Abstract
Using satellite data to extract forest structure mapping parameters assists forest management. In this research, structural parameters including species, density, canopy, and gaps were extracted from SPOT-7 satellite data over Hyrcanian forests (Iran). A detailed ground inventory was initially conducted, over 12 × 1 ha (100 m × 100 m) plots, in which tree coordinates were plotted, using a differential global positioning system (DGPS), along with data on tree species, diameter-at-breast-height and height, as well as canopy dimensions, and canopy gap shapes, sizes, and positions, for each plot. Then, spectral transformations, vegetation indices, and simple spectral ratios were extracted from SPOT-7 data, and a supervised, pixel-based classification method and a support-vector machine algorithm were used to classify and determine tree species types. In addition, canopy tree borders and gaps were classified, using an object-based method, and tree densities per unit area were determined, using the canopy gravity center. Finally, the original ground data was used to perform an accuracy assessment on the extracted information, with the results showing that forest type could be determined with 95% accuracy and a Kappa coefficient of 0.8. Canopy and gap coverage achieved an overall accuracy of 91% (Kappa coefficient: 0.7), and tree densities per hectare were determined, on average, to be 47 trees fewer than reality. In conclusion, we have shown that forest structural parameters could be extracted, with good accuracy, using a combination of pixel- and object-based methods applied to SPOT-7 imaging.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Current forest conditions and future forestry operations have been determined by identifying forest structure (Koch et al. 2006). In fact, selecting suitable forestry operations and mapping forest stand structures are important methods for preserving biological diversity and the dynamics and sustainability of the forest (Awad 2018; Lu et al. 2017; Molinier et al. 2016; Pratihast et al. 2014; Hudak et al. 2006; Soenen et al. 2009). There are many forest structure mapping parameters, including density, type of species, height, diameter-at-breast-height (DBH), the spatial pattern of trees, canopies, gaps, etc., and using traditional methods for their accurate calculation is both costly and time-consuming (Bayat et al. 2019; Piermattei et al. 2019). Measurement has also been mainly limited to particular plots (sample plots) and thus could not provide continuous spatial and temporal information about the forest structure at different dimensions or scales (Vashum and Jayakumar 2012; Fatehi et al. 2015).
Satellite data could potentially be an effective source for collecting and providing broad coverage information (Lu et al. 2004; Meng et al. 2016) and could provide the processing ease and data supply timeliness required by managers (Makela and Pekkarinen 2004). On the other hand, with significant usage of this data, improvements in the detail and accuracy of the measured specifications have always been challenged (Couturier et al. 2009; Moisen et al. 2006). For example, in a study by Wallner et al. (2015), forest structural characteristics, including the density of tree bases, were determined using Rapid Eye images, which achieved acceptable results. In another study on forest type (Wolter and Townsend 2011), Landsat, Radar 1, Alos Palsar, and SPOT-5 data were used, which resulted in a map of dominant forest species with 78% accuracy. In another study (Schneider et al. 2013), canopy, gap dimensions, and single tree data were extracted, using Rapid Eye and Terra-X Data optical and radar data synchronization, while Wolter et al. (2009) and Meng et al. (2016) used SPOT Satellite data to determine structural characteristics.
Different methods of extracting satellite data have been evaluated, and their accuracy and ease of processing have been tested. For example, in a study by Wallner et al. (2015), the object-based method was used for determining the structural characteristics of a forest. In some cases, methods have been combined to determine their capabilities in extracting information. In the research of Salah (2014), the combination of pixel- and object-based data from IKONOS was used for land cover classification, and in the research of Wang et al. (2004), object- and pixel-based methods, also using IKONOS images, were applied to produce a mangrove forest classification map, with 91% precision. In another study, forest species were classified using World View 2 data interpreted using pixel- and object-based methods (Immitzer et al. 2012). Although there are many studies have been implemented in order to extract the forest characteristic information. However, there are some remain problems such as lack of research to determination of the ability of one specific data satellite for different forest structure parameters in multi-storied and dense forest; also to increase accuracy and synergies in extracting forest structure mapping parameters from satellite data, using different classification methods, this study was carried out. In the other hand, forest in Iran is different from other forests. There is no sufficient study that has been implemented, and studies in forests of Iran are still missing in order to test the suitable data satellite and different methods to forest structure extraction; therefore, we conduct this study. The parameters for this research included species, density, size, and shape of forest tree canopies and canopy gaps. For this purpose, we needed high-resolution satellite data with the potential to show the tree canopy and multiple bands, and after considering different satellite data characteristics, SPOT-7 data, with four original bands (with 1.5 m panchromatic and 6 m multispectral resolution), seemed to be the most appropriate.
From the different classification methods available for parameter extraction in forest, we used pixel-based and object-based methods because these methods show the potential application to extract forest information using satellite image and training area (Rafieyan et al. 2011b). Therefore, in present study to extract parameters of forest structure, an area located in Hyrcanian natural mixed and dense forest of Iran was chosen to test the suitability of SPOT-7 data with these two mentioned methods.
