Abstract
Americano 44 is a landrace derived wheat cultivar from Uruguay that has had long lasting resistance to the leaf rust pathogen Puccinia triticina. A population of 92 recombinant inbred lines derived from the cross ‘Thatcher*3/Americano 44d’ was evaluated for segregation of adult plant leaf rust resistance in four field plot tests and two greenhouse tests. A genetic map was constructed with 381 Diversity Array Technology markers, five simple sequence repeat markers, and seven kompetitive allele specific PCR markers. Significant effects for reduction of leaf rust severity were found for quantitative trait loci (QTL) on chromosomes 3AS, 3DS and 6DS. Individually the 3AS and 3DS regions did not decrease leaf rust severity in any of the tests compared to the susceptible parent, yet strongly interacted when present in the same genotype to decrease leaf rust severity equal to the resistant parent. Genotypes with only the 6DS region had significantly lower leaf rust severity compared to the susceptible parent in three of the field plot tests. The 6DS QTL also interacted with the 3DS QTL to decrease leaf rust severity. Major QTLs on chromosome 3AS and 3DS are both located in the distal region and may be encoded by homoeoalleles.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leaf rust of wheat (Triticum aestivum L.) caused by Puccinia triticina Eriks. is a widely distributed disease that occurs on an annual basis in many major wheat growing regions of the world. The leaf rust fungus is highly variable with 50–70 races described every year in the U.S. (Kolmer and Hughes 2015). To date over 70 genes for leaf rust resistance in wheat have been mapped to a chromosome location (McIntosh et al. 2017). Many of these genes condition highly effective resistance to specific races of the leaf rust fungus in wheat seedlings and in adult plants, but do not provide resistance to other races. In the Great Plains region of the U.S., the continual release of cultivars with race specific leaf rust resistance genes Lr21, Lr24, Lr37 and Lr39 have placed a constant selection pressure for virulence in the P. triticina population. As a result of the broad environmental adaption of P. triticina (Roelfs et al. 1992) and the presence of many races with virulence to important leaf rust resistance genes, stable and long lasting resistance in wheat to leaf rust has been difficult to achieve.
Other genes, most notably Lr34, Lr46, Lr67 and Lr68 (Lagudah 2011) are most effective in the adult plant stage and are effective to all current races of P. triticina. These genes are optimally expressed in the adult plant stage and usually condition an intermediate level of resistance with the production of fewer and smaller uredinia, compared to susceptible genotypes, when present in a single gene wheat line. Combinations of these genes, or genes with race specific and race nonspecific resistance have provided long lasting resistance to leaf rust in released cultivars (Oelke and Kolmer 2005).
Cultivar Americano 44d was derived from a landrace population in Uruguay and released in 1918. This cultivar was noted to have long lasting resistance to leaf rust (Roelfs 1988). A previous study (Kolmer et al. 2007) determined that this cultivar had at least two genes that conditioned adult plant resistance. Adult plants of backcross lines of ‘Thatcher’ (Tc) wheat with leaf rust resistance derived from Americano 44d had a different leaf rust resistance phenotype compared to Tc lines with Lr13, an adult plant gene that conditions race specific resistance, and Tc lines with Lr34. The objective of this research was to characterize and map the adult plant resistance derived from Americano 44d in a partial Thatcher background to enable the direct comparison of this resistance with other Thatcher lines that are near-isogenic for adult plant and seedling leaf rust resistance genes (McIntosh et al. 2017). Since Americano 44d is a spring wheat with late maturity, a backcross line of Thatcher with adult plant leaf rust resistance derived from Americano 44d was crossed with Thatcher in order to develop a population of recombinant inbred lines that were relatively uniform for maturity and plant height.
