Abstract
Conservation breeding programs are increasingly used as recovery actions for wild animals; bringing founders into captivity to rear captive populations for future reintroduction into the wild. The International Union for the Conservation of Nature recommends that founders should come from genetically close populations and should have sufficient genetic diversity to avoid mating among relatives. Genomic data are highly informative for evaluating founders due to their high resolution and ability to capture adaptive divergence, yet, their application in that context remains limited. Woodland caribou are federally listed as a Species at Risk in Canada, with several populations facing extirpation, such as those in the Rocky Mountains of Alberta and British Columbia (BC). To prevent local extirpation, Jasper National Park (JNP) is proposing a conservation breeding program. We examined single nucleotide polymorphisms for 144 caribou from 11 populations encompassing a 200,0002 km area surrounding JNP to provide information useful for identifying appropriate founders for this program. We found that this area likely hosts a caribou metapopulation historically characterized by high levels of gene flow, which indicates that multiple sources of founders would be appropriate for initiating a breeding program. However, population structure and adaptive divergence analyses indicate that JNP caribou are closest to populations in the BC Columbia range, which also have suitable genetic diversity for conservation breeding. We suggest that collaboration among jurisdictions would be beneficial to implement the program to promote recovery of JNP caribou and possibly other caribou populations in the surrounding area, which is strategically at the periphery of the distribution of this endangered species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As the rate of biodiversity loss caused by anthropogenic activities dramatically increases, numerous species and populations are now extirpated or face the threat of extirpation (Pimm et al. 2014). Consequently, recovery actions like conservation breeding programs are sometimes used to reintroduce species to areas where they used to occur, or to augment populations (Seddon et al. 2007; Brichieri-Colombi and Moehrenschlager 2016; Bubac et al. 2019). Conservation breeding programs encompass the act of bringing rare or endangered animals into captivity with the aim of rearing captive populations for eventual reintroduction of their progeny into the wild (Seddon et al. 2014). Examples of such programs for ungulates include those conducted for the European bison (Bison bonasus) in Poland (Tokarska et al. 2009), the Przewalski’s horse in Mongolia and China (Der Sarkissian et al. 2015), the American bison (Bison bison) in Oklahoma (Kleiman 1989), USA, the Arabian oryx (Oryx leucoryx) in Arabia (Arif et al. 2010), and the Pére David's deer (Elaphurus davidianu) in China (Dayuan et al. 2022).
Despite multiple examples of successful conservation breeding programs, their execution is challenging (Fischer and Lindenmayer 2000; Armstrong and Seddon 2008). To improve their success, guidelines and best practices are continuously improved (Frankham 2010). For example, the International Union for the Conservation of Nature (IUCN) guidelines indicate that reintroductions and other conservation translocations should be based on plans that evaluate economic, social, and ecological aspects, such as a species’ current and historical distribution, and the removal or substantial reduction, of extinction threat(s) (IUCN/SSC 2013). The IUCN guidelines emphasize that selection of appropriate founders is a critical factor affecting reintroduction success (IUCN/SSC 2013; Forsman 2014). Ideally, founders should have appropriate demographic characteristics (e.g., number, sex, and age), come from geographically close populations, and have characteristics (i.e., ecology, behaviour, etc.) similar to the populations being restored (Robert 2009; Soorae 2018).
Considering the genetics of founders is crucial for the success of conservation breeding programs (Araki et al. 2007; Witzenberger and Hochkirch 2011). From a genetic perspective, the best source populations are those that are genetically similar to the populations that had been extirpated and have sufficient genetic diversity to reduce mating among relatives of captive individuals (Frankham 2010; Pelletier et al. 2009; Miller et al. 2010). However, recent work has also indicated that genetic diversity is important and matching genetic groups may be overstated if there is not a biological difference between the groups, i.e. (Frankham et al. 2017; Ralls et al. 2020). Accounting for proper genetic characteristics of founders can increase the survival probability of released individuals and improve the chance of having healthy and self-sustaining reintroduced populations in the long term (Robert 2009; Williams and Hoffman 2009).
Until recently, genetic studies aimed at identifying founders for conservation breeding programs have been based on small sets of putatively neutral molecular markers (i.e., regions of the genome that are non-coding) such as microsatellites and/or mitochondrial DNA (mtDNA) (Russello and Jensen 2018). However, these markers have known limitations, including limited resolution and ability to capture adaptive differentiation. Single nucleotide polymorphisms (SNPs), have the potential to overcome such limitations and be more informative for the selection of founders (Allendorf et al. 2010; He et al. 2016). The higher resolution of SNPs can provide more precise estimates of genetic parameters (e.g., genetic diversity and inbreeding) of candidate founders when compared to microsatellites and mtDNA (Luikart et al. 2003; Ivy et al. 2009; Galla et al. 2020). In addition, as SNPs may be located in genes, they can be used to identify specific loci of interest in founders that might be desirable to maintain (e.g., loci under selection) in the captive and reintroduced populations (Laikre 1999; Luikart et al. 2003; Weeks et al. 2011; Flanagan et al. 2018; Wright et al. 2021).
Caribou (Rangifer tarandus), also known as reindeer, are a Holarctic species that is globally declining, and listed as a Species at Risk in Canada (Environment Canada 2014). Woodland caribou populations located at the southern edge of the species’ distribution, such as those in the study area in the Rocky Mountains of Alberta (AB) and British Columbia (BC) (Fig. 1), are particularly at risk of disappearing, also due to climate change (Vors and Boyce 2009). These caribou populations belong to one subspecies (Woodland Caribou) and two Designatable Units (DU; Central and Southern Mountain), which largely overlap in meaning with Evolutionary Significant Units (ESUs; Crandall et al. 2000; Green 2005; COSEWIC 2011). DUs are further divided into subpopulations, also known as “herds” (although herds of Woodland Caribou are not social groups; Bergerud 1988), delineated for management and conservation purposes (Environment Canada 2014).