Materials and methods
Study area
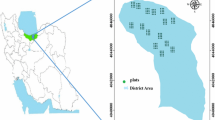
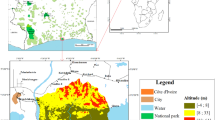
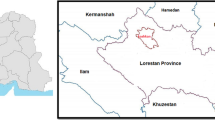
Hyrcanian natural forest is located south of the Caspian Sea in Iran. Most of Hyrcania is in current Iran and Turkmenistan. In Iran, it covers in the provinces of Golestan, Mazandaran, and Gilan that included deciduous, broadleaved, and uneven-aged forest, in a semi-Mediterranean, temperate, and humid climate (Bourque et al. 2019). The dense forests of the SANGDEH region in Mazandaran Province (part of the Hyrcanian forests) were selected as the study area (Fig. 1), for the identification and extraction of forest structure mapping parameters, using satellite data and analysis. This study area is ideally suited for this purpose because there is a typical sample of the Hyrcanian forest that has various forest stands structure; also we could find a suitable satellite data for this research.
The area was divided to seven compartments in two districts in one forestry plan and was located between 36° 03′ 22.89″ N, 53° 13′ 55.12 ″ E and 36° 01′ 33.90 ″ N, 53° 11′ 20.24 ″ E. The total study area was ~377 ha, located at elevations between 1320 and 2000 m above sea level. The annual temperature in this forest varies between 2.9 and 19.8 °C and has an annual rainfall average of 692 mm. Ground inventories were compiled for 12 × 1 ha (100 m × 100 m) sample plots, with each one located in different ecological and structural conditions.
Satellite data
SPOT-7 image sub-scenes with the panchromatic spatial resolution of 1.5 m and multispectral resolution of 6 m were used in this research. The multi-images were cloud-free and contained four bands (red, green, blue, and near infra-red (NIR), with 12-bit radiometric resolution and 9.3° nadir angle, and the data characteristics have been summarized in Table 1. The satellite data was acquired at the same time as the ground inventory was conducted (September 2016).
Methods
In the method flowchart (Fig. 2), two main phases can be seen, with Phase 1 covering the ground data collection procedure and Phase 2 showing how information was extracted from satellite data.
Ground data collection and analysis methods
The ground inventory was performed at the end of summer and in autumn 2016 and stored for use as ground-truth data. At the first stage, using preliminary maps for the 377 ha study area, basic information, such as 1: 25000 topographic maps, was secured. After producing a digital elevation model (DEM), ground layer maps (a digital terrain model with 10 m contours), including slope, aspect, and hypsometry, were produced, and then, by overlapping these layers, physiographic land units were determined, in order to reduce and remove any land shape variable effects. Then, homogeneous environmental units were identified, using ARC GIS10 (Ghanbari Motlagh et al. 2018). In the next step, areas with slope > 35% were eliminated, for error reduction of extracted information from data satellite in the mountainous forest, and finally, 29 homogeneous environmental land units were identified, from which 12 land units with area > 1 ha were selected. From each 1 ha (100 m × 100 m) sample plot taken selectively from homogeneous areas (Fig. 3), parameters such as trees position, species, DBH (> 7.5 cm), the largest and smallest canopy diameter (Noorian et al. 2016), and the size, shape, and position of forest gaps were measured, with 100% coverage of each 1 ha sample plot. At the next stage, a DGPS device (RTK model), with an RMSE accuracy of ~ 1–5 cm, was used to digitize all sample plot location data (Bettingera et al. 2019). The position of each tree was determined by recording the coordinates of the first tree in each plot and then measuring the azimuth and distance between them. It should be noted that measuring the position of all trees using DGPS would have been excessively costly and time-consuming, so we included some ground control points (GCPs) in order to correct for measurement errors.
The shapes and sizes of all large canopy gaps (> 20 m2) were measured by taking the coordinates of the gap center and then recording the azimuth and distances to the corners of the gap, using the eight-directional Brokaw method (Brokaw 1982) (Ferreira de Lima 2005)(Fig. 4).
Gap measurement methods (Brokaw 1982)
After calculating the latitude and longitude of all trees, using trigonometric methods, all information was entered into the ARC GIS software. Statistical analyses were also performed, to identify the stand structural status for each sample plot, including the frequency of species, stand tree types, tree densities, tree heights, number distributions for each diameter class, and parameter values for tree canopies and gaps.
Satellite data analysis
Firstly, radiometric and atmospheric corrections were performed on image bands (Richards 1995; Castillo-Santiago et al. 2010; Meng et al. 2016); then, in order to facilitate geometric correction and to increase multispectral data resolution, panchromatic and multispectral images were fused in PCI-Geomatics software, using the algorithms of the Pansharp process (Geomatica orthoEngine orthorectifying 2013; Zhang et al. 2012). At the next stage, 1: 25000 topography maps (Hidayat and Wiweka 2013) were used for ground control point collection, and the images were corrected to orthorectified level (Kalbi et al. 2014; Noorian et al. 2016; Ghanbari Motlagh et al. 2018). Finally, the orthorectified images were controlled using actual tree locations in each sample plot which showed that, in some areas, there was evident inaccuracy in the orthorectified image.
In order to solve this problem and increase image geometry accuracy, the DGPS tool was again used, this time in late January when trees were leafless and there was a less probability for GPS error, and 30 additional GCPs were logged. The geometric correction was performed again in the PCI-Geomatics software, after which the root mean square error was equal to 0.7 pixels, and since each pixel had a 1.5 m resolution, this meant that the orthorectification accuracy was now ~ 1 m, which was acceptable for this study. As in Rafieyan et al. (2011b), the DGPS tool was used as a tool for the secondary preprocessing of the image, to correct the geometry, which improved the results.
