJ) (Jarriault et al. 1995). When the NICD binds to suh, the gene switches from being a repressor to an activator of transcription and regulates hairy and enhancer of split (Hes) and Hes-related YRPW motif TF (Hey), which are Notch target genes (Struhl and Greenwald 2001; Jarriault et al. 1995). These target genes function as transcriptional factors to regulate the expression of other genes in different cells. In previous studies, Notch signaling was observed to be involved in gonad development and assembly of primordial follicle (Vanorny et al. 2014; Wang et al. 2015), and suh knockout mice exhibited testicular failure (Garcia et al. 2014). Furthermore, in neonatal murine ovary, Notch ligands, jagged1 and jagged2, were noted to be expressed in germ cells, whereas Notch receptors, notch1 and notch2, were found to be expressed in pregranulosa cells (Johnson et al. 2001; Trombly et al. 2009), and blocking of Notch signal transduction was observed to affect ovarian development (Vanorny et al. 2014; Trombly et al. 2009; Manosalva et al. 2013; Chen et al. 2014; Feng et al. 2014). In our previous study, many differentially expressed genes were riched in Notch signaling pathway duringthe early stage of gonadal development (Jia et al. 2018), so we speculated that this pathway was involved in gonadal differentiation of carp.
As sex determination mechanism in fish is the most diversified among vertebrates, fish is an excellent model organism for studying the molecular mechanisms of sex determination and gonad development. Yellow River carp is the longest cultured and most widely domesticated fish species and has significant economic importance in China. As female carp could grow significantly faster than male carp after gonad differentiation (Gui 2007), mass production of female carp has always attracted increasing attention owing to its economic value. However, the genetic information for sex determination and gonad differentiation in carp remains unclear. In the present study, Yellow River carp suh1 and suh2 genes were verified and characterized, and N-[N-(3,5-difluorophenacetyl-L-alanyl)]-(S)-phenylglycine t-butyl ester (DAPT), which blocks Notch receptor proteolysis and is widely used in studying Notch signaling pathway, was employed for blocking the Notch signaling pathway in primordial gonad. The findings of this study could help in developing a new approach to regulate gonad development in Yellow River carp using Notch signaling pathway, as well as provide a novel method to study the molecular mechanisms of sex determination and gonad development in fish.
Materials and methods
Animals and sample collection
Yellow River carp were obtained from the Henan Academy of Fishery Science (Zhengzhou, Henan Province, China) and maintained at the Genetic Laboratory (Henan Normal University, Henan Province, China) in flow-through water tanks at 23 ± 2 °C under natural photoperiod for an initial acclimation period. Carp embryos were obtained by natural spawning and cultured in embryo medium following standard procedures. The staging of embryos was performed as described in a previous study (Chen 1960). The larvae were fed Artemia nauplii twice a day for the duration of the experiment, and no larvae died during the experiment. To study the expression profiles of suh, the embryos at blastula stage (2.5 h post-fertilization, hpf), gastrulae stage (7 hpf), neurula stage (13 hpf), tailbud stage (24 hpf), and hatching stage (3 days post-fertilization, dpf) were collected. Furthermore, tissues from heart, liver, spleen, kidney, forebrain, hindbrain, foregut, hindgut, gonads (ovary or testis), gill, scale, fin, eye, and skeletal muscle of adult male and female carp were obtained. Each sample was collected in triplicate. The collected embryo and tissue samples were immediately frozen in liquid nitrogen until RNA isolation. Before sample collection, the embryos and larvae were euthanized with 200 mg/L tricaine methanesulfonate (MS 222). This study was approved by the Committee of Laboratory Animal Experimentation at Henan Normal University (HNULSC101201). The DAPT (Sigma, USA) was dissolved in DMSO (Solarbio, China).
Sequence analysis of carp suh genes
Carp suh1 and suh2 sequences were obtained by RNA-Seq (Jia et al. 2018), and the open reading fragment (ORF)sequences were verified by reverse transcription-PCR (RT-PCR) using the sequence-specific primers (Supplementary Table 1). The amplification conditions were as follows: 95 °C for 3 min, followed by 30 cycles of 94 °C for 60 s, 56 °C for 60 s, and 72 °C for 90 s, and a final extension at 72 °C for 10 min. The RT-PCR products were separated by electrophoresis on 1% agarose gel and visualized using a UV imaging system (Bio-Rad, USA). The resulting sequences were confirmed on the NCBI Blast Server. The deduced amino acid sequences of carp suh1 and suh2 were aligned with those from other vertebrates available in GenBank using ClustalW multiple alignment program software (http://www.ebi.ac.uk/Tools/msa/clustalo/). The molecular mass (MM) and isoelectric point of carp suh1 and suh2 were predicted by ProtParam tool (http://web.expasy.org/protparam/). A phylogenetic tree was constructed by neighbor-joining algorithms using MEGA6. The data of chromosome synteny of suh1 and suh2 among vertebrates were obtained from NCBI and carp genome sequences. Comparison of the flanking regions of suh among fish and mammals was performed with Dialign software from the Genomatix suite (http://www.genomatix.de/cgi-bin/dialign/dialign.pl). Bioinformatics analyses of the potential TF-binding sites within the 5′-regulatory region of carp suh1 and suh2 were performed using the online program MatInspector (http://www.genomatix.de/matinspector.html) (Gao et al. 2015).