Caribou sampled in the southern Rocky Mountains of Canada for genomic analyses (n = 137; sampling locations in Fig. 3). Black lettered circles indicate sampled herds (i.e., a term used to indicate subpopulations, although herds of Woodland caribou are not social groups) with circle size proportional to sample size (mean = 12.45, SD = 7.03, range 2–20). Grey-scale polygons show the distribution of Designatable Units (DUs) encompassing multiple herds
In the southern Rocky Mountains of Canada, caribou herds occur on public lands available for multiple land uses including resource extraction, and in provincially and federally protected areas, however even in protected areas some have been extirpated and others are declining (Environment Canada 2014; Serrouya et al. 2019). In the mountain national parks, the last 9 caribou were extirpated from Banff in 2009 (Hebblewhite et al. 2010), and caribou now only occur in Revelstoke and Jasper (JNP) national parks. Caribou in JNP are either extirpated or facing near-extirpation, a status that calls for conservation actions (Parks Canada Agency 2022a, b). Caribou declines in JNP are due to multiple factors. Decades ago the JNP caribou lost migratory access to traditional forested foothills outside of the national park, due to high levels of habitat change and loss from industrial land uses. Loss of annual migration has been demonstrated to compromise the population viability of central mountain caribou (Williams et al. 2021). In addition, caribou remaining within JNP have been negatively affected by apparent competition with elk (Holt 1977) mediated by wolves (Bradley and Neufeld 2012). A rapid increase of elk density, which occurred after their reintroduction in 1960, resulted in higher wolf densities and an increase in predation rate on elk and caribou. Recently, lower wolf and elk densities have resulted in more favourable ecological conditions for caribou, and potential for recovery (Parks Canada Agency 2022a, b). Despite these current more favourable conditions, caribou within JNP continue to struggle with low survival rates, and the population is now too small to recover without population augmentation (Parks Canada Agency 2022a, b). Following IUCN guidelines, JNP is taking actions to aid recovery by exploring conservation breeding of JNP caribou. As all caribou herds within JNP are either extirpated, near extirpated or declining and as the habitat for caribou has been considered either favourable or improving, conservation breeding was considered a viable option by conservation planners and stakeholders (Foundations of Success and Parks Canada 2021; Salafsky et al. 2022). This exploration includes assessment of genetic information to identify options for selection of founder animals (Parks Canada Agency 2022a, b).
Caribou genetics studies conducted in the southern Rocky Mountains of AB and BC have primarily used neutral molecular markers (i.e., mitochondrial DNA haplotypes and autosomal microsatellites) (McDevitt et al. 2009; Serrouya et al. 2012; Weckworth et al. 2012 Yannic et al. 2014), although more recent work has used genomic data (Cavedon et al. 2019; Taylor et al. 2022). MtDNA indicated the presence of a hybrid zone where sympatric individuals have either Beringian/Northern or Southern mitochondrial lineage (McDevitt et al. 2009; Yannic et al. 2014). Studies based on microsatellites and SNPs also indicated the presence of shared genetic traits (McDevitt et al. 2009; Serrouya et al. 2012; Weckworth et al. 2012; Cavedon et al. 2019; Taylor et al. 2022). For example, population structure studies indicated that JNP caribou had genetic characteristics of both Central and Southern Mountain DU herds but that it was not possible to ascertain which characteristics predominated (McDevitt et al. 2009; Serrouya et al. 2012; Weckworth et al. 2012). However, past studies did not survey an adequate number of herds in the immediate JNP area to clarify local genetic structure and inform founder selection.
We conducted a genomic study to inform JNP’s planned conservation breeding program. We characterised genomic diversity in 144 individuals from 11 herds in a 200,000 km2 area using a newly developed caribou SNP array (Carrier et al. 2022), and sampling equally from the Central and Southern Mountain DUs. Our aims were to: (i) define levels of genetic differentiation, (ii) evaluate population structure, and (iii) detect signatures of adaptive divergence between population groups determined with population structure analyses. The goal of our study was to identify suitable founder populations for the establishment of a JNP conservation breeding program, and to interpret the findings in view of genetic and ecological traits of caribou in the southern Rocky Mountains of AB and BC.
Materials and methods
Sampling and genomic data
Blood and tissue samples were collected from both live and deceased animals as part of government agencies’ caribou monitoring activities between 2012 and 2020, corresponding to ≥ one generation for caribou (following McLoughlin et al. 2003).The collected samples spanned a 200,000 km2 study area located in the mountainous region along the southern AB and BC border in proximity to JNP, Canada (Fig. 1). Caribou in this area belong to either the Central Mountain DU or the Southern Mountain DU (COSEWIC 2011) and are from the montane cordillera ecozone, which includes habitats ranging from alpine tundra, to dense coniferous forests, to dry sagebrush and grasslands (Ecological Stratification Working Group 1995).
We extracted DNA from samples using the Qiagen kits following manufacturer protocols for both single spin columns (DNeasy Blood & Tissue Kit) and 96-well plates (QIAamp 96 DNA QIAcube HT kit). DNA samples were then quantified using either the BioTek Synergy LX Multimode Reader or the Thermo Fisher Qubit 3 Fluorometer following Thermo Fisher with Qubit and Quant-iT 1X dsDNA high sensitivity (HS) or broad range (BR) Kits. One hundred and forty-four samples that yielded ≥ 400 ng of DNA were selected for analyses. Samples were normalized to a quantity of 400 ng, dried on a Thermo Scientific Savant SpeedVac DNA 130 Integrated Vacuum Concentrator System, and shipped at room temperature to Genome Québec Ltd (Montréal, Québec) where genotyping using the Illumina Caribou 60 K SNP array (Carrier et al. 2022) was outsourced. The array accounts for SNPs evenly distributed across the entire genome (~ every 50 Kb) with known minor alleles across populations world-wide. In addition, a subset of SNPs was selected to represent rare and local alleles which could be used for ecotype and population assignments—information urgently needed for conservation planning.