Image classification
For classification, different types of image transformation, plus common vegetation indices and simple spectral ratios, were used, in addition to the original bands resulting in there being the four original bands, plus 23 band ratios, indices, and image transformations, as listed in Tables 2, 3, and 4.
The pixel-based method was used to classify forest types, tree species, and other classes of the area, such as non-forested areas and gaps. Training samples are made up of the dominant tree species (based on ground studies) that affected area typology, non-forest areas (including agricultural, residential, gardens, and roads), and forest gaps or open space. Generally, two pixel-based classification methods are available for this purpose. Supervised classification is a method in which the analyst determines the classes to extract from the image; in this way, the analyst determines the areas to be nominated as training areas and shows each class of information, and then, after correcting class specifications, the computer categorizes the images (Aronoff 2005; Lillesand et al. 2004; Li et al. 2014). In contrast, unsupervised classification is determined automatically, based on statistical clustering of the value of image pixels, regardless of training samples (Lillesand et al. 2004; Puletti et al. 2014; Li et al. 2014). Out of the several different processes and algorithms applicable for each of these two methods (Li et al. 2014), we selected one of the most popular and accurate supervised classification algorithms the support vector machine (SVM) algorithm (Thanh Noi and Kappas 2018; Tavangar et al. 2019) to determine the type of forest species (Gualtieri and Cromp 1999; Huang et al. 2002; Pal and Mather 2005; Marconcini et al. 2009). The accuracy of the classifications (total accuracy, Kappa coefficient) was calculated using the previously prepared ground-truth mapping.
Determining the crown canopy and forest canopy gaps is the most important parameter for analyzing ecosystem dynamics (Bagaram et al. 2018). Object-based classification extracts phenomena from the image instead of pixels (Petropoulos et al. 2011). So for in this research, the object-based method was used to identify the crown canopy of trees, forest gaps (Malahlelaa et al. 2014), and tree density (Fadaie et al. 2012). Currently, there are several methods for performing object-based classification methods, including region growing, the Markovian method, the Watershed method, the Hierarchy algorithm, and multi-resolution segmentation (Li et al. 2014); so in this research, we used multi-resolution segmentation for extraction of the canopy and gaps, to increase classification accuracy of information. Object-based classification is often performed in two stages: (1) image segmentation based on compactness, shape, and scale and (2) classifying the segments for phenomena extraction (Dwivedi et al. 2004; Hay et al. 2005; Amini et al. 2018). Based on above-mentioned, at first, spectral value histograms were plotted, using the training sample obtained from the forest canopy and gaps, in all 27 bands (produced as explained before), and then the best bands were selected, to determine canopy and gap borders. The bands were weighted according to the histogram results and by determining compactness, scale, and shape (Sohlbach et al. 2004; Frauman and Wolff 2005; Hay and Castilla 2006; Amini et al. 2018). The plots (segments) were classified by multi-resolution segmentation, using trial and error, to find the best separation of canopy and forest gaps, and then the produced plots were classified by averaging the values of the Red band (which gave the best gap and canopy separation in histograms) across the best forest and gap class in the image. Also, to determine the density of trees in each sample plot, the geographic coordinates of the center of canopies that extracted from the images were used. Finally, the accuracy (overall accuracy and Kappa coefficient) of the produced classes was reviewed, using the ground-truth mapping (Malahlelaa et al. 2014).
Results and discussion
Sample plots analysis results
The ground inventory identified 11 species (Fagus orientalis, Carpinus betulus, Acer velutinum, Alnus subcordata, Acer cappadocicum, Prunus avium, Ulmus glabra, Ulmus carpinifolia, Quercus castaneifolia, Sorbus torminalis, and Tilia begonifolia) in the sample plots. Fagus Orientalis was the most frequent species, and then Carpinus betulus, Acer velutinum, and Alnus subcordata species were the most common species in the area. The results of sample plots’ typology mainly showed pure Fagus orientalis types, followed by Fagus orientalis-Carpinus betulus, Fagus orientalis-Acer velutinum, and Fagus orientalis-Alnus subcordata types in the area as listed in Table 5.
All characteristics were measured for 3558 trees, and the average number of trees in the stands was found to be 296 trees per hectare. The minimum number of trees (122) was in Plot 1, while the maximum (513) was in Plot 12. The ground inventory results have been listed in Tables 6 and 7. We were able to produce the gap layer (see example in Fig. 5) and tree species layers, using ArcGIS with shapefile format. The results showed that the forest was uneven-edged forest. There were around 61 gaps with an average 81.90 (m2) area.
Multispectral image information extraction results
The results of pixel-based classification
The forest species mix is an important factor in describing forest ecosystems (Immitzer et al. 2012). The orthorectified images, with area borders, were classified into six categories, using the SVM algorithm, with examples shown in Fig. 6 and Table 8.
Pixel-based classification for one sample area can be seen in Fig. 7, which shows that the species types in this particular sample plot were Fagus orientalis and Carpinus betulus and comparison with ground-truth data showed that this was correct and accurate. Canopy gaps were also accurately recognized using this method, as also seen in Fig. 7. It was noted though that the pixel-based method could not recognize Alnus subcordata in this sample plot, although the ground-truth data showed that it was present, and it is thought that this was because Alnus subcordata was present as a middle story and was therefore not so visible on satellite imagery.
