Abstract
Septoria tritici blotch (STB), caused by the ascomycete fungus Zymoseptoria tritici (also known as Mycosphaerella graminicola), is one of the most devastating foliar wheat diseases worldwide. Host resistance is the most effective strategy for management of the disease. A factor that complicates the determination of resistance is its reported interaction with heading date (Hd) and plant height (Ph). In this study, we report findings from a genome-wide association study of resistance to STB in a world-wide collection of 96 wheat accessions. The collection was evaluated under conditions of artificial infection for seedling and adult plant STB resistance, Hd and Ph in field trials. Marker-trait associations (MTAs) were detected using a mixed linear model. STB disease severities showed significant phenotypic variation. In total, 73 MTAs involving STB resistance were detected. The chromosomal locations of some of them were similar to known Stb genes or quantitative trait loci; whereas others were detected in new genomic regions. The field experiment showed evidence of genetic association between STB resistance and Hd, but only for a few genotypes. This was corroborated at the molecular level, where a total of eight genomic regions associated with STB resistance were located in similar positions to MTAs for Hd. New genomic regions associated with STB resistance found here could be useful in wheat breeding aimed at controlling STB after validation in relevant genetic backgrounds
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Septoria tritici blotch (STB), caused by the ascomycete fungus Zymoseptoria tritici (also known as Mycosphaerella graminicola), is one of the most devastating foliar wheat diseases (King et al. 1983). It is considered one of the top two or three diseases in most wheat-growing regions worldwide, including Europe, North America, South America and Australia (Eyal et al. 1987). Control of STB relies to a large extent on the use of fungicides. In Europe, more than 70% of the fungicides applied to wheat are used to control this disease (Goodwin 2007). The excessive use of fungicides to control the disease has led to resistance and/or reduced sensitivity against the widely used chemical groups such as the strobilurins, triazoles and most recently succinate dehydrogenase inhibitors (SDHIs) (Cools and Fraaije 2008; Torriani et al. 2009; Dooley et al. 2016). As a result, the identification and use of resistance genes is the most effective, economical and environmentally safe strategy for successful management of the disease. As described by Brown et al. (2015) resistance to STB may be isolate-specific or quantitative, polygenic and isolate-nonspecific. Specific interactions between wheat cultivars and Z. tritici isolates may occur in both seedling tests and under field conditions (Arraiano et al. 2001).
During the last decade, 21 major resistance genes (Stb genes) were identified and characterized (McIntosh et al. 2013). Nevertheless, this number is limited mainly because most of them exhibit isolate-specific resistance, thus limiting their use (Arraiano and Browm 2006). The pathogen population is highly diverse (Goodwin et al. 2011) because Z. tritici undergoes several sexual cycles during the wheat-growing season (Suffert and Sache 2011). Genetic recombination increases the likelihood of emergence of new variants and the risk of adaptation to resistance genes deployed in the host population. This emphasizes the need to identify new genes and to combine them in the same genetic background. Such combinations could improve the level and durability of resistance under field conditions, a strategy that has sometimes been effective in controlling other crop diseases (Mundt 2014). As well as reports on mapping single Stb genes there are several recent studies on mapping quantitative trait loci (QTL) (Brown et al. 2015). Complete and partial resistance to STB may be also combined; there are several quantitative studies that reveal the presence of general and specific combining ability of resistance to STB (Van Ginkel and Scharen 1987; Simón and Cordo 1998).
Many STB resistance factors have been identified in classical bi-parental mapping populations. Although linkage analysis in bi-parental crosses has been successful in identifying QTL associated with traits of agronomic importance the approach has major disadvantages. It demands considerable time and cost in the development of segregating populations, besides being able to identify the actual resistance alleles in the two parents (Adhikari et al. 2011). Association mapping (AM) or linkage disequilibrium mapping is an alternative method of QTL detection that allows identification of direct associations between genotype and phenotype. Numerous marker-trait associations (MTAs) in wheat were detected in earlier GWAS studies exploiting DArT marker technology (Crossa et al. 2007; Adhikari et al. 2011; Neumann et al. 2011).
One of the most confounding factors in determining resistance to STB is the reported correlation between resistance, plant height (Ph), and heading date (Hd) (Simón et al. 2005). Many phenotypic studies assumed that this relationship was due to genetic linkage (Rosielle and Brown 1979; Eyal 1981; Baltazar et al. 1990). This was confirmed in genetic studies by the co-localization of QTL for STB resistance and Hd loci as well as Ph QTL (Risser et al. 2011; Goudemand et al. 2013; Kollers et al. 2013). However, in these studies disease development was measured on the same day for all cultivars. According to Arama et al. (1999) and Simón et al. (2004, 2005) disease severity should be measured at the same stage of development in order to compare the resistance of a group of cultivars and its correlation with earliness and height. Covariance analysis was proposed by Van Beuningen and Kohli (1990) and Simón et al. (2004) to identify resistant genotypes independently of Hd and Ph. Otherwise, differences in disease levels between earlier and later, or shorter and taller, cultivars may be due to differences in leaf age as well as to differences in the duration of exposure of leaves to the disease. Phenotypic studies that consider this aspect (Arama et al. 1999; Simón et al. 2005) conclude that this correlation is due to epidemiological or environmental factors rather than genetic linkage. However, the presence of genetic linkage may depend on the genetic material.