We used PLINK v1.9 (Purcell et al. 2007) to perform data quality control, which included filtering out individuals and SNPs with call rates < 0.98 and SNPs with a minor allele frequency (MAF) < 0.01. After filtering there were 44,112 SNPs remaining and 137 individuals from both the Central Mountain DU (nherds = 6, individuals = 77) and Southern Mountain DU (nherds = 5, individuals = 60) (Fig. 1). For population structure analyses (below), we used PLINK to further exclude SNPs exhibiting strong linkage disequilibrium (“–indep-pairwise 50 5 0.5”) and those not in Hardy–Weinberg equilibrium (“–hwe 0.001”), leaving 36,053 SNPs for each individual (Purcell et al. 2007). In addition, for population structure analyses, we removed 42 animals which had a close relative in the dataset (28 and 14 belonging to the Central Mountain DU and Southern Mountain DUs, respectively) based on an identity by descent (IBD; –genome) degree of recent shared ancestry threshold of 0.25 (second-degree relatives) (see for example Kominakis et al. 2021).
Assessing and comparing diversity of caribou
We used the R package DARTR (Gruber et al. 2018) to estimate observed (Ho) and expected heterozygosity (He) and inbreeding coefficients (FIS, with 95% Confidence Intervals (CI) of bootstrap values) for each herd as well as pairwise fixation index values (FST) between herds. We then used FST values to derive the number of migrants per generation (Nm; i.e., an estimate of gene flow; Wright 1931). The applied metrics of FST are sensitive to a species’ heterozygosity, and FST as well as the population STRUCTURE analyses (below) also to sample sizes. Due to low sample size, we therefore combined individuals from the Banff herd (n = 3) with those from JNP (n = 15). We also combined individuals from the Purcell South herd (n = 4) with those from South Selkirk herd (n = 3). The pooling of these samples was warranted due to known genetic similarities (McDevitt et al. 2009; Serrouya et al. 2012; Weckworth et al. 2012) and geographic proximity. We estimated a kinship matrix between caribou individuals using the R package POPKIN (Ochoa and Storey 2021), and from this, we also obtained the inbreeding coefficients for each caribou. To avoid a bias and potential underestimation of kinship and inbreeding, we used a SNP dataset including all caribou individuals (i.e., also including relatives), and where SNPs were filtered by LD and HWE as is standard practice.
Population structure analyses
To assess isolation by distance (IBD), we performed a Mantel test to detect the potential correlation between the geographic and genetic distances calculated between individuals pairwise. We calculated the geographic distances with the function “distGeo” within the R package GEOSPHERE (Karney 2013), whereas we calculated the genetic distances with the function “gl.dist.ind” within the R package DARTR (Gruber et al. 2018). Lastly, we performed the mantel test with the function “gl.ibd” also within the dartR package.
To visualize patterns of population structure, we calculated pairwise Nei’s genetic distances (Nei 1972) between all individuals using the R package StAMPP v1.6.1 (Pembleton et al. 2013) and constructed a neighbour-joining tree based on the genetic distances using the R package APE 5.2 (Paradis et al. 2004). We also evaluated population structure using STRUCTURE v2.3.4, which uses a Bayesian iterative algorithm to place samples into clusters (K) whose members share similar patterns of genetic variation (Pritchard et al. 2000; Falush et al. 2003). We ran STRUCTURE with 20,000 burn-in iterations followed by 50,000 sampling iterations for K = 1 through 10 (Schweizer et al. 2016; Cavedon et al. 2022a, b, c). Each run was performed 10 times, and the ΔK statistic of Evanno et al. (2005) was calculated using STRUCTURE HARVESTER to help determine the most appropriate number of genetic clusters (Earl and von Holdt 2012). For comparison, we also assessed population structure with a maximum likelihood approach implemented in ADMIXTURE v1.3 and the most appropriate number of genetic clusters was detected by examining the cross-validation errors for K varying from 1 to 10 (Alexander et al. 2009). Lastly, as is common in genetic studies examining population structure (see for example Cavedon et al. 2022a, b, c), we performed a principal component analysis (PCA) to determine population groups using the program SMARTPCA within EIGENSTRAT v3.0 (Price et al. 2006).
Detection of adaptive divergence between caribou population groups
We identified putatively adaptatively divergent SNPs between population groups determined with population structure analyses (see results) with BAYESCAN v.2.1 (Foll and Gaggiotti 2008). BAYESCAN tests whether population group-specific allele frequencies, measured by an FST coefficient, are significantly different from the allele frequency within the common gene pool. It also assigns a posterior probability (alpha) to a model in which selection explains the difference in allele frequencies better than a null model. A positive alpha indicates population group-specific directional selection, while a negative alpha suggests balancing or purifying selection. We ran analyses using a prior odd of 10, where SNPs were considered to be under selection when below the false discovery rate (FDR) threshold of 0.05.
