Abstract
Background
Ion channels and transporters are potential markers and therapeutic targets for several cancers. However, their expression during hepatocellular carcinoma (HCC) development remains unclear.
Aim
To investigate the mRNA expression of Na+, K+ and Ca2+ channels and ABC transporters during rat HCC development, as well as Abcc3 protein in human liver biopsies.
Methods
Wistar rats were treated with diethylnitrosamine (DEN) and developed both cirrhosis (12 weeks of treatment) and either pre-neoplastic lesions (16 weeks of treatment) or multinodular HCC (16 weeks of treatment plus 2 weeks DEN-free). The mRNA expression of 12 ion channels and two ABC transporters was studied using real-time RT-PCR. Tumor-containing or tumor-free liver sections were isolated by laser-capture microdissection. Abcc3 protein expression was studied by immunohistochemistry in healthy, cirrhotic and HCC human biopsies.
Results
We observed expression changes in seven genes. Kcna3, Kcnn4, Kcnrg and Kcnj11 potassium channel mRNA expression reached peak values at the end of DEN treatment, while Scn2a1 sodium channel, Trpc6 calcium channel and Abcc3 transporter mRNA expression reached their highest levels in the presence of HCC (18 weeks). Whereas Kcnn4 and Scn2a1 channel expression was similar in non-tumor and tumor tissue, the Abcc3 transporter and Kcna3 potassium channels were preferentially overexpressed in the tumor sections. We observed differential Abcc3 protein subcellular localization and expression in human samples.
Conclusions
The ion channel/transporter expression profile observed suggests that these genes are potential early markers or therapeutic targets of HCC. The differential localization of Abcc3 may be useful in the diagnosis of cirrhosis and HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) has a very high mortality-to-incidence ratio and is a major cause of cancer-related death worldwide [1]. Because HCC has a very poor prognosis and is often diagnosed at an advanced stage, new early markers and therapeutic targets to prevent, diagnose or treat this disease are urgently needed. Most cases of HCC develop within a background of chronic liver damage, including cirrhosis or chronic hepatitis [2]. Taken together, viral and toxic agents (alcohol and aflatoxin B1) account for approximately 80 % of all HCC cases in humans [2]. Hepatocarcinogenesis is a slow and multistep process involving different genetic alterations that ultimately lead to the malignant transformation of hepatocytes [2–4]. Diethylnitrosamine (DEN)-induced hepatocarcinogenesis is one of the most commonly used in vivo HCC models because the histological and genetic signatures observed in this model resemble those of human HCC, including the progression of cirrhosis to HCC [5, 6].
A great limitation in the treatment of HCC is the development of drug resistance. Cancer cells might be either insensitive to drug treatment from the beginning of therapy (intrinsic drug resistance) or become resistant after chemotherapy (acquired drug resistance), leaving the patient with no effective therapeutic options and a very poor prognosis. Therefore, in addition to the HCC markers reported (for instance HepPar1, alpha-fetoprotein, GPC3 and CK19), it is very important to study the expression of the transporters involved in multidrug resistance. This might serve to identify biomarkers predicting the potential response to chemotherapy and lead to more efficient personalized medicine in HCC [7–9].
Several ion channels, including voltage-gated and transient receptor potential channels (TRP) as well as ATP-binding cassette (ABC) transporters, have been suggested to play relevant roles in cancer biology and may be promising novel markers and targets for clinical intervention [10–15]. The expression of several ion channels is regulated by cancer-associated factors, including hormones, chemical carcinogens or human papilloma virus oncogenes [16–18]. Accordingly, the potassium channel Kv10.1 has been found to be overexpressed in pre-malignant cervical lesions [19] and in diverticulitis, which has the potential to develop into colonic cancer [17]. Notably, the overexpression of several ion channels has been observed at different times and tissue locations in a colonic cancer model induced by dimethylhydrazine and N-methyl-N-nitrosourea treatment in mice [17]. Based on the channel expression reported for other cancers, we selected 12 ion channels to evaluate in this study. In addition, unpublished results from our group revealed that Abcc3 expression is increased in an HCC model. Thus, we also studied the expression of two transporters, including Abcc3.
Nevertheless, despite the rapidly increasing number of studies in the ion channel field and the poor prognosis of HCC, the expression of ion channels and transporters during HCC development remains uncharacterized. Therefore, we here investigated the expression of several ion channels and transporters during HCC development in whole liver tissue in rats, as well as in tumor-containing and tumor-free tissue isolated by laser capture microdissection [20–22]. We also studied Abcc3 protein expression in human liver samples. Our results suggest that some of these genes may be potential early markers and therapeutic targets of HCC.