The goals of the current study were to (i) assess seedling and adult stage STB responses of a core collection of 96 winter wheat accessions of diverse origin, (ii) identify MTAs for resistance to Z. tritici expressed as necrosis percentage (NP) and pycnidial coverage (PC) through GWAS, (iii) compare the results with the chromosomal locations of known Stb resistance genes and QTL, and (iv) determine phenotypic and genetic relationships between STB response, Ph and Hd.
Materials and methods
Plant materials
In this study, a core collection of 96 winter wheat accessions (mainly cultivars and advanced breeding lines) from 20 countries across five continents was used as an AM population (Table 1). These genotypes were carefully targeted from a larger collection maintained at the Institute of Field and Vegetable Crops (Novi Sad, Serbia), on the basis of contrasting phenotypic expression of traits of agronomic importance, including those tested in this study (Kobiljski et al. 2002; Quarrie et al. 2003). The collection included a number of important “founder genotypes” widely used as parents in breeding programs across the world.
Experimental design and phenotypic trait evaluation
Seedling stage
Two field experiments were carried out at the J. Hirschhorn Experimental Station, Faculty of Agricultural and Forestry Sciences, National University of La Plata, Argentina during 2012 and 2013 under artificial inoculation in a split–split–plot design with two blocks. Main plots were years (2012 and 2013) and subplots were inoculations with two isolates (FALP0112 and FALP0212) within each year. The sub-subplots were 98 genotypes (two Argentinean spring wheat cultivars: BioINTA 3005 and Buck SY 110, susceptible to Z. tritici used as controls, and the 96 wheat accessions). Because many genotypes had winter growth habit seeds were germinated in Petri dishes and vernalized for 3 weeks at 4–8 °C in a growth chamber before planting in the field. The experiments 1 and 2 were sown on 21 June 2012 and 31 July 2013, respectively. Ten to 15 seedlings of each genotype were planted in each row. Plots were 0.5 m single rows separated by one row of oat to reduce interplot interference; rows were 0.2 m apart. Weather data (monthly precipitation, relative humidity, and minimum, maximum and mean temperature) were recorded at a Davis Meteorological Station situated 300 m from the experiments.
The entire collection was inoculated with Z. tritici isolates FALP0112 and FALP0212 from the Argentinean locations Pla and Nueve de Julio, respectively. Inoculum was prepared by aseptically scraping sporulating colonies with a scalpel and suspending the conidia in deionized water. The conidial suspension was adjusted to 5 × 106 spores mL−1 using a Newbauer counting chamber. Tween 20 (Polysorbate 20) was added at 0.5 mL L−1. The plant material was inoculated at growth stage (GS) 12 (Zadoks et al. 1974) until runoff with a manually operated sprayer, which was calibrated at a rate of 10 L 100 m−2. After inoculation, plants were kept moist by sprinkling water several times a day over a 3 days period to maintain humid conditions necessary for disease development. In order to estimate seedling resistance, disease severity was assessed 28–30 days after the first inoculation at the same time for all cultivars by estimating the NP and PC on the inoculated three first leaves of seven plants of every plot for each replication. Evaluations started when STB was well expressed in the two susceptible Argentinean cultivars used as a control.
Adult plant stage
Three field experiments (1, 2 and 3), sown on 21 June 2012, 14 June and 31 July 2013, respectively, under artificial infection with the above isolates were conducted at the J. Hirschhorn Experimental Station. The experiments were performed as described above with inoculations being made at tillering (GS 23). For evaluation of adult-stage response disease severities were assessed 30 d after flag leaf emergence by estimating the NP and PC on the three upper leaves (flag leaf, flag leaf-1, flag leaf-2) of seven plants of every plot for each replication. Average responses of the three leaves of each plot were calculated for both growth stages.
Ph was recorded as the average of 10–15 plants per plot, measured in centimeters from the soil surface to the tip of the spike. Days to heading were recorded as the number of days from planting until 50% spike emergence.
Phenotypic data analysis
The phenotypic data collected and calculated across trials (NP, PC, Ph and Hd) were analyzed by combined analysis of variance (ANOVA) for split–split–plot design in Genstat 12 Ed (VSN International 2009). Heritability (h2) was calculated as the ratio of genotypic to phenotypic variance: \({\text{h}}^{2} = \, \sigma^{2}_{\text{g}} / \, \left( {\sigma^{2}_{\text{g}} + \, \sigma^{2}_{\text{ge}} /{\text{E }} + \, \sigma^{2} /{\text{ER}}} \right).\) Where \(\sigma^{2}_{\text{g}}\) denotes the genotypic variance, \(\sigma^{2}_{\text{ge}}\) the genotype × environment interaction variance and σ2 the error variance. E and R are the number of environments and blocks, respectively. On the other hand, multiple linear regression analyses, considering disease severity (NP and PC) as the dependent variable and Hd and Ph as the independent variables were performed to determine the relationship and percentage of variation in severity accounting for the morpho-physiological traits.
