Abstract
The commercial interest in pigmented wheat grain flows from an understanding that they are nutritionally superior to white kernels. The pigment of purple coloured bread and durum wheat grains results from the accumulation of anthocyanins in the pericarp; its genetic basis is the action of Pp-1 and Pp3 genes. Here, the development of a set of bread wheat near isogenic lines (NILs) carrying various combinations of Pp alleles is described, along with a demonstration of their utility for the genetic dissection of the purple pericarp trait. A marker-assisted backcrossing strategy was based on the use of microsatellite markers linked to Pp3 (chromosome 2A), Pp-A1 (7A) and Pp-D1 (7D). Pp-A1 is a newly uncovered gene of weak effect. A qRT-PCR-based analysis of the anthocyanin synthesis structural genes [Chi (chalcone-flavanone isomerase) and F3h (flavanone 3-hydroxylase)] transcript abundance in the pericarp of the NILs suggested that the Pp genes up-regulate their transcription in contrasting ways. These NILs represent a resource for studying the effect of grain pigmentation on other wheat traits and end products.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Descriptions of grain colour in wheat are generally qualitative in nature: white, red, blue or purple. Red pigmentation is associated with the deposition of proanthocyanidin in the testa, whereas both blue and purple pigmentation flows from the accumulation of anthocyanin in respectively the aleurone and the pericarp (reviewed by Zeven 1991; Khlestkina 2013). As the consumption of anthocyanins is associated with a number of health benefits (Lila 2004), wheat grains enriched for these compounds are an attractive product.
The mode of inheritance of the purple pericarp has been known for many years: in some situations it appears to be monogenic (Sharman 1958; Dobrovolskaya et al. 2006), while in others it is digenic (Bolton 1970; Piech and Evans 1979; Arbuzova et al. 1998; Dobrovolskaya et al. 2006; Khlestkina et al. 2010a). According to Dobrovolskaya et al. (2006), the Pp3 locus maps to a marker-defined region of chromosome 2A of bread wheat, and the corresponding region of tetraploid (durum) wheat chromosome arm 2AL also harbours a probable Pp3 orthologue (Khlestkina et al. 2010a). Durum wheat also houses Pp-B1 (complementary to Pp3), mapping to chromosome arm 7BS (Khlestkina et al. 2010a), while its D-genomic homoeologue in bread wheat (Pp-D1) lies on chromosome arm 7DS (Tereshchenko et al. 2012a). Based on an analysis of introgression lines, Tereshchenko et al. (2012b) have shown that a functional allele of at least one purple pericarp gene is retained in both Aegilops speltoides and Triticum timopheevii, although in the latter species, the purple grain trait has to date never been described (Tereshchenko et al. 2012b).
Analysis of the transcription of various structural genes encoding enzymes active in anthocyanin synthesis in near-isogenic lines (NILs) varying for their Pp gene content has suggested that Pp-D1 and Pp3 are both transcriptional regulators (Tereshchenko et al. 2013). However, the Pp-1 and Pp3 genes have been not sequenced yet. The goal of the current study was to develop a set of NILs carrying various combinations of Pp alleles as a tool for genetically dissecting the purple pericarp trait, including analysis of the transcription of the key anthocyanin biosynthesis structural genes Chi (encoding chalcone-flavanone isomerase and cloned in wheat by Shoeva et al. 2014) and F3h (flavanone 3-hydroxylase; Khlestkina et al. 2008, 2013) in the pericarp of newly developed NILs.
To accelerate development of NILs microsatellite markers were utilized for genotyping of plant material obtained in crosses. Microsatellite (or SSR—simple sequence repeats) markers are widely used for marker-assistant selection in wheat (Reviewed by Leonova 2013), since they are abundant, convenient, reliable and characterized by precise positions in wheat genome (Röder et al. 1998; Ganal and Röder 2007).
