Abstract
In recent years, a devastating fungal disease characterized by small black spots (<1 mm) on sandy pear (Pyrus pyrifolia Nakai) leaves has occurred in sudden outbreaks in southern China, resulting in severe defoliation and a loss of fruit quality and yield. To identify the etiology of the disease, 147 fungal isolates from sandy pear leaves showing typical black spots were collected from 30 orchards; 10 fungal species were included among the isolates, and the prevailing 124 Colletotrichum isolates were subjected to morphological and molecular characterization. Based on differences in colony morphology, the isolates were separated into five groups (I to V); the groups were specifically separated according to a combination of colony color, pigment, sporulation, perfect stage, and spore size. In addition to their morphological features, the five groups were assessed for their molecular and taxon statuses based on ITS sequences, multilocus regions (ITS, ACT, TUB, CHS-1, GAPDH), and the Apn2/MAT locus, which provided molecular proof for the identification of these isolates as Colletotrichum fructicola Prihastuti. The result also indicated different resolution of the three loci in delimiting the species. Koch’s postulates were fulfilled by inoculating conidium suspensions of the representative isolates on attached and detached leaves of sandy pear cvs. Cuiguan, Xueqing and Huali No. 2 with analogous conditions (under saturated humidity and with tiny wounds) to those that appeared in the field, and similar symptoms were elicited by the isolates, suggesting that C. fructicola was causing the disease. To our knowledge, this is the first report of C. fructicola causing leaf black spot of pear. The etiological clarification and characterization of the C. fructicola isolates provide useful information to aid in the understanding of leaf black spot disease in pears and for designing management strategies to control this economically significant disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since 2009, a new disease characterized by small black spots on the leaves of sandy pear (Pyrus pyrifolia Nakai) has been observed in many provinces in the southern Yangtze valley in China, especially in Fujian, Zhejiang and Jiangxi provinces, which are located along the coast. The disease occurs most severely after typhoons and storms have passed over these areas, and the incidence has varied for different sandy pear cultivars (Chen et al. 2011). With severe defoliation, the vigor, fruit quality and yield of the infected trees decreased remarkably, some trees did not produce any yield in the current year, and the formation of flower buds was also reduced (Chen et al. 2011). The disease was widespread in P. pyrifolia cv. Cuiguan, causing a devastating loss for growers. For example, in Fujian province, the disease spread throughout 15–30 % of the pear-cultivated areas and caused more than 30 million kg in fruit loss and an estimated economic loss of more than 30 million US dollars each year (Huang et al. 2010). The causal agent has thus far not been identified.
A leaf spot disease on apples (Malus pumila Mill. cv. Gala) called Glomerella leaf spot (GLS) was initially described in the United States in 1998 (Sutton and Sanhueza 1998). Subsequently, the same symptoms were described in apple fruits in Brazil, and it was demonstrated that the disease was caused by G. cingulata (anamorph: Colletotrichum gloeosporioides (Penzig) Penzig & Saccardo). The symptoms of the disease were defoliation of up to 75 % of the leaves, weakened vigor and reduction in fruit yield from infected trees (González et al. 2006). The disease has also been reported in Japanese pear (P. pyrifolia Nakai cv. culta Nakai), and C. gloeosporioides was identified as the causal agent in cvs. Housui and Niitaka (Morita et al. 1994; Tashiro et al. 2012) and by C. acutatum in cv. Kousui (Fukaya 2004), showing severe defoliation and reduced vigor of infected trees. Although black spot disease of sandy pear in China showed the symptomatic black spots on their leaves and defoliation, G. cingulata, C. gloeosporioides or C. acutatum were not consistently isolated from the symptomatic tissues in our previous work.
Identification of the responsible pathogen will provide useful information for the prevention and control of the disease. Initially, we sought to identify this disease, but without success. We hypothesized that unsuccessful inoculation might be due to the use of an unsuitable method. Considering that the disease occurred after typhoons and storms, it was likely that the conidia are disseminated in the air, landing on host leaves, germinating at saturated relative humidity, and the disease becoming manifested as black spots in the invaded locations. Simultaneously, tiny sand particles flying up along with the conidia during typhoons and storms likely wound the leaves, rendering them more vulnerable to pathogen invasion. This cycle of weakened leaves and vulnerability to infection may result in the appearance of increasing numbers of black spots. We therefore designed protocols with analogous conditions; in an environment of saturated humidity, we used conidia to inoculate wounded and unwounded sandy leaves (with or without carborundum particles, respectively) to clarify the responsible etiology.
Materials and methods
Sample collection and survey of field symptoms
A field symptom survey was carried out on sandy pear trees of P. pyrifolia (cvs. Cuiguan, Xuefang, Xueqing, Xingao, Qingxiang, Xiuyu, Jinshui No. 2, Yushui, Zaomeisu, Deshengxiang, Huanghua, Longquan No. 1, Ningmenghuang, Xueying, Longquan No. 19, Jin Twenty Century, Mixueli, and Longquan No. 23) in thirty orchards in Jianning, Mingxi, Qingliu and Jianou counties in the Fujian province of China, from August 2013 to May 2014. Approximately 60 leaves showing small black spots were collected from the sandy pear trees of cv. Cuiguan in 20 of the surveyed orchards.