Results
Genetic diversity and gene flow
Observed heterozygosity (Ho) of caribou herds in the study area ranged from 0.33 to 0.39 (mean 0.37 + SD 0.02; Table 1) and expected heterozygosity (He) ranged from 0.35 to 0.39 (mean 0.37 + SD 0.01). Inbreeding coefficients (FIS) ranged from 0.003 to 0.047 (mean 0.019 + SD 0.015). Pairwise fixation index (FST) between herds ranged from 0.01 to 0.09 (mean 0.06 + SD 0.02). FST values between JNP and the other herds varied from 0.06 to 0.09 (mean 0.07 + SD 0.04; Table 2). The estimated number of migrants per generation (Nm values) between herds assessed pairwise were all > 1 (mean 5.38 + SD 4.10). The estimated Kinship matrix indicated moderate relatedness among caribou individuals within the same herd (Fig. 2), with the exception of caribou within the Hart South and North Caribou herds. The matrix indicated lower relatedness among caribou individuals belonging to different herds. The inbreeding coefficients for caribou individuals calculated from the kinship matrix followed a gaussian distribution (Fig. S1).
Kinship matrix of caribou individuals. Colors indicate the degree of relatedness between individuals (darker color indicate higher relatedness). Individuals were grouped by herd: MOB = Moberly; QUI = Quintette: RED = Redrock-Prairie Creek; ALP = À La Pêche: JNP = Jasper + Banff; HS = Hart South; NC = North Caribou; COL = Columbia North; PS = South Selkirk and Purcell South
Population structure of caribou individuals
We found a significant correlation between geographic distance and genetic distance, across the 137-individual SNP data set (r = 0.499; Mantel test P = 0.001; Fig. S2). JNP and Banff individuals grouped together in the neighbor-joining tree and were most similar to caribou from Columbia North, Purcell South, and South Selkirk. JNP and Banff individuals then grouped together with caribou from À La Pêche, and then caribou from Redrock-Prairie Creek (a group that was admixed with some Hart South individuals) (Fig. 3). The remaining groups (Hart South, North Cariboo, Quintette, and Moberly) were admixed and branched together separately from JNP and Banff (Fig. 3).
STRUCTURE and ADMIXTURE analyses both supported the presence of two genetic clusters (K = 2, Fig. 4). Results also indicated that JNP and Banff individuals had similar assignment to individuals belonging to the Columbia North, Purcell South, and South Selkirk populations, but were different from individuals from À La Pêche, Redrock-Prairie Creek, Quintette, Moberly, Hart South, and North Cariboo.
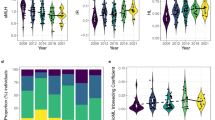
Population structure of caribou sampled in the southern Rocky Mountains of Canada. a Bar plots from STRUCTURE and ADMIXTURE analyses indicate individual proportions of assignment into two main genetic clusters (red or blue color). The most likely number of clusters (K) obtained with STRUCTURE (higher values best) and ADMIXTURE (lower values best) is indicated with circles on the respective scatter plots. b Map showing distribution of sampled caribou (capture coordinates) with proportion of belonging to either cluster for each individual obtained with STRUCTURE (pie chart)
A principal component analysis indicated that individuals belonging to Central Mountain or Southern Mountain DUs could not be obviously separated along the two principal axes (Fig. 5a). It also indicated that some Central and Southern Mountain DU caribou were genetically close, including an assemblage formed by À La Pêche, Redrock-Prairie Creek, Quintette, Moberly, Hart South, and North Cariboo individuals, and including another assemblage formed by Columbia North, South Selkirk, Purcell South, JNP, and Banff individuals (Fig. 5b). JNP and Banff caribou could not be distinguished from Columbia North, Purcell South, and South Selkirk along the PC1 axis, whereas they could be distinguished from other caribou along the PC2 axis. The first and second axis accounted for 3.37 and 2.79% of the observed genetic variation, respectively (the 3rd axis accounted for 1.88%). Our analyses therefore determined four distinguishable groups (Fig. 5b), including caribou individuals with varying proportions of assignment to the two major genetic clusters detected with STRUCTURE (Fig. 5c). A population group (called “JNP”) was formed by individuals from the JNP and Banff herds. One more population group was formed by individuals from À La Pêche (“ALP”), one additional by individuals from Columbia North, Purcell South, and South Selkirk (“COL”). The fourth and final population group (“CR”) included individuals from the remaining sampled herds, which were also close on the PCA plot (Hart South, North Cariboo, Redrock-Prairie Creek, Quintette, and Moberly).
Principal Component Analysis (PCA) of caribou sampled in the southern Rocky Mountains of Canada. Circles represent caribou individuals, and colors represent DUs (in Panel a), herds (in Panel b) or proportion of assignment to the two major genetic clusters detected with STRUCTURE (in Panel c). Dashed circles in panel B indicate the caribou population groups we detected, which were then used in analyses of adaptive divergence: ALP = À La Pêche; COL = Columbia North, Purcell South, and South Selkirk; JNP = Jasper and Banff; CR = Moberly, Quintette, Redrock-Prairie Creek, Hart South, and North Cariboo
Divergence between caribou population groups
To assess putatively adaptive divergence, we ran pairwise BAYESCAN analyses between the four caribou population groups identified with PCA analyses (see above). We identified 89 outlier SNPs divergent between JNP and ALP, with FST values ranging between 0.20 and 0.36 (average 0.25). We also detected 68 SNPs diverging between JNP and CR, with FST values ranging between 0.15 and 0.24 (average 0.19). Lastly, we found only 53 outlier SNPs between JNP and COL with FST values ranging between 0.21 and 0.29 (average 0.25; Fig. 6).
