Abstract
The brace-root is a crucial part of the whole root system in maize and plays an important role in the maintenance of lodging resistance. In this study, a BC2F1 population with three replicates, derived from the cross between Yi17 (well-developed brace-root) and Yi16 (poorly developed brace-root) was used for quantitative trait locus (QTL) detection. The total lengths of the genetic linkage map for the three replicates were 659.6, 662.9 and 936.8 cM, respectively, and the average distances between adjacent markers were 3.43, 3.68 and 4.61 cM, respectively. In total, 21 QTLs were detected in the BC2F1 population. The detected QTLs were mainly located at bin 3.05 (four QTLs) and 8.04–8.05 (three QTLs). Bin 3.05 was first detected to be rich in QTLs for brace-root traits in all three replicates, and QTLs detected in bin 8.04–8.05 were consistent with our previous results in the F2:3 population. The QTLs qW3a and qVA3 were coincident QTLs; of these, qW3a was a major effect QTL. These results may provide important information for maize breeders to pyramid favorable chromosome fragments or QTL in breeding programmes targeted at well-developed brace-root materials.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
An important food and feed crop, corn is the world’s highest yielding crop (Khush 2005; Craufurd and Wheeler 2009). Maize (Zea mays L.) has a complex root system, including underground roots and brace-roots (Hochholdinger et al. 2004). Brace-roots emerge from the stem nodes of successive basal phytomers and have a crucial role in plant development. Moreover, the brace-roots not only have the ability to improve lodging resistance but also exert a substantial influence on grain yield through the absorption of nutrients and water under soil water-limited conditions (Lynch 1995; Aiken and Smucker 1996; De Dorlodot et al. 2007; Li et al. 2015).
Maize researchers have devoted many efforts to study brace-root traits and to develop maize inbred lines and hybrids with well-developed brace-roots (Ku et al. 2012). Previous studies have shown that the maize brace-root number was positively correlated with maize yield (Stupar et al. 2008), and that brace-root number, root weight, and total root length were highly heritable (Trachsel et al. 2010). Moreover, a negative correlation between diameter and length of the main axis of the crown roots was observed in IBM (Intermated B73 × Mo 17 population) and OhW [Oh43 (Ames 19288) and W64A (PI 587152)] populations (Burton et al. 2014). However, only a few quantitative trait loci (QTLs) for brace-root traits have been reported in maize. Four seminal root number QTLs and seven root pulling force QTLs have been identified (Lebreton et al. 1995). Later, the QTLs for root angle and diameter in the seventh layer root were detected (Guingo et al. 1998). In addition, several QTLs for maize brace-root tier number (BRTN) were identified by two research groups using different populations (Ku et al. 2012; Jiao et al. 2014). Owing to the limitations of the experimental conditions and measurement tools, previous studies mainly focused on the early stage of root development or easily measured root traits, such as seminal root number, BRTN, and brace-root number. More important brace-root traits, such as brace-root surface area (BRSA), brace-root average diameter (BRAD), brace-root volume (BRV), and brace-root length were not studied in detail.
For the detection of QTLs related to maize brace-root traits, a BC2F1 population was constructed. For phenotype evaluation, we used a root scanner (WinRHIZO root system, http://www.regentinstruments.com/assets/winrhizo_about.html) to determine maize brace-root traits. We analyzed QTLs for seven important brace-root traits, including root dry weight (RDW), brace root dry weight (BRDW), BRTN, brace-root surface area (BRSA), brace-root average diameter (BRAD), brace-root volume (BRV) and brace root total length (BRTL). The purpose of this study was to provide references for the identification of brace-root related genes and for pyramiding favorable brace-root alleles in maize.