Materials and methods
Population development
Tc*2/Americano 44d F4 lines that had leaf rust severity of 10–20% and response of moderately resistant to moderately susceptible in field plots (Kolmer et al. 2007) were advanced to F6 generation by single seed descent. These lines lacked any seedling resistance to P. triticina isolates with race designations BBBDB and SBDGG (Kolmer et al. 2007; Long and Kolmer 1989). The F6 lines were also tested for resistance as adult plants in greenhouse tests with an isolate of P. triticina with race designation THBJG (Supplemental Table 1). Seed from a single F6 plant designated as Tc*2/Americano 44d 39A, that had clearly discernable adult plant resistance in a greenhouse test, was chosen for crossing with seed derived from a single plant selection of Tc. Tc*3/Americano 44d 39A recombinant inbred lines (RIL) were advanced to the F6:7 to F6:8 generations in a greenhouse.
Leaf rust assessment
One hundred RILs and the two parental lines were evaluated for field resistance in St. Paul MN, in 2011, 2012, 2013, and 2014 and in adult plant greenhouse tests in 2009 and 2011. In the greenhouse test in 2009, F6 RILs were used; in the field tests in 2011, 2012, 2013 and the greenhouse test in 2011, F6:7 lines were used; and in the field test in 2014, F6:8 lines were used. In the field plot tests, 50–60 seed of the parents and each RIL were planted in 2-m rows without replication spaced 30 cm apart perpendicular to spreader rows of the wheat cultivars Tc, ‘Morocco’, ‘Max’, and ‘Little Club’, that are all susceptible to leaf rust. A mixture of P. triticina isolates with race designations MLDSD, TDBGG, MFPSB, THBJG, TNRJJ, and MCRKG in 2011; MHDSB, MFPSB, MLDSD, TBBGJ, and TFBGQ in 2012 and 2013; and MHDSB, MFPSB, MLDSD, TBBGJ, TFBJQ, TFBGQ, and TBBGS in 2014 (Supplemental Table 1), were inoculated to the spreader rows approximately 30 days after planting, when the plants were at jointing stage. Urediniospores were mixed with Soltrol 170 oil (Chevron Phillips, The Woodlands, TX) and the suspension was atomized onto the spreader rows with an ultralow volume applicator. The parents and RILs were evaluated once for leaf rust severity and response approximately 60 days after planting using the modified Cobb scale (Peterson et al. 1948). Leaf rust resistance response was rated R = necrosis surrounding small uredinia; MR = necrosis surrounding moderate size uredinia; MS = distinct chlorosis surrounding moderate to large uredinia; S = large uredinia lacking necrosis or chlorosis. The plots were rated when the susceptible parent Tc had leaf rust severity of 60–80% with a susceptible response. Five flag leaves collected from different parts of the row were assessed for severity and response. In the greenhouse tests, four seeds of the parents and each RIL were planted in 15-cm pots filled with steamed topsoil. The plants were grown in a greenhouse set at 20° C with a 16 h light period. The RIL population was inoculated at anthesis with the same mixture of P. triticina isolates used in the field plot tests. The flag leaves were inoculated individually with a mixture of Soltrol 170 oil and urediniospores. Eight to 10 flag leaves of each RIL and parent were inoculated with an atomized mixture of 5 mg of rust spores and 0.750 ml of oil. After inoculation the plants were dried for at least one hour and then placed in a closed chamber with a humidifier filled with distilled water running for 60 s every five minutes. The adult plants were removed the following morning and placed on a greenhouse bench. The RILs and parents were evaluated for infection type using the standard 0–4 scale (Long and Kolmer 1989) 14 days after inoculation. Infection type scores were converted to a 0–100 score for quantitative trait analysis with a scale similar to that used in Turner et al. (2016). Leaf rust severity in the greenhouse tests were also recorded, however there were little differences in severity among the RILs, therefore only the converted infection type scores were used for subsequent analysis. The leaf rust severity (percentage of leaf area covered) of the RILs in the four field tests in St. Paul and the converted infection type in the two greenhouse tests were tested for correlation using Pearson’s coefficient with PROC CORR in Statistical Analysis Software (SAS, Cary NC).