Signatures of adaptive divergence between caribou population groups determined by outlier analyses. The horizontal axis indicates the BAYESCAN-assessed log10 of the q value (the false discovery rate (FDR) analog to the p-value) and the vertical axis is the mean genetic differentiation (FST). Each point represents a SNP and significant outliers are visible right of the grey vertical line. Identifiers represent population groups: ALP = À La Pêche; COL = Columbia North, Purcell South, and South Selkirk; JNP = Jasper and Banff; CR = Moberly, Quintette, Redrock-Prairie Creek, Hart South, and North Cariboo
Discussion
We conducted a genomic study to identify candidate founders for a conservation breeding program currently planned for caribou, an approach suggested as a best practice for endangered species in general, but requiring substantial preparation and research to be successful (Russello and Jensen 2018). Our work relied on a newly developed SNP-array specifically produced for caribou (Carrier et al. 2022), which in this study produced around 40,000 SNPs that were successfully genotyped in 95% of the sampled individuals. Across the 200,000 km2 study area we found that, despite wide-spread recent declines (Serrouya et al. 2019), caribou subpopulations (herds) have retained levels of genetic diversity and natural connectivity (e.g. more than one migrant per generation [Wright 1931; Slatkin 1987; Weeks et al 2011] genetically estimated by this study between any pair of herds) which in other studies have been considered adequate for conservation interventions including translocations of individuals. However, we found two major genetic clusters and additional population groups, which should inform conservation planning. If a conservation breeding program is established in JNP, founders could be selected from clusters or population groups most similar to JNP as a first priority, while also taking into consideration information on potentially-adaptive divergence (see discussion below on results of outlier analyses).
The levels of heterozygosity we identified (including He and Ho) were aligned with other reported values for caribou and approaching levels considered as “high” in wild mammals (see for example Cavedon et al. 2019). The levels of inbreeding coefficients we also determined (FIS), which provide information on relatedness among individuals (Crow and Kimura 1970; Caballero et al. 2021), were low to moderate (see Solmundson et al. 2020). Therefore, our findings indicate that the study area’s wild caribou have retained levels of genetic diversity, which if properly maintained in captive populations could perhaps circumvent risk of inbreeding depression. Also valuable to avoid inbreeding depression, in this study, we identified close relatives (those with IBD > 0.25 corresponding to second-degree relatives), which should be discarded when selecting source founders. Kinship is an important tool used to identify breeders for captive populations and can be used to effectively evaluate the potential for future inbreeding in wild populations (Ballou and Lacy 1995; Frankham et al. 2017; Flesch et al. 2022). Our kinship findings indicated moderate relatedness only among caribou individuals within the same herd and suggested that inbreeding could be minimized by relying on founders from different source herds, should a captive population be established. Our results also indicated that herds known to overlap (example, Hart South and North Caribou with other neighbouring herds; Environment Canada 2014) naturally had lower relatedness. Future studies could examine inbreeding and relatedness in more detail, such as by examining “runs of homozygosity” (see Broman and Weber 1999) across the genome, an approach that may show signatures of natural and/or human mediated selection (Kardos et al. 2015; Caballero et al. 2021; Solmundson et al. 2020).
We estimated that the number of migrants per generation (Nm) were all greater than one, indicating significant historical and/or recent gene flow among all herd pairs. One migrant per generation, as a minimum, is typically considered sufficient to offset genetic deterioration within subunits (Wright 1931; Slatkin 1987; Weeks et al 2011). It is therefore likely that caribou herds in the study area were connected until recently, with barriers to gene flow likely arising in the last decades. Consistent with this interpretation, barriers to dispersal have been identified in studies of radio-collared southern mountain caribou (Van Oort et al. 2011), which likely reflect contemporary, non-historical patterns (i.e., those observed during the 2–3 years lifespan of a GPS collar), and which were deployed just on females (i.e., the least vagile sex in caribou, as discussed in Cavedon et al. 2022a, b, c). By contrast, presence of barriers between population ranges was not as consequential: habitat suitability followed by predation risk was associated with overall gene flow in a caribou study conducted previously (Gubili et al. 2017). Importantly,, levels of Nm exceeding one have been historically used to manage wildlife populations as one unit (Mills and Allendorf 1996; Vucetich and Waite 2000; Wang 2004), and perhaps the same threshold could be applied to caribou (manageable in the future as a unit, similar to other wildlife populations). The levels of Nm we detected could be suitable for conservation translocations among these populations. This study’s populations are also within natural dispersal distances characteristic of the species, whereas dispersals themselves might be infrequent recently, likely due to habitat fragmentation (Van Oort et al. 2011). Nonetheless, further analyses on gene flow are recommended, as both FST and Nm are known to also depend on genetic diversity and on sample sizes (Holsinger and Weir 2009), which were limited in this study.
Population structure results indicate that caribou individuals could be grouped in two major genetic clusters, which is consistent with both past and current studies conducted in western Canada (see McDevitt et al. 2009; Cavedon et al. 2022a, b, c). In this study, we targeted common variants (SNPs with MAF > 0.01), which are suited to examine the deep evolutionary history of species (Gibson 2012). Our findings may therefore have captured similar diversification patterns as those detected with mtDNA analyses, which indicated the presence of either a Beringian/Northern or a Southern lineage in the study area (McDevitt et al. 2009; Serrouya et al. 2012; Weckworth et al. 2012; Yannic et al. 2014; Taylor et al. 2021).
The two genetic clusters we identified did not overlap fully with currently recognized DUs. We found that although caribou from some Central Mountain DU herds were genetically similar to one another, they were also similar to caribou from some Southern Mountain herds (note the mainly blue circles in Fig. 4b), and this information could also be used to plan future translocations. Our findings indicating the presence of two genetic clusters and potentially two DUs in the area are consistent with other recent studies also relying on genomic data for caribou (Cavedon et al. 2022a, b, c; Taylor et al. 2020, 2021). Our findings, however, partially contrast with past studies that rely on neutral markers (autosomal microsatellites and mtDNA), which have also indicated the presence of two main clusters and DUs, but with slightly different boundaries (McDevitt et al. 2009; Serrouya et al. 2012; Weckworth et al. 2012). This can be explained by the fact that these neutral genetic markers cannot detect local adaptation, unlike genomic SNPs (Luikart et al. 2003; Allendorf et al. 2010).