Genotyping, population structure and linkage disequilibrium analysis
DArT technology was provided by Triticarte Pty Ltd (Canberra, Australia; http://www.triticarte.com.au), a whole-genome profiling service laboratory. Each of the 874 polymorphic markers was named using the prefix ‘‘wPt’’ followed by a unique numerical identifier. Thirty-nine DArT markers with <5% allele frequencies were excluded from all analyses. The integrated map of DArT markers developed by Crossa et al. (2007) was used to assign 525 trait-associated markers to chromosome arms. The chromosomal locations of previously unmapped markers were provided by Triticarte. The methodology and results concerning population structure and DL of this population were described earlier (Neumann et al. 2011). ANOVA was performed for all phenotypic traits using population subgroups to determine the significance of the confounding effect of population structure on the phenotypic traits.
Marker-trait association (MTA) analysis
For MTA analysis, each trait was represented by the means of the two blocks in each experiment and analyses were performed separately for every experiment. Associations between the markers and each trait were calculated using a mixed linear model (MLM) based on both Q-matrix and kinship-matrix, suggested by Yu et al. (2006), with the software program TASSEL 2.1. The Q-matrix was derived from STRUCTURE and kinship-matrix was calculated in TASSEL. The efficient mixed model analysis (Kang et al. 2008) was chosen to reduce computing time and the MLM parameters were left at the default settings from TASSEL. In all cases, MTAs were considered significant when the P value was < 0.05 and highly significant at P < 0.01. Consistency through at least two experiments was taken into account as an additional criterion to reduce the risk of false MTAs. Maps and significantly associated regions with each of the traits were represented with the MapChart version 2.3 software (Voorrips 2002). Genetic distances are provided in centimorgans (cM).
Results
Weather conditions
Considering the whole growing season (June to December), trends of temperature and relative humidity were similar for both years (2012 and 2013) (Fig. 1a, b). In contrast, the rainfall pattern and total precipitation varied considerably between years. Total precipitation in 2012 was 687.6 mm and in 2013, 414.1 mm compared to the regional mean of 519.5 mm. The most important difference in monthly rainfall was for August, where precipitation was much higher in 2012 (222.8 mm) compared to 2013 (7.2 mm) and therefore more conducive to disease development (Fig. 1c).
Phenotypic data
Analysis of variance revealed highly significant differences (P < 0.001) among genotypes regarding response to STB, Hd and Ph in the combined data analysis across experiments (Table 2). NP and PC were highly significant (P < 0.001) at both the seedling and adult stages, for genotypes, experiments × genotypes, isolates × genotypes and experiments × isolates × genotypes interactions. There were also significant differences (P < 0.001) between genotypes and experiments × genotypes in days to Hd and Ph. Moreover, there were significant differences for Hd (P < 0.001) but not for Ph (P = 0.49). Some lines showed a differential response at both the seedling (Fig. 2a, b) and adult (Fig. 2c, d) stages. The level of resistance of some genotypes also differed between seedlings and adult plants (Fig. 2e, f) for PC and NP, respectively.
The resistance components (NP and PC) at the seedling and adult stages showed only moderate correlations across experiments (Fig. S1). This was due to the considerable variation in rainfall between 2012 and 2013, and consistently reflected in the disease severity values. Consequently, mean values of NP and PC were higher in 2012 than in 2013. Heritability was 0.40 and 0.76 for NP and PC at seedling stage, whereas at adult stage it was 0.75 and 0.78 for each resistance component, respectively. Finally, Hd and Ph as expected, had higher heritabilities of 0.95 and 0.98, respectively.
Regarding the two structural groups previously determined in this mapping population by Neumann et al. (2011) there were significant differences for STB response expressed as PC and NP at the seedling and adult stages (P < 0.01) and Hd (P < 0.01). Considering PC at both stages, for example, group Q1 comprising genotypes of diverse origin was more resistant with a mean of 2.40 and 7.76% compared to Q2, which contained mainly Serbian elite lines and cultivars, with means of 2.96 and 11.78%, respectively, for the averages of isolates and experiments. At both stages, the more resistant genotypes in Q2 originated mainly from countries like Tibet, USA, Ukraine and Great Britain. The average Hd of genotypes within group Q1 was about three days earlier than genotypes in group Q2.
Marker-trait association (MTA) analysis
Considering all assessed traits 100 MTAs involving 78 DArT markers were identified. Sixty-one markers were associated with a single trait and were therefore called trait-specific markers; whereas the others involving two or more traits were considered multi-trait markers. For a given trait two associated markers were considered as belonging to the same associated locus/region when they were separated by a distance of less than 2 cM or shared at least one associated group and were less than 5 cM apart. In this way, association analysis revealed several genomic regions conferring resistance to STB among the collection of 96 mainly winter wheat accessions.