Materials and Methods
Animals and Treatments
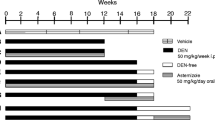
Male Wistar adult rats (210–250 g, 7 weeks old) were used. All animal care and experimentation conformed to the Institutional Animal Care and Use Committee Guidelines. The experimental protocol was previously revised and approved by the local Committee for Animal Experimentation. The animals were fed Purina rat chow and water ad libitum. HCC was induced with DEN (diethylnitrosamine, Sigma Chemical Co., St Louis, MO, USA) following the procedure described by Schiffer et al. [6]. The rats were divided into control and DEN-treated groups. DEN was i.p. administered (50 mg/kg) once a week. The first treatment group received DEN for 12 weeks; at this time, the animals develop liver cirrhosis. The second treated group received DEN for 16 weeks; at this time, the animals develop liver nodular lesions. Finally, the third treated group received DEN for 16 weeks and was left 2 additional weeks without DEN (DEN-free); at this time, the animals develop multinodular HCC. The body weight was recorded every week for each animal. The rats were killed by decapitation at the end of the corresponding treatments. The livers were frozen in 2-methyl butane (St Louis, MO, USA) with liquid nitrogen and stored at −80 °C.
Real Time-RT-PCR
Frozen liver slices were homogenized in RLT buffer (RNeasy kit, Qiagen, Germany) supplemented with β-mercaptoethanol, as detailed in the manufacturer’s instructions. RNA was extracted using Qiagen RNeasy Mini Kit spin columns (Qiagen, Germany) following the manufacturer’s instructions. The RNA concentration was determined spectrophotometrically at 260 and 280 nm with a NanoDrop ND1000 spectrophotometer (Thermo Fisher Scientific, Wilmington, DE, USA). The RNA integrity was confirmed using 1.5 % agarose gel electrophoresis. The total RNA (750 ng) was reverse transcribed with the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) following the manufacturer’s instructions. The concentration and integrity of the RNA obtained from the tissue sections isolated with the Laser Capture Microdissection technique were analyzed with the 2100 Bioanalyzer (Agilent Technologies, Inc., Santa Clara, CA, USA). Real-time quantitative PCR was performed using TaqMan Gene Expression Assays (Applied Biosystems, Foster City, CA, USA). The following probes were used: (ion channels) KCNA3 (Kv1.3, Rn00570552_s1), KCNH2 (Kv11.1, Rn00588515_m1), KCNN4 (KCa3.1, Rn00576373_m1), KCNJ11 (Kir6.2, Rn01764077_s1), KCNC4 (Kv3.4, Rn01748431_m1), KCNRG (Rn04244668_m1), CACNA1H (Cav3.2, Rn01460348_m1), Scn2a1 (Nav1.2, Rn00561862_m1), Scn9a (Nav1.7, Rn00591020_m1), TRPV6 (Rn00586673_m1), TRPC6 (Rn00677559_m1), and TRPM8 (Rn00592665_m1); (ABC transporters) MRP3 (Abcc3, Rn01452854_m1) and MRP8 (Abcc8, Rn00564778_m1); Gstp (Rn00561378_gH); these probes were labeled with a FAM dye reporter on the 5′ end (Applied Biosystems, Foster City, CA, USA). The cycling conditions were as follows: 50 °C for 5 min, 95 °C for 10 min, followed by 45 cycles of 95 °C for 15 s and 60 °C for 1 min. All PCR reactions were performed on the 7900HT FAST system (Applied Biosystems, Foster City, CA, USA). The housekeeping genes used for normalization were glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (Rn99999916_s1) and ribosomal 18 s (Rn03928990); these probes were labeled with a VIC dye reporter on the 5′ end (Applied Biosystems, Foster City, CA, USA). Data analysis was performed using the 2−ΔΔCt method.
Follow-Up of Tumor Development and Laser-Capture Microdissection
Tumor development was evaluated by counting the number of macroscopic pre-neoplastic nodules and histological examination, as well as by studying γ-glutamyl-transferase (GGT, a commonly used liver tumor marker) activity in the histological sections and glutathione S-transferase pi 1 (Gstp1) gene expression. Gstp1 gene expression was investigated by real-time RT-PCR as described above. Liver tissues embedded in paraffin were sliced into 3-µm sections. The slides were stained with hematoxylin and eosin (H&E) for routine histologic examination. GGT activity was detected by staining methods as previously described [23]. Histological images of GGT activity were analyzed with ImageJ software (v. 1.43, National Institutes of Health, USA). Laser capture microdissection (LCM) was used to isolate either tumor-containing (GGT-positive) or tumor-free (GGT-negative) liver sections from the heterogeneous tissue using a low-power infrared laser (Veritas™ Microdissection Instrument, Arcturus Bioscience, Inc., Mountain View, CA) as previously described [22]. The tumor-containing or tumor-free areas removed by LCM were placed directly into a microcentrifuge tube for RNA extraction.