Expression profiles of carp suh genes
The expression profiles of carp suh1 and suh2 were investigated with quantitative real-time PCR (qPCR) using Roche real-time PCR system (LightCycler96 Roche) with SYBR green fluorescent label. The primers of carp suh1 and suh2 were designed outside the conserved domains to prevent any non-specific amplification (Supplementary Table 1). The qPCR was performed under the following conditions: 95 °C for 10 s, followed by 40 cycles of 95 °C for 15 s, and 60 °C for 1 min. The 2−ΔΔCt method was used to analyze the expression level. Each sample was amplified in triplicate to normalize the system and reduce pipetting error by using the standard curve method with 40S (40S ribosomal protein S11) and gapdh (glyceraldehyde-3-phosphate dehydrogenase) genes as reference genes (Zhang et al. 2016). Negative controls were included to confirm the absence of DNA contamination.
Effect of DAPT on gene expression in primordial gonad in vitro
The larvae at 40 days post-hatching (dph) were euthanized and their primordial gonads were isolated under a stereomicroscope. The gonads were washed thrice in PBS, transferred to a 24-well plate (BD Biosciences, USA), and maintained in a complex medium at 28 °C and 3% CO2 in a humidified incubator. The complex medium contained 50% L15, 35% DMEM-high glucose, and 15% Ham’s F-12 supplemented with 10% fetal bovine serum, 0.15 g/L sodium bicarbonate, 50 ng/mL mouse epidermal growth factor (EGF), and 0.01 mg/mL bovine insulin. The explants were cultured in parallel and divided into three groups: control group (without any added reagents to the medium), DMSO group (addition of 1% DMSO to the medium), and DAPT group (addition of 20 μM DAPT to the medium). Half of the medium was replaced with fresh medium every 48 h, and the experiments were repeated at least three times. Each group comprised six explants. After incubation for 1, 3, 5, and 7 days, the RNA of the explants was extracted using Total RNA Extraction Kit (RNAiso reagent, TaKaRa, Japan), and the cDNA was synthesized by using Prime Script Reverse Transcriptase (TaKaRa).
Long-term effect of DAPT on gonad development in larvae in vivo
A total of 300 larvae at 40 dph were randomly divided into three groups: control group in which the larvae were cultured in water without intervention, DMSO group in which the larvae were treated with 0.001‰ DMSO in water, and DAPT group in which the larvae were treated with 20 μM DAPT in water. The water was changed every 3 days, and after 30 days, the larvae were euthanized and dissected under a stereomicroscope. One side of the gonads was isolated, snap-frozen in liquid nitrogen, and stored at − 80 °C until further use. The other side of the gonads was pre-fixed in 4% PFA for 20 h at 4 °C, dehydrated through graded ethanol, embedded in paraffin, sectioned at 6 μm thickness, stained with hematoxylin-eosin, and observed under microscope to determine the stages of gonad and sex ratios. Three gonads of the same sex were selected for RNA extraction, and the cDNA was synthesized using Prime Script Reverse Transcriptase (TaKaRa).
Effect of DAPT on gene expression in primordial gonad in vivo
To confirm the effect of DAPT on Notch signaling pathway in the gonad of Yellow River carp, genes related to Notch signaling pathway (suh1, suh2, notch1, notch2, notch3, jagged1, jagged2, aph1, pen2, her6, hey1, and hey2 genes) and gonad development (amh, dax1, sf1, dmrt1, sox9a, nobox, foxl2, zp2, piwil, nanos, gdf9, figla, inhba, and inhbb genes) were detected with qPCR. Some primers used were designed based on the known genes in NCBI, while the others were based on common carp genome database (http://www.carpbase.org/download_home.php). All the primer sequences used in this study are listed in Supplementary Table 1. The qPCR conditions were similar to those mentioned earlier, and 2−ΔΔCt method was used to analyze the expression level. Each sample was amplified in triplicate to normalize the system, and all the reactions were independently repeated thrice to ensure reproducibility using the standard curve method with 40S and gapdh as reference genes (Zhang et al. 2016). Negative controls were included to confirm the absence of DNA contamination.
Statistical methods
For each set of data, independent experiments were repeated at least three times, and the data were analyzed using SPSS 16.0 and presented as mean ± SD. Furthermore, multiple comparisons were performed, and all the parameters were tested for normality using unpaired Student’s t test and analyzed using one-way ANOVA and Duncan’s. Significant difference was accepted at P < 0.05.
Results
Sequence analysis of carp suh genes
Initially, the sequences of carp suh1 and suh2 were identified from the carp genomic data and found to be homologous to the suh1 and suh2 genes of other species, respectively, by BLAST search on NCBI. The full-length suh1 cDNA was 6671-bp long, with a 1494-bp ORF, which encoded a protein of 497 amino acid residues (Supplementary Fig. 1). Comparison of genomic and cDNA sequences showed that suh1 contained 11 exons and 10 introns, similar to the structure of zebrafish suh1 and suh2 (Supplementary Fig. 2). The estimated MM of the predicted amino acids was 55.64 kDa and the theoretical isoelectric point was 8.42. With regard to carp suh2, the full-length cDNA sequence was 4251 bp, with a 1473-bp ORF (Supplementary Fig. 3). Moreover, suh2 was found to comprise 11 exons and 10 introns (Supplementary Fig. 2), and the putative suh2 protein of 491 amino acids had an estimated MM of 54.60 kDa and a theoretical isoelectric point of 8.28.