Materials and methods
Population development
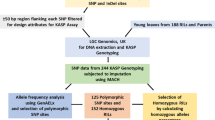
The parental lines Yi17, with a well-developed root system, and Yi16, with a poorly-developed root system, were derived from landrace ‘Baimaya’ and tropical germplasm ‘PWP’, respectively (Fig. 1). Both lines were developed by China’s Southwest University and used in maize breeding in the southwest area of China. The cross Yi17 × Yi16 (F1) was made in spring 2012 at Xiema, Chongqing, China. The F1 plants were backcrossed to Yi16 to produce the BC1F1 population in spring 2013. After phenotype screening, the individuals with well-developed brace-roots were continuously backcrossed to Yi16 to produce BC2F1 in spring 2015 (Fig. S1).
Field experiments and trait evaluation
In the BC2F1 population, three replicates, Q511 (198 plants from one ear), Q535 (252 plants from one ear), and DQT (586 plants from four ears), and both parental lines were planted at Xiema in spring 2016. The row spacing and plant spacing were 70 cm and 50 cm, respectively. All the plants were used for evaluation of the brace-root traits. At the R6 stage (physiological maturity), the whole root system of each plant was carefully pulled and washed out. After the soil was washed away and the moisture on the surface of roots was absorbed by tissue paper, the roots were manually dissected to determine the BRTN. The BRSA, BRAD, BRV, and BRTL were scanned and analyzed by using a root scanner (WinRHIZO root system). The fresh roots were dried at 80 °C in an oven until they reached a constant weight, and subsequently, BRDW, and RDW were determined.
Statistical analysis
The significance of traits between parents was determined by using a t test. Variation analysis of traits, skewness and kurtosis tests of traits distribution, and correlation analysis between brace-root traits for three replicates were computed using IBM SPSS Statistics 19.0 (IBM, USA).
DNA extraction and linkage map construction
DNA was extracted from the leaves of the BC2F1 plants and parent plants by using the CTAB method (Chen and Ronald 1999). Of 1020 SSR markers covering the entire maize genome (http://www.maizegdb.org), 224 polymorphic markers were used to construct genetic linkage maps by using JoinMap v.4.0 (Stam 1993). The recombination frequency between linked loci was transformed into genetic distance by using Kosambi’s function (Kosambi 1943). The total length was 659.6 cM for the Q511 replicate, 662.9 cM for Q535, and 936.8 cM for DQT, and the average distances were 3.43 cM, 3.68 cM, and 4.61 cM, respectively.
QTL analysis
QTL mapping was performed using the integrated software MAPQTL 6.0 (https://www.kyazma.nl/) based on interval mapping. The parameters for forward regression analysis were set with a window size of 10 cM and a walk speed of 1 cM. The significance threshold for declaring a QTL was determined by 1000 random permutations of the phenotypic data at the 5% significance level according to Stuber et al. (1987) and Tuberosa et al. (1998a, b). The averages of all the brace-root traits were used for QTL analysis. The logarithm of odds (LOD) threshold used in the study was averaged from the threshold value calculated for each trait. The closest QTL peak to the marker locus position, the additive effect, and the proportion of phenotypic variance explained by the QTL (R2) were obtained by using MAPQTL 6.0. Map Chart Vision 2.2 was used to mark the QTL location on the map. The QTL was named in the following way, by taking qW3a as an example; ‘q’ is the abbreviation of QTL; ‘W’ is an abbreviation for trait, namely root weight; the number ‘3’ is the serial number of the chromosome; and ‘a’ is the symbol used for differentiating the QTLs that controlled the same traits but were located at different chromosome fragments in the same linkage group.
Results
Phenotypic variation
The phenotypic analyses of the parents and BC2F1 plants are shown in Table 1. All the trait values of Yi17 were significantly higher than those of Yi16 (P < 0.01), which indicated that genetic differences existed between the two parents for all the traits. The mean of each trait in BC2F1 was between the two parents’ phenotypic values, and transgressive segregation was also observed for each trait. The skewness and kurtosis showed that all the traits were normally or nearly normally distributed. Therefore, these traits were controlled by polygenes and suitable for QTL analysis.