The principal component analysis detected four distinguishable groups, which we used for further analyses of potentially adaptive divergence. Overall, we found more outlier SNPs between JNP and ALP than between JNP and CR and especially between JNP and COL. Outlier SNPs may indicate divergence due to demographic events or selection processes related to local adaptation (Luikart et al. 2003; Allendorf et al. 2010). Previous genomic studies conducted for caribou on a larger scale also identified candidate loci under selection that were associated with ecological, behavioral, and climatic factors (Cavedon et al. 2019, 2022a, b, c). Together, the results of our current and previous studies (Cavedon et al. 2019, 2022a, b, c) indicate that caribou population groups in the study area may be differently adapted due to selective forces (e.g., environmental and climatic conditions), an important detail that should be accounted for in any conservation plan (Des Roches et al. 2021).
Our study of caribou genomics and distribution should inform selection of founders for the proposed JNP conservation breeding program. We found that the study area likely hosted a historic caribou metapopulation, characterized by high levels of gene flow which has led to high levels of genetic diversity. Our results showed that JNP could potentially acquire founders from any of the sampled herds, some of which, thanks to recovery actions, have recently started to increase and might be able to sustain the removal of some individuals (Government of Alberta 2017; Eacker et al. 2019; McNay et al. 2022). However, population structure analyses and adaptive divergence analyses indicate that JNP caribou are most genetically related to caribou in the Columbia range of BC. Selecting founders from the Columbia range would provide the best chance of maintaining genetic traits most similar to JNP caribou. Herds in the Columbia range are however numerically low compared to others (e.g., 184 individuals in Columbia North vs. 405 in Hart; see also Serrouya et al. 2021; McNay et al. 2022) and might not tolerate the removal of individuals. However, admixture of two caribou genetic clusters has also been consistently documented throughout the study area (see McDevitt et al. 2009, this study), therefore indicating that any conservation program should aim at maintaining such diversity (sensu Frankham et al. 2017, Ralls et al. 2020) too. It should also be taken into consideration that caribou genetic diversity in the study area is correlated with ecological and behavioural diversity also including seasonal migration (McDevitt et al. 2009; Cavedon et al. 2022a, b, c), and any conservation program should aim at maintaining all diversities in these partially migrating populations (Cavedon et al. 2019).
In addition to genetic analyses, it will be important to conduct population viability analyses, or an equivalent assessment approach, to understand the conservation implications of caribou removals for source populations of this species at risk (Hoban et al. 2012).
Similarly to caribou, other endangered species can benefit from the evaluation of genomic data applied to conservation breeding programs (Russello and Jensen 2018), as a key step to identify suitable units or groups for sourcing founder animals beyond arbitrary, human-created boundaries. Preserving and restoring such intraspecies diversity, though the path we indicated for caribou, is key to maintaining a species’ evolutionary potential, and in turn its critical ecological functions (Des Roches et al. 2021).
Data Availability
Caribou sample locations and population and DU range maps are available from the Government of British Columbia –see https://catalogue.data.gov.bc.ca/dataset/caribou-herd-locations-for-bc. Restrictions apply to the availability of this Species At Risk’s raw SNP data, which were used under license for this study’s academic personnel.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19(9):1655–1664. https://doi.org/10.1101/gr.094052.109
Allendorf FW, Hohenlohe PA, Luikart G (2010) Genomics and the future of conservation genetics. Nat Rev Genet 11(10):697–709. https://doi.org/10.1038/nrg2844
Araki H, Cooper B, Blouin MS (2007) Genetic effects of captive breeding cause a rapid, cumulative fitness decline in the wild. Science 318(5847):100–103. https://doi.org/10.1126/science.1145621
Arif IA, Khan HA, Shobrak M, Al Homaidan AA, Al Sadoon M, Al Farhan AH (2010) Measuring the genetic diversity of Arabian Oryx using microsatellite markers: implication for captive breeding. Genes Genet Syst 85(2):141–145
Armstrong DP, Seddon PJ (2008) Directions in reintroduction biology. Trends Ecol Evol 23(1):20–25. https://doi.org/10.1016/j.tree.2007.10.003
Ballou JD, Lacy RC (1995) Identifying genetically important individuals for management of genetic variation in pedigreed populations. In: Ballou JD, Gilpin M, Foose TJ (eds) Population management for survival and recovery: analytical methods and strategies in small population conservation. Columbia University Press, New York, pp 76–111
Bergerud AT (1988) Caribou, wolves and man. Trends Ecol Evol 3(3):68–72. https://doi.org/10.1016/0169-5347(88)90019-5
Bradley M, Neufeld L (2012) Climate and management interact to explain the decline of woodland caribou (Rangifer tarandus caribou) in Jasper National Park. Rangifer 32:183–191. https://doi.org/10.7557/2.32.2.2268