Thirty-eight MTAs involving 30 DArT markers were significantly associated with STB response of seedlings. Eight and 10 of them were effective against the FALP0112 isolate, whereas six and 14 were effective against isolate FALP0212 for NP and PC, respectively. Seven markers were associated with both isolates. The 30 DArT markers corresponded to 26 distinct genetic loci/regions on chromosomes 1A (two), 1B (two), 2A (one), 2B (three), 2D (one), 3A (one), 3B (two), 3D (one), 4B (two), 5A (one), 5B (one), 6A (three), 6B (two) and 7A (four). The P values of significant MTAs and phenotypic variations explained by DArT markers across experiments are presented in Table 3.
Thirty-five MTAs (32 DArT markers) in 24 genomic regions were associated with STB response at the adult plant stage. These regions involved 14 chromosomes: 1B (two), 1D (one), 2A (one), 2B (three), 2D (one), 3A (two), 3B (one), 4A (one), 4B (one), 5D (two), 6A (two), 6B (one), 7A (four), and 7B (three). Fourteen and five markers were effective against FALP0112, whereas seven and 10 were effective against FALP0212 for variables NP and PC, respectively. Among all significant markers three were common for both isolates. P values of significant MTAs and phenotypic variations explained by the DArT markers across of the experiments are presented in Table 4. Markers significant for PC at the adult stage, including wPt7094, wPt8168, wPt7062, wPt3883, and wPt7734 located on chromosomes 1B (position 11.9 cM), 1B (44.7), 4B (109.8), 7A (88.8), and 7A (89.6), respectively, were also significant at the seedling stage.
Twelve MTAs (11 markers) were found for Ph on chromosomes 1B (one), 2B (one), 3A (two), 4A (one), 6A (one), 6B (two), 7A (two), and 7B (two). Nine markers were associated only with this trait, whereas the other three were involved in other trait-associations (two with Hd and one with STB response). For Hd 18 MTAs were located on chromosomes 1A (one), 1B (two), 2B (three), 3B (three), 4B (one), 5D (two), 6A (two), and 6B (four). Seven were associated only with Hd, nine with STB response, and two with Ph (Table 5).
Relationships between STB resistance, Hd and Ph
At both the seedling and adult stages multiple linear regression analysis between PC as the dependent variable and Ph and Hd as independent variables, yielded significant R 2 values for most isolates and experiments. In general, regression coefficients of Hd were negative, significant and very consistent. By contrast coefficients for Ph were rarely significant and inconsistent. Multiple linear regression analysis using NP as the dependent variable and Hd and Ph as independent variables was also significant for most isolates and experiments at both growth stages. Regression coefficients for Hd were also significant, but the relationship was negative for seedling assays and positive for adult plant tests. Again, when NP was used as the dependent variable, regression coefficients for Ph were inconsistent. For the average of all experiments and isolates, models including both independent variables accounted for 11.95 and 6.35% and 8.8 and 32.3% of the variation in PC and NP, in seedlings and adult plant stage, respectively (Table 6).
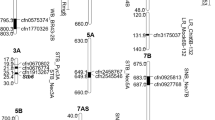
In addition, genetic linkage among markers significantly associated with STB resistance, Hd and Ph was determined following the classification proposed by Maccaferri et al. (2005), who defined four classes of marker pairs: 1—tight linkage (distance < 10 cM); class 2—moderate linkage (10–20 cM); class 3—loose linkage (20–50 cM); and class 4—independence (>50 cM). Considering both growth stages, a total of eight genomic regions (one at the seedling, two at the adult, and five at both growth stages) on 1BS (two), 2BS, 3BS, 4BS, 5DS and 6AS and three genomic regions (one at the seedling, one at the adult and one at both growth stages) on 1BS, 3AS and 6AS respectively) associated with STB resistance were found in a similar position and close linkage with markers associated with Hd and Ph, respectively (Fig. 3).
Genetic map including marker-trait associations (MTAs) identified for measured phenotypic traits. Markers in bold represent MTAs detected at the seedling stage. Underlined and not underlined markers correspond to PC and NP, respectively. Symbols diamond and triangle represent isolates FALP0112 and FALP0212, respectively. Genetic distances between markers are in centimorgans (cM)
Discussion
We report the results of a GWAS aimed at identifying genomic regions conditioning resistance to STB, Hd and Ph in a core collection of 96 mainly winter wheat accessions, employing 525 polymorphic DArT markers. Considerable phenotypic variation was observed in all measured traits across the 96 accessions. This allowed to study of the genetic basis of phenotypic variation under a wide range of environmental conditions. Regarding STB severity, the higher levels observed in 2012 compared to 2013 were attributed to more favorable environmental conditions for disease development. The main source of Z. tritici inoculum during the spring and summer are pycnidiospores (Eriksen and Munk 2003; Suffert and Sache 2011). The pycnidiospores are spread from the base and disperse upwards in the crop canopy through rain splash. Significant differences between the two subgroups generated by STRUCTURE were also observed for STB response at the seedling and adult stages and for Ph. These results are in agreement with Edae et al. (2014), who reported significant effects of population structure in winter wheat on Ph and stem-related traits.