Human Liver Biopsies
We obtained 30 liver biopsies from the tumor bank of the Mexico City General Hospital “Dr. Eduardo Liceaga” following local ethical considerations, including the patients' written informed consent. This is a retrospective study; thus, following the jornal requirements, we state that formal consent is not required for this type of study. The biopsies were histologically diagnosed as normal liver (n = 10), cirrhosis (n = 10) or HCC (n = 10). It is worth mentioning that the normal livers obtained from the tumor bank were from patients whose deaths were not related to any liver disease.
Immunohistochemistry
Immunostaining was performed using formalin-fixed, paraffin-embedded tissue sections of surgically resected liver specimens from 30 patients. Serial sections (4 µm) were mounted on charged glass slides and deparaffinized using xylene and a decreasing series of ethanol. After washing with TBS/Tween (pH 7.4), antigen retrieval was performed by microwaving in 10 mmol/l citrate buffer (pH = 6.0) for CK19 and Tris/EDTA buffer (pH = 9.0) for GPC3 and Abcc3. The sections were incubated with protein block solution (DakoCytomation). Next, the sections were incubated for either 1 h with the corresponding primary antibodies against CK19 (1:250, rabbit monoclonal, Bio SB) and GPC3 (1:250, mouse monoclonal, Bio SB) or for 2 h for Abcc3 (1:20, mouse monoclonal, Abcam Inc.). Afterwards, the sections were incubated with secondary antibodies conjugated to peroxidase-labeled polymer (EnVision system, DakoCytomation) for GPC3 and Abcc3 and with SignalStain Boost IHC Detection Reagent (HRP Rabbit, Cell Signaling) for CK19. The specific staining reaction was completed by incubating the slides in the presence of 3,3′-diaminobenzidine (DAB) in buffer reaction solution (Dako) and observed as a brown staining. The sections were counterstained with hematoxylin. The negative controls were carried out by substitution of the primary antibodies with non-immunized serum, resulting in no immunostaining signal detection. Immunohistochemical staining was interpreted by a specialized pathologist.
Statistical Analysis
One-way ANOVA followed by Tukey’s test was performed using GraphPad Prism (version 5.00 for Windows, GraphPad Software, San Diego, CA, USA). All results are shown as the mean ± SEM. Differences were considered statistically significant when p values were ≤0.05.
Results
HCC Development Evaluation
Figure 1 shows the major histological changes during HCC development. The livers from untreated control animals showed completely normal appearances and histology (Fig. 1A), while the livers from animals treated for 12 weeks with DEN were clearly fibrotic (Fig. 1B). The presence of neoplastic nodules became evident in the animals treated for 16 weeks with the carcinogen (Fig. 1C), and the number of nodules increased in both number and size in the animals also treated for 16 weeks with DEN but allowed to progress for 2 more weeks in the absence of DEN (Fig. 1D). Eight animals were studied in each group. While no tumors were detected in the control group, the percentage of animals displaying pre-neoplastic nodules increased to 25 and 87.5 % in the groups treated with DEN for 12 and 16 weeks, respectively. The group treated with DEN for 16 weeks and left for 2 more weeks with no DEN showed neoplastic nodules in 100 % of the animals. The relative liver weight was 3 % in the control group and increased to 4.3 and 4.5 % in the groups treated with DEN for 16 and 16 + 2 weeks DEN-free, respectively. Tumor development was also followed studying two known liver tumor markers, namely GGT activity [24] and Gstp1 gene expression [11]. GGT activity was detected in the liver sections by the appearance of round red areas. GGT activity clearly increased with the progress of HCC (Fig. 2A, B). Although GGT is mainly used as a tumor marker, it is also found in small amounts in the bile ducts of normal liver (not visible in this figure). Another liver tumor marker commonly used in rat experimental hepatocarcinogenesis is Gstp1 gene expression, which noticeably increased during HCC development (Fig. 2C).