Brichieri-Colombi TA, Moehrenschlager A (2016) Alignment of threat, effort, and perceived success in North American conservation translocations. Conserv Biol 30(6):1159–1172. https://doi.org/10.1111/cobi.12743
Broman KW, Weber JL (1999) Long homozygous chromosomal segments in reference families from the Centre d’Etude du Polymorphisme Humain. Am J Hum Genet 65(6):1493–1500. https://doi.org/10.1086/302661
Bubac CM, Johnson AC, Fox JA, Cullingham CI (2019) Conservation translocations and post-release monitoring: identifying trends in failures, biases, and challenges from around the world. Biol Cons 238:108239
Caballero A, Villanueva B, Druet T (2021) On the estimation of inbreeding depression using different measures of inbreeding from molecular markers. Evol Appl 14(2):416–428. https://doi.org/10.1111/eva.13126
Carrier A, Prunier J, Poisson W, Trottier-Lavoie M, Gilbert I, Cavedon M et al (2022) Design and validation of a 63K genome-wide SNP-genotyping platform for caribou/reindeer (Rangifer tarandus). BMC Genomics 23(1):1–14. https://doi.org/10.1186/s12864-022-08899-6
Cavedon M, Gubili C, Heppenheimer E, vonHoldt B, Mariani S, Hebblewhite M et al (2019) Genomics, environment and balancing selection in behaviourally bimodal populations: The caribou case. Mol Ecol 28(8):1946–1963. https://doi.org/10.1111/mec.15039
Cavedon M, Poissant J, vonHoldt B, Michalak A, Hegel T, Heppenheimer E, Hervieux D, Schwantje H, Neufeld L, Polfus J, Schwantje H, Steenweg R, Musiani M (2022a) Population structure of threatened caribou in western Canada inferred from genome-wide SNP data. Conserv Genet 23:1089–1103
Cavedon M, vonHoldt B, Hebblewhite M, Hegel T, Heppenheimer E, Hervieux D et al (2022b) Genomic legacy of migration in endangered caribou. PLoS Genetics 18(2):e1009974. https://doi.org/10.1371/journal.pgen.1009974
Cavedon M, VonHoldt B, Hebblewhite M, Hegel T, Heppenheimer E, Hervieux D et al (2022c) Selection of both habitat and genes in specialized and endangered caribou. Conserv Biol. https://doi.org/10.1111/cobi.13900
Committee on the Status of Endangered Wildlife in Canada (COSEWIC) (2011) Designatable units for caribou (Rangifer tarandus) in Canada. Committee on the Status of Endangered Wildlife in Canada, Ottawa
Crandall KA, Bininda-Emonds OR, Mace GM, Wayne RK (2000) Considering evolutionary processes in conservation biology. Trends Ecol Evol 15(7):290–295. https://doi.org/10.1016/S0169-5347(00)01876-0
Crow JF, Kimura M (1970) An introduction to population genetics theory. Harper & Row Publishers, New York
Dayuan X, Yuanyuan Z, Zhibin C, Zhenyu Z, Ming C, Mengdi F et al (2022) Père David’s deer (Elaphurus davidianus) in China: population dynamics and challenges. J Resour Ecol 13(1):41–50. https://doi.org/10.5814/j.issn.1674-764x.2022.01.005
Der Sarkissian C, Ermini L, Schubert M, Yang MA, Librado P, Fumagalli M et al (2015) Evolutionary genomics and conservation of the endangered Przewalski’s horse. Curr Biol 25(19):2577–2583. https://doi.org/10.1016/j.cub.2015.08.032
Des Roches S, Pendleton LH, Shapiro B, Palkovacs EP (2021) Conserving intraspecific variation for nature’s contributions to people. Nat Ecol Evol 5(5):574–582. https://doi.org/10.1038/s41559-021-01403-5
Eacker DR, Hebblewhite M, Steenweg R, Russell M, Flasko A, Hervieux D (2019) Web-based application for threatened woodland caribou population modeling. Wildl Soc Bull 43(1):167–177. https://doi.org/10.1002/wsb.950
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361. https://doi.org/10.1007/s12686-011-9548-7
Ecological Stratification Working Group (1995) A national ecological framework for Canada. Agriculture and Agri-Food Canada, Research Branch, Centre for Land and Biological Resources Research and Environment Canada, State of the Environment Directorate, Ecozone Analysis Branch, Ottawa
Environment Canada (2014) Recovery strategy for the Woodland Caribou, Southern Mountain population (Rangifer tarandus caribou) in Canada. Species at Risk Act Recovery Strategy Series, Environment Canada, Ottawa
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164(4):1567–1587. https://doi.org/10.1093/genetics/164.4.1567
Fischer J, Lindenmayer DB (2000) An assessment of the published results of animal relocations. Biol Cons 96(1):1–11. https://doi.org/10.1016/S0006-3207(00)00048-3
Flanagan SP, Forester BR, Latch EK, Aitken SN, Hoban S (2018) Guidelines for planning genomic assessment and monitoring of locally adaptive variation to inform species conservation. Evol Appl 11(7):1035–1052. https://doi.org/10.1111/eva.12569
Flesch EP, Graves T, Thomson J, Proffitt K, Garrott R (2022) Average kinship within bighorn sheep populations is associated with connectivity, augmentation, and bottlenecks. Ecosphere 13:e3972
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180(2):977–993. https://doi.org/10.1534/genetics.108.092221
Forsman A (2014) Effects of genotypic and phenotypic variation on establishment are important for conservation, invasion, and infection biology. Proc Natl Acad Sci USA 111(1):302–307. https://doi.org/10.1073/pnas.1317745111
Foundations of Success and Parks Canada (2021) Assessing the Evidence for Adoption of a Conservation Breeding Strategy to Enable Recovery of Southern Mountain Caribou Populations in Jasper National Park, Canada.