DNA genotyping
High quality DNA of the parents and 92 of the F6 RILs was isolated using the CTAB buffer method recommended by Diversity Arrays Technology (DArT) (Triticarte Pty Ltd, Canberra, Australia) (Akbari et al. 2006). DNA samples of the RILs and parents were genotyped with the DArT methodology. Polymorphic markers were indicated as present (1) or absent (0).
Simple sequence repeat (SSR) markers were amplified from DNA of the mapping population. PCR assays were conducted in a 10 μl reaction volume containing 15 ng DNA, 0.25 U Taq DNA polymerase, 1× PCR buffer, 100 nM each of forward and reverse primer, 200 μM dNTP and 1.5 mM MgCl2. Amplifications were performed in a PTC-200 thermocycler (MJ Research, Waltham MA) using the following cycling conditions: 93 °C initial denaturation followed by 35–38 cycles (depending on primer pair) of denaturation at 93 °C for 30 s, annealing at various temperatures depending on the primer pair for 40 s, and extension at 72 °C for 1 min, followed by 5 min at 72 °C. The amplification products were separated on vertical polyacrylamide gels and visualized via silver staining using the method of Liu and Anderson (2003).
To develop breeder-friendly markers for genomic regions conferring leaf rust resistance, selective genotyping was performed on 44 selected F6 RILs using the Infinium iSelect 90 K wheat bead chip array described in (Wang et al. 2014). Based on DArT marker genotypes and leaf rust disease response scores, 12 RILs were selected for high rust severity and response, and absence of resistance alleles at three QTLs identified, while the remainder were selected for low rust response and fixation for one or more resistance alleles of the identified QTLs. Analysis of the chip array data identified 265 markers linked to the 6D region, 43 markers linked to the 3D region and 30 markers linked to the 3A region. A subset of 20 SNPs were selected that were located in one of three QTL regions based on the consensus 90 K SNP map (Wang et al. 2014). KASP primer sequences were downloaded from Polymarker (www.polymarker.tgac.ac.uk) and markers were assayed following the manufactures’ instructions on a CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad). Eleven KASP markers produced good cluster patterns of allele calls that fell into two discrete groups and were genotyped in the entire RIL population. Seven KASP markers with confirmed chromosome locations based on the physical map of ‘Chinese Spring’ (CS) wheat in the RefSeq v1.0 database (http://www.wheatgenome.org/News/Latest-news/RefSeq-v1.0-URGI) were used in the final map.
Quantitative trait locus (QTL) analysis
Since the parental lines were highly related, a reduced number of segregating DArT and SNP markers would be expected in chromosomal regions not associated with leaf rust resistance. However, because the resistant parent was selected for leaf rust resistance in comparison with Thatcher, it would be expected to find segregating markers in regions associated with adult plant leaf rust resistance. Linkage groups of segregating DArT, SSR and KASP marker loci were assembled with Mapmaker v2.0 (Lander et al. 1987) for MacIntosh using the Kosambi mapping function with a logarithm of odds (LOD) of 4 and r = 0.3. Marker order was confirmed using R/qtl (Broman et al. 2003). Composite interval mapping (CIM) was performed with QGENE (Nelson 1997) to calculate the coefficient of determination (R2) and LOD scores for each marker interval at a significance level of α = 0.05 with 1000 permutations of the dataset. The marker closest to the LOD peak was selected by QGENE as a cofactor. Markers associated with leaf rust severity were also identified by single factor regression with SAS. The datasets were evaluated with PROC GLM in SAS to evaluate interactions between tests and marker regions associated with leaf rust resistance. Differences in leaf rust severity between RIL genotypes were assessed with Tukey’s honest significant difference in PROC GLM at p < 0.05.
Confirmation of the chromosome locations of mapped SSR markers was completed by aneuploid analysis. DNA of CS, and select CS chromosome 3A and 3D nullisomic-tetrasomic and ditelosomic genetic stocks was isolated using Qiagen DNeasy (Qiagen, Hilden, Germany) kits according to the manufacturers’ instructions. Subsequently, the DNA was used for PCR amplification of SSR markers, gel electrophoresis and fragment visualization as described for SSR marker acquisition in the mapping population.