Fungal isolation
A small piece (approximately 5 mm2) containing only one black spot was cut from each of the collected leaf samples, surface-sterilized with 75 % ethanol, air-dried, placed on potato dextrose agar (PDA) in Petri dishes, and incubated at 25 °C for 7 days in darkness. When colony growth started, each colony was transferred to a new PDA Petri dish and was designated as a specific isolate. Each isolate was further purified by culturing from a single conidium. Briefly, an isolate was cultured on PDA for 7 days at 25 °C in darkness for sporulation; the formulated conidia were then rinsed with sterile water and diluted to a low concentration with one or two conidia under the microscope (viewed at 100× magnification). One hundred microliters of this conidial solution was transferred to PDA for single colony formulation, and four single colonies of each isolate were selected and stored in 25 % glycerol at −70 °C until use. The isolates that did not sporulate were purified by the single-mycelium tipping method, which has been described before (Yaegashi et al. 2012).
Morphological observation
Mycelial plugs were taken from areas of active growth close to the margin of a colony, freshly cultured for 3 days, transferred to a new PDA plate, and incubated at 25 °C for 7 days in darkness for morphological observation (performed in triplicate). The growth rate and appearance of the colonies were recorded daily for 5 days post inoculation (dpi).
The shape, color and size of fruiting bodies, conidia, ascospores and appressoria were observed under a light microscope (Nikon Eclipse E600 FN, Nikon, Japan). Fifty conidia or ascospores were measured to determine the average size unless fewer spores were produced. Appressoria were produced using a slide culture technique (Johnston and Jones 1997; Cai et al. 2009) and subjected to microscopic inspection after incubating the conidia in sterile water at 25 °C in darkness for 12 h (Liu et al. 2014).
Pathogenicity analysis
Pathogenicity analysis was performed on attached and detached leaves (1 to 2 months old) from sandy pear trees of P. pyrifolia cvs. Cuiguan, Xueqing and Huali No. 2 using conidial suspensions of 1.0 × 106 or 1.5 × 106 conidia per ml. Each leaf was surface-sterilized with 75 % ethanol, washed with sterile water, air-dried, and inoculated with 40 μl of conidial suspensions after first gently rubbing the leaves with conidial solution with carborundum particles (74 μm in diameter) or by spraying the conidial suspensions with a hand held sprayer over the leaves without carborundum particles. For the attached leaves, six leaves on each of three potted trees (1 year old) were inoculated. Control leaves were inoculated in parallel with sterile water. The inoculated attached leaves were covered with closed translucent plastic bags to maintain water saturated air for 24 to 48 h and were cultured outdoors in Wuhan, Hubei province of China, from April to May 2014. During the 2 months, the average daily lowest temperature was 13 °C and 18 °C, and the highest temperature was 21 °C and 26 °C, with an average monthly rainfall of 137 and 166 mm after inoculation, respectively. The leaves were sprayed with water as a drought view was observed in bags to maintain a high humidity each day. The experiments were carried out two times with 10 days between start of each. The inoculated detached leaves were placed on sterilized wet clothes in a box, covered with a plastic membrane to secure water saturated air, and incubated at 25 °C with a 12/12 h light/dark photoperiod for 12 days.
Black spots were counted at 7 dpi and subjected to statistical analysis. The severity of the disease was quantified using a disease index (McKinney 1923) (\( DI={\displaystyle \sum_{k=1}^k{F}_k{x}_k/n{x}_k} \), Fk, the number of leaves with certain degree of infection xk, k = 0, 1, …,k; n, the number of leaves evaluated; xk, the highest number from the adopted scale). The values of disease severity were assigned according to the number of black spots on the inoculated leaves as 0 (0 black spots), 1 (less than 10), 2 (10 to 50), 3 (51 to 120), 4 (121 to 200) and 5 (more than 200).
DNA extraction, PCR amplification and sequencing
Fungal genomic DNA was extracted with cetyltrimethyl ammonium bromide (CTAB) buffer [2 % w/v CTAB, 1.42 M NaCl, 20 mM EDTA,100 mM Tris · HCl, pH 8.0, 0.2 % (w/v) β-mercaptoethanol] according to a previously described method (Freeman et al. 1996) and dissolved in Tris-EDTA (TE) buffer (10 mM Tris–HCl, 1 mM EDTA, pH 8.0). PCR amplification was carried out in a Thermal Cycler (Model PTC-100, MJ Research, Inc., Watertown, MA, USA) using the primers listed in Table 1 with the reported programs (Su et al. 2011; Silva et al. 2012; Weir et al. 2012). Amplified PCR products were gel-purified using a MoreBio PCR Purification Kit (Wuhan More Biotechnology Corporation, Wuhan, China), cDNA fragments less than 1000 bp were directly sequenced, and fragments greater than 1000 bp were sequenced after they were cloned into the pMD18-T vector (TaKaRa Biotech. Co., Dalian, China) at GenScript Co., Ltd, Nanjing, China.
Phylogenetic analysis
Sequences were aligned with those extracted from the GenBank database using the BLASTn program (http://blast.ncbi.nlm.nih.gov). Sequence alignment was carried out using CLUSTALX (version 1.83) (Larkin et al. 2007). A phylogenetic tree was constructed using the MEGA 5.2.2 software (Tamura et al. 2011) with the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) or Maximum Likelihood (ML) method, with default parameters and bootstrap values of 1000 replicates.