Materials and methods
Plant material and phenotyping
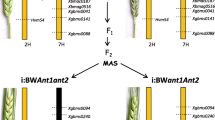
A description of the parental materials is given in Table 1. All these lines were developed in the genetic background of spring bread wheat ‘Saratovskaya 29’ (i:S29Pp-A1pp-D1pp3). The pp-A1 donor was spring bread wheat ‘Janetzkis Probat’, the Pp3 and the Pp-D1 donors were spring wheats ‘Purple’ (k-46990) and ‘Purple Feed’ (k-49426). Derivatives carrying both Pp3 and Pp-D1 were generated by Arbuzova et al. (1998). The line i:S29pp-A1pp-D1pp3 was selected from a set of doubled haploid lines described by Khlestkina et al. (2010b). The crossing scheme and the marker-assisted selection interventions are illustrated in Fig. 1. Since the same scheme was used to produce lines from both i:S29Pp-A1Pp-D1Pp3 PF and i:S29Pp-A1Pp-D1Pp3 P, only the one used for the first line has been illustrated. To evaluate anthocyanin pigmentation in the coleoptile (which was exploited as an additional marker for the selection of Pp-A1 and Pp-D1), F2 populations bred from each cross were evaluated according to Khlestkina et al. (2011). Pericarp pigmentation was scored in developing seeds within 55th–75th day after sowing. Pericarp samples for RNA extraction were collected from immature grains. Three biological replicates (from a bulk of 3–5 plants) were collected for each entry line marked in Table 1 with asterisks. The plants were grown using resources of ICG Greenhouse Core Facilities (Novosibirsk, Russia) under 12 h of light per day at 20–25 °C.
Crossing scheme and marker interventions used to obtain and validate NILs carrying various combination of Pp alleles. Chromosomal segments inherited from Purple Feed is marked in black, from Janetzkis Probat in white and from Saratovskaya 29 in grey. MAS marker-assisted selection. a–c Crosses designations
DNA extraction and microsatellite analysis
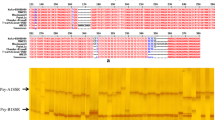
DNA was extracted from leaf material harvested from each segregant and the parental lines using a procedure described by Plaschke et al. (1995). A set of informative microsatellites chosen from the GWM series (Röder et al. 1998; Ganal and Röder 2007) was assembled for marker assisted selection purposes (Fig. 1). The PCR-conditions were as described in Röder et al. (1998). Amplicons were separated either through 5 % ACTGene agarose gels (ACTGene, Inc., Piscataway, NJ, USA) or by capillary electrophoresis using an ABI PRISM 3100 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). In the latter case, GeneScan v2.1 0 software was used to determine fragment sizes. The capillary electrophoresis was performed using resources of SB RAS Genomics Core Facilities (Novosibirsk, Russia, http://www.niboch.nsc.ru/doku.php/corefacility).
RNA extraction, reverse transcription and qRT-PCR
RNA was extracted from the pericarp of immature grains using a Plant RNA MiniPrep™ kit (Zymo Research Corporation, Irvine, CA, USA), then treated with DNAse. Each entry was represented by three biological replicates. A 0.7 μg aliquot of RNA was used to prepare single-stranded cDNA by reverse transcription, based on a RevertAid™ kit (Thermo Fisher Scientific Inc., Waltham, MA, USA) and a (dT)15 primer. The subsequent qRT-PCR was based on a SYNTOL SYBR Green I kit (Syntol, Moscow, Russia). Chi and F3h transcript abundance was assessed using the respective primer pairs 5-CTCGCCGCCAAGTGGG/5-TTCTCGAACTCGCCGGTGAC and 5-AAGGGCGGCTTCATCGTCTC/5-CCCTCCAGTCCTGCACCGC. The primers were designed using OLIGO software (Offerman and Rychlik 2003) based on multiple alignment of the Chi (Khlestkina and Shoeva 2014; Shoeva et al. 2014) and F3h (Khlestkina et al. 2008, 2013) sequences. MULTALIN software (Corpet 1988) was used to obtain multiple sequence alignments. The reference sequence used was Ubc (ubiquitin), assayed using primers suggested by Himi et al. (2005). Three technical replicates of each reaction were run. The significance of differences in transcript abundance between entries was tested using the Mann–Whitney U-test, with a p = 0.05 significance level.