Chromosome location and map data for the DArT, SSR, and KASP markers for the RILS and the leaf rust severity data for all greenhouse and field plots tests are available in the supplemental materials.
Results
Inheritance of leaf rust resistance
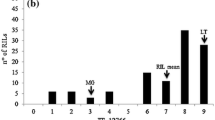
The Tc*3/Americano 44d 39A RILs showed a wide range of infection types for leaf rust resistance in the two greenhouse tests and in the four field plot tests (Fig. 1). In the two greenhouse tests, the most resistant RILs and the Tc*2/Americano 44d 39A parent had flecks with small necrotic uredinia mixed with moderate to large size uredinia that lacked chlorosis or necrosis that translated to a score of 10 on the 0–100 scale. The susceptible parent Thatcher and the most susceptible RILs had moderate to large size uredinia that lacked any chlorosis or necrosis which translated to scores of 60–70. The Tc*2/Americano 44d 39A parent had between 10 and 40% leaf rust severity in the four field tests and the susceptible parent Thatcher had between 50 and 70% leaf rust severity (Fig. 1). In the 2013 field plot test, the Tc*2/Americano 44d 39A parental line had a response of 40MR-MS. The higher severity for that year may have been due to heavy inoculum pressure in that part of the field plot. There was a strong correlation between severity and resistance response in the RIL population in the field plot tests. Lines that had low severity also had a stronger resistance response, while lines with high severity had a more susceptible response. The leaf rust severity ratings of the Tc*3/Americano 44d 39A RILs had a good correlation overall in the six tests. The two greenhouse tests had a correlation of 0.68, and correlations between 0.39 and 0.66 with the four field tests (Table 1). The four field tests had higher correlations that varied from 0.75–0.82.
Genetic mapping and QTL analysis
A total of 397 polymorphic markers in 92 F6 RILs were grouped into 20 linkage groups that covered 759 cM. Three hundred eighty-one DArT markers were used to construct the initial map. Single factor regression and composite interval mapping of the genotyped RILs identified regions on chromosomes 3A, 3D and 6A that were associated with leaf rust severity. Five SSR markers cfd79U, cfd79L, wmc532, barc12 and barc321, and seven KASP markers on chromosomes 3AS, 3DS, and 6DS were added to the linkage map (Supplemental Table 2). The putative chromosome arm locations of the linkage groups on chromosomes 3A and 3D were confirmed by aneuploid analysis of the mapped SSR markers. The linkage group on chromosome 3A was found to be located on the short arm of this chromosome based upon the absence of the SSR markers in both CS nulli3A-tetra3D and ditelo 3AL aneuploid genetic stocks. Similarly, the linkage group on chromosome 3D was localized to the short arm of that chromosome based on the lack of SSR makers in 3D aneuploid lines.
The region on chromosome 3A associated with leaf rust severity was designated as QLr.cdl.-3A, with the SSR marker barc321 closest to the LOD peak (Figure 2a; Table 2). In the two greenhouse tests this region accounted for 27% and 19% of the total variation with LOD scores of 5.9 (p < 0.01) and 4.1 (p < 0.01), in 2009 and 2011 respectively. In the field tests, the LOD peak was 2.7 (p < 0.10) with an R2 of 0.13 in 2012, and the LOD peak was less than 1.0 in the other field tests, with low values of R2. In the single factor regression over all the tests, this region had an R2 value of 0.09 (p < 0.0001). KASP marker K74975 was 2.2 cM distal to barc321 on chromosome 3AS in the physical map of CS chromosome 3AS based on the alignment of the 90 K SNP probe in the NRGene RefSeq v1.0 assembly (Supplemental Table 2).