Statistical analysis
Data (growth rate, conidium and ascospore size, and pathogenicity analysis) were analyzed with SPSS Statistics 21.0 (WinWrap Basic; http://www.winwrap.com) by one-way ANOVA, and means were compared using Tukey’s test at P = 0.05. Homogeneity of variance was tested before analysis.
Results
Field symptom survey
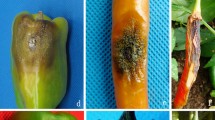
On all of the 18 surveyed cultivars of sandy pear, leaf black spot disease showed the same symptomatic small black spots (<1 mm) on the leaves. A few small black spots initially appeared on the leaves (Fig. 1a). Later black spots rapidly increased in numbers until most of the leaf was covered (Fig. 1b). The infected leaf formed small “green island” regions restricted to the areas where the black spots were located (Fig. 1c), gradually turning to a yellowish color (Fig. 1d), and finally fell from the tree 4–5 days after the first symptoms had emerged (Fig. 1e). From the initial emergence of the black spots up to the point of defoliation, the size of black spots remained unchanged.
Typical symptoms of leaf black spot caused by Colletotrichum fructicola observed in sandy pear cv. Cuiguan in the field. a initial leaf symptoms; b–c gradually developing leaf symptoms showing numerous black spots and the light yellowing of infected tissues; d yellowish discoloration of the leaf at a later stage; e defoliation of an infected tree
Collection of Colletotrichum species and appearance of colonies
Out of 60 leaf samples of P. pyrifolia cv. Cuiguan exhibiting symptomatic black spots collected from 30 orchards in the Fujian province, a total of 147 single colonies (i.e., isolates) were obtained. Of those, nine fungal species were isolated with a low incidence and identified based on their ITS sequences and morphologies, including Guignardia mangiferae (five colonies), Nigrospora sphaerica (5), Phomopsis sp. (4), Pestalotiopsis sp. (2), Fusarium sp. (2), Neurosporocrassa sp. (2), Alternaria sp. (1), Aspergillus sp. (1), and Trametes versicolor (1). In addition to the nine species, sparse, cotton-like colonies with white edges, characteristic of fungi belonging to the Colletotrichum genus, appeared on 124 of the isolates.
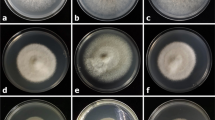
The 124 Colletotrichum isolates were further examined via the single conidium technique or single-mycelium tipping for the isolates lacking sporulation. After 5 days of culturing at 25 °C in darkness, differences in colony morphologies of the Colletotrichum isolates allowed for the formation of five groups (temporarily termed groups I to V), which contained 81, 24, 17, 1 and 1 isolate(s), respectively (Fig. 2a and Table 2).
Colony appearance and perfect and imperfect stages of representative isolates of Colletotrichum fructicola of five groups (FJ-85, FJ-110, FJ-116, FJ-103 and FJ-42) after 5 days on PDA. Numerals I-V of panel a refer to the colony morphology of group I (FJ-85), II (FJ-110), III (FJ-116), IV (FJ-103) and V (FJ-42) from the upper and reverse sides, respectively. Numerals I to VIII of panel b indicate conidiophores and conidia (I), conidia (II), germinated conidia (III), germlings with appressoria (IV), agar-embedded stromas (V), perithecium (VI), cross-section of a perithecium (VII), and ascospores (VIII). Scale bar = 10 μm
Group I isolates showed off-white colonies with gray centers of sizes less than half of that of the Petri plates (Fig. 2aI, top). On the reverse side gray pigments were discretely distributed in the center and spread to variable extents, ranging from 1/3 to 1/2 of the colonies, and a tiny pinkish pigment was faintly observed (Fig. 2aI, reverse). Group II isolates showed off-white colonies with one or two faintly depressed concentric rings (Fig. 2aII, top). On the reverse side tiny gray pigments were spread within the concentric rings, and pink pigments were faintly dispersed throughout the center (Fig. 2aII, reverse). Group III isolates showed colonies that were checkered with black and white mycelia and had gray fan-shaped regions (Fig. 2aIII, top). On the reverse side remarkable dark-brown pigments were spread over approximately 2/3 of the colonies (Fig. 2aIII, reverse). The Group IV isolate showed a snow-white colony embodied with a black mycelium (Fig. 2aIV, top). On the reverse side black or dark-brown pigments were deposited in the region corresponding to the black mycelium region observed from the top (Fig. 2aIV, reverse). The Group V isolate showed snow-white colonies with one sunken concentric ring (Fig. 2aV, top), and no pigment was produced within the mycelia (Fig. 2aV).
Micromorphological characterization
For all of the isolates, the hyphae were hyaline, septate and branched, and setae were absent. Conidia were produced in abundance on short conidiophores (Fig. 2bI) after 7 days of culture. The conidia were hyaline, cylindrical, straight, and had rounded ends (Fig. 2bII). When germinated, the conidia produced one or two long round hyaline germ tubes from the end(s) or occasionally from the side(s) (Fig. 2bIII). The appressoria were oval to fusoid with smooth edges, not lobed, brown to dark brown in color, and contained gray granular particles (Fig. 2bIV). The isolates were cultured on PDA for 12–20 days and readily formed abundant stromata that were brown, globose, often immersed in the agar (Fig. 2bV), and in fewer numbers on the surfaces or amongst the aerial mycelia. As the stromata were crushed, brown subglobose perithecia were observed (Fig. 2bVI). The asci were unitunicate, clavate to cymbiform, and vertically arranged in the bottom of the perithecium (Fig. 2bVII); each contained eight hyaline guttulate fusiform-to-slightly curved ascospores with rounded ends (Fig. 2bVIII).