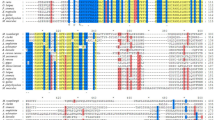
BLAST and ClustalW analyses showed that the deduced amino acid sequences of carp suh1 shared higher identities with zebrafish suh1 (97.0%), and lower identities with human, chicken, alligator, and frog suh (88.3%-89.9%). When compared with suh2 homologs, carp suh2 presented total amino acid identities of 87.5% with carp suh1, and a much higher identity with zebrafish suh2 (95.1%) (Table 1). It must be noted that carp suh1 and suh2 had three domains that were highly conserved among species (Supplementary Fig. 4).
To evaluate the evolutionary relationships between the Yellow River Carp suh1/2 and other vertebrates suh, a phylogenetic tree was constructed based on the full-length amino acid sequences. The suh proteins were grouped into two distinct clades. The first clade was composed with most of the fish Suh, such as carp and zebrafish Suh2, S. grahami. Carp and zebrafish suh1, mammals, chicken, turtle and frog suh gene clustered into the other clade, and carp suh1 was first clustered with zebrafish suh1, then with the tetrapod suh gene (Fig. 1).
Phylogenetic analysis of carp suh1 and suh2 in comparison with other representative vertebrates based on amino acid sequences. The tree was constructed by MEGA (version 6.0) using Poisson Correction distance based upon the neighbor-joining method with 1000 bootstrap replicates. The scale bar is 0.01. For GenBank accession numbers of sequences, see Supplementary Fig. 4
Chromosome synteny and genomic analysis of carp suh genes
The homologous relationship between carp suh1/2 and other species suh was determined by cross-species comparison of chromosome locations. Based on current C. carpio whole-genome sequencing data (http://www.carpbase.org/index.php), carp suh1 and tandem genes (chic and fip1) were found to be repeated on LG2, whereas carp suh2 was on LG36 and flanked by cckar and itih5 (Fig. 2). The chromosome syntenic relationship of suh1 and suh12 was evolutionarily conserved between fish and mammals..
Chromosome syntenic relationship of the fish suh1 and suh2 genes with their vertebrate orthologs. Conserved syntenies are shown for chromosomal segments containing suh1, suh2, and suh. Rectangles represent genes in chromosome/scaffold and arrows represent gene-coding direction. Suh, smim20, cckar, tbc1d19, stim2, and slc34a2a orthologs are shown in red, pink, green, light blue, yellow, and purple, respectively. Chr, chromosome; Sca, scaffold
For promoter analysis, the transcriptional initiation site (ATG) was designated as + 1 and 2000-bp upstream flanking sequences of carp suh1 and suh2 were examined. The upstream flanking sequence of carp suh1 presented higher sequence conservation with zebrafish suh1 at their 3′-terminus (Supplementary Fig. 5). Analysis using MatInspector predicted numerous essential TF-binding sites within the 5′-regulatory region, and those with a matrix score higher than 0.9 are illustrated in the schematic diagram presented in Fig. 3. Some of these TFs, including sf1, dmrt1, smad3, smad4, nobox, ar, sox5, sox6, gata3, and foxl1, were found to be involved in gonad differentiation and reproductive system development. Besides, some TFs closely related to various pluripotency or stem cell properties were identified, including oct4, nanog, and foxi1a. With regard to carp suh1, Nfκb signal pathway-related genes and liver-enriched and muscle-specific TFs such as P65, foxa1 (hnf3α), foxa2 (hnf3β), hnf6, and mtbf were detected. Besides, ubiquitous binding sites for ap1, cebpb, usf, hsf2, and sp1 were also identified. The predicted TF-binding sites of carp suh2 not only included sf1, sox6, and nanog genes, but also comprised nobox, esr2, and sox30, which were involved in gonad differentiation.
Schematic diagram of putative regulatory motifs in the promoter of the suh1 and suh2 in Yellow River carp. The scale is given above and the full names of the potential TF binding sites are provided at the bottom. The plus-minus sign indicates the TF binding strand. The gray blocks represent the conserved upstream region among species (details can be seen in Supplementary Fig. 5). Transcriptional initiate site (ATG) is designated as + 1
Expression pattern of carp suh genes
The expression pattern of carp suh1 and suh2 mRNA during embryonic development was analyzed with qPCR using 40S and gapdh as reference genes. The highest level of suh1 transcript was detected in the blastula stage. With the progression of embryonic development, the expression of suh1 followed a declining trend, reaching the lowest level at hatching (Fig. 4). Likewise, suh2 exhibited a similar expression pattern, with the peak suh2 expression noted in the early blastula stage. Besides, the amount of suh1 transcript was lower than that of suh2 transcript (Fig. 4).
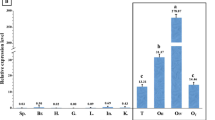
Furthermore, expression analysis in various adult tissues revealed that suh1 and suh2 were predominantly expressed in the brain (Fig. 5). In particular, suh1 transcript exhibited sexual dimorphism expression pattern in gonad, hindbrain, spleen, and gill (P < 0.05) (Fig. 5a), whereas suh2 transcript was found at the highest level in the brain and at low level in fin and liver (Fig. 5b).