Correlation between different traits
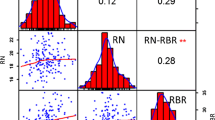
To further elucidate the relationships between QTLs and traits, it was necessary to perform a correlation analysis. As shown in Table 2, the correlations varied with different traits. Generally, RDW, BRDW, BRSA, BRV, and BRTL were all highly correlated with each other (r = 0.720–0.984). RDW, BRDW, BRSA, BRV and BRTL were all moderately correlated with BRTN (r = 0.381–0.610). BRAD was moderately correlated with BRV (r = 0.411–0.634). In comparison with other traits, BRAD showed the weakest correlation with BRTN and BRTL (r = − 0.076–0.196). Overall, BRTL was the most important trait in all three simple traits, including BRTN, BRAD and BRTL.
QTL mapping using the Q511 replicate
In the Q511 replicate, eight QTLs for all traits were detected in total (Table 3; Fig. 2). qW3a, located in the mark interval mmc0022–umc1973 on chromosome 3, was relevant to root weight traits, and explained 11.4–12.6% of phenotypic variation. qWAVL5, explaining 8.3–15.7% of phenotypic variation, was relevant to BRDW, BRV, and BRSA traits. qT7, qAL8, qD8, and qV8, explaining 7.1–12.9% of phenotypic variation, were located in the mark intervals umc1713–zct657 on chromosome 7, and umc1460–umc1858, umc2367–umc1562 and bnlg2046–umc2367 on chromosome 8, respectively.
Distribution of QTLs for all traits on a linkage map in three replicates. (1) RDW root dry weight; BRDW brace-root dry weight; BRTN brace-root tier number; BRSA brace-root surface area; BRAD brace-root average diameter; BRV brace-root volume; BRTL brace-root total length. (2) Red represents QTLs together detected in Q511 and Q535 replicates. Green represents QTLs together detected in DQT and Q535 replicates. (3) a Q511 replicate; b Q535 replicate; c DQT replicate. (Color figure online)
QTL mapping using the Q535 replicate
In the Q535 replicate, 11 QTLs were identified in total (Table 3; Fig. 2). These QTLs were distributed on chromosomes 1 (3 QTLs), 3 (3 QTLs), 4 (1 QTL), 6 (1 QTLs), 8 (1 QTL), 9 (1 QTL), and 10 (1 QTL). Among the 11 QTLs, only two were major effect QTLs. In bin 3.04–3.05, three QTLs explaining 7.4–13.8% of phenotypic variation were identified; of these, qW3a and qW3b for root weight traits, explained 10.8–13.8% of phenotypic variation and were located in the mark intervals mmc0022–umc1973, and umc2002–bnlg1035, respectively; qVA3, located in the mark interval bnlg1035–umc2155 within a distance of 0.9 cM, was relevant to BRSA and BRV.
QTL mapping using the DQT replicate
In the DQT replicate, four QTLs were identified (Table 3; Fig. 2). qVA3, located in the interval umc2155–bnlg1035 at bin 3.05, was relevant to BRDW, BRV, and BRSA traits. In the interval of bnlg339–umc1718 on chromosome 7, one major effect QTL, qD7, was detected with an LOD of 5.55.
Discussion
BRDW and BRSA are both complex traits, composed of BRTN, BRTL and BRAD. They are closely related to the ability to uptake water and nutrients from soil, and lodging resistance. Among the three simple traits, BRTL was the most correlated with BRDW and BRSA as shown by correlation analysis. Therefore, efforts should be made to increase the BRTL in the maize breeding program.