Moderately high heritabilities of combined and separate seedling and adult responses were observed for the variables NP and PC, ranging from 0.40 to 0.79, due to resistance genes that were effective across environments. Thus, repeatable and reliable phenotypic data for GWAS were obtained. Moderately high heritabilities of STB response were also reported in other studies (Goudemand et al. 2013; Kollers et al. 2013; Simón and Cordo 1998) indicating that resistance to STB can be improved by selection.
Specific interactions among wheat cultivars and Z. tritici isolates may occur in combined, and separate seedling and adult-plant stage tests (Arraiano et al. 2001; Simón et al. 2005). In our study, correlations between seedling and adult plants for PC showed moderate values for both isolates, indicating that there were genotypes with combined seedling and adult stage resistance and others displaying resistance only at specific growth stages. Simón et al. (2005) proposed that selection at both stages might be done to achieve acceptable levels of resistance for the entire growing period. Additionally, several genotypes in the core collection showed similar reactions to both isolates. Although more isolates should be tested this suggests that non-isolate-specific horizontal resistance may be also present. This type of resistance is an important genetic resource for the improvement of wheat against STB as reported in earlier studies (Jlibene et al. 1994; Simón and Cordo 1998).
Despite the high economic importance of STB only 21 major resistance genes have been named (McIntosh et al. 2013). However, many QTL for partial resistance have been reported, and these have greater significance for breeding (Brown et al. 2015). Here we report a total of 73 MTAs related to 57 DArT markers significantly associated with STB resistance. The 53 DArT markers were distributed over 42 chromosomal regions on 18 of the 21 chromosomes. Among these regions, 18 and 16 were specifically associated with resistance to STB at the seedling and adult stages. The remaining 8 regions showed DArT markers simultaneously associated with resistance at both growth stages. These results are consistent with phenotypic performance and indicate that resistance at both the seedling and adult stages is controlled by some common regions but also by different regions.
Of all the chromosomal regions associated with STB response those located on 1BS, 1DS, 2BL, 3AS, 3BS, 5AL, 6AS, 7AS and 7BL, were identified at similar regions to known genes Stb11/Stb2, Stb10, Stb9, Stb6, Stb14, Stb17, Stb15, Stb3 and Stb8, respectively (Brown et al. 2015). However, further work is needed to confirm whether the genomic regions found here correspond to known Stb genes. It would be important to use DArT markers found in our study for marker-assisted selection (MAS). As far as we know no Stb gene is located on chromosome arms 1AS, 1BL, 2AS, 2BS, 3BL 2DS, 4AS, 4BS/L, 5BL, 5DS, 6AL, 6BS/L, 7AL and 7BS where genomic regions associated with STB response were found in the present population. This infers that new sources of STB resistance were detected in our study.
In addition to identified genomic regions suggestive of Stb genes DArT markers wPt7030 (1AS), wPt5672 (2BS), wPt8650 (4BS), wPt7062 (4BL), and wPt7777 located in the same position as wPt1856 were previously reported as significant for STB by Risser et al. (2011) and Goudemand et al. (2013). On the other hand, the genomic regions on chromosomes 1AS (markers wPt5374 and wPt4765), 1BS (markers wPt1328 and wPt7094) and 6AL (markers wPt0959 and wPt4229) were reported as conferring resistance to tan spot (caused by Pyrenophora tritici-repentis), leaf rust (Puccinia triticina) and yellow rust (Puccinia striiformis) (Crossa et al. 2007; Singh et al. 2015). Broad-spectrum resistance (BSR) has been widely reported in wheat, including detection in non-adapted germplasm with resistance to multiple diseases and clustering of resistance genes in distinct genomic regions (Gurung et al. 2009). These results are also in line with those reported by Miedaner et al. (2012, 2013) and Zwart et al. (2010), who found genomic regions conferring BSR to Fusarium head blight (Gibberella zeae), tan spot, STB and leaf rust. Stacking QTL that affect response to traits of interest from different chromosome regions into one background is a challenging and time consuming task in plant breeding. The use of multi-trait markers in MAS may increase QTL pyramiding efficiency (Edae et al. 2014).
Regarding the relationship between STB resistance and traits Hd and Ph there was a significant negative association between earliness and STB response expressed as PC, a finding that agrees with previous results (Risser et al. 2011; Goudemand et al. 2013). Although not always significant NP also showed a negative trend with earliness. Ph by contrast displayed no consistent relationship between variables (NP and PC) and experiments. These results are in agreement with Simón et al. (2004), who only found associations between susceptibility and reduced Ph in very short wheats, indicating that moderately short wheats are not necessarily more susceptible to STB. At seedling all genotypes were inoculated and scored at the same growth stage and at the same time and therefore disease development was subjected to the same weather conditions. We assume that the relationship found between earliness and STB infection at the seedling stage is due to a genetic association between the two traits. However, the low correlation values could be attributed to only a few genotypes (e.g. Tibet Dwarf, Tom Thumb, Brigand, Ana, Mexico 3, Helios, Hira) that showed this association, contrary to many others where this relationship did not occur. In the latter group, for example, the genotypes Lambriego Inia and Tr. Compactum with moderate-long growth cycles (131 and 134 d to Hd) showed higher values (56.62 and 3.95%, respectively) for NP and PC than the averages (51.23 and 2.53%) for isolates and experiments. Genotypes like Triple Dirk ‘S’ and Ai-Bian with moderately short growth cycles (103 and 105 d to Hd) showed lower values (41.43 and 0.6%) for NP and PC, respectively, than the average (51.23 and 2.53%) for both isolates and experiments.