In order to classify the obtained images accurately, the spectral values of the training samples were compared in the classes. The overall classification accuracy, achieved using the pixel-based method, was 95%, with a Kappa coefficient of 0.8884 (Table 9).
Among other distant sensing classifiers, SVM classification has been successfully used in recent decades (Camps-Valls and Bruzzone 2009; Izquierdo-Verdiguier et al. 2013). In comparison with Wolter and Townsend (2011), in a study on forest type using Landsat, Radar 1, Palsar, and SPOT-5 imagery, the data achieved 78% accuracy. In another study, on mangrove mapping in the Artificial Mangrove Species Mapping Using Pleiades study (Wang et al. 2018), pixel-based classification and SVM algorithm application recognized forest types with 79.63% accuracy. Based on these method accuracy confirmations, the method was also supported by use of the SVM algorithm on another forest classification study (Lu et al. 2014) and again achieved good results.
Regarding the overall accuracy (95%) of SVM method classifications, it is worth noting that the first classification is performed solely using the main bands (red, green, blue, and NIR), applying only the algorithm that identifies three classes of forest, non-forest, and gap which is not an acceptable result for stand typology. After adding the spectral indices, simple spectral ratios, and image transformations and repeating the classification operations, six classes (Fagus orientalis, Carpinus betulus, Alnus subcordata, Acer velutinum, gaps, and non-forest areas), with a total accuracy of 95% and Kappa coefficient of 0.8, were determined. These results showed that the multispectral image main bands showed similar behavior towards different study site species, in terms of their spectral reflection, but the differences in the same bands derived from simple spectral ratios (inter-band differential and division) were tangible and that their usage was very important in classification operations.
The results here cause us to highly recommend using simple inter-band ratios in forest typology studies. In another study of Rafieyan et al. (2011b) Identification of Tree Species in the Mixed Planted forest using Object-Based Classification of UltraCamD Imagery, classification accuracy was improved by using transformed images. The high overall accuracy of this classification method may also have been because the study area mainly consisted of pure Fagus orientalis stands, so the lack of species diversity and training samples probably facilitated easy, accurate, and widespread identification of F. orientalis and thus increased accuracy.
The results of object-based classification
According to the results from the inter-band spectral histograms and from the main and produced bands (27 bands), 14 bands (red, green, blue, NIR, PCA1, GEMI , Savi1, HSV2, B1 / B2, B1 / B3, B1 / B4, B2 / B3, B3-B1, and B3-B2) were distinguished for the separation of the tree canopy and gaps. The object-based method was used to weigh the bands in segmentation, and the results for compactness, scale, and shape criteria were 0.5, 6, and 0.2, respectively. The results of drawing spectral values histograms were used to classify the created plots for canopies and gap separation. Given that the best bands for canopy and gap separation were red, blue, and green and that the GEMI index determined that the most suitable bands were red and green, the red band was used for this separation, with the separation threshold for the red band taken as 150, using the histogram results. Finally, classification was performed using the object-based method. The segmentation results, and the canopy and gap classifications for the area, took three output forms: (1) the vector polygon layer of the canopy border for determining the border of the canopy and gap, (2) the point layer of the canopy center of gravity (as the tree bases), and (3) the raster layer of the canopy and gap. The results from one of sample plots are shown in Fig. 8 a) and b).
These results included gaps <20 m2, identified through the object-based method, that we did not measure during the ground inventory (which was restricted to gaps >20m2), and so the number of gaps identified for each plot exceeded the ground inventory result. This supports results from other research (Nyamgeroh et al. 2018), where detection of forest canopy gaps from very high-resolution aerial images using the object-based method also resulted in gap overestimation. This result showed the advantage of using this method for gap measurement and indicated that the object-based method could separate canopy and gaps with 91% accuracy and a Kappa coefficient of 0.7. The confusion matrix accuracy has been documented in Table 10.
In another study (Malahlela et al. 2014) on gap and canopy classification using Worldview 2 and object-based methods, the highest accuracy (93%) was obtained when the modified plant senescence reflectance index involved the red-edge band. So, our results with these available bands showed acceptable accuracy, and we deduced that errors were mostly the result of shadows (Zielewska-Büttner et al. 2016) and reduced ability to separate gaps from shadows.
It was noted that the shapes of the gaps extracted from satellite data looked different to those measured using the Brokaw method during the ground inventory. We concluded that this was because the satellite data incidence angle gave a different perspective to each gaps from that when observation was from directly below.
The density of trees in each sample plot that extracted from the data satellite in comparison with the ground inventory results is shown in Fig. 9.
Overall, in the tree density comparisons obtained using the two methods, satellite image interpretations resulted in underestimations of 47 trees per plot, on average (SD = 84), due mainly to three-storied stands and by having tree crowns covered by—or interlaced with neighboring trees (Kansanen et al. 2019). The largest error was in plot 12, which had many young, thin, short trees in its lower story. In fact, existing satellite image characteristics seem quite logical, as they simply extracted information from the forest upper layer, or from lower flat areas with a high density (such as in plot 12) and were not able to capture information relating to lower stories (Parma and Shataee-Joybari Sh 2010). In another study, a relatively low tree density determination ability was found, in comparison with this study, when mapping Zagros forest canopies using ETM+ images. Other work (Kahriman et al. 2014), which resulted in a vegetation index being produced from satellite TM data, an acceptable forest density result was achieved.