In this study, in total, 21 QTLs, with only two coincident QTLs, were detected in all three replicates. This suggested that the genes controlling the brace-root traits largely varied with the different environments. Coincident and major effect QTLs were the most important for plant breeding. qW3a, related to BRDW traits, and qVA3, related to BRDW and BRAD traits, were coincident QTLs in this study. In particular, qW3a explained 10.8% to 12.9% of the phenotypic variance. Furthermore, two regions (bin 3.05 and bin 8.04–8.05) were detected to be rich in QTLs for brace-root traits. QTLs in bin 1.06 and 7.04 were reported to influence the maize root system architecture, such as primary root length, primary root diameter, primary root weight, and root volume. (Hund et al. 2011; Tuberosa et al. 2002; Liu et al. 2017). A QTL region in bin 6.02 within the interval umc1006-umc1257 was found to control vertical root pulling resistance in the BC4F3 and RIL populations (Liu et al. 2010). Based on the different stages of root development, most QTLs for root traits were located in bin 6.02–6.04 on chromosome 6 in stage I, and the QTL for axial root number was located in bin 10.04 on chromosome 10 at all three developmental stages (Cai et al. 2012). Gu et al. (2017) detected two coincident QTLs for BRTN and BRAD in bin 8.05 in the F2 and F2:3 populations in multiple environments. This showed that the region bin 8.05 harbored QTLs for brace-root traits in previous studies, as well as in our study. The region bin 3.05 was newly detected to harbor QTLs for brace-root traits. This new coincident region bin 3.05 was associated with many brace-root traits, such as BRDW, BRSA, BRTN, BRV, and root dry weight. This revealed that these traits were probably controlled by a single pleotropic effect QTL. Therefore, the regions bin 3.05 and bin 8.05, and the coincident and major effect QTL qW3a are of great importance in exploring brace-root-related genes and for breeding inbred maize lines and hybrids with well-developed brace-roots.
Abbreviations
- Add:
-
Additive effect
- BRAD:
-
Brace-root average diameter
- BRDW:
-
Brace-root dry weight
- BRSA:
-
Brace-root surface area
- BRTL:
-
Brace-root total length
- BRTN:
-
Brace-root tier number
- BRV:
-
Brace-root volume
- cM:
-
CentiMorgan
- IF2 :
-
Immortalized F2
- LOD:
-
Logarithm of odds
- RIL:
-
Recombinant inbred line
- QTL:
-
Quantitative trait locus
- R 2 :
-
Phenotypic variance explained
- SSR:
-
Simple sequence repeat
References
Aiken RM, Smucker AJM (1996) Root system regulation of whole plant growth. Annu Rev Phytopathol 34:325–346
Burton AL, Johnson JM, Foerster JM, Hirsch CN, Buell CR, Hanlon MT (2014) QTL mapping and phenotypic variation for root architectural traits in maize (Zea mays L.). Theor Appl Genet 127:2293–2311
Cai HG, Chen FJ, Mi GH, Zhang FS, Maurer HP, Liu WX, Reif JC, Yuan LX (2012) Mapping QTL for root system architecture of maize (Zea mays L.) in the field at different developmental stages. Theor Appl Genet 125:1313–1324
Chen DH, Ronald PC (1999) A rapid DNA minipreparation method suitable for AFLP and other PCR applications. Plant Mol Biol Rep 17:53–57
Craufurd PQ, Wheeler TR (2009) Climate change and the flowering time of annual crops. J Exp Bot 60:2529–2539
De Dorlodot S, Forster BP, Pages L, Price A, Tuberosa R, Draye X (2007) Root system architecture: opportunities and constraints for genetic improvement of crops. Trends Plant Sci 12:474–481
Gu D, Mei X, Yu T, Sun NN, Xu D, Liu CX, Cai YL (2017) QTL identification for brace-root traits of maize in different generations and environments. Crop Sci 57:13–21
Guingo E, Hebert Y, Charcosset A (1998) Genetic analysis of root traits in maize. Agronomie 18:225–235