PC maintained a negative relationship with Hd at the adult stage, but this was not always significant. All genotypes were scored at the same growth stage (30 days after flag leaf emergence) but at different dates. Weather conditions (temperature, precipitation and humidity) during the period of exposure of the leaves to the pathogen obviously varied within and between experiments. Temperatures were higher for later genotypes (20.75, 19.04 and 19.68 °C) than for earlier ones (17.9, 14.34 and 17.62 °C, respectively) in all three experiments. For PC, an optimum temperature of 25 °C was reported (Hess and Shaner 1987). Hence, temperatures were more favorable for the expression of the disease in later genotypes than in earlier ones, but in general, this was not reflected in the final levels of PC. Conversely, humidity was slightly lower for later genotypes (82.05, 82.91 and 79.66%) than for earlier ones (88.63; 84.68 and 81.89%) in the three experiments, respectively. Finally, precipitation was higher for the later (2.82, 4.48 and 3.24 mm) than for the earlier (5.49, 1.51 and 2.46 mm) genotypes in two of the three experiments. These results assume the presence of genetic association, which is supported by the fact that the relationship between earliness and STB infection expressed as PC was always negative despite the fluctuations in the weather conditions. In contrast, there was a significant and positive association between earliness and NP. Apparently, this could be due to the greater values of radiation and temperature to which later genotypes were exposed. This could have generated higher levels of senescence due to other causes.
Interestingly, at both stages, all the genotypes that were significant for the association between Hd and PC belonged to sub-group 1 (Q1), except one (NS 46/90) that belonged to sub-group 2 (Q2). This finding as well as low r values indicated that presence or absence of genetic linkage between Hd and STB resistance depended on genotype. Thus, as reported by Simón et al. (2004) some STB resistance factors are subject to this complication (association between resistance and Hd or Ph) but are not. Hence it should be possible to improve the STB resistance independently of Hd.
The genetic associations between STB resistance and Hd were confirmed by the presence of genetic linkage or co-localization of markers associated with both traits. Eight genomic regions associated with STB resistance were located in similar positions to Hd MTAs. One region for seedling stage STB resistance, two for adult stage response and five identified at both stages were in similar positions to genes for Hd. Associations of resistance genes on chromosomes 2BL and 3BL and Hd found in this study were also reported by Risser et al. (2011). This indicates that the genetic association between STB response and Hd not only depends on the evaluated material, but also the growth stage.
In conclusion, this study identified new germplasm with moderate to high levels of STB resistance to two pathogen isolates. Some genomic locations seem to be new whereas others were at similar positions to known Stb genes and QTL. Confirmation of these results is required and it would be important to use DArT markers found in our study for MAS. The STB resistances found in our study involve several loci, each explaining only a small part of the total phenotypic variance. Conventional selection of QTLs with small effects on phenotypic variation could be improved in a breeding population through cycles of MAS for multiple QTLs. Although there were significant associations of STB resistance and Hd this applied to only relatively few genotypes.
References
Adhikari TB, Jackson EW, Gurung S, Hansen JM, Bonman JM (2011) Association mapping of quantitative resistance to Phaeosphaeria nodorum in spring wheat landraces from the USDA National Small Grains Collection. Phytopathology 101:1301–1310. doi:10.1094/phyto-03-11-0076
Arama PF, Parlevliet JE, van Silfhout CH (1999) Heading date and resistance to Septoria tritici blotch in wheat not genetically associated. Euphytica 106:63–68. doi:10.1023/A:1003593218761
Arraiano LS, Browm JKM (2006) Identification of isolate-specific and partial resistance to Septoria tritici blotch in 238 European wheat cultivars and breeding lines. Plant Pathol 55:726–738. doi:10.1111/j.1365-3059.2006.01444.x
Arraiano LS, Brading PA, Brown JKM (2001) A detached seedling leaf technique to study resistance to Mycosphaerella graminicola (anamorph Septoria tritici) in wheat. Plant Pathol 50:339–346. doi:10.1046/j.1365-3059.2001.00570.x