Conclusion
In this study, we have shown that the species mix and forest type could be accurately determined, with acceptable error rates, using a combination of the main SPOT-7 image bands, with produced indices, spectral transformations, and simple ratios, and using the pixel-based method and the SVM algorithm.
Separation of canopy and gaps using the object-based method and multi-resolution segmentation was also successful. Gap shapes extracted from SPOT-7 data differed somewhat from those established using the Brokaw method during the ground inventory, but gap size and number estimates were considered to be acceptable.
Tree numbers/ha (forest density) established with the object-based method consistently resulted in underestimates, but we considered this to be an understandable and acceptable result for uneven-aged, three-storied forests.
When determining exact tree locations, in either terrestrial or temporal situations, image segmentation and establishing GCPs, through a DGPS tool, will help to significantly increase the accuracy of the extracted information.
We have concluded therefore that the multispectral data used in this study, and the combination of object- and pixel-based methods, were suitable for determining forest structural components, such as the predominant species, tree density, canopy extent, and forest gaps.
References
Amini, S., Homayounib, S., Safari, A., & Darvishsefat, A. A. (2018). Object-based classification of hyperspectral data using Random Forest algorithm. Geo-spatial Information Science, 21(2), 127–138. https://doi.org/10.1080/10095020.2017.1399674.
Aronoff, S. (2005). Remote sensing for GIS managers. Redlands: ESRI Press xiiii and 487 pp., diagrams, photos, images, appendices, index. ISBN 1-58948-081-3, https://trove.nla.gov.au/version/40025360.
Attarchi, S., & Gloaguen, R. (2014). Classifying complex mountainous forests with L-band SAR and Landsat data integration: A comparison among different machine learning methods in the Hyrcanian Forest. Remote Sensing, 6(5), 3624–3647. https://doi.org/10.3390/rs6053624.
Awad. M. M. (2018) Forest mapping: a comparison between hyperspectral and multispectral images and technologies. Journal of Forestry Research, 29 (5):1395-1405, https://doi.org/10.1007/s11676-017-0528-y
Bagaram, M., Giuliarelli, D., Chirici, G., Giannetti, F., & Barbati, A. (2018). UAV remote sensing for biodiversity monitoring: Are forest canopy gaps good covariates? Remote Sensing, 10(9), 1397. https://doi.org/10.3390/rs10091397.
Bayat, M., Thanh Noi, P., Zare, R., & Tien, B. D. (2019). A semi-empirical approach based on genetic programming for the study of biophysical controls on diameter-growth of Fagus orientalis in northern Iran. Remote Sensing, 11, 1680. https://doi.org/10.3390/rs11141680.
Bettingera, P., Merry, K., Bayat, M., & Tomaštíkc, J. (2019). GNSS use in forestry – a multi-national survey from Iran, Slovakia and southern USA. Computers and Electronics in Agriculture, 158, 369–383. https://doi.org/10.1016/j.compag.2019.02.015.
Bourque, C. P. A., Bayat, M., & Zhang, C. (2019). An assessment of height–diameter growth variation in an unmanaged Fagus orientalis-dominated forest. European Journal of Forest Research, 1-15, https://doi.org/10.1007/s10342-019-01193-3.
Brokaw, N. V. L. (1982). The definition of treefall gap and its effect on measures of forest dynamics. Biotropica, 14. NO, 2, 158–160. https://doi.org/10.2307/2387750 https://www.jstor.org/stable/2387750.
Camps-Valls, G., & Bruzzone, L. (2009). Methods for remote sensing data analysis. Hoboken: Wiley https://www.wiley.com/en-us/Kernel+Methods+for+Remote+Sensing+Data+Analysis+-p-9780470722114.
Castillo-Santiago, M. A., Ricker, M., & de Jong, B. H. (2010). Estimation of tropical forest structure from spot-5 satellite images. International Journal of Remote Sensing, 31, 2767–2782. https://doi.org/10.1080/01431160903095460.
Couturier, S., Gastellu-Etchegorry, J., Patiño, P., & Emmanuel, M. (2009). A model-based performance test for forest classifiers on remote-sensing imagery. Forest Ecology and Management, 257, 23–37. https://doi.org/10.1016/j.foreco.2008.08.017.
Dwivedi, R. S., Kandrika, S., & Ramana, K. V. (2004). Comparison of classifiers of remote-sensing data for land-use/land-cover mapping. Current Science, 86(2), 328–335 https://www.jstor.org/stable/24107878.
Fadaie, H., Suzuki, R., & Avtar, R. (2012). Estimation tree density as object-based in arid and semi-arid regions using ALOS, Proceedings of the 4th GEOBIA, May 7–9 - Rio de Janeiro - Brazil. p. 668.
Fatehi, P., Damm, A., Schaepman, M. E., & Kneubühler, M. (2015). Estimation of Alpine Forest structural variables from imaging spectrometer data. Remote Sensing, 7(12), 16315–16338. https://doi.org/10.3390/rs71215830.
Ferreira de Lima, R. A. (2005). Gap size measurement: The proposal of a new field method. Forest Ecology and Management, 214, 413–419. https://doi.org/10.1016/j.foreco.2005.04.011.