Conidia of the representative isolates of the five groups had average lengths ranging from 11.8 to 14.1 μm and widths ranging from 4.3 to 6.0 μm. For ascospores, lengths varied from 17.2 to 20.8 μm and widths from 4.9 to 5.6 μm. The conidia or ascospores had significant differences in the lengths (p < 0.001) and widths (p < 0.001) among isolates. No ascospores were observed for the isolate from group V.
In accordance with previous characterization of Colletotrichum spp. (Simmonds 1968) and other descriptions (Prihastuti et al. 2009; Weir et al. 2012; Huang et al. 2013; Peng et al. 2013), the 124 fungal isolates were identified as members of the Colletotrichum genus based on morphological characteristics, including the appearance of colonies, appressoria, conidia and ascospores.
Growth rate
Measurements of the growth rates of the isolates used for micromorphological characterization revealed growth rates ranging from 10.0 to 13.4 mm/d. There was a clear difference between the growth rates of different isolates (P < 0.001), with the lowest growth rates of 10.0 and 10.2 mm/d exhibited by isolates FJ-119 (group I) and FJ-103 (group IV), respectively; a medium growth rate of 10.8 for FJ-110 (group II); and highest growth rates of 12.6 and 13.4 mm/d for FJ-1 (group II) and FJ-116 (group III), respectively (Table 2).
Pathogenicity analysis
Because the isolates from groups IV and V produced insufficient conidia for inoculation, only seven isolates belonging to groups I through III (i.e., group I: FJ-73-2, FJ-85, FJ-119, FJ-122; group II: FJ-1, FJ-110; group III: FJ-116) were subjected to fulfillment of Koch’s postulates by inoculating conidia on attached leaves of sandy pear cv. Cuiguan. The simulation of the wounded leaf conditions (performed by grinding the conidia with carborundum particles) showed that all of the isolates except FJ-122 had produced small black spots (with an approximate size of 1 mm) on the inoculated leaves at 4–6 dpi (Fig. 3I, wounded) and elicited symptoms matching those observed in the field. The exception, FJ-122, showed larger black spots (4–7 mm) as a result of wounding. Control leaves inoculated with water developed no disease symptoms. If not wounded, four isolates (group I: FJ-85, FJ-119; group II: FJ-1, FJ-110) caused small black spots on inoculated leaves at 6–10 dpi (Fig. 3I, unwounded).
Representative symptoms induced by inoculation of conidial suspensions of Colletotrichum fructicola isolate FJ-1 onto attached leaves of sandy pear under wounded and unwounded conditions. Symptoms at 10–15 dpi on detached sandy pear leaves of cvs. Cuiguan (I), XueQing (II), and Huali No. 2 (III), which were induced by gently rubbing the leaves with conidia mixed with carborundum (wounded) or spraying the conidia on the leaves without carborundum (unwounded)
The incidence of disease symptoms for all of the isolates was higher for wounded compared to unwounded leaves (P < 0.001) (Fig. 3I), with average differences of 618 versus 126 for FJ-1 (P = 0.029), 308 versus 84 for FJ-85, 94 versus 30 for FJ-110, 308 versus 63 for FJ-116, and 106 versus 0 for FJ-119. The incidence corresponded to disease indices of 65.0, 80.0, 50.0, 100 and 46.7 % under wounded conditions and 40.0, 46.7, 33.3, 50.0 and 0.0 % under unwounded conditions for these isolates, respectively. Control leaves inoculated with water showed no disease symptoms.
Other sandy pear cultivars, including Xueqing and Huali No. 2, were also inoculated with FJ-1 conidia on attached leaves under wounded and unwounded conditions, respectively, and cv. Cuiguan was inoculated in parallel for comparison. The same black spot symptoms were observed on all of the inoculated leaves, with higher numbers of black spots (P = 0.009) for wounded than unwounded leaves, with average amounts of 94 vs. 24 for cv. Xueqing and 293 vs. 127 for cv. Huali No. 2, respectively (Fig. 3II-III).
Re-isolated colonies showed similar morphologies and ITS sequences as the colonies that were used for inoculation (data not shown).
The four Phomopsis sp. isolates were also selected and subjected to pathogenic analysis by inoculating them with conidium suspensions on attached sandy pear leaves (cv. Cuiguan), but no black spot symptoms had developed until 7 dpi.