Liver morphology and histology during hepatocellular carcinoma development. A Untreated animals displayed normal liver tissue, characterized by liver parenchyma organized into plates, with homogeneous cell size and uninucleated cells. B Fibrosis and loss of the normal architecture of the liver as well as an increasing number of binucleated hepatocytes was observed in the rats treated with diethylnitrosamine (DEN) for 12 weeks (black arrows, right panel). C Dysplastic nodules (black arrows, left panel) and disorganized liver histology showing large and abundant cytoplasm as well as an irregular nucleus (black arrow, right panel) were observed in rats treated for 16 weeks with DEN. D Large and abundant nodules (black arrows, left panel) were displayed in the livers from animals treated for 16 weeks with DEN and left DEN-free for 2 weeks as well as neoplastic cells with a large nucleus (black arrows, right panel). Scale bar = 400 μm
Characterization of hepatocellular carcinoma (HCC) with tumor markers. A Histologic gamma-glutamyl-transferase (GGT) activity was determined as described in “Methods” and visualized as red-colored, circle-like areas (black arrows), indicating the presence of preneoplastic and tumor lesions. a Control group, b group treated with diethylnitrosamine (DEN) for 12 weeks, c group treated with DEN for 16 weeks and d group treated with DEN for 16 weeks plus 2 weeks DEN-free. B Percentage of GGT-positive areas in the livers during HCC development (groups a–d). Mean ± SEM. *p < 0.005 versus group a; **p < 0.05 versus group b; ***p < 0.05 versus group c. C mRNA expression of Gstp1 determined by real-time RT-PCR clearly increased during HCC. Mean ± SEM. *p < 0.05 versus control group a
Differential Expression of Ion Channels and Transporters During HCC Development
Because of the role and gene expression changes of several ion channels and transporters in other types of cancer [10–15, 25, 26], we investigated the mRNA expression of 12 ion channels and two transporters by real-time RT-PCR. No mRNA expression changes during HCC development were observed for the ion channels Kcnh2, Kcnc4, Cacna1h, Scn9a, Trpv6 and Trpm8 and for the Abcc8 transporter. In contrast, statistically significant increased expression changes during HCC development were found for the rest of the genes studied (Abcc3, Trpc6, Scn2a1, Kcnn4, Kcna3, Kcnj11 and Kcnrg). Two different expression patterns were found (Fig. 3). In comparison with control untreated animals, mRNA expression of the Scn2a1 sodium channel, Trpc6 calcium channel and Abcc3 transporter displayed a gradual increase and reached the highest levels in the presence of HCC, that is, in the group treated for 16 weeks with DEN and left DEN-free for 2 weeks. The second expression pattern found was for the Kcna3, Kcnn4, Kcnrg (potassium channel-regulating protein) and Kcnj11 potassium channels, where the mRNA expression reached peak values at the end of DEN treatment (16 weeks) but returned to almost normal levels after 2 weeks in the absence of DEN, despite the presence of HCC. This analysis was performed by studying the whole liver tissue, where the combined presence of normal hepatocytes, fibrosis, preneoplastic lesions and HCC-type tumors can be found. We wondered whether some of these genes were expressed in either the tumor-containing (GGT-positive) or tumor-free (GGT-negative) tissue sections. Thus, we isolated the GGT-positive areas by laser-capture microdissection and studied the expression of several ion channels and transporters.
Differential mRNA expression of ion channels and transporters during hepatocellular carcinoma (HCC) development. Abcc3 transporter and Scn2a1 and Trpc6 ion channel expression increased with diethylnitrosamine (DEN) treatment and reached the highest levels in the group displaying clear HCC. Potassium channel expression reached the highest level in the group treated for 16 weeks with DEN, but almost normal values were detected in the presence of HCC. Mean ± SEM. *p < 0.05 group treated with DEN for 16 weeks plus 2 weeks DEN-free versus control group; **p < 0.05 group treated with DEN for 12 weeks versus group treated with DEN for 16 weeks. Eight animals were studied in each group
Gene Expression in Tumor-Containing and Tumor-Free Liver Sections
Laser-capture microdissection is a very useful tool to select specific cell populations. This technique allows the isolation of neoplastic cell regions (GGT-positive, red-color areas) from neoplastic cell-free areas (GGT-negative) [22]. Figure 4A shows the liver tissue sections before and after the isolation of tumor and tumor-free areas by laser-capture microdissection. Once isolated, we studied the mRNA expression of several ion channels and transporters in these different sections. There was substantial overlap in the data, and no statistically significant differences were found. Nevertheless, the Abcc3 transporter and Kcna3 channel were preferentially overexpressed in the tumor-containing (GGT-positive) sections (Fig. 4B). The expression of the Kcnn4 channel was more consistent in the GGT-positive and GGT-negative sections, while the Scn2a1 channel expression was more extensively expressed in the non-tumor regions.
Ion channel and transporter expression in tumor-containing and tumor-free liver sections. A Representative laser-capture microdissection from liver tissue. a Captured gamma-glutamyl-transferase (GGT)-negative (non-tumor) liver tissue; b captured GGT-positive (tumor) liver tissue. B mRNA expression studied by real-time RT-PCR in the laser-captured sections. Magnification = ×2, scale bar = 6 mm
Abcc3 and Tumor Marker Expression in Human Liver Biopsies
We studied the protein expression of the Abcc3 transporter, which displayed the most dramatic changes in mRNA expression during HCC development. Abcc3 protein expression was studied in 30 human liver biopsies from normal, cirrhotic or HCC tissue. The expression of the other already proposed markers, namely CK19 and GPC3, was also investigated (Fig. 5).