Results
Marker-assisted development of NILs
Each of i:S29Pp-A1Pp-D1Pp3 PF and i:S29Pp-A1Pp-D1Pp3 P was crossed with i:S29Pp-A1pp-D1pp3 (Fig. 1a); pericarps of the F1 hybrid plants were dark purple in both cases. F2 progeny expressing a dark red coleoptile (carriers of Rc-D1 on chromosome 7D) were genotyped using microsatellites flanking Pp-D1 and Pp3 (Fig. 1) to select homozygous Pp-A1Pp-A1Pp-D1Pp-D1pp3pp3 plants, while those with light red coleoptiles (genotype Rc-A1Rc-A1rc-D1rc-D1) were used to select homozygous Pp-A1Pp-A1pp-D1pp-D1Pp3Pp3 plants. F2 segregants homozygous for the Purple Feed and Purple chromosome 7D microsatellite alleles and homozygous for the Saratovskaya 29 chromosome 2A microsatellite alleles were designated i:S29Pp-A1Pp-D1pp3 PF and i:S29Pp-A1Pp-D1pp3 P, respectively. F2 segregants homozygous for the Purple Feed and Purple chromosome 2A microsatellite alleles and homozygous for the Saratovskaya 29 chromosome 7D microsatellite alleles were designated i:S29Pp-A1pp-D1Pp3 PF and i:S29Pp-A1pp-D1Pp3 P, respectively. Both i:S29Pp-A1Pp-D1pp3 PF and i:S29Pp-A1Pp-D1pp3 P exhibited strongly pigmented coleoptiles and an uncolored pericarp, while i:S29Pp-A1pp-D1Pp3 PF and i:S29Pp-A1pp-D1Pp3 P plants produced weakly pigmented coleoptiles (due to the presence of Rc-A1) and a light purple pericarp.
Each of i:S29Pp-A1pp-D1Pp3 PF and i:S29Pp-A1pp-D1Pp3 P was crossed with i:S29pp-A1pp-D1pp3 (Fig. 1c); pericarps of the F1 hybrid plants were light purple in both cases. F2 segregants from each cross were selected on the basis of an uncolored coleoptile (genotype rc-A1/rc-A1), and were then genotyped using microsatellite markers flanking Pp-A1 and Pp3 (Fig. 1). Segregants homozygous for Janetzkis Probat chromosome 7A microsatellite alleles and Purple Feed or Purple chromosome 2A microsatellite alleles were designated i:S29pp-A1pp-D1Pp3 PF and i:S29pp-A1pp-D1Pp3 P, respectively. These lines produced uncolored coleoptiles and an uncolored pericarp.
Test crosses
The crosses i:S29Pp-A1Pp-D1pp3 PF/i:S29Pp-A1pp-D1Pp3 PF and i:S29Pp-A1Pp-D1pp3 P/i:S29Pp-A1pp-D1Pp3 P were made to verify the presence of the dominant alleles at the Pp-D1 and Pp3 loci in the lines, which had been selected either for Pp-D1 or Pp3 exclusively using markers (Fig. 1b). The purple pericarp trait was restored in the F1 plants in each cross. This confirmed the presence of the dominant allele Pp-D1 in i:S29Pp-A1Pp-D1pp3 PF/P and Pp3 allele in i:S29Pp-A1pp-D1Pp3 PF/P.
Lines i:S29Pp-A1Pp-D1pp3 PF/P and i:S29Pp-A1pp-D1Pp3 PF/P are suitable as testers to reveal the presence of the dominant allele at, respectively, Pp3 and Pp-D1. When i:S29Pp-A1pp-D1Pp3 PF was crossed with i:S29Ra (Table 1), the F1 plants bore grains exhibiting a purple pericarp, meaning that i:S29Ra carries the Pp-1 allele.
Transcription of Chi and F3h in the pericarp of the NILs
The qRT-PCR-based evaluation of transcription of Chi and F3h in the pericarp of the NILs is summarized in Fig. 2. Chi transcript was detectable even in plants lacking dominant alleles in the Pp-1 and Pp3 loci. The presence of Pp-A1 increased the transcript abundance by five fold, and a further increase (1.5–2 fold) was induced by the additional presence of either Pp-D1 or Pp3. In plants carrying both Pp-D1 and Pp3, the abundance of Chi transcript was increased eight fold compared to i:S29Pp-A1pp-D1pp3 and 40 fold compared to i:S29pp-A1pp-D1pp3 (Fig. 2). The level of F3h transcription was 2,000 fold higher in i:S29Pp-A1Pp-D1Pp3 than in i:S29pp-A1pp-D1pp3. Other combinations of the Pp alleles had no effect on the abundance of F3h transcript (Fig. 2).