The region on chromosome 3D associated with leaf rust severity was designated as QLr.cdl-3D, with wPt-741949 the marker closest to the LOD peak at 38 cM (Fig. 2b, Table 2). In the two greenhouse tests and in the four field tests, this region accounted for 17–32% of the total variation. In greenhouse tests in 2009, 2011, and field tests in 2012 and 2014 this region had LOD scores > 4.0 with p < 0.01. In the field tests in 2011 and 2013 this region had LOD scores > 3.0 with p < 0.05. In the single factor regression over all the tests, this region had an R2 value of 0.19 (p < 0.0001). The QTL was detected distal to KASP marker K16928 that was located at 5.6 Mbp on the physical map of chromosome 3DS in CS wheat. (Supplemental Table 2). Given the distal location of both QTL on chromosome 3AS and 3DS, it is possible that genes in homoeologous regions carry leaf rust resistance QTLs.
The region on chromosome 6D associated with leaf rust severity was designated as QLr.cdl-6D, with wPt-664670 as the closest marker to the LOD peak at 0 cM (Fig. 2c, Table 2). This region had LOD scores greater than 3.0 with p < 0.01 in the field 2013 test, and p < 0.05 in the field 2012 and 2014 tests. The QLr.cdl-6D region did not have a LOD score greater than 3.0 in the two greenhouse tests or in the field 2011 tests. This region had R2 values between 0.15 and 0.23. In the single factor regression over all the tests, this region had an R2 value of 0.10 (p < 0.0001). This QTL was 3.4 cM distal to KASP marker K37229, which was located at 42.0 Mbp on the physical map of chromosome 6DS in CS wheat (Supplemental Table 2).
The 92 F6 RIL genotypes were placed into eight groups based on the presence of resistance alleles at QLr.cdl-3A, QLr.cdl-3D and QLr.cdl-6D (Table 3). The genotype × environment interaction was non-significant for the both the two greenhouse tests (p = 0.24) and four field tests (p = 0.06), which allowed the severity scores to be averaged over the two types of tests. Lines with missing data at these loci or lines that were heterozygous were excluded. In both greenhouse and field tests, RILs with resistance alleles at QLr.cdl-3A and QLr.cdl-3D, and lines with resistance alleles at all three QTLs had the lowest leaf rust severity scores compared to all other genotypes. When considered singly, RILs with resistance alleles at QLr.cdl-3A or QLr.cdl-3D did not differ significantly for lower leaf rust severity from lines that lacked resistance alleles at all three QTLs. RILs with resistance at only QLr.cdl-6D had significantly lower leaf rust severity scores averaged over the four field tests compared to lines that lacked resistance alleles at all three QTLs, but in the greenhouse tests were not significantly different compared to lines that lacked all resistance alleles. RILS with resistance alleles at QLr.cdl-3A and QLr.cdl-6D did not differ significantly for leaf rust severity compared to lines that lacked resistance at all three QTLs in the greenhouse and field tests. RILs with resistance alleles at QLr.cdl-3D and QLr.cdl-6D had significantly less rust severity compared to lines that lacked resistance alleles at all three QTLs over the four field tests, but were not significantly different in the greenhouse tests.
The ANOVA (Supplemental Table 3) of leaf rust severity at the three QTLs over the six tests showed a strong interaction between resistance alleles at QLr.cdl-3A and QLr.cdl-3D (p < 0.001), and between resistance alleles at QLr.cdl-3A, QLr.cdl-3D and QLr.cdl-6D for decreased leaf rust severity (p < 0.001). There was also a significant interaction (p < 0.02) for decreased leaf rust severity between resistance alleles at QLr.cdl-3D and QLr.cdl-6D. There were significant effects for Test × QLr.cdl-3A (p < 0.001), and for Test × QLr.cdl-3A × QLr.cdl-3D (p = 0.04). The QLr.cdl-6D region also had a significant Test × genotype effect (p = 0.03).
Discussion
The inheritance of adult plant leaf rust resistance derived from Americano 44d was complex, with three genomic regions (QLr.cdl-3A, QLr.cdl-3D and QLr.cdl-6D) found to interact for lower leaf rust severity when present in a single genotype.