Hochholdinger F, Park W, Sauer M, Woll K (2004) From weeds to crops: genetic analysis of root development in cereals. Trends Plant Sci 9:42–48
Hund A, Reimer R, Messmer R (2011) A consensus map of QTL controlling the root length of maize. Plant Soil 344:143–158
Jiao FC, Li YX, Chen L, Liu ZZ, Shi YS (2014) Genetic dissection for kernel row number in the specific maize germplasm four-rowed waxy corn. Sci Agric Sin 47:1256–1264
Khush GS (2005) What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol 59:1–6
Ku LX, Sun ZH, Wang CL, Zhang J, Zhao RF, Liu HY, Tai GQ, Chen YH (2012) QTL mapping and epistasis analysis of brace root traits in maize. Mol Breed 30:697–708
Kosambi DD (1943) Statistics in function space. J Ind Math Soc 7:76–88
Lebreton C, Lazicjancic V, Steed A, Pekic S, Quarrie SA (1995) Identification of QTL for drought responses in maize and their use in testing causal relationships between traits. J Exp Bot 46:853–865
Li PC, Chen FJ, Cai HG, Liu JC, Pan QC, Liu ZG, Gu RL (2015) A genetic relationship between nitrogen use efficiency and seedling root traits in maize as revealed by QTL analysis. J Exp Bot 66:3175–3188
Liu JC, Cai HG, Chu Q, Chen XH, Chen FJ, Yuan LX, Mi GH, Zhang FS (2010) Genetic analysis of vertical root pulling resistance (VRPR) in maize using two genetic populations. Mol Breed 28:463–474
Liu ZG, Gao K, Shan SC, Gu RL, Wang ZK, Craft EJ, Mi GH, Yuan LX, Chen FJ (2017) Comparative analysis of root traits and the associated QTL for maize seedlings grown in paper roll, hydroponics and vermiculite culture system. Plant Sci 8:436
Lynch J (1995) Root architecture and plant productivity. Plant Physiol 109:7–13
Stam P (1993) Construction of integrated genetic linkage maps by means of a new computer package: Join Map. Plant J 3:739–744
Stuber CW, Edwards MD, Wendel JF (1987) Molecular marker-facilitated investigations of quantitative trait loci in maize. II. Factors influencing yield and its component traits. Crop Sci 27:639–648
Stupar RM, Gardiner JM, Oldre AG, Haun WJ, Chandlcr VL, Springer NM (2008) Gene expression analyses in maize inbreds and hybrids with varying levels of heterosis. BMC Plant Biol 8(1):33
Trachsel S, Kaeppler SM, Brown KM, Lynch JP (2010) Shovelomics: high throughput phenotyping of maize (Zea mays L.) root architecture in the field. Plant Soil 341:75–87
Tuberosa R, Parentoni S, Kim TS, Sanguineti MC, Phillips RL (1998a) Mapping QTL for ABA concentration in leaves of a maize cross segregating for anthesis date. Maize Genet Coop Newsl 72:72
Tuberosa R, Sanguineti MC, Landi P, Salvi S, Casarini E, Conti S (1998b) RFLP mapping of quantitative trait loci controlling abscisic acid concentration in leaves of drought-stressed maize (Zea mays L.). Theor Appl Genet 97:744–755
Tuberosa R, Sanguineti MC, Landi P, Giuliani MM, Salvi S, Conti S (2002) Identification of QTL for root characteristics in maize grown in hydroponics and analysis of their overlap with QTL for grain yield in the field at two water regimes. Plant Mol Biol 48:697–712
Acknowledgements
The authors would like to thank the key Research Projects of Chongqing (cstc2016shms-ztzx80016, cstc2016shms-ztzx80017), Research Fund for the Doctoral Program of Southwest University (SWU114035), and Fundamental Research Funds for the Central Universities (XDJK2018C052) for providing financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, N., Liu, C., Mei, X. et al. QTL identification in backcross population for brace-root-related traits in maize. Euphytica 216, 32 (2020). https://doi.org/10.1007/s10681-020-2561-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-020-2561-8