Relative expression levels of carp suh1 and suh2 genes in different adult tissues by qPCR analysis. The relative expression variance is given as ratio (the amounts of carp suh1 and suh2 mRNA normalized to the values of the corresponding reference genes). Data are shown as mean ± SD (n = 3). Values with different letters indicate statistically significant difference (P < 0.05)
Histological characteristics of Yellow River carp gonads treated with DAPT
Histologically, primordial germ cells appeared in the primordial gonad of larvae at 40 dph (Fig. 6a). In the control group, the larvae completed sex differentiation at 70 dph, and the structural features of ovary or testis appeared (Fig. 6b, c). Furthermore, differentiated spermatogonium, primary spermatocyte, and spermatids were observed in the testis of the control group (Fig. 6c). In the DMSO group, the structural features of ovary and testis were similar to those in the control group. In the DAPT group, although ovary differentiation was noted (Fig. 6d), testis development stagnation occurred, germ cells continued to undergo mitosis, and a cyst including two or four dividing germ cells and composed of somatic cells was observed. Moreover, no spermatogonium and primary spermatocyte appeared in the testis (Fig. 6e). At the end of the treatment (70 dph), the proportion of ovary in the control, DMSO, and DAPT groups was 49, 51, and 61%, respectively. These results revealed that DAPT might play a role in the inhibition of spermatogenesis at a crucial stage of gonadal development in Yellow River carp.
Sections of Yellow River carp gonad. The same section was stained by hematoxylin and eosin (thickness, 6 μm). a Primordial gonad (40 dph). b Ovary of control group (70 dph). c Testis of control group (70 dph). d Ovary of DAPT group (70 dph). e Testis of DAPT group (70 dph). PGC: primordial germ cell, SC: somatic cell, OG: oogonium, PO: primary oocyte, SG: spermatogonium, SM: spermatocyte, ST: spermatid, GC: germ cell
Effects of short-term DAPT exposure on gene expression in primordial gonad in vitro
The expression of Notch components and their target genes in primordial gonad was effectively inhibited by DAPT (Supplementary Fig. 6). Suh1 were downregulated from day 1 to day 3 (P < 0.05). With regard to aph1 and pen2, the target genes of DAPT, severe downregulation of them were noted after DAPT exposure (P < 0.05).The Notch components, notch1, notch2, notch3, jagged1, and jagged2, were significantly downregulated after DAPT treatment (P < 0.05), and the downregulation of Notch target genes her6 (hes1), hey1, and hey2 persisted during the 7-day culture of primordial gonad with DAPT (P < 0.05, Supplementary Fig. 6).
It must be noted that amh, dax1, sf1, dmr1t, sox9a, inhba, and inhbb are related to the development of testis in fish. The expression of sf1, dmrt1, sox9a, dax1, and inhbb in the DAPT group was significantly lower, when compared with that in the control and DMSO groups (P < 0.05, Supplementary Fig. 7).
The genes foxl2, cyp19a, nobox, zp2, piwil, nanos, gdf9, and figla are critical for ovary development. The expression of foxl2, piwil, gdf9, nobox, and zp2 in the DAPT group was significantly higher, when compared with that in the control and DMSO groups throughout the experiment period (P < 0.05). However, the expression of cyp19a, cyp19b, nonos, and figla in the DAPT group was not significantly different from that in the control and DMSO groups (Supplementary Fig. 7). These results suggested that DAPT/Notch signaling pathway affected ovary development in Yellow River carp by regulating the expression of foxl2, piwil, gdf9, nobox, and zp2, but not cyp19a, cyp19b, nonos, and figla.
Effects of long-term DAPT treatment on gene expression in primordial gonad in vivo
The expression of Notch genes was downregulated at various levels after 30 days of DAPT exposure. All Notch-related genes in the testis of the DAPT group were significantly downregulated, when compared with those in the testis of the control and DMSO groups (P < 0.05). In addition, most of thesegenes were also downregulated in the ovary of the DAPT group, and in particular, suh1, pen2, jagged1, notch2, and her6 genes showed significant decrease in the ovary (P < 0.05, Supplementary Fig. 8).
In the control and DMSO groups, the gonads completed sex differentiation at 70 dph. Among the testis development-related genes, amh, sf1 dax1, sox9a, dmrt1, inhba, and inhbb were significantly downregulated in the testis of the DAPT group, when compared with those in the control and DMSO groups (P < 0.05). With regard to gene expression in ovary, the expression of sf1 and inhba was lower in the ovary of the DAPT group, when compared with that in the control and DMSO groups (P < 0.05, Supplementary Fig. 9). Furthermore, the expression of foxl2, gdf9, and nobox in the ovary of the DAPT group was significantly increased, when compared with that in the control and DMSO groups (P < 0.05) (Supplementary Fig. 9).
Discussion
Sex determination in fish is extremely complex and can be affected by environmental factors and genetic system (Devlin and Nagahama 2002). As one of the most important signaling pathways, Notch signaling pathway plays an important role in regulating cell differentiation and gonad development in mammals (Trombly et al. 2009; Johnson et al. 2001; Chen et al. 2014); however, it is still unclear whether Notch signaling is involved in gonad development in carp. In the present study, the suh genes of carp were examined because they are crucial in Notch signaling pathway and sex dimorphism pattern in carp.
The results confirmed that suh is duplicated in Yellow River carp, similar to that observed in other fish species such as zebrafish (O’Brien et al. 2011). The notion that carp suh1 and suh2 are orthologs derived from a gene duplication event was also supported by their chromosomal localizations: different chromosomes contained duplicated genomic sequences; the structure, chromosome synteny, and genomic analyses of suh1 and suh2 suggested that they are conserved.