Resistance alleles at the QLr.cdl-3A and QLr.cdl-3D regions in particular strongly interacted to condition an intermediate level of leaf rust resistance. Neither of these regions conditioned effective resistance when present singly. Only the QLr.cdl-3DS region had consistently significant (p < 0.05) LOD scores in all of the tests. The RILs with the resistance allele at QLr.cdl-3A singly had the highest severity scores in all the tests. RILs with the resistance allele at only QLr.cdl-6D had lower average leaf rust severity over the four field tests. This region also interacted with the 3D region for decreased leaf rust severity. In contrast, lines with resistance alleles at both QLr.cdl.3A and QLr.cdl.6D did not differ significantly for severity compared to lines that lacked resistance alleles at all three QTLs.
The leaf rust resistant QTL expressed very differently in the greenhouse and field plot tests. The QLr.cdl-3A region had significant LOD scores (p < 0.05) only in the field plot tests. This was reflected in the overall high genotype × environment interaction for this region. The QLr.cdl-6D region had significant LOD scores (p < 0.05) in the four field tests, but not in the two greenhouse tests. This was also seen in the significant overall genotype × environment interaction for this region. The QLr.cdl-3D region expressed resistance in both field and greenhouse tests, and did not have an overall significant genotype × environment interaction.
The three QTL that interact for increased resistance may have different modes of action that are complementary. The QLr.cdl-3A region was effective mostly in the greenhouse tests that measured infection type, while the QLr.cdl-6D region was effective only in field plot tests that were rated using severity. It is likely that different resistance mechanisms affect the infection process that the QLr.cdl-3A region affected, compared to the mechanisms that slow the growth and sporulation of the leaf rust fungus, that were affected by the QLr.cd.-6D region. The QLr.cdl-3D region was effective in both field and greenhouse tests, and may provide resistance during the infection process and may also restrict growth and sporulation of the leaf rust fungus.
The different races of P. triticina used in the field and greenhouse experiments over the years may have also effected the expression of the resistance QTL. There is a possibility of some degree of race specificity among the three QTL, which may account for some of the inconsistent expression of QLr.cdl-3A and QLr.cdl-6A. The consistent effectiveness of QLr.cdl-3D with the different mixtures of races, suggests that this QTL may provide non-specific race resistance.
In a previous study, Tc*2/Americano 44d BC1F2 families segregated in a 3:1 ratio in a field plot test for segregating and homozygous susceptible families respectively (Kolmer et al. 2007). This suggested that at least two genes conditioned the adult plant resistance derived from Americano 44d. Since Americano 44d has much later maturity compared to Thatcher, the BC1F2 families varied considerably within and between families for maturity, making a uniform assessment of leaf rust severity difficult. In the current study, the Tc*3/Americano 44d 39A RILs had relatively uniform maturity, thus making assessment of leaf rust severity more straightforward. Because a single Tc*2/Americano 44d line was chosen as the resistant parent, it is possible that other adult plant leaf rust resistance genes or QTLs that are present in the original source of resistance, Americano 44d, were not detected in this study.
QTLs for leaf rust resistance have been studied extensively in numerous studies. Soriano and Royo (2015) listed three meta-QTL on chromosome 3A that included QTLs from the cultivars Forno and Oberkulmer (Messmer et al. 2000); line TA4152-60 (Chu et al. 2009); and the durum cultivar Lloyd (Maccaferri et al. 2008). Lan et al. (2014) described a QTL for leaf rust resistance on 3DS, however none of the markers in their study were in common with the markers associated with QLr.cdl-3D. In two recent summaries (Li et al. 2014; Soriano and Royo 2015), no QTLs for leaf rust resistance were described on chromosome 6D. The chromosome arm location of the three QTL described in this paper were confirmed by markers in common between this study and other mapping populations. Liu et al. (2013) mapped barc321, which was the closest marker for QLr.cdl-3A, to chromosome 3AS. Cui et al. (2014) mapped wPt-730651 and wPt-667139 to 3DS. Singh et al. (2013) mapped wPt-664670, the closest marker for QLr.cdl-6D, to the distal region of 6DS. The chromosomal assignment for the three QTLs was further supported by ditelosomic assignment, and genetic mapping performed for the linked SNPs in Wang et al. (2014). QLr.cdl-3A is unlikely to be the same QTL mapped in Forno and Oberkulmer (Messmer et al. 2000), or TA-4152-60 (Chu et al. 2009), since they were mapped to chromosome 3AL. The QTL from the durum wheat Lloyd (Maccaferri et al. 2008) mapped to 3AS in the same region as QLr.cdl-3A based on the consensus map of Li et al. (2014).