Expression of Abcc3, CK19 and GPC3 in human liver biopsies. Serial sections of the same tissue sample (normal liver, cirrhosis or HCC) are shown. Abcc3: A Normal liver showing granular pattern staining (Gr) and diffuse staining in the cytoplasm. B Cirrhotic tissue also showed diffuse cytoplasmic staining, but notably, it displayed plasma membrane (PM) immunoreactivity. C HCC showing strong and diffuse cytoplasmic staining; the arrows indicate the nuclei of the neoplastic cells (Nu) (insets show immunoreactivity for Abcc3 in a lower magnification). CK19: A Normal liver showed no signal in hepatocytes, but focal staining was observed in the bile duct of the periportal zone. B Cirrhotic tissue showed mainly focal expression in the cholangiocytes. C HCC showed no signal in this case, but focal staining was observed in 70 % of the cases. GPC3: Normal liver (A) and cirrhotic tissue (B) were negative, as expected. C HCC showed focal positive staining. The inset shows a complete view of the liver biopsy displaying immunonegativity (“−”) for some nodules and immunoreactivity (“+”) for another nodule in the same tissue. Original magnification, ×400; inset original magnification ×100 (scale bar = 100 μm)
Abcc3 protein expression (Fig. 5A) was found in 10/10 (100 %) of the normal liver cases, with faint and diffuse cytoplasmic staining, mainly in the periportal area. Abcc3 staining was detected in granular form. This result was expected because Abcc3 expression in normal human hepatocytes has been reported [27]. Liver cirrhosis biopsies also displayed Abcc3 expression in 10/10 (100 %) of the cases. Notably, in all of these cases, we observed Abcc3 expression at the plasma membrane of the hepatocytes. Abcc3 immunoreactivity was observed in 7/10 (70 %) of the HCC samples in the cytoplasm; this result was also expected because Abcc3 expression in human HCC has been reported [27]. The alanine aminotransferase (ALT) enzyme levels were far above the normal range, and the liver was significantly increased in size and weight (3 kg) in all of the Abcc3-positive HCC patients. HCC patients negative for Abcc3 expression displayed smaller liver size and weight (approximately 1 kg). Chronic history of alcoholism was reported for six out of the seven Abcc3-positive cases from HCC patients.
CK19 immunohistochemical staining (Fig. 5B) was negative in the normal liver tissue, as expected, but positive in 7/10 (70 %) of the cases of cirrhotic tissue. CK19 was found in the bile duct of the cirrhotic tissue but absent in the hepatocytes, as expected. Five out of ten (50 %) of the HCC cases were positive for CK19. In these five samples, staining was found to be cytoplasmic.
Finally, GPC3 immunostaining (Fig. 5C) was negative in the normal and cirrhotic tissues, as expected, but positive in 6/10 (60 %) of the HCC cases. The inset in Fig. 5C shows a complete view of the liver biopsy detecting heterogeneous GPC3 expression in HCC, that is, positive or negative staining in the nodules in the same biopsy. All three markers were identified in only one HCC case. Five out of ten HCC patients including this one were diagnosed with liver cirrhosis, multicenter HCC and lung metastasis. However, this was the only case showing Glisson capsule breakdown; therefore, it will be very interesting to evaluate more biopsies to determine the potential association of Abcc3-CK19-GPC3 coexpression with the Glisson capsule breakdown.
Discussion
HCC has one of the highest cancer mortality-to-incidence ratios [1]. Thus, new early markers are needed to treat the disease at potentially curable stages. In addition, new therapeutic markers are needed to treat the disease at stages in which the patients do not respond to the available therapies. Although several ion channels and transporters have been suggested as potential markers and therapeutic targets for different types of cancers [10–15, 25, 26, 28], the expression of ion channels and transporters during HCC development has not been studied. We observed the differential expression of ion channels and transporters during HCC development induced by DEN.
DEN is a representative chemical carcinogen with a mechanism of action of forming ethyl adducts at N- and O-atoms in DNA bases [29]. Despite this nonspecific mechanism of action, we observed the overexpression of only 7 out of the 14 genes studied. The mechanism of the regulation of these genes by DEN during HCC remains elusive. However, it has been shown that the specific overexpression of certain ion channels also occurs in a colonic cancer model induced by dimethylhydrazine and N-methyl-N-nitrosourea treatment in mice [17].