Molecular characterization and phylogenetic analysis
For further molecular characterization and determination of taxon status of the isolates, the ribosomal internal transcribed spacer (ITS, including a partial 18S ribosomal RNA gene, internal transcribed spacer 1, 5.8S ribosomal RNA gene, internal transcribed spacer 2, and partial 28S ribosomal RNA gene) of the 124 isolates were amplified, and the target bands of 542 to 563 bp in length were sequenced. A sequence alignment showed that 117 isolates had the same ITS sequence, while the remaining seven isolates (group I: FJ-60, FJ-71, FJ-74-2, FJ-79 and FJ-109; group II: FJ-76; group V: FJ-42) shared similarities of 94.1to 99.8 % among themselves. BLASTn searches showed high sequence identities of more than 96.0 % (e = 0.0) among the sequences of the species belonging to the C. gloeosporioides complex. Phylogenetic analysis of the ITS sequences from all of the species of C. gloeosporioides, using the UPGMA method, revealed that the 117 isolates clustered together with C. aeschynomenes and C. fructicola isolates, while the other isolates clustered together with other species, including C. siamense and C. fructicola (FJ-42, FJ-109, FJ-60, and FJ-71), G. cingulata and C. kahawae (FJ-76), and C. kahawae and C. boninense (FJ-79 and FJ-74-2) (Fig. S1A). When phylogenetic analysis was performed with the ML method, all of the isolates clustered together with C. aeschynomenes and C. fructicola, within which FJ-74-2, FJ-76 and FJ-79 clustered together far from the other isolates, with FJ-74-2 at the most divergent position (Fig. S1B). Therefore, phylogenetic analysis based on the ITS sequences showed limited species distinctions for these isolates. Twenty-five representative isolates, including 22 isolates showing identical ITS sequences, isolates FJ-42 and FJ-109 in a separate branch, and isolate FJ-74-2 at the most divergent site, were further subjected to sequencing analysis in their gene regions, including those encoding glyceraldehyde-3-phosphate dehydrogenase (GAPDH), calmodulin (Smit et al. 1996), actin (ACT), chitin synthase (CHS-1), β-tubulin 2 (TUB2), and glutamine synthetase (GS). The CAL and GS regions were not amplified for fourteen and nine isolates, respectively, and only one other gene region failed to be amplified for four isolates. Therefore, gene-region sequences were successfully obtained for most isolates, and only eighteen isolates were further subjected to phylogenetic analysis by the UPGMA method based on concatenated sequences, which included ITS, ACT, TUB, CHS-1 and GAPDH. All isolates clustered together with the representative C. fructicola isolate, except for FJ-74-2, which formed a separated branch between C. salsolae and C. gloeosporioides (Fig. 4a). When the phylogenetic analysis was performed with the ML method, all of the isolates were clustered together with C. fructicola (data not shown). Considering that seven isolates were not involved in the multilocus phylogenetic tree and that isolate FJ-74-2 remained in an indefinite taxon position, the taxon status of the 25 isolates was re-assessed by sequencing and phylogenetic analysis of the Apn2/MAT (ApMat) loci. Unexpectedly, all of the isolates showed identical sequences in this gene region. Phylogenetic analysis was performed based on the sequences of this gene region from 20 species of the C. gloeosporioides complex (the other four species were not included because their ApMat loci were not reported). The 25 isolates clearly clustered together with the representative isolate of C. fructicola (Fig. 4b).
Phylogenetic analysis of Colletotrichum fructicola isolates from this study and other species belonging to the C. gloeosporioides species complex (type strains are marked with *) using the MEGA 5.2.2 software and the UPGMA method with 1000 bootstrap replicates and other default parameters. a and b Phylogenetic trees based on the nucleotide sequences of concatenated regions ITS, ACT, TUB, CHS-1, and GAPDH or on the ApMat region, respectively. The isolates of C. boninense and C. hippeastri are designated as outgroups. Bootstrap support values higher than 60 % are shown at the nodes. The isolates obtained in this study are indicated by a triangle (Black up-pointing triangle), and the isolates showing identical multilocus or ApMat sequences are represented by isolate FJ-1 and indicated as “FJ-1 and other isolates”
According to the phylogenetic analysis of the multilocus, ApMat sequences and analysis of morphological features, the obtained 124 Colletotrichum isolates in this study were conclusively identified as C. fructicola.
Discussion
To identify a new black spot disease with sudden outbreaks in sandy pear in the south of China, a wide isolation of fungi was carried out from symptomatic leaves from the Fujian province, which was severely affected by the disease. Based on the obtained fungal isolates, Koch’s postulates were fulfilled by inoculating conidium suspensions of the representative isolates on attached and detached leaves of sandy pear, and successfully verified that C. fructicola as the causal agent of black spot disease. The obtained 124 Colletotrichum isolates were separated into five groups depending on their macroscopic features, which were mainly reflected in the color, amount and distribution of the pigments (Fig. 2). The C. fructicola species had been reported to be highly divergent (see Fig. 23E Weir et al. 2012; Jiang et al. 2014), a characteristic that was also observed in isolates belonging to groups I and II after sub-culturing or between repeats of the same culture. The divergence was mainly notified as fan-shaped regions formed with variable areas and shapes, diverse amounts and distributions of pigments, and numbers of concentric rings (data not shown). Further characterization of these isolates revealed that these characteristic types had no obvious correlation with virulence or the growth rate, while a distinct correlation existed with their microscopic characters, including sporulation, sizes and amount of conidia, and presence of the perfect stage. In comparison with the colony appearances of C. fructicola specimens that had been described previously (Weir et al. 2012), it was not found that the isolates from groups I to IV matched any known specimens, likely due in part to the fact that most of the latter produced remarkable pinkish pigments (Fig. 2). However, the group V isolate looked similar to specimen ICMP 18646, a Tetragastris panamensis leaf endophyte isolated from Panama (Weir et al. 2012). Furthermore, the conidium and ascospore sizes of the isolates were obviously different from those of the specimens and isolates previously described (See Fig. 23E in Weir et al. 2012) and the isolates from cultivated citrus in China (Huang et al. 2013). In addition, the growth rates of all of the groups are obviously higher (10.0–13.0 mm/d) than those (7 mm/d) of the C. fructicola isolates from cultivated citrus in China (Huang et al. 2013). These results show that the obtained isolates in this study are possibly different strains of C. fructicola.