Chi and F3h transcription in the pericarp of NILs carrying various combinations of Pp alleles. The lines illustrated are those which inherited the dominant allele at Pp3 and Pp-D1 from Purple Feed. Similar results were obtained for those which had inherited the dominant alleles from Purple. A statistical analysis is given in Tables S1 and S2
Discussion
The use of molecular markers can accelerate the selection process, lead to a greater accuracy of selection, reduce the acreage occupied by breeding material, and save labour and material resources (Moose and Mumm 2008; Leonova 2013; Khlestkina 2014a). Their use has halved the time needed to split i:S29Pp-A1Pp-D1Pp3 into the two homozygous lines i:S29Pp-A1pp-D1Pp3 and i:S29Pp-A1Pp-D1pp3, since the process took just three growing seasons, rather than the six which would have been required relying only on phenotypic selection. The volume of plant material needed (and hence the planting area required) was reduced by some 70 fold.
The NILs harbouring Pp-D1 and pp3 produced an uncolored pericarp, while those with pp-D1 and Pp3 had a light purple pericarp. The incomplete inhibition of anthocyanin production in the pericarp of the latter plants may be due to the continuing presence of the putative Pp-A1 gene lying within the cluster of anthocyanin synthesis regulatory genes present on cv. Saratovskaya 29 chromosome 7A (Khlestkina et al. 2010b). The suggestion is that the effect of Pp-A1 is much weaker than that of its homoeologue Pp-D1, in the same way that the effect of cv. Saratovskaya 29 Rc-A1 is less than that of Rc-D1, a gene which is quite widely distributed (Khlestkina et al. 2002, 2009, 2014). The introgression of the critical part of chromosome 7A from a non-pigmented cultivar such as cv. Janetzkis Probat into the lines i:S29Pp-A1pp-D1Pp3 PF and i:S29Pp-A1pp-D1Pp3 P having light purple pericarp resulted in an uncolored pericarp (in the lines i:S29pp-A1pp-D1Pp3 PF and i:S29pp-A1pp-D1Pp3 P; Table 1), thereby confirming the location of Pp-A1 in the genetic interval defined by Xgwm0060 and Xgwm0974 (Fig. 1). This location fits well those of Pp-D1 on chromosome 7D (Tereshchenko et al. 2012a) and Pp-B1 on chromosome 7B (Khlestkina et al. 2010a).
The utility of the NILs as testers for the presence of Pp genes has been successfully demonstrated. When i:S29Pp-A1pp-D1Pp3 PF was crossed with i:S29Ra (a line which expresses intense anthocyanin pigmentation of its coleoptile, auricles, leaf blades and leaf sheaths, but develops a non-pigmented pericarp), the resulting F1 hybrids bore grains exhibiting a dark purple pericarp, implying that i:S29Ra harbours a dominant allele at a Pp-1 gene. This gene is likely Pp-D1, a member of a complex of pigmentation genes (Rc-D1, Pc-D1, Pls-D1, Plb-D1 and Ra-D1) present on chromosome 7D (Khlestkina et al. 2014).
A previous analysis has concluded that the Pp genes act as regulators of anthocyanin synthesis in the pericarp (Tereshchenko et al. 2013). Here, the qRT-PCR method was exploited to analyze transcript abundances of Chi and F3h, the key structural genes with respect to anthocyanin synthesis, in the pericarp of the NILs carrying the various combinations of Pp alleles. The outcome of the analysis confirmed that the Pp genes indeed acted to up-regulate these anthocyanin synthesis genes. However, the conversion of chalcone to flavanone (catalyzed by CHI) did not require the presence of both Pp-1 and Pp3; one of these genes (even the weak Pp-A1) was sufficient to allow Chi transcription activation (Fig. 2). In contrast, the F3H-enabled conversion of naringenin to dihydroflavonol relied on the presence of both Pp-D1 and Pp3, since the presence of just one of these without the other was ineffective (Fig. 2). Pp-1 and Pp3 have been not sequenced, but some evidence (based on comparative mapping) points to their belonging to, respectively, the myb and myc families of transcription factors (reviewed by Khlestkina 2013). Unlike the synthesis of anthocyanin in the pericarp, that of anthocyanin in the coleoptile (and some other organs) requires the presence of a single dominant myb-like transcription factor (Himi et al. 2005; Khlestkina et al. 2008; Khlestkina 2013; Tereshchenko et al. 2013). The set of the lines developed here is suitable for clarifying the mechanisms underpinning the regulation of tissue-specific (and species-specific; Shoeva and Khlestkina 2014) anthocyanin synthesis in wheat.