Frauman, E., & Wolff, E. (2005). Segmentation of very high spatial resolution satellite images in urban areas for segments-based classification. In In proceedings for 3rd international symposium remote sensing and data fusion over urban areas. Tempe: Arizona. https://www.semanticscholar.org/paper/Segmentation-of-very-high-spatial-resolution-images-Frauman-Wolff/95680b7a45ed8425e592e069e06674033f257f94.
Geomatica OrthoEngine Orthorectifying SPOT6 data, Tutorial (2013) http://www.pcigeomatics.com/pdf/SPOT6_OE_Ortho_Pan_2013.pdf.
Ghanbari Motlagh, M., Babaie Kafaky, S., Mataji, A., & Akhavan, R. (2018). Estimating and mapping forest biomass using regression models and Spot-6 images (case study: Hyrcanian forests of north of Iran). Environmental Monitoring and Assessment, 190, 352–314. https://doi.org/10.1007/s10661-018-6725-0.
Gualtieri, J. A., & Cromp, R. F. (1999). Support vector machines for hyperspectral remote sensing classification. In the 27th AIPR workshop: Advances in computer-assisted recognition. International Society for Optics and Photonics, 3584, 221–232. https://doi.org/10.1117/12.339824.
Hay, G. J., & Castilla, G. (2006). Object-based image analysis: Strengths, weaknesses, opportunities and threats (SWOT). Paper presented at the International Archives of the Photogrammetry. Remote Sensing and Spatial Information Sciences, http://www.isprs.org/proceedings/XXXVI/4-C42/Papers/01_Opening%20Session/OBIA2006_Hay_Castilla.pdf.
Hay, G. J., Castilla, G., Wulde, M. A., & Ruiz, J. R. (2005). An automated object-based approach for the multiscale image segmentation of forest scenes. International Journal of Applied Earth Observation and Geoinformation, 7(4), 339–359. https://doi.org/10.1016/j.jag.2005.06.005.
Hidayat, S. A., & Wiweka. (2013). Accuracy evaluation orthorectification of SPOT image. International Conference on ICT for Smart Society, 13–14 June, Jakarta, Indonesia, https://doi.org/10.1109/ICTSS.2013.6588096.
Huang, C., Davis, L. S., & Townshend, J. R. G. (2002). An assessment of support vector machines for land cover classification. International Journal of Remote Sensing, 23, 725–749. https://doi.org/10.1080/01431160110040323.
Hudak, A. T., Crookston, N. L., Evans, J. S., Falkowski, M. J., Smith, A. M., Gessler, P. E., et al. (2006). Regression modeling and mapping of coniferous forest basal area and tree density from discrete-return LiDAR and multispectral satellite data. Canadian Journal of Remote Sensing, 32, 126–138 https://www.fs.usda.gov/treesearch/pubs/24612.
Immitzer, M., Atzberger, C., & Koukal, T. (2012). Tree species classification with random forest using very high spatial resolution 8-band WorldView-2 satellite data. Remote Sensing, 4(9), 2661–2693. https://doi.org/10.3390/rs4092661.
Izquierdo-Verdiguier, E., Laparra, V., Gómez-Chova, L., & Camps-Valls, G. (2013). Encoding invariances in remote sensing image classification with SVM. Geoscience and Remote Sensing Letters, 10(5. 1 page). https://doi.org/10.1109/LGRS.2012.2227297.
Jensen, J R. (2000). Remote sensing of the environment (an earth resource perspective). Prentice hall upper saddle river, nj 07458.
Jiang, Y., Carrow, R. N., & Duncan, R. R. (2003). Correlation analysis procedures for canopy spectral reflectance data of seashore paspalum under traffic stress. Journal of American Society, 13, 187–208. https://doi.org/10.21273/jashs.128.3.0343.
Kahriman, A., Gunlu, A., & Karahalil, U. (2014). Estimation of crown closure and tree density using Landsat TM satellite images in mixed forest stands. Journal of the Indian Society of Remote Sensing, 42, 559–567. https://doi.org/10.1007/s12524-013-0355-3.
Kalbi, S., Fallah, A., & Shataee, S. (2014). Estimation of forest attributes in the Hyrcanian forests, comparison of advanced space-borne thermal emission and reflection radiometer and satellite poure I’observation de la terre high resolution grounding data by multiple linear, and classification and regression tree regression models. Journal of Applied Remote Sensing, 8, 083632–083632. https://doi.org/10.1117/1.JRS.8.083632.
Kansanen, K., Vauhkonen, J., Lähivaara, T., Seppänen, A., Maltamo, M., & Mehtätalo, L. (2019). Estimating forest stand density and structure using Bayesian individual tree detection, stochastic geometry, and distribution matching. ISPRS Journal of Photogrammetry and Remote Sensing, 152, 66–78. https://doi.org/10.1016/j.isprsjprs.2019.04.007.
Koch, B., Heyder, U., & Weinacker, H. (2006). Detection of individual tree crowns in airborne LIDAR data. Photogrammetric Engineering and Remote Sensing, 72(4), 357–363. https://doi.org/10.14358/PERS.72.4.357.
Li, M., Zang, S., Zhang, B., Li, S., & Changshan, W. (2014). A review of remote sensing image classification techniques: The role of Spatio-contextual information. European Journal of Remote Sensing, 47, 389–411. https://doi.org/10.5721/EuJRS20144723.