Baltazar BM, Scharen AL, Kronstad WE (1990) Association between dwarfing genes Rht1 and Rht2 and resistance to Septoria tritici blotch in winter wheat (Triticum aestivum L.). Theor Appl Genet 79:422–426. doi:10.1007/bf01186089
Brown JK, Chartrain L, Lasserre-Zuber P, Saintenac C (2015) Genetics of resistance to Zymoseptoria tritici and applications to wheat breeding. Fungal Genet Biol 79:33–41. doi:10.1016/j.fgb.2015.04.017
Cools HJ, Fraaije BA (2008) Are azole fungicides losing ground against Septoria wheat disease resistance mechanisms in Mycosphaerella graminicola. Pest Manag Sci 64:681–684. doi:10.1002/ps.1568
Crossa J, Burgueno J, Dreisigacker S, Vargas M, Herrera- Foessel SA, Lillemo M, Singh RP, Trethowan R, Warburton M, Franco J, Reynolds M, Crouch JH, Ortiz R (2007) Association analysis of historical bread wheat germplasm using additive genetic covariance of relatives and population structure. Genetics 177:1889–1913. doi:10.1534/genetics.107.078659
Dooley H, Shaw MW, Mehenni-Ciz J, Spink J, Kildea S (2016) Detection of Zymoseptoria tritici SDHI-insensitive field isolates carrying the SdhC-H152R and SdhD-R47W substitutions. Pest Manag Sci 72:2203–2207. doi:10.1002/ps.4269
Edae EA, Byrne PF, Haley S, Lopes MS, Reynolds MP (2014) Genome-wide association mapping of yield and yield components of spring wheat under contrasting moisture regimes. Theor Appl Genet 127:791–807. doi:10.1007/s00122-013-2257-8
Eriksen L, Munk L (2003) The occurrence of Mycosphaerella graminicola and its anamorph Septoria tritici in winter wheat during the growing season. Eur J Plant Pathol 109:253–259. doi:10.1023/a:1022851502770
Eyal Z (1981) Integrated control of Septoria diseases of wheat. Plant Dis 65:763–768. doi:10.1094/pd-65-763
Eyal Z, Scharen AL, Prescott JM, van Ginkel M (1987) The Septoria diseases of wheat: concepts and methods of disease management. International Maize and Wheat Improvement Centre (CIMMYT), Mexico
GenStat for Windows 12 Th Edition (2009) Copyright VSN International Ltd
Goodwin SB (2007) Back to basics and beyond: increasing the level of resistance to Septoria tritici blotch in wheat. Aust Plant Pathol 36:532–538. doi:10.1071/ap07068
Goodwin SB, Ben M’Barek S, Dhillon B, Wittenberg AHJ, Crane CF et al (2011) Finished genome of the fungal wheat pathogen Mycosphaerella graminicola reveals dispensome structure, chromosome plasticity, and stealth pathogenesis. PLoS Genet 7(6):e1002070. doi:10.1371/journal.pgen.1002070
Goudemand E, Laurent V, Duchalais L, Tabib Ghaffary SM, Kema GHJ, Lonnet P, Margale E, Robert O (2013) Association mapping and meta-analysis: two complementary approaches for the detection of reliable Septoria tritici blotch quantitative resistance in bread wheat (Triticum aestivum L.). Mol Breed 32:563–584. doi:10.1007/s11032-013-9890-4
Gurung S, Bonman JM, Ali S, Patel J, Myrfield M, Mergoum M, Singh PK, Adhikari TB (2009) New and diverse sources of multiple disease resistance in wheat. Crop Sci 49:1655–1666. doi:10.2135/cropsci2008.10.0633
Hess DE, Shaner G (1987) Effect of moisture and temperature on development of Septoria tritici blotch in wheat. Phytopathology 77:215–219
Jlibene M, Gustafson JP, Rajaram S (1994) Inheritance of resistance to Mycosphaerella graminicola in hexaploid wheat. Plant Breed 112:301–310. doi:10.1111/j.1439-0523.1994.tb00688.x
Kang HM, Zaitlen NA, Wade CM, Kirby A, Heckermann D, Daly MJ, Eskin E (2008) Efficient control of population structure in model organism association mapping. Genetics 178:1709–1723. doi:10.1534/genetics.107.080101
King JE, Cook RJ, Melville SC (1983) A review of Septoria diseases of wheat and barley. Ann Appl Biol 103:345–373. doi:10.1111/j.1744-7348.1983.tb02773.x
Kobiljski B, Quarrie SA, Dencic S, Kirby J, Iveges M (2002) Genetic diversity of the Novi Sad wheat core collection revealed by microsatellites. Cell Mol Biol Lett 7:685–694
Kollers S, Rodemann B, Ling J, Korzun V, Ebmeyer E, Argillier O, Hinze M, Plieske J, Kulosa D, Ganal MW, Roder MS (2013) Genetic architecture of resistance to Septoria tritici blotch (Mycosphaerella graminicola) in European winter wheat. Mol Breed 32:411–423. doi:10.1007/s11032-013-9880-6
Maccaferri M, Sanguineti MC, Noli E, Tuberosa R (2005) Population structure and long-range linkage disequilibrium in a durum wheat elite collection. Mol Breed 15:271–289. doi:10.1007/s11032-004-7012-z
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers WJ, Morris CF et al (2013) Catalogue of gene symbols for wheat in 12th International Wheat Genetics Symposium, edited by R.A. McIntosh, Yokohama, Japan. http://wheat.pw.usda.gov/GG2/Triticum/wgc/2013/GeneCatalogueIntroduction.pdf. Accessed 23 Dec 2015