The NILs developed here may aid in elucidating the physiological role of anthocyanins in the wheat pericarp. NILs represent a powerful means of establishing gene function, since they allow contrasts between a set of closely related genotypes which differ from each other largely only in and around a known target gene (reviewed by Khlestkina 2014b). Furthermore, the NILs developed here may have breeding value as donors of particular Pp alleles. The commercial interest in pigmented wheat grain flows from an understanding that they are nutritionally superior to white kernels. A combination of functional regulatory genes underlying both purple and blue grained materials may be particularly attractive. Syed Jaafar et al. (2013) have demonstrated that anthocyanin content can be boosted by stacking the purple pericarp and blue aleurone trait in a number of genetic backgrounds. The anthocyanin composition of pigmented grains (Abdel-Aal et al. 2006; Ficco et al. 2014) and the contribution of anthocyanin to the grain’s antioxidant potential (Abdel-Aal et al. 2008) are well studied. In combination with an understanding of the genetic basis of the trait and associated mapping data (Dobrovolskaya et al. 2006; Khlestkina et al. 2010a; Arbuzova et al. 2012; Tereshchenko et al. 2012a, 2013; current study), there is now a strong basis for using marker-assisted selection to increase the antioxidant content of the wheat grain.
References
Abdel-Aal E-SM, Young JC, Rabalski I (2006) Anthocyanin composition in black, blue, pink, purple, and red cereal grains. J Agric Food Chem 54:4696–4704
Abdel-Aal E-SM, Abou-Arab AA, Gamel TH, Hucl P, Young JC, Rabalski I (2008) Fractionation of blue wheat anthocyanin compounds and their contribution to antioxidant properties. J Agric Food Chem 56:11171–11177
Arbuzova VS, Maystrenko OI, Popova OM (1998) Development of near-isogenic lines of the common wheat cultivar ‘Saratovskaya 29’. Cereal Res Commun 26:39–46
Arbuzova VS, Badaeva ED, Efremova TT, Osadchaya TS, Trubacheeva NV, Dobrovolskaya OB (2012) A cytogenetic study of the blue-grain line of the common wheat cultivar Saratovskaya 29. Russ J Genet 48:785–791
Bolton FE (1970) Inheritance of blue aleurone and purple pericarp in hexaploid wheat. Plant Breed Abstr 40:2684
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 6:10881–10890
Dobrovolskaya OB, Arbuzova VS, Lohwasser U, Röder MS, Börner A (2006) Microsatellite mapping of complementary genes for purple grain colour in bread wheat (Triticum aestivum L.). Euphytica 150:355–364
Ficco DBM, De Simone V, Nigro VF, Finocchiaro F, Papa R, De Vita P (2014) Genetic variability in anthocyanin composition and nutritional properties of blue, purple and red bread (Triticum aestivum L.) and durum (Triticum turgidum L. spp. turgidum var. durum) wheats. J Agric Food Chem. doi:10.1021/jf5003683
Ganal M, Röder MS (2007) Microsatellite and SNP markers in wheat breeding. In: Varshney RK, Tuberosa R (eds) Genomics-assisted crop improvement. Vol. 2. Genomics applications in crops. Springer, Dordrecht, pp 1–24
Himi E, Nisar A, Noda K (2005) Colour genes (R and Rc) for grain and coleoptile upregulate flavonoid biosynthesis genes in wheat. Genome 48:747–754
Khlestkina EK (2013) Genes determining the coloration of different organs in wheat. Russ J Genet Appl Res 3:54–65
Khlestkina EK (2014a) Molecular markers in genetic studies and breeding. Russ J Genet Appl Res 4:236–244
Khlestkina EK (2014b) Current applications of wheat and wheat-alien precise genetic stocks. Mol Breed 34:273–281
Khlestkina EK, Shoeva OY (2014) Intron loss in the chalcone-flavanone isomerase gene of rye. Mol Breed 33:953–959
Khlestkina EK, Pestsova EG, Röder MS, Börner A (2002) Molecular mapping, phenotypic expression and geographical distribution of genes determining anthocyanin pigmentation of coleoptiles in wheat (Triticum aestivum L.). Theor Appl Genet 104:632–637
Khlestkina EK, Röder MS, Salina EA (2008) Relationship between homoeologous regulatory and structural genes in allopolyploid genome—a case study in bread wheat. BMC Plant Biol 8:88
Khlestkina EK, Pshenichnikova TA, Röder MS, Börner A (2009) Clustering anthocyanin pigmentation genes in wheat group 7 chromosomes. Cereal Res Commun 37:391–398