In addition to C. fructicola, nine other fungal species were isolated in a much lower incidence. Considering the characteristic symptoms that may develop by each fungus, Phomopsis sp. was considered the most likely to cause black spots on sandy pear leaves. Therefore, the Phomopsis sp. isolates were also subjected to pathogenic analysis as was done with C. fructicola; however, no black spot symptoms were found after inoculation. In addition, no other Colletotrichum species were isolated, suggesting that C. fructicola represents the sole etiological agent responsible for black spot disease. The quantity of black spots elicited after inoculation was lower under unwounded than wounded conditions. This result is in agreement with the initial hypothesis that tiny sand particles blown along with the conidia may enforce the infection and result in more black spots on the leaves.
The Colletotrichum gloeosporioides species complex represents a large group of plant pathogens (Weir et al. 2012) that have led to great economic losses in many different crops (Sutton 1992; Phoulivong 2011; Liu et al. 2014). The limit of the C. gloeosporioides species complex is defined genetically and is based on a strongly supported clade within the Colletotrichum ITS gene tree (Weir et al. 2012). Based on multi-gene phylogenies, 22 species plus one subspecies have been accepted within the C. gloeosporioides complex. Of these species, the sets that are genetically closest to C. musae or C. kahawae were assigned as the Musae clade or the Kahawae clade, respectively (Cannon et al. 2008; Phoulivong et al. 2010; Weir et al. 2012; Huang et al. 2013). In this study, phylogenetic analysis based on the ITS sequences was too insufficient to resolve the species delimitations of the obtained 124 isolates, similar to many C. gloeosporioide complex species that cannot be reliably distinguished using ITS alone (Weir et al. 2012). Therefore, 25 isolates were chosen for further phylogenetic analysis based on the multilocus regions ACT, TUB2, CAL, CAL, CHS-1, GAPDH, ITS; however, some regions were unsuccessfully amplified in certain isolates, limiting the means of identification for these isolates. Finally, the taxon status was re-assessed by sequencing and performing phylogenetic analysis of the ApMAT gene region, which was 100 % identical for all 25 isolates. This result provided molecular proof, in addition to the identifying morphological features, that these isolates are C. fructicola and belong to the C. gloeosporioide complex in the Musae clade. The sum of the results provides an insightful molecular characterization of these C. fructicola isolates and indicates that the ApMAT was most informative for C. fructicola species identification. It is worth noting that, despite remarkable divergence in the macroscopic and microscopic characteristics, we did not detect diversity among the multiple loci ITS, ACT, TUB, CHS-1, GAPDH and ApMAT regions among the 124 isolates; these regions were 100 % identical for the representative isolates. This suggests that the isolates responsible for leaf black spot disease should have a common origin.
Recently, C. fructicola reportedly caused black spots on crisp pear (P. bretschneideri cv. Suli) fruits, leading to severe bitter rot in the Anhui province of China (Li et al. 2013; Jiang et al. 2014). Isolates from diseased fruits in that study showed higher biological and molecular variability compared to the ones obtained in the present work. First, the isolates infecting on different pear species and organs, reflected a potentially different host or organ adaptation. A similar situation was illustrated by G. cingulata, in which isolates from apple fruits (cv. Gala) were only capable of infecting fruit tissue and not the leaves, while isolates from apple leaves could infect both fruits and leaves (González et al. 2006). Second, based on their ITS sequences, the C. fructicola isolates that caused black spots on crisp pear showed a branch in the phylogenetic tree distinct from the isolates obtained in this study (Fig. S2). Colletotrichum fructicola isolates were also reported to infect leaves of P. pyrifolia in Japan (Weir et al. 2012) and other plants, including Coffea arabica in Thailand, Limonium sinuatum in Israel, Malus domestica in Brazil, M. domestica and Fragaria × ananassa in the USA, Persea americana in Australia, Ficus edulis in Germany, Dioscorea in Nigeria, Theobroma and Tetragastris in Panama (Weir et al. 2012), and Citrus reticulate (Huang et al. 2013), Camellia sinensis (Liu et al. 2014) and Vitis vinifera (Peng et al. 2013) in China. Phylogenetic analysis of the ITS sequences of these isolates revealed no obvious divergence, nor did their host plants or geographic locations (Fig. S2).
To our knowledge, this is the first report of C. fructicola causing leaf black spot in pear. The combined morphological and molecular characterization of the isolates showed that C. fructicola was the causal agent of leaf anthracnose of sandy pear in the Fujian province of China. From other countries, it is known that C. gloeosporioides and C. acutatum may cause black spot of sandy pear (Morita et al. 1994; Tashiro et al. 2001, 2012; Fukaya 2004). Clarification of the etiology and characterization of the C. fructicola isolates provides useful information for understanding the leaf black spot disease of pear and will help to improve management strategies and to control this economically important disease, e.g., in selecting an effective biocide against the disease.
References
Cai, L., Hyde, K. D., Taylor, P. W. J., Weir, B., Waller, J., Abang, M. M., Zhang, J. Z., Yang, Y. L., Phoulivong, S., & Liu, Z. Y. (2009). A polyphasic approach for studying Colletotrichum. Fungal Diversity, 39, 183–204.