Although suh gene is the main effector in Notch pathway, its functions and TF-binding sites have not yet been reported. The present study examined the TF-binding sites of carp suh1 and suh2, which might participate in the regulation of gene expression and function, and is the first to predict the TF-binding sites of carp suh genes (Supplementary Tables 2 and 3). Some TF-binding sites relevant to gonad differentiation, including sf1, sox5, sox6, nobox, esr2, and GATA3, in vertebrates (Espinosa et al. 2015; Mu et al. 2013; Connor et al. 1995; Denny et al. 1992) may interact with carp suh1 and suh2 to regulate their expression in gonad development in carp.
The suh gene is a Notch transcriptional mediator, and previous studies have suggested that this gene is critical in Notch pathway and early processes in embryogenesis and testis development (Zhang et al. 2014; Garcia et al. 2014). However, to date, there are only few reports on teleost suh (O’Brien et al. 2011). The sexual dimorphism expression pattern of carp suh1 in gonad observed in the present study indicated its involvement in gonad differentiation. Furthermore, DAPT has been observed to block the Notch signal transduction in the ovary of mammals in vitro and inhibit gonad development (Chen et al. 2014) and has been widely used to study the development of tissues and organs by blocking the Notch signaling pathway (Chen et al. 2014; Feng et al. 2014; Zhang et al. 2011). However, the molecular mechanisms of gonad development in carp are still unclear (Chen et al. 2015; Gui 2007), and the signal pathways in the carp gonad, which regulate gonad development and sex differentiation, have not been extensively studied. In the present study, after short-term treatment with DAPT in vitro, the expression of Notch genes, such as suh1, notch1, notch2, notch3, jagged1, jagged2, apha1, and pen2, was significantly downregulated in primordial gonad. Moreover, following long-term DAPT exposure, suh1, suh2, and other Notch-related genes were significantly downregulated in the testis of the DAPT group, when compared with that in the control and DMSO groups (P < 0.05). In addition, pen2, notch2, jagged1, and her6 in the ovary of the DAPT group were also significantly downregulated (P < 0.05). These results revealed that Notch signaling was suppressed by DAPT during the early development of gonad in Yellow River carp.
It has been reported that amh, dmrt1, sox9a, sf1, dax1, inhba, and inhbb play roles in testis development in fish (Zhou et al. 2014l Rodriguez-Mari et al. 2005; Hattori et al. 2012; Herpin and Schartl 2011; Herpin et al. 2013; Hornung et al. 2007; Tada et al. 2002), and the expression of these genes was detected with qPCR in the present study. Furthermore, these genes were downregulated, especially in testis, after DAPT treatment both in vitro and in vivo (P < 0.05). Besides, DAPT delayed the development of testis, germ cells continued to undergo mitosis in cyst, and spermatogonium and primary spermatocyte did not appear in the testis at 70 dph. These findings suggest that transcription factors amh, dax1, dmrt1, sox9a are sensitive to DAPT, and their decreased expression in testis could affect the development of testis. Therefore, it can be assumed that DAPT could suppress testis differentiation by regulating the expression of amh, dax1, dmrt1, sox9a, and sf1.
The foxl2, figla, gdf9, piwil, zp2, nanos, and nobox genes are critical for the development of ovary (Zhou et al. 2014; Herpin et al. 2013; Liu and Ge 2007; Chen et al. 2015; Draper et al. 2007; Rajkovic et al. 2004; Huang et al. 2014; Jia et al. 2016). In the present study, the expression pattern of these genes was enhanced in the ovary, when compared with that in the testis, and their levels of expression varied in gonads subjected to DAPT treatment. Previous studies had shown that primordial follicle formation was regulated by Notch pathway by affecting the expression of nobox gene (Chen et al. 2014). In the present study, the expression of foxl2, gdf9, and nobox was higher in the primordial gonad after DAPT treated in vitro, indicating that DAPT could increase the expression of foxl2, gdf9, and nobox. Similar results were noted in ovary following long-term DAPT exposure. Taken together, these results revealed that DAPT participates in ovary differentiation by regulating the expression of foxl2, gdf9, and nobox. It has been reported that piwil gene is crucial for the differentiation and development of germ cells (Zhou et al. 2014; Zhou et al. 2012). After long-term DAPT exposure, the differentiation and meiosis of germ cells in testis were suppressed by downregulated expression of piwil. The testis section of the DAPT group revealed that the germ cell differentiation was not initiated and the cells remained in the germ cell syncytia, whereas spermatids already appeared in the testis of the control group. In contrast, higher expression of piwil in ovary helped in the differentiation of oocytes, although its regulation mechanism is not clear and requires further investigation. Moreover, the higher proportions of ovary at 70 dph suggest that the increased expression of foxl2, gdf9, nobox, and piwil positively regulates the early development of ovary. Sex differentiation is a complex process, including changes in early genetic levels and subsequent morphological changes. Amh, dax1, dmrt1, sox9a, foxl2, gdf9, nobox, and piwil are crucial transcription factors during early gonad differentiation. The expression of them was changed significantly after DAPT treatment IN primordial gonad suggested notch signal/DAPT should be involved in the early differentiation of the gonads by regulating these upstream genes in the process of gonad differentiation.