Miedaner T, Risser P, Paillard S, Schnurbusch T, Keller B, Hartl L, Holzapfel J, Korzun V, Ebmeyer E, Utz HF (2012) Broad-spectrum resistance loci for three quantitatively inherited diseases in two winter wheat populations. Mol Breed 29:731–742. doi:10.1007/s11032-011-9586-6
Miedaner T, Zha Y, Gowda M, Longin CFH, Korzun V, Ebmeyer E, Kazman E, Reif JC (2013) Genetic architecture of resistance to Septoria tritici blotch in European wheat. BMC Genom 14:858. doi:10.1186/1471-2164-14-858
Mundt CC (2014) Durable resistance: a key to sustainable management of pathogens and pests. Infect Genet Evol 27:446–455. doi:10.1016/j.meegid.2014.01.011
Neumann K, Kobiljski B, Dencic S, Varshney RK, Borner A (2011) Genome-wide association mapping: a case study in bread wheat (Triticum aestivum L.). Mol Breed 27:37–58. doi:10.1007/s11032-010-9411-7
Quarrie SA, Dodig D, Pekic S, Kirby J, Kobiljski B (2003) Prospects for marker-assisted selection of improved drought responses in wheat. Bulg J Plant Physiol 28:83–95
Risser P, Ebmeyer E, Korzun V, Hartl L, Miedaner T (2011) Quantitative trait loci for adult-plant resistance to Mycosphaerella graminicola in two winter wheat populations. Phytopathology 101:1209–1216. doi:10.1094/phyto-08-10-0203
Rosielle AA, Brown AGP (1979) Inheritance, heritability and breeding behaviour of resistance to Septoria tritici in wheat. Euphytica 21:152–161
Simón MR, Cordo CA (1998) Diallel analysis of four resistance components to Septoria tritici in six crosses of wheat (Triticum aestivum). Plant Breed 117:123–126. doi:10.1111/j.1439-0523.1998.tb01464.x
Simón MR, Worland AJ, Struik PC (2004) Influence of plant height and heading date on the expression of the resistance to Septoria tritici blotch in near isogenic lines of wheat. Crop Sci 44:2078–2085. doi:10.2135/cropsci2004.2078
Simón MR, Perello AE, Cordo CA, Larran S, van der Putten PEL, Struik PC (2005) Association between Septoria tritici blotch, plant height, and heading date in wheat. Agron J 97:1072–1081. doi:10.2134/agronj2004.0126
Singh PK, Crossa J, Duveiller E, Singh RP, Djurle A (2015) Association mapping for resistance to tan spot induced by Pyrenophora tritici-repentis race 1 in CIMMYTs historical bread wheat set. Euphytica 206:1–11. doi:10.1007/s10681-015-1528-7
Suffert F, Sache I (2011) Relative importance of different types of inoculum to the establishment of Mycosphaerella graminicola in wheat crops in north-west Europe. Plant Pathol 60:878–889
Torriani SF, Brunner PC, McDonald BA, Sierotzki H (2009) QoI resistance emerged independently at least 4 times in European populations of Mycosphaerella graminicola. Pest Manag Sci 65:155–162. doi:10.1002/ps.1662
Van Beuningen LT, Kohli MM (1990) Deviation from the regression of infection on heading and height as a measure of resistance to Septoria tritici blotch in wheat. Plant Dis 74:488–493. doi:10.1094/pd-74-0488
Van Ginkel M, Scharen AL (1987) Generation mean analysis and heritabilities of resistance to Septoria tritici in durum wheat. Phytopathology 77:1629–1633. doi:10.1094/phyto-77-1629
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Heredity 93:77–78. doi:10.1093/jhered/93.1.77
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB et al (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208. doi:10.1038/ng1702
Zadoks JC, Chang TT, Konzak CF (1974) A decimal code for the growth stages of cereals. Weed Res 14:415–421. doi:10.1111/j.1365-3180.1974.tb01084.x
Zwart RS, Thompson JP, Milgate AW, Bansal UK, Williamson PM, Raman H, Bariana HS (2010) QTL mapping of multiple foliar disease and root-lesion nematode resistances in wheat. Mol Breed 26:107–124. doi:10.1007/s11032-009-9381-9
Acknowledgements
This study was funded by ANPCYT (Agencia Nacional de Promoción Científica y Tecnológica, Argentina) PICT 2181/2010. We wish to thank the staff from the J. Hirschhorn Experimental Station, Faculty of Agricultural and Forestry Sciences, National University of La Plata, Argentina. In addition, we thank Boris Kobiljski, Novi Sad, Serbia for developing and providing the original seed stocks. We also thank to the reviewers and editor for valuable comments and suggestions for improving our manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gerard, G.S., Börner, A., Lohwasser, U. et al. Genome-wide association mapping of genetic factors controlling Septoria tritici blotch resistance and their associations with plant height and heading date in wheat. Euphytica 213, 27 (2017). https://doi.org/10.1007/s10681-016-1820-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-016-1820-1