Ion channels and transporters have been suggested to play integral roles in cancer biology, including uncontrolled growth, decreased apoptosis, disorganized angiogenesis, aggressive migration, invasion and metastasis [10–15, 25, 26, 28]. We observed two different expression patterns of ion channels and transporters during HCC development. Potassium channel expression reached its highest level at the end of DEN treatment and returned to almost normal levels in the absence of DEN despite the presence of HCC. These channels likely serve as indicators of exposure to chemical carcinogens and early markers of the disease as well as potential targets to prevent HCC development. In contrast, the genes reaching higher expression in the presence of HCC might serve as targets to treat patients not responding to conventional treatments.
In particular, Kcnn4 (KCa3.1) might have an important role in diseases characterized by excessive cell proliferation [30]. The selective pharmacological inhibition or knockdown of Kcnn4 suppresses mitogen-driven cell proliferation and ameliorates disease progression [31]. Kcna3 (Kv1.3) seems to be involved in cancer cell proliferation and apoptosis in several types of tumors. Additionally, Kcna3 expression varies depending on the cancer stage [32], and knockdown of Kcna3 significantly suppressed cell proliferation and increased apoptosis [33]. Kcnrg encodes a potassium channel-regulating protein and was proposed as a new tumor suppressor gene [34], located in the 13q14.3 human chromosome. Several cytogenetic studies have reported that chromosome 13q is one of the most common deleted regions and contains one or more of the genes associated with the development or progression of HCC. Previous results suggest that genetic alterations and the expression of Kcnrg might play an important role in the development and/or progression of a subset of HCC [35].
Trpc6 belongs to the family of transient receptor potential channel proteins and can be activated by receptor tyrosine kinases or G protein-coupled receptors, which are involved in human cancer cell proliferation [25, 28]. Trpc6 channels contribute to cell cycle progression, and blockade of Trpc6 induces G2/M phase arrest and inhibits the development of xenografted gastric tumors [36].
Increased expression of voltage-gated Na+ channels has been reported in several cancer types, including breast, prostate and lung cancer, and has been associated with the metastatic properties of cancer cells [37]. Increased invasion of cancer cells is proposed to be mediated by increasing H+ efflux [38] and by stimulating cysteine cathepsin activity [39]. The gradual increase in Scn2a1 (Nav1.2) channel expression during HCC development suggests this channel as a novel HCC biomarker.
Abcc3 transports organic anions and phase II metabolic products and is mainly expressed in the kidney, liver and gut. Additionally, Abcc3 expression has been reported in cancer tissues, and a correlation with doxorubicin resistance in lung cancer has been reported [40–42]. Abcc3 mRNA expression displayed a very high increase in carcinoma tissue compared to normal tissue, probably as a response to eliminate the toxic agent (DEN) from the cells. Notably, we observed increased Abcc3 expression in the tumor tissue (GGT+), suggesting that it might be involved in cell proliferation or multidrug resistance, which is a common phenomenon in HCC. Although Abcc3 expression was consistent with the hepatocellular lineage in human liver samples, we observed a differential subcellular localization and expression pattern in each type of liver pathology: granular in normal liver, at the plasma membrane in cirrhotic liver and only cytoplasmic in HCC. This particular localization might be useful in the diagnosis of cirrhosis and HCC, especially if used in combination with CK19 and GPC3 expression.
The HCC model used here resembles human HCC development, namely progression from cirrhosis to HCC [5, 6]. Insulin-like growth factor 1 (IGF-1) and alpha-fetoprotein (AFP) have been proposed as potential liver tumor markers. IGF-1 and its receptor (IGF-1R) were increased in rat liver fibrosis induced by carbon tetrachloride [43]. Increased serum AFP was observed in the Solt-Farber carcinogenesis model over the first 2 months of treatment, then declined but increased again after 12 months when HCC appeared [44, 45]. In this model (also known as “the resistant hepatocyte model”), the rats are initiated with DEN, then treated with 2-acetylaminofluorene for 3 consecutive days; a 70 % partial hepatectomy is then performed [44, 45]. Although AFP is not a specific marker, it will be very important to evaluate IGF and AFP expression in the HCC model used in the present study to know whether such expression precedes ion channel and transporter expression. It will also be relevant to study ion channel/transporter expression in other liver cirrhosis and HCC models, including those associated with hepatitis virus infection, as well as in HCC biopsies with different etiologies.
The role of the proteins and the corresponding genes studied here in HCC development, as well as the potential HCC-preventive and therapeutic effects of ion channel inhibitors, should continue to be studied further. Nevertheless, although further studies are needed, the ion channel/transporter expression profile reported here suggests the possibility of using these genes as potential early markers and therapeutic targets in HCC.
References
Globocan. International Agency for Research on Cancer, 2012. https://globocan.iarc.fr. Accessed 22 June 2013.