Conclusions
In this study, the structure of Yellow River carp suh1 and suh2 genes was described and their expression profiles during embryo development as well as in adult tissues were examined. In addition, the potential regulatory TF-binding sites in the upstream flanking sequences of suh1 and suh2 were analyzed. The sexual dimorphism expression pattern of suh1 suggested its potential functions in the regulation of gonadogenesis in Yellow River carp. Chromosome synteny analysis indicated that the carp suh1 and suh2 are highly conserved during evolution. These results could help in further understanding of the functions of suh gene in teleost fish. In our previous study, results showed that DAPT could affect the sex ratio during the early development of gonads in carp. The mechanism should be related to the notch signal/DAPT is involved in early gonadal differentiation by regulating the expression of transcription factors upstream of the genetic regulation.
References
Artavanis-Tsakonas S, Fau - Muskavitch MA, Muskavitch MA, Fau - Yedvobnick B, Yedvobnick B (1983) Molecular cloning of notch, a locus affecting neurogenesis in Drosophila melanogaster. Proc Natl Acad Sci U S A 80:1977–1981
Chen CL (1960) The study on embryonic development of Carp (Cyprinus carpio L). Chin J Zool 4:165–168
Chen C, Fu X, Wang L, Wang J, Ma H, Cheng S, Hou Z, Ma J, Quan G, Shen W, Li L (2014) Primordial follicle assembly was regulated by notch signaling pathway in the mice. Mol Biol Rep 41:1891–1899
Chen JJ, Xia XH, Wang LF, Jia YF, Nan P, Li L, Chang ZJ (2015) Identification and comparison of gonadal transcripts of testis and ovary of adult common carp Cyprinus carpio using suppression subtractive hybridization. Theriogenology 83:1416–1427
Connor F, Wright E, Denny P, Koopman P, Ashworth A (1995) The Sry-related HMG box-containing gene Sox6 is expressed in the adult testis and developing nervous system of the mouse. Nucleic Acids Res 23:3365–3372
Denny P, Swift S, Connor F, Ashworth A (1992) An SRY-related gene expressed during spermatogenesis in the mouse encodes a sequence-specific DNA-binding protein. EMBO J 11:3705–3712
Devlin R, Nagahama Y (2002) Sex determination and sex differentiation in fish. Aquaculture 208:191–364
Draper BW, McCallum CM, Moens CB (2007) Nanos1 is required to maintain oocyte production in adult zebrafish. Dev Biol 305:589–598
Espinosa I, Gallardo A, D'Angelo E, Mozos A, Lerma E, Prat J (2015) Simultaneous carcinomas of the breast and ovary: utility of Pax-8, WT-1, and GATA3 for distinguishing independent primary tumors from metastases. Int J Gynecol Pathol 34:257–265
Feng YM, Liang GJ, Pan B, Qin XS, Zhang XF, Chen CL, Li L, Cheng SF, De Felici M, Shen W (2014) Notch pathway regulates female germ cell meiosis progression and early oogenesis events in fetal mouse. Cell Cycle 13:782–791
Gao J, Zhang W, Li P, Liu J, Song H, Wang X, Zhang Q (2015) Identification, molecular characterization and gene expression analysis of sox1a and sox1b genes in Japanese flounder, Paralichthys olivaceus. Gene 574:225–234
Garcia TX, Farmaha JK, Kow S, Hofmann MC (2014) RBPJ in mouse Sertoli cells is required for proper regulation of the testis stem cell niche. Development 141:4468–4478
Greenwald I (2012) Notch and the awesome power of genetics. Genetics 191:655–669
GuiJ (2007) Genetic basis and artificial control of sexuality and reproduction in fish. Science Press, Beijing, 125–126
Hattori RS, Murai Y, Oura M, Masuda S, Majhi SK, Sakamoto T, Fernandino JI, Somoza GM, Yokota M, Strussmann CA (2012) A Y-linked anti-Mullerian hormone duplication takes over a critical role in sex determination. Proc Natl Acad Sci U S A 109:2955–2959
Herpin A, Schartl M (2011) Dmrt1 genes at the crossroads: a widespread and central class of sexual development factors in fish. FEBS J 278:1010–1019
Herpin A, Adolfi MC, Nicol B, Hinzmann M, Schmidt C, Klughammer J, Engel M, Tanaka M, Guiguen Y, Schartl M (2013) Divergent expression regulation of gonad development genes in medaka shows incomplete conservation of the downstream regulatory network of vertebrate sex determination. Mol Biol Evol 30:2328–2346
Hornung U, Herpin A, Schartl M (2007) Expression of the male determining gene dmrt1bY and its autosomal coorthologue dmrt1a in medaka. Sex Dev 1:197–206
Huang CX, Wei XL, Chen N, Zhang J, Chen LP, Wang WM, Li JY, Wang HL (2014) Growth differentiation factor 9 of Megalobrama amblycephala: molecular characterization and expression analysis during the development of early embryos and growing ovaries. Fish Physiol Biochem 40:193–203