Lillesand, T. M., Kiefer, R. W., & Chipman, J. W. (2004). Remote sensing and image interpretation (5th ed.). Hoboken: Wiley.
Lu, D., Mausel, P., Brondízio, E. S., & Moran, E. (2004). Relationships between forest stand parameters and Landsat TM spectral responses in the Brazilian Amazon Basin. Forest Ecology and Management, 198, 149–167. https://doi.org/10.1016/j.foreco.2004.03.048.
Lu, M., Chen, B., Liao, X., Yue, T., Yue, H., Ren, S., et al. (2017). Forest types classification based on multi-source data fusion. Remote Sensing, 9(11), 1153. https://doi.org/10.3390/rs9111153.
Lu, D., Chen, Q., Wang, G., Liu, L., Li, G., Moran, E. (2014). A survey of remote sensing-based aboveground biomass estimation methods in forest ecosystems, International Journal of Digital Earth, 9 (1), pages 63- 105, http://dx.doi.org/10.1080/17538947.2014.990526
Makela, H., & Pekkarinen, A. (2004). Estimation of forest stand volumes by Landsat TM imagery and stand-level field-inventory data. Forest Ecology and Management, 196, 245–255. https://doi.org/10.1016/j.foreco.2004.02.049.
Malahlelaa, O., Azong Choa, M., & Mutanga, O. (2014). Mapping canopy gaps in an indigenous subtropical coastal forest using high-resolution WorldView-2 data. International Journal of Remote Sensing, 35(17), 6397–6417. https://doi.org/10.1080/01431161.2014.954061.
Marconcini, M., Camps-Valls, G., & Bruzzone, L. (2009). A composite semisupervised SVM for classification of hyperspectral images. IEEE Geoscience and Remote Sensing Letters, 6, 234–238. https://doi.org/10.1109/LGRS.2008.2009324.
Meng, J., Li, S., Wang, W., Liu, Q., Xie, S., & Ma, W. (2016). Estimation of forest structural diversity using the spectral and textural information derived from SPOT-5 satellite images. Remote Sensing, 8(2), 125. https://doi.org/10.3390/rs8020125.
Moisen, G. G., Freeman, E. A., Blackard, J. A., Frescino, T. S., Zimmermann, N. E., & Edwards Jr., T. C. (2006). Predicting tree species presence and basal area in Utah: A. comparison of stochastic gradient boosting, generalized additive models, and tree based methods. Ecological Modelling, 199, 176–187. https://doi.org/10.1016/j.ecolmodel.2006.05.021.
Molinier, M., Lopez-Sanchez, C. A., Toivanen, T., Korpela, I., Corral-Rivas, J. J., Tergujeff, R., & Häme, T. (2016). Relasphone- Mobile and participative in situ forest biomass measurements supporting satellite image mapping. Remote Sensing, 8(10), 869. https://doi.org/10.3390/rs8100869.
Noorian, N., Shataee-Jouibary, S., & Mohammadi, J. (2016). Assessment of different remote sensing data for forest structural attributes estimation in the Hyrcanian forests. Forest Systems, 25(3), e074, 11 pages. https://doi.org/10.5424/fs/2016253-08682%20.
Nyamgeroh, B. B., Groen, T. A., Weir, M. J. C., Dimov, P., & Zlatanov, T. (2018). Detection of forest canopy gaps from very high resolution aerial images. Ecological Indicators, 95, 629–636. https://doi.org/10.1016/j.ecolind.2018.08.011.
Pal, M., & Mather, P. M. (2005). Support vector machines for classification in remote sensing. International Journal of Remote Sensing, 26, 1007–1011. https://doi.org/10.1080/01431160512331314083.
Parma, R. A., & Shataee- Joybari, Sh. (2010). Capability study on mapping the diversity and canopy cover density in Zagros forests using ETM+ images (case study Ghalajeh forests, Kirmanshah province). Iranian Journal of Forest, 2(3) https://www.sid.ir/En/Journal/ViewPaper.aspx?ID=181568.
Petropoulos, G. P., Kontoes, C., & Keramitsoglou, I. (2011). Burnt area delineation from a Uni-temporal perspective based on Landsat TM imagery classification using support vector machines. International Journal of Applied Earth Observation and Geoinformation, 13(1), 70–80. https://doi.org/10.1016/j.jag.2010.06.008.
Piermattei, L., Karel, W., Wang, D., Wieser, M., Mokroš, M., Surový, P., Koreň, M., Tomaštík, J., Pfeifer, N., & Hollaus, M. (2019). Terrestrial structure from motion photogrammetry for deriving Forest inventory data. Remote Sensing, 11(8), 950. https://doi.org/10.3390/rs11080950.
Pratihast, A. K., DeVries, B., Avitabile, V., Bruin, S., Kooistra, L., Tekle, M., et al. (2014). Combining satellite data and community-based observations for forest monitoring. Forests, 5, 2464–2489. https://doi.org/10.3390/f5102464.
Puletti, N., Perria, R., & Storchi, P. (2014). Unsupervised classification of very high remotely sensed images for grapevine rows detection. European Journal of Remote Sensing, 47, 45–54. https://doi.org/10.5721/EuJRS20144704.