Khlestkina EK, Röder MS, Börner A (2010a) Mapping genes controlling anthocyanin pigmentation on the glume and pericarp in tetraploid wheat (Triticum durum L.). Euphytica 171:65–69
Khlestkina EK, Röder MS, Pshenichnikova TA, Börner A (2010b) Functional diversity at Rc (red coleoptile) locus in wheat (Triticum aestivum L.). Mol Breed 25:125–132
Khlestkina EK, Antonova EV, Pershina LA, Soloviev AA, Badaeva ED, Börner A, Salina EA (2011) Variability of Rc (red coleoptile) alleles in wheat and wheat-alien genetic stock collections. Cereal Res Commun 39:465–474
Khlestkina EK, Dobrovolskaya OB, Leonova IN, Salina EA (2013) Diversification of the duplicated F3h genes in Triticeae. J Mol Evol 76:261–266
Khlestkina EK, Gordeeva EI, Arbuzova VS (2014) Molecular and functional characterization of wheat near-isogenic line ‘i:S29Ra’ having intensive anthocyanin pigmentation of the coleoptile, culm, leaves and auricles. Plant Breed 133:454–458
Leonova IN (2013) Molecular markers: implementation in crop plant breeding for identification, introgression and gene pyramiding. Russ J Genet Appl Res 3:464–473
Lila AM (2004) Anthocyanins and human health: an in vitro investigative approach. J Biomed Biotechnol 5:306–313
Moose SP, Mumm RH (2008) Molecular plant breeding as the foundation for 21st century crop improvement. Plant Physiol 147:969–977
Offerman JD, Rychlik W (2003) Oligo primer analysis software. In: Krawetz SA, Womble DD (eds) Introduction to bioinformatics: a theoretical and practical approach. Humana Press Inc, New Jersey, pp 345–361
Piech J, Evans LE (1979) Monosomic analysis of purple grain colour in hexaploid wheat. Z Pflanzenzucht 82:212–217
Plaschke J, Ganal MW, Röder MS (1995) Detection of genetic diversity in closely related bread wheat using microsatellite markers. Theor Appl Genet 91:1001–1007
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier M-H, Leroy P, Ganal MW (1998) Microsatellite map of wheat. Genetics 149:2007–2023
Sharman BC (1958) Purple pericarp: a monofactorial dominant gene in tetraploid wheats. Nature 181:929
Shoeva OY, Khlestkina EK (2014) The specific features of anthocyanin biosynthesis regulation in wheat. In: Nasuda S, Takumi S, Matsuoka Y (eds) Wheat genetics: from genome to field. Springer, Japan
Shoeva OY, Khlestkina EK, Berges H, Salina EA (2014) The homoeologous genes encoding chalcone-flavanone isomerase in Triticum aestivum L.: structural characterization and expression in different parts of wheat plant. Gene 538:334–341
Syed Jaafar SNS, Baron J, Siebenhandl-Ehn S, Rosenau T, Böhmdorfer S, Grausgruber H (2013) Increased anthocyanin content in purple pericarp × blue aleurone wheat crosses. Plant Breed 132:546–552
Tereshchenko OY, Gordeeva EI, Arbuzova VS, Börner A, Khlestkina EK (2012a) The D genome carries a gene determining purple grain colour in wheat. Cereal Res Commun 40:334–341
Tereshchenko OY, Pshenichnikova TA, Salina EA, Khlestkina EK (2012b) Development and molecular characterization of a novel wheat genotype having purple grain colour. Cereal Res Commun 40:210–214
Tereshchenko OY, Arbuzova VS, Khlestkina EK (2013) Allelic state of the genes conferring purple pigmentation in different wheat organs predetermines transcriptional activity of the anthocyanin biosynthesis structural genes. J Cereal Sci 57:10–13
Zeven AC (1991) Wheats with purple and blue grains: a review. Euphytica 56:243–258
Acknowledgments
This study was partially supported by RFBR (Grant no. 14-04-31637), Grant from the President of the Russian Federation (MK-4252.2015.4), and the State Budget Programme (Project No. VI.53.1.5.). We thank Ms Galina Generalova for technical assistance, Dr. Valentina Arbuzova for the parental lines used in the crosses, and Dr. Robert Koebner (www.smartenglish.co.uk) for linguistic advice during the preparation of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gordeeva, E.I., Shoeva, O.Y. & Khlestkina, E.K. Marker-assisted development of bread wheat near-isogenic lines carrying various combinations of purple pericarp (Pp) alleles. Euphytica 203, 469–476 (2015). https://doi.org/10.1007/s10681-014-1317-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-014-1317-8