Jarriault S, Brou C, Logeat F, Schroeter EH, Kopan R, Israel A (1995) Signalling downstream of activated mammalian notch. Nature 377:355–358
Jia Y, Zhang W, Zhang R, Liang T, Du Q, Chang Z (2016) Timing of sex determination and expression analysis of genes in Yellow River carp (Cyprinus carpio Var.) Acta Hydrobiologica sinica 40:1121–1127
Jia Y, Nan P, Zhang W, Wang F, Zhang R, Liang T, Ji X, Du Q, Chang Z (2018) Transcriptome analysis of three critical periods of ovarian development in Yellow River carp (Cyprinus carpio). Theriogenology 105:15–26
Johnson J, Espinoza T, McGaughey RW, Rawls A, Wilson-Rawls J (2001) Notch pathway genes are expressed in mammalian ovarian follicles. Mech Dev 109:355–361
Liu L, Ge W (2007) Growth differentiation factor 9 and its spatiotemporal expression and regulation in the zebrafish ovary. Biol Reprod 76:294–302
Manosalva I, Gonzalez A, Kageyama R (2013) Hes1 in the somatic cells of the murine ovary is necessary for oocyte survival and maturation. Dev Biol 375:140–151
Mu WJ, Wen HS, Shi D, Yang YP (2013) Molecular cloning and expression analysis of estrogen receptor betas (ERbeta1 and ERbeta2) during gonad development in the Korean rockfish, Sebastes schlegeli. Gene 523:39–49
O'Brien LL, Grimaldi M, Kostun Z, Wingert RA, Selleck R, Davidson AJ (2011) Wt1a, Foxc1a, and the notch mediator Rbpj physically interact and regulate the formation of podocytes in zebrafish. Dev Biol 358:318–330
Rajkovic A, Pangas SA, Ballow D, Suzumori N, Matzuk MM (2004) NOBOX deficiency disrupts early folliculogenesis and oocyte-specific gene expression. Science 305:1157–1159
Rodriguez-Mari A, Yan YL, Bremiller RA, Wilson C, Canestro C, Postlethwait JH (2005) Characterization and expression pattern of zebrafish anti-Mullerian hormone (Amh) relative to sox9a, sox9b, and cyp19a1a, during gonad development. Gene Expr Patterns 5:655–667
Struhl G, Greenwald I (2001) Presenilin-mediated transmembrane cleavage is required for notch signal transduction in drosophila. Proc Natl Acad Sci U S A 98:229–234
Tada T, Endo M, Hirono I, Takashima F, Aoki T (2002) Differential expression and cellular localization of activin and inhibin mRNA in the rainbow trout ovary and testis. Gen Comp Endocrinol 125:142–149
Trombly DJ, Woodruff TK, Mayo KE (2009) Suppression of notch signaling in the neonatal mouse ovary decreases primordial follicle formation. Endocrinology 150:1014–1024
Vanorny DA, Prasasya RD, Chalpe AJ, Kilen SM, Mayo KE (2014) Notch signaling regulates ovarian follicle formation and coordinates follicular growth. Mol Endocrinol 28:499–511
Wang Z, Niu W, Wang Y, Teng Z, Wen J, Xia G, Wang C (2015) Follistatin288 regulates germ cell cyst breakdown and primordial follicle assembly in the mouse ovary. PLoS One 10:e0129643
Zhang CP, Yang JL, Zhang J, Li L, Huang L, Ji SY, ZY H, Gao F, Liu YX (2011) Notch signaling is involved in ovarian follicle development by regulating granulosa cell proliferation. Endocrinology 152:2437–2447
Zhang S, Kong SB, Wang BY, Cheng XH, Chen YJ, WW W, Wang Q, Shi JC, Zhang Y, Wang SM, JH L, Lydon JP, DeMayo F, Pear WS, Han H, Lin HY, Li L, Wang HM, Wang YL, Li B, Chen Q, Duan EK, Wang HB (2014) Uterine Rbpj is required for embryonic-uterine orientation and decidual remodeling via notch pathway-independent and -dependent mechanisms. Cell Res 24:925–942
Zhang W, Jia Y, Ji X, Zhang R, Liang T, Du Q, Chang Z (2016) Optimal reference genes in different tissues, gender,and gonad of Yellow River Carp (Cyprinus carpio var) at various developmental periods. Pakistan J Zool 48:1615–1622
Zhou Y, Wang F, Liu S, Zhong H, Liu Z, Tao M, Zhang C, Liu Y (2012) Human chorionic gonadotropin suppresses expression of Piwis in common carp (Cyprinus carpio) ovaries. Gen Comp Endocrinol 176:126–131
Zhou Y, Zhong H, Liu S, Yu F, Hu J, Zhang C, Tao M, Liu Y (2014) Elevated expression of Piwi and piRNAs in ovaries of triploid crucian carp. Mol Cell Endocrinol 383:1–9
Acknowledgments
This work was partially supported by the following financing: Innovative Research Team (in Science and Technology) in University of Henan Province (No. 17IRTSTHN017) and the Scientific and Technological Research Projects of Henan Province (No. 182300410302).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This study was approved by the Committee of Laboratory Animal Experimentation at Henan Normal University (HNULSC101201).
ᅟ
Conflict of interests
The authors declare that they have no conflict of interest.