Rafieyan, O., Darvishsefat, A. A., Babaii, S., & Mataji, A. (2011b). Object-based classification of UltraCamD imagery for identification of tree species in the mixed planted forest. Caspian Journal of Environmental Sciences, 9(1), 67–79 https://cjes.guilan.ac.ir/article_1051.html.
Richards, J. A. (1995). Remote sensing digital image analysis: An introduction. Springer.
Salah, M. (2014). Combining pixel-based and object- based support vector machines using Bayesian probability theory. ISPRS Annals of the Photogrammetry, Remote Sensing and Spatial Information Sciences, volume II-7, ISPRS technical commission VII symposium, 29 September – 2 October, Istanbul, Turkey, https://www.isprs-ann-photogramm-remote-sens-spatial-inf-sci.net/II-7/67/2014/isprsannals-II-7-67-2014.pdf.
Schneider, T., Elatawneh, A., Rahlf, J., Kindu, M., Rappl, A., Thiele, A., Boldt, M., & Hinz, S. (2013). Parameter determination by RapidEye and TerraSAR-X Data: A Step Toward a Remote Sensing Based Inventory, Monitoring and Fast Reaction System on Forest Enterprise Level. Earth Observation of Global Changes. https://doi.org/10.1007/978-3-642-32714-8_6.
Soenen, S. A., Peddle, D. R., Coburn, C. A., Hall, R. J., & Hall, F. G. (2009). Canopy reflectance model inversion in multiple forward model: Forest structural information retrieval from solution set distributions. Photogramm. Engineering & Remote Sensing, 75, 361–374. https://doi.org/10.14358/PERS.75.4.361.
Sohlbach, M., Weber, M., & Willhauck, G. (2004). eCognition professional: User guide 5. München: Definiens Imaging GmbH.
Tavangar, S., Moradi, H., Massah Bavani, A., & Gholamalifard, M. (2019). A futuristic survey of the effects of LU/LC change on stream flow by CA–Markov model: A case of the Nekarood watershed. Iran. Geocarto International. https://doi.org/10.1080/10106049.2019.1633419.
Thanh Noi, P., & Kappas, M. (2018). Comparison of random Forest, k-nearest neighbor, and support vector machine classifiers for land cover classification using Sentinel-2 imagery. Remote Sensing, 18(1), 18. https://doi.org/10.3390/s18010018.
Tucker, C. J. (1979). Red and photographic infrared linear combinations for monitoring vegetation. Remote Sensing of Environment, 8, 127–150. https://doi.org/10.1016/0034-4257(79)90013-0.
Vashum, K. T., & Jayakumar, S. (2012). Methods to estimate above-ground biomass and carbon stock in natural forests: A review. Journal of Ecosystem & Ecography, 2, 1–7. https://doi.org/10.4172/2157-7625.1000116.
Wallner, A., Elatawneh, A., Schneider, T., & Knoke, T. (2015). Estimation of forest structural information using RapidEye satellite data. Forestry: An International Journal of Forest Research, 88(1), 96–107. https://doi.org/10.1093/forestry/cpu032.
Wang, L., Sousa, W. P., & Gong, P. (2004). Integration of object-based and pixel-based classification for mapping mangroves with IKONOS imagery. International Journal of Remote Sensing, 25(24). https://doi.org/10.1080/014311602331291215.
Wang, D., Wan, B., Qiu, P., Su, Y., Guo, Q., & Wu, X. (2018). Artificial mangrove species mapping using Pléiades-1: An evaluation of pixel-based and object-based classifications with selected machine learning algorithms. Remote Sensing, 10(2), 294. https://doi.org/10.3390/rs10020294.
Wolter, P. T., & Townsend, P. A. (2011). Multi-sensor data fusion for estimating forest species composition and abundance in northern Minnesota. Remote Sensing of Environment, 115, 671–691. https://doi.org/10.1016/j.rse.2010.10.010.
Wolter, P. T., Townsend, P. A., & Sturtevant, B. R. (2009). Estimation of forest structural parameters using 5 and 10 m SPOT-5 satellite data. Remote Sensing of Environment, 113, 2019–2036. https://doi.org/10.1016/j.rse.2009.05.009.
Zhang, L., Shen, H., Gong, W., & Zhang, H. (2012). Adjustable model-based fusion method for multispectral and panchromatic images. IEEE Transactions on system, man, and cybernetics-part B, 42, 1693–1704. https://doi.org/10.1109/TSMCB.2012.2198810.
Zielewska-Büttner, K., Adler, P., Michaela Ehmann, M., & Braunisch, V. (2016). Automated detection of forest gaps in spruce dominated stands using canopy height models derived from stereo aerial imagery. Remote Sensing, 8, 175. https://doi.org/10.3390/rs8030175.
Acknowledgments
The authors would like to appreciate those who assisted in conducting this work. Also thanks to Remote sensing Institute of K.N. Toosi University of Tech. for providing the opportunity to use the remote sensing lab for this research. We thank anonymous reviewers who provided many helpful comments and suggestions for improving this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rahimizadeh, N., Babaie Kafaky, S., Sahebi, M.R. et al. Forest structure parameter extraction using SPOT-7 satellite data by object- and pixel-based classification methods. Environ Monit Assess 192, 43 (2020). https://doi.org/10.1007/s10661-019-8015-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-019-8015-x