Thorgeirsson SS, Grisham JW. Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet. 2002;31:339–346.
Heindryckx F, Colle I, Van Vlierberghe H. Experimental mouse models for hepatocellular carcinoma research. Int J Exp Pathol. 2009;90:367–386.
Farber E. The multistep nature of cancer development. Cancer Res. 1984;44:4217–4223.
Liu YF, Zha BS, Zhang HL, et al. Characteristic gene expression profiles in the progression from liver cirrhosis to carcinoma induced by diethylnitrosamine in a rat model. J Exp Clin Cancer Res. 2009;28:107.
Schiffer E, Housset C, Cacheux W, et al. Gefitinib, an EGFR inhibitor, prevents hepatocellular carcinoma development in the rat liver with cirrhosis. Hepatology. 2005;41:307–314.
Paradis V. Histopathology of hepatocellular carcinoma. Rec Results Cancer Res. 2013;190:21–32.
Filmus J, Capurro M. Glypican-3: a marker and a therapeutic target in hepatocellular carcinoma. FEBS J. 2013;280:2471–2476.
Lee JI, Lee JW, Kim JM, et al. Prognosis of hepatocellular carcinoma expressing cytokeratin 19: comparison with other liver cancers. World J Gastroenterol. 2012;18:4751–4757.
Pardo LA, del Camino D, Sanchez A, et al. Oncogenic potential of EAG K(+) channels. EMBO J. 1999;18:5540–5547.
Cuddapah VA, Sontheimer H. Ion channels and transporters in cancer. 2. Ion channels and the control of cancer cell migration. Am J Physiol Cell Physiol. 2011;301:C541–C549.
Camacho J. Ether a go-go potassium channels and cancer. Cancer Lett. 2006;233:1–9.
Rodriguez-Rasgado JA, Acuna-Macias I, Camacho J. Eag1 channels as potential cancer biomarkers. Sensors (Basel). 2012;12:5986–5995.
Hemmerlein B, Weseloh RM, Mello de Queiroz F, et al. Overexpression of Eag1 potassium channels in clinical tumours. Mol Cancer. 2006;5:41.
Pardo LA, Stuhmer W. The roles of K(+) channels in cancer. Nat Rev Cancer. 2014;14:39–48.
Diaz L, Ceja-Ochoa I, Restrepo-Angulo I, et al. Estrogens and human papilloma virus oncogenes regulate human ether-a-go-go-1 potassium channel expression. Cancer Res. 2009;69:3300–3307.
Ousingsawat J, Spitzner M, Puntheeranurak S, et al. Expression of voltage-gated potassium channels in human and mouse colonic carcinoma. Clin Cancer Res. 2007;13:824–831.
Zhang L, Barritt GJ. Evidence that TRPM8 is an androgen-dependent Ca2+ channel required for the survival of prostate cancer cells. Cancer Res. 2004;64:8365–8373.
Ortiz CS, Montante-Montes D, Saqui-Salces M, et al. Eag1 potassium channels as markers of cervical dysplasia. Oncol Rep. 2011;26:1377–1383.
Makino H, Uetake H, Danenberg K, et al. Efficacy of laser capture microdissection plus RT-PCR technique in analyzing gene expression levels in human gastric cancer and colon cancer. BMC Cancer. 2008;8:210.
Curran S, McKay JA, McLeod HL, et al. Laser capture microscopy. Mol Pathol. 2000;53:64–68.
Torres Mena JE, Sanchez Rodriguez R, Quintanar Jurado V, et al. Laser capture microdissection after gamma-glutamyl transferase histochemistry: an optimization for gene expression analysis. Anal Biochem. 2014;447:126–132.
Rutenburg AM, Kim H, Fischbein JW, et al. Histochemical and ultrastructural demonstration of gamma-glutamyl transpeptidase activity. J Histochem Cytochem. 1969;17:517–526.
Pompella A, De Tata V, Paolicchi A, et al. Expression of gamma-glutamyltransferase in cancer cells and its significance in drug resistance. Biochem Pharmacol. 2006;71:231–238.
Djamgoz MB, Coombes RC, Schwab A. Ion transport and cancer: from initiation to metastasis. Philos Trans R Soc Lond B Biol Sci. 2014;369:20130092.
Arcangeli A, Crociani O, Bencini L. Interaction of tumour cells with their microenvironment: ion channels and cell adhesion molecules. A focus on pancreatic cancer. Philos Trans R Soc Lond B Biol Sci. 2014;369:20130101.
Nies AT, Konig J, Pfannschmidt M, et al. Expression of the multidrug resistance proteins MRP2 and MRP3 in human hepatocellular carcinoma. Int J Cancer. 2001;94:492–499.