Electronic supplementary material
Supplementary Fig. 1
Nucleotide sequences of the suh1 gene in Cyprinus carpio. The deduced amino acid sequence is shown underneath the CDS (coding sequence). The LAG1-DNA binding domain is indicated with single line, Beta-trefoil DNA-binding domain shaded in gray and the IPT_RBP-J kappa domain is boxed. The start and stop codons are labeled in red color. Nucleotides and amino acids are numbered at the right end of the lines. (JPEG 5279 kb)
Supplementary Fig. 2
Comparisons of genomic organizations of suh gene between carp and zebrafish. Exons are shown in dark box while introns are shown in straight line. The sizes of primary transcripts and each part are indicated. Notably, the homeodomain is located between the exons with arrows above. (JPEG 204 kb)
Supplementary Fig. 3
Nucleotide sequences of the suh2 gene in Cyprinus carpio. The deduced amino acid sequence is shown underneath the CDS (coding sequence). The LAG1-DNA binding domain is indicated with single line, Beta-trefoil DNA-binding domain shaded in gray and the IPT_RBP-J kappa domain is boxed. The start and stop codons are labeled in red. Nucleotides and amino acids are numbered at the right end of the lines. (JPEG 3395 kb)
Supplementary Fig. 4
Multiple alignment of Suh proteins in different species. All the sequences of Suh homologs were retrieved from NCBI. The LAG1-DNA binding domain is marked with black single line, Beta-trefoil DNA-binding domain indicated green single line and the IPT_RBP-J kappa domain is labelled in blue single line. Nucleotides and amino acids are numbered at the right end of the lines. The alignment was generated with Clustal W. The GenBank accession numbers of sequences are as follows: C.carpio-Suh1: Cyprinus carpio suh1, KX904343; C.carpio-Suh2: Cyprinus carpio suh2, KX904344; B.pectinirostris-Suh: Boleophthalmus pectinirostris suh, XM_020937781; C.variegatus-Suh: Cyprinodon variegatus suh, XM_015391780; L.bergylta-Suh: Labrus bergylta suh, XM_020629233; S.partitus-Suh: Stegastes partitus suh, XM_008287419; L.crocea-Suh: Larimichthys crocea suh, XM_010741712; C.semilaevis-Suh: Cynoglossus semilaevis suh, XM_008322828; O.latipes-Suh: Oryzias latipes suh, XM_020711680.1; O.niloticus-Suh: Oreochromis niloticus suh, XM_003449374; E.lucius-Suh: Esox lucius suh, XM_013140478; D.rerio-Suh2: Danio rerio suh2, NM_001083551.1; D.rerio-Suh1: Danio rerio suh1, NM_198878.2; M.musculus-Suh: Mus musculus suh, BC051387; M.musculus-Suh: Human sapiens suh, BC051387; G.gallus-SuhX1: X.laevis-Suh1: Xenopus laevis Suh1, BC133204; Gallus gallus suh, XM_004936138; P.sinensis-Suh: Pelodiscus sinensis suh, XM_006138171. (JPEG 3012 kb)
Supplementary Fig. 5
Partial alignment of the suh1 and suh2 5′-flanking regions sequences in different species with Genomatix suite. The species-conserved region were shown. The sequences were obtained from the NCBI database and the start codon ATG were on the right side of each sequence. Gene IDs: carp suh1, suh2: this study; zebrafish suh1: 386772; suh2, 563306;Sinocyclocheilus grahami suh, 107597867; human suh, 3516; mouse suh, 19664. (JPEG 7567 kb)
Supplementary Fig. 6
Effect of DAPT on the expression of notch member and target genes of primordial gonad in vitro. When the primordial gonads (40 dph) were cultured for 1, 3, 5 and 7 days with DAPTin vitro, notch member and target genes were down regulated. The results present as Mean ± SD. Compared to control group, *P < 0.05, **P < 0.01. (JPEG 591 kb)
Supplementary Fig.7
Effect of DAPT on the expression of gonads development related genes of primordial gonad in vitro. When the primordial gonad (40 dph) were cultured for 1, 3, 5 and 7 days with DAPTin vitro, amh, dax1, sf1, dmrt1, sox9a,inhba and inhbb were down-regulated, foxl2, piwil, gdf9, nobox and zp2 were up-regulated. The results present as Mean ± SD. Compared to control group, *P < 0.05, **P < 0.01. (JPEG 741 kb)
Supplementary Fig. 8
Effect of DAPT on the expression of notch member and target genes in vivo. When the primordial gonads (40dph) were treated with DAPT for 30 days, notch member and target genes were down regulated. The results present as Mean ± SD. Compared to control group, *P < 0.05, **P < 0.01. (JPEG 513 kb)
Supplementary Fig. 9
Effect of DAPT on the expression of gonad development related genes of gonad in vivo. When the primordial gonad (40dph) were treated with DAPT for 30 days, amh, dax1, sf1, dmrt1 sox9a inhba and inhbb were down-regulated in testis of DAPT group compare to control group, foxl2, gdf9 and nobox were up-regulated in ovary of DAPT group. The results present as Mean ± SD. *P < 0.05, **P < 0.01. (JPEG 663 kb)
Supplementary Table 1
Oligonucleotide primers used for PCR. Some primers designed for amplifying were based on the known gene of NCBI, the number presenting the accession number, the others were based on common carp genome database. (DOCX 18 kb)
ESM 1
(XLS 129 kb)
ESM 2
(XLS 122 kb)
Rights and permissions
About this article
Cite this article
Jia, Y., Wang, F., Zhang, R. et al. Identification of suh gene and evidence for involvement of notch signaling pathway on gonadal differentiation of Yellow River carp (Cyprinus carpio). Fish Physiol Biochem 44, 375–386 (2018). https://doi.org/10.1007/s10695-017-0441-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10695-017-0441-5