One interesting result of our study was the finding that separate SSR amplicons for cfd79 were located in proximity to the QTLs detected on chromosome arms 3AS and 3DS. Amplification of wheat DNA with cfd79 primers results in three amplicons. The largest amplicon, designated cfd79U, mapped to a locus approximately 51.2 cM from the best QTL marker on chromosome 3A. Similarly, the smallest amplicon detected, cfd79L, mapped approximately 37.8 cM from the closest QTL marker on chromosome 3D. Fine mapping and isolating genes that confer resistance on both chromosomes will ultimately determine if these QTLs are encoded by homoeoalleles.
Use of Americano 44d as a leaf rust resistant parent would be complicated by late maturity and by the relatively complex inheritance and small effects of the individual QTLs. However the three identified QTLs may be useful in combination with major adult plant resistance genes such as Lr34. The winter wheat Forno that had broadly effective durable resistance was determined to have Lr34 plus a number of other QTLs with small effects (Schnurbusch et al. 2004). Combinations of other less effective adult plant resistance genes, such as Lr46, with QTLs that have small effect can also be useful in breeding projects. Kolmer (2015) found that a QTL on chromosome 5BL with small effects additively enhanced the resistance of Lr46, derived from the wheat landrace derived cultivar Americano 25e. The single-marker KASP assays developed in this study for SNPs closely linked to each of the three QTLs will facilitate the pyramiding of rust resistance loci from Americano 44d into future cultivars.
References
Akbari M, Wenzl P, Caig V, Carling J, Xia L, Yang S, Hayden MJ (2006) Diversity arrays technology (DArT) for high-throughput profiling of the hexaploid wheat genome. Theor Appl Genet 113(8):1409–1420
Broman K, Wu H, Sen S, Churchill GA (2003) QTL mapping in experimental crosses. Bioinformatics 19:889–890
Chu CG, Friesen TL, Xu SS, Faris JD, Kolmer JA (2009) Identification of novel QTLs for seedling and adult plant leaf rust resistance in a wheat double haploid population. Theor Appl Genet 119:263–269
Cui F, Fan X, Zhao C, Zhang W, Chen M, Ji J, Li J (2014) A novel genetic map of wheat: utility for mapping QTL for yield under different nitrogen treatments. BMC Genet 15:57–73
Kolmer JA (2015) A QTL on chromosome 5BL in wheat enhances leaf rust resistance of Lr46. Mol Breed 35:74
Kolmer JA, Hughes ME (2015) Physiologic specialization of Puccinia triticina on wheat in the United States in 2013. Plant Dis 99:1261–1267
Kolmer JA, Oelke LM, Liu JQ (2007) Genetics of leaf rust resistance in three Americano landrace-derived wheat cultivars from Uruguay. Plant Breed 126:152–157
Lagudah ES (2011) Molecular genetics of race non-specific resistance in wheat. Euphytica 179:81–91
Lan C, Rosewarne GM, Singh RP, Herrera-Foessel SA, Huerta-Espino J, Basnet BJ, Zhang Y, Yang E (2014) QTL characterization of resistance to leaf rust and stipe rust in the spring wheat line Francolin #1. Mol Breed 34:789–803
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Li Z, Lan C, He Z, Singh RP, Rosewarne GM, Chen X, Xia X (2014) Overview and application of QTL for adult plant resistance to leaf rust and powdery mildew in wheat. Crop Sci 54:1907–1925
Liu S, Anderson JA (2003) Marker-assisted evaluation of Fusarium head blight resistant wheat germplasm. Crop Sci 43:760–766
Liu S, Sehgal SK, Li J, Lin M, Trick HN, Yu J, Gill BS, Bai G (2013) Cloning and characterization of a critical regulator for preharvest sprouting in wheat. Genetics 195:263–273