Cannon, P. F., Buddie, A. G., & Bridge, P. D. (2008). The typification of Colletotrichum gloeosporioides. Mycotaxon, 104, 189–204.
Carbone, I., & Kohn, L. M. (1999). A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia, 91, 553–556.
Chen, Y. T., Liu, X. M., Chen, X. M., Chen, T., Wang, W., Zhang, C. H., Bao, J. P., & Huang, X. Z. (2011). Preliminary investigation and analysis of abnormal early defoliation of pear in Fujian. 29, 43–45.
Freeman, S., Katan, T., & Shabi, E. (1996). Characterization of Colletotrichum gloeosporioides isolates from avocado and almond fruits with molecular and pathogenicity tests. Applied and Environmental Microbiology, 62, 1014–1020.
Fukaya, M. (2004). First report of Japanese pear anthracnose disease caused by Colletotrichum acutatum and its chemical control (in Japanese). Japanese Journal of Phytopathology, 70, 184–189.
Glass, N. L., & Donaldson, G. C. (1995). Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Applied and Environmental Microbiology, 61, 1323–1330.
González, E., Sutton, T. B., & Correll, J. C. (2006). Clarification of the etiology of Glomerella leaf spot and bitter rot of apple caused by Colletotrichum spp. based on morphology and genetic, molecular, and pathogenicity tests. Phytopathology, 96, 982–992.
Huang, F., Chen, G. Q., Hou, X., Fu, Y. S., Cai, L., Hyde, K. D., & Li, H. Y. (2013). Colletotrichum species associated with cultivated citrus in China. Fungal Diversity, 61, 61–74.
Huang, X. Z., Chen, Y. T., Lei, Y., Cai, S. H., & Chen, X. M. (2010). Causes and control strategies of a large number of early falling leaves of pear in Fujian (in Chinese). Chinese Agricultural Science Bulletin, 26, 91–95.
Jiang, J. J., Zhai, H. Y., Li, H. N., Wang, Z. H., Chen, Y. S., Hong, N., Wang, G. P., Gilbert, N. C., & Xu, W. X. (2014). Identification and characterization of Colletotrichum fructicola causing black spots on young fruits related to bitter rot of pear (Pyrus bretschneideri Rehd.) in China. Crop Protection, 58, 41–48.
Johnston, P. R., & Jones, D. (1997). Relationships among Colletotrichum isolates from fruit-rots assessed using rDNA sequences. Mycologia, 89, 420–430.
Larkin, M. A., Blackshields, G., Brown, N. P., Chenna, R., McGettigan, P. A., Mcwilliam, H., Valentin, F., Wallace, I. M., Wilm, A., Lopez, R., Thompson, J. D., Gibson, T. J., & Higgins, D. G. (2007). Clustal W and Clustal X version 2.0. Bioinformatics, 23, 2947–2948.
Li, H. N., Jiang, J. J., Hong, N., Wang, G. P., & Xu, W. X. (2013). First report of Colletotrichum fructicola causing bitter rot of pear (Pyrus bretschneideri) in China. Plant Disease, 97, 1000.
Liu, W., Ye, N. X., Chen, Y. S., Lian, L. L., Jin, S., Lai, J. D., & Xie, Y. H. (2014). Identification and phylogenetic analysis of anthracnose pathogen Colletotrichum fructicola isolated from Camellia sinensis (in Chinese). Journal of Tea Science, 34, 95–104.
Mckinney, H. H. (1923). Influence of soil, temperature and moisture on infection of wheat seedlings by Helminthosporium sativum. Journal of Agricultural Research, 26, 195–217.
Morita, Y., Yano, K., Matsumoto, K., Kotani, S., & Kurata, M. (1994). Occurrence and control of anthracnose on leaves of Japanese pear (in Japanese). Bulletin of the Kochi Agricultural Research Center, 3, 1–10.
O’donnell, K., Nirenberg, H. I., Aoki, T., & Cigelnik, E. (2000). A multigene phylogeny of the Gibberella fujikuroi species complex: detection of additional phylogenetically distinct species. Mycoscience, 41, 61–78.
Peng, L. J., Sun, T., Yang, Y. L., Cai, L., Hydef, K. D., Bahkali, A. H., & Liu, Z. Y. (2013). Colletotrichum species on grape in Guizhou and Yunnan provinces, China. Mycoscience, 54, 29–41.
Phoulivong, S. (2011). Colletotrichum, naming, control, resistance, biocontrol of weeds and current challenges. Current Research in Environmental & Applied Mycology, 1, 53–73.
Phoulivong, S., Cai, L., Parinn, N., Chen, H., Abd-Elsalam, K. A., Chukeatirote, E., & Hyde, K. D. (2010). A new species of Colletotrichum from Cordyline fruticosa and Eugenia javanica causing anthracnose disease. Mycotaxon, 114, 247–257.
Prihastuti, H., Cai, L., Chen, H., McKenzie, E. H. C., & Hyde, K. D. (2009). Characterisation of Colletotrichum species associated with coffee berries in northern Thailand. Fungal Diversity, 39, 89–109.
Silva, D. N., Talhinas, P., Várzea, V., Cai, L., Paulo, O. S., & Batista, D. (2012). Application of the Apn2/MAT locus to improve the systematics of the Colletotrichum gloeosporioides complex: an example from coffee (Coffea spp.) hosts. Mycologia, 104, 396–409.