Urrego D, Tomczak AP, Zahed F, et al. Potassium channels in cell cycle and cell proliferation. Philos Trans R Soc Lond B Biol Sci. 2014;369:20130094.
Oliveira PA, Colaco A, Chaves R, et al. Chemical carcinogenesis. An Acad Bras Cienc. 2007;79:593–616.
Ohya S, Niwa S, Kojima Y, et al. Intermediate-conductance Ca2+-activated K+ channel, KCa3.1, as a novel therapeutic target for benign prostatic hyperplasia. J Pharmacol Exp Ther. 2011;338:528–536.
Freise C, Ruehl M, Seehofer D, et al. The inhibitor of Ca(2+)-dependent K+ channels TRAM-34 blocks growth of hepatocellular carcinoma cells via downregulation of estrogen receptor alpha mRNA and nuclear factor-kappaB. Invest New Drugs. 2013;31:452–457.
Jang SH, Kang KS, Ryu PD, et al. Kv1.3 voltage-gated K(+) channel subunit as a potential diagnostic marker and therapeutic target for breast cancer. BMB Rep. 2009;42:535–539.
Wu J, Zhong D, Wu X, et al. Voltage-gated potassium channel Kv1.3 is highly expressed in human osteosarcoma and promotes osteosarcoma growth. Int J Mol Sci. 2013;14:19245–19256.
Ivanov DV, Tyazhelova TV, Lemonnier L, et al. A new human gene KCNRG encoding potassium channel regulating protein is a cancer suppressor gene candidate located in 13q14.3. FEBS Lett. 2003;539:156–160.
Cho YG, Kim CJ, Song JH, et al. Genetic and expression analysis of the KCNRG gene in hepatocellular carcinomas. Exp Mol Med. 2006;38:247–255.
Cai R, Ding X, Zhou K, et al. Blockade of TRPC6 channels induced G2/M phase arrest and suppressed growth in human gastric cancer cells. Int J Cancer. 2009;125:2281–2287.
Fraser SP, Ozerlat-Gunduz I, Brackenbury WJ, et al. Regulation of voltage-gated sodium channel expression in cancer: hormones, growth factors and auto-regulation. Philos Trans R Soc Lond B Biol Sci. 2014;369:20130105.
Brisson L, Gillet L, Calaghan S, et al. Na(V)1.5 enhances breast cancer cell invasiveness by increasing NHE1-dependent H(+) efflux in caveolae. Oncogene. 2011;30:2070–2076.
Gillet L, Roger S, Besson P, et al. Voltage-gated sodium channel activity promotes cysteine cathepsin-dependent invasiveness and colony growth of human cancer cells. J Biol Chem. 2009;284:8680–8691.
Szakacs G, Paterson JK, Ludwig JA, et al. Targeting multidrug resistance in cancer. Nat Rev Drug Discov. 2006;5:219–234.
Scheffer GL, Kool M, de Haas M, et al. Tissue distribution and induction of human multidrug resistant protein 3. Lab Invest. 2002;82:193–201.
Young LC, Campling BG, Cole SP, et al. Multidrug resistance proteins MRP3, MRP1, and MRP2 in lung cancer: correlation of protein levels with drug response and messenger RNA levels. Clin Cancer Res. 2001;7:1798–1804.
Wang XZ, Chen ZX, Zhang LJ, et al. Expression of insulin-like growth factor 1 and insulin-like growth factor 1 receptor and its intervention by interleukin-10 in experimental hepatic fibrosis. World J Gastroenterol. 2003;9:1287–1291.
Anilkumar TV, Golding M, Edwards RJ, et al. The resistant hepatocyte model of carcinogenesis in the rat: the apparent independent development of oval cell proliferation and early nodules. Carcinogenesis. 1995;16:845–853.
Sell S. Heterogeneity and plasticity of hepatocyte lineage cells. Hepatology. 2001;33:738–750.
Acknowledgments
We thank Eduardo García Osornio, M.V.Z. Ricardo Gaxiola and Rafael Leyva for their technical assistance. This work was partially supported by the Consejo Nacional de Ciencia y Tecnología grant 168102 to JC and grant 115431 to JIP-C.
Ethical approval
All applicable international, national and/or institutional guidelines for the care and use of animals were followed. All procedures performed in studies involving animals were in accordance with the ethical standards of the institution or practice at which the studies were conducted. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This is a retrospective study, and for this type of study, formal consent is not required.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zúñiga-García, V., Chávez-López, M.d.G., Quintanar-Jurado, V. et al. Differential Expression of Ion Channels and Transporters During Hepatocellular Carcinoma Development. Dig Dis Sci 60, 2373–2383 (2015). https://doi.org/10.1007/s10620-015-3633-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-015-3633-9