Long DL, Kolmer JA (1989) A North American system of nomenclature for Puccinia recondita f.sp. tritici. Phytopathology 79:525–529
Maccaferri M, Mantovani P, Tubersoa R, Deambrogia E, Giuliani S, Demontis A, Massi A, Sanguineti MC (2008) A major QTL for durable leaf rust resistance widely exploited in durum wheat breeding programs maps on the distal region of chromosome arm 7BL. Theor Appl Genet 117:1225–1240
Messmer MM, Seyfarth R, Keller M, Schachermayr G, Winzler M, Zanetti S, Feuillet C, Keller B (2000) Genetic analysis of durable leaf rust resistance in winter wheat. Theor Appl Genet 100:419–431
McIntosh RA, Dubcovsky J, Rogers WJ, Morris, C 2017. Catalogue of gene symbols for wheat: 2017 supplement. Komugi Wheat Genetic Resources Database http://shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp. Accessed Apr 4 2017
Nelson JC (1997) QGENE: software for marker-based genomic analysis and breeding. Mol Breed 3:239–245
Oelke LM, Kolmer JA (2005) Genetics of leaf rust resistance in spring wheat cultivars Norm and Alsen. Phytopathology 95:773–778
Peterson RF, Campbell AB, Hannah AE (1948) A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can J Res 26(5):496–500
Roelfs AP (1988) Resistance to leaf and stem rusts in wheat. In: Simmonds NW, Rajaram S (eds) Breeding strategies for resistance to the rusts of wheat. CIMMYT, Mexico, pp 10–22
Roelfs AP, Singh RP, Saari EE (1992) Rust diseases of wheat: concepts and methods of disease management. CIMMYT, Mexico
Schnurbusch T, Paillard S, Schori A, Messmer M, Schachermayr G, Winzler M, Keller B (2004) Dissection of quantitative and durable leaf rust resistance in Swiss winter wheat reveals a major resistance QTL in the Lr34 chromosomal region. Theor App Genet 108:477–484
Singh A, Knox RE, DePauw RM, Singh AK, Cuthbert RD, Campbell HL, Singh D, Bhavani S, Fetch T, Clarke F (2013) Identification and mapping in spring wheat of genetic factors controlling stem rust resistance and the study of their interactions across multiple environmnents. Theor Appl Genet 126:1951–1964
Soriano JM, Royo C (2015) Dissecting the genetic architecture of leaf rust resistance in wheat by QTL meta-analysis. Phytopathology 105:1585–1593
Turner KM, Kolmer JA, Pumphrey MO, Bulli R, Chao S, Anderson JA (2016) Association mapping of leaf rust resistance in a spring wheat core collection. Theor Appl Genet 130:345–361
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, International Wheat Genome Sequencing Consortium, Lillemo M, Mather D, Appels R, Dolferus R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo M-C, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden MJ, Akhunov E (2014) Characterization of polyploid wheat genomic diversity using a high-density 90 000 single nucleotide polymorphism array. Plant Biotechnol 12:787–796
Acknowledgements
We thank K. Xiao and M. Hughes for excellent technical assistance. The authors would like to thank the IWGSC (www.wheatgenome.org) as well as project leaders Nils Stein, Curtis Pozniak, and Jesse Poland, for providing pre-publication access to IWGSC WGA v1.0. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kolmer, J.A., Garvin, D.F., Hayden, M. et al. Adult plant leaf rust resistance derived from the wheat landrace cultivar Americano 44d is conditioned by interaction of three QTL. Euphytica 214, 59 (2018). https://doi.org/10.1007/s10681-018-2141-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-018-2141-3