Simmonds, H. J. (1968). Type specimens of Colletotrichum var minor and Colletotrichum acutatum. Queensland Journal of Agricultural & Animal Sciences, 25, 17–37.
Smit, W. A., Viljoen, C. D., Wingfield, B. D., Wingfield, M. J., & Calitz, F. J. (1996). A new canker disease of apple, pear, and plum rootstocks caused by Diaporthe ambigua in South Africa. Plant Disease, 80, 1331–1335.
Stephenson, S. A., Green, J. R., Manners, J. M., & Maclean, D. J. (1997). Cloning and characterisation of glutamine synthetase from Colletotrichum gloeosporioides and demonstration of elevated expression during pathogenesis on Stylosanthes guianensis. Current Genetics, 31, 447–454.
Su, Y. Y., Noireung, P., Liu, F., Hyde, K. D., Moslem, M. A., Bahkali, A. H., Abdelsalam, K. A., & Cai, L. (2011). Epitypification of Colletotrichum musae, the causative agent of banana anthracnose Mycoscience, 1–7.
Sutton, B. C. (1992). The genus Glomerella and its anamorph Colletotrichum. Wallingford: CAB international.
Sutton, T. B., & Sanhueza, R. M. (1998). Necrotic leaf blotch of Golden Delicious-Glomerella leaf spot: a resolution of common names. Plant Disease, 82, 267–268.
Tamura, T., Peterson, D., Peterson, N., Stecher, G., Nei, M., & Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony method. Molecular Biology and Evolution, 28, 2731–2739.
Tashiro, N., Manabe, K., & Ide, Y. J. (2012). Emergence and frequency of highly benzimidazole-resistant Colletotrichum gloeosporioides, pathogen of Japanese pear anthracnose, after discontinued use of benzimidazole. Journal of General Plant Pathology, 78, 221–226.
Tashiro, N., Idem, Y., & Etho, T. (2001). Occurrence of benzimidazole-tolerant isolate of Colletotrichum gloeosporioides, a causal fungus of anthracnose of Japanese pear and its effective fungicides (in Japanese). Kyushu Agricultural Research, 63, 81.
Templeton, M. D., Rikkerink, E. H. A., Solon, S. L., & Crowhurst, R. N. (1992). Cloning and molecular characterization of the glyceraldehyde-3-phosphate dehydrogenase encoding gene and cDNA from the plant pathogenic fungus Glomerella cingulata. Gene, 122, 225–230.
Weir, B. S., Johnston, P. R., & Damm, U. (2012). The Colletotrichum gloeosporioides species complex. Studies in Mycology, 73, 115–180.
White, T. J., Bruns, T., Lee, S., & Taylor, J. W. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. New York: Academic Press.
Yaegashi, H., Kanematsu, S., & Ito, T. (2012). Molecular characterization of a new hypovirus infecting a phytopathogenic fungus, Valsa ceratosperma. Virus Research, 165, 143–150.
Acknowledgments
This study was financially supported by the earmarked fund for Pear Modern Agro-industry Technology Research System (CARS-29-10) and the Chinese Ministry of Agriculture, Industry Technology Research Project (200903004–05). The authors would like to thank Professor Yangdou Wei University of Saskatchewan, Canada, and Chaoxi Luo, Huazhong Agricultural University, China, for critical revisions of the manuscript.
Compliance with ethical standards
The authors declare no conflict of interest. This research work does not include any animal studies.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Phylogenetic analysis of Colletotrichum fructicola isolates and other species belonging to the C. gloeosporioides species complex based on the ITS sequences using the MEGA 5.2.2 software with 1,000 bootstrap replicates and other default parameters. Type strains are marked with symbol “*”. A and B, Phylogenetic tree constructed using the UPGMA and ML methods, respectively. The isolates of C. boninense and C. hippeastri are designated as outgroups. Bootstrap support values of more than 60 % are shown at the nodes. The isolates obtained in this study are indicated by a triangle (▲). Of these isolates, the 117 isolates showing identical ITS sequences are represented by FJ-1 and are indicated as “FJ-1 and other isolates”. (JPEG 642 kb)
Fig. S2
Phylogenetic tree based on the ITS sequences of the 25 Colletotrichum fructicola isolates obtained in this study and those of other reported isolates. The isolates of C. boninense, C. siamense, and C. gloeosporioides are shown as outgroups. Bootstrap support values greater than 60 % are shown at the nodes. The isolates obtained in this study are indicated by a triangle (▲). Out of these isolates, the 20 isolates showing identical ITS sequences are represented by FJ-1 and are indicated as “FJ-1 and other isolates.” The accession number, host and original country of each isolate are successively presented in the phylogenetic position of each C. fructicola isolate. An isolate obtained from Pyrus bretschneideri fruit in China is indicated by a blank triangle (Δ). (JPEG 655 kb)
Rights and permissions
About this article
Cite this article
Zhang, P.F., Zhai, L.F., Zhang, X.K. et al. Characterization of Colletotrichum fructicola, a new causal agent of leaf black spot disease of sandy pear (Pyrus pyrifolia). Eur J Plant Pathol 143, 651–662 (2015). https://doi.org/10.1007/s10658-015-0715-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-015-0715-7