Frankham R (2010) Where are we in conservation genetics and where do we need to go? Conserv Genet 11(2):661–663. https://doi.org/10.1007/s10592-009-0010-2
Frankham R, Ballou JD, Ralls K, Eldridge M, Dudash MR, Fenster CB et al (2017) Genetic management of fragmented animal and plant populations. Oxford University Press, Oxford
Galla SJ, Moraga R, Brown L, Cleland S, Hoeppner MP, Maloney RF et al (2020) A comparison of pedigree, genetic and genomic estimates of relatedness for informing pairing decisions in two critically endangered birds: Implications for conservation breeding programmes worldwide. Evol Appl 13(5):991–1008. https://doi.org/10.1111/eva.12916
Gibson G (2012) Rare and common variants: twenty arguments. Nat Rev Genet 13(2):135–145. https://doi.org/10.1038/nrg3118
Government of Alberta (2017) DRAFT: provincial woodland caribou range plan. https://open.alberta.ca/dataset/932d6c22-a32a-4b4e-a3f5-cb2703c53280/resource/3fc3f63a-0924-44d0-b178-82da34db1f37/download/draft-caribourangeplanandappendices-dec2017.pdf. Accessed 25 May 2023
Green DM (2005) Designatable units for status assessment of endangered species. Conserv Biol 19(6):1813–1820. https://doi.org/10.1111/j.1523-1739.2005.00284.x
Gruber B, Unmack PJ, Berry OF, Georges A (2018) dartr: An r package to facilitate analysis of SNP data generated from reduced representation genome sequencing. Mol Ecol Resour 18(3):691–699. https://doi.org/10.1111/1755-0998.12745
Gubili C, Mariani S, Weckworth BV, Galpern P, McDevitt AD, Hebblewhite M et al (2017) Environmental and anthropogenic drivers of connectivity patterns: a basis for prioritizing conservation efforts for threatened populations. Evol Appl 10(2):199–211
He X, Johansson ML, Heath DD (2016) Role of genomics and transcriptomics in selection of reintroduction source populations. Conserv Biol 30(5):1010–1018. https://doi.org/10.1111/cobi.12674
Hebblewhite M, White C, Musiani M (2010) Revisiting extinction in national parks: mountain caribou in Banff. Conserv Biol 24(1):341–344
Hoban S, Bertorelle G, Gaggiotti OE (2012) Computer simulations: tools for population and evolutionary genetics. Nat Rev Genet 13(2):110–122. https://doi.org/10.1038/nrg3130
Holsinger KE, Weir BS (2009) Genetics in geographically structured populations: defining, estimating and interpreting FST. Nat Rev Genet 10(9):639–650. https://doi.org/10.1038/nrg2611
Holt RD (1977) Predation, apparent competition, and the structure of prey communities. Theor Popul Biol 12(2):197–229. https://doi.org/10.1016/0040-5809(77)90042-9
IUCN, SSC (2013) Guidelines for reintroductions and other conservation translocations. Version 1.0. IUCN Species Survival Commission, Gland
Ivy JA, Miller A, Lacy RC, DeWoody JA (2009) Methods and prospects for using molecular data in captive breeding programs: an empirical example using parma wallabies (Macropus parma). J Hered 100(4):441–454. https://doi.org/10.1093/jhered/esp019
Kardos M, Luikart G, Allendorf FW (2015) Measuring individual inbreeding in the age of genomics: marker-based measures are better than pedigrees. Heredity 115(1):63–72. https://doi.org/10.1038/hdy.2015.17
Karney CFF (2013) Algorithms for geodesics. J Geod 87:43–55. https://doi.org/10.1007/s00190-012-0578-z
Kleiman DG (1989) Reintroduction of captive mammals for conservation. Bioscience 39(3):152–161. https://doi.org/10.2307/1311025
Kominakis A, Tarsani E, Hager-Theodorides AL, Mastranestasis I, Gkelia D, Hadjigeorgiou I (2021) Genetic differentiation of mainland-island sheep of Greece: implications for identifying candidate genes for long-term local adaptation. PLoS One 16(9):e0257461. https://doi.org/10.1371/journal.pone.0257461
Laikre L (1999) Hereditary defects and conservation genetic management of captive populations. Zoo Biol 18(2):81–99. https://doi.org/10.1002/(SICI)1098-2361(1999)18:2%3c81::AID-ZOO1%3e3.0.CO;2-2
Luikart G, England PR, Tallmon D, Jordan S, Taberlet P (2003) The power and promise of population genomics: from genotyping to genome typing. Nat Rev Genet 4(12):981–994. https://doi.org/10.1038/nrg1226
McDevitt AD, Mariani S, Hebblewhite M, Decesare NJ, Morgantini L, Seip D et al (2009) Survival in the Rockies of an endangered hybrid swarm from diverged caribou (Rangifer tarandus) lineages. Mol Ecol 18(4):665–679. https://doi.org/10.1111/j.1365-294X.2008.04050.x
McLoughlin PD, Dzus E, Wynes BOB, Boutin S (2003) Declines in populations of woodland caribou. J Wildl Manag 67:755–761
McNay RS, Lamb CT, Giguere L, Williams SH, Martin H, Sutherland GD, Hebblewhite M (2022) Demographic responses of nearly extirpated endangered mountain caribou to recovery actions in central British Columbia. Ecol Appl 32:e2580. https://doi.org/10.1002/eap.2580
Miller W, Wright SJ, Zhang Y, Schuster SC, Hayes VM (2010) Optimization methods for selecting founder individuals for captive breeding or reintroduction of endangered species. Pac Symp Biocomput 15:43–53. https://doi.org/10.1142/9789814295291_0006
Mills LS, Allendorf FW (1996) The one-migrant-per-generation rule in conservation and management. Conserv Biol 10(6):1509–1518. https://doi.org/10.1046/j.1523-1739.1996.10061509.x
Nei M (1972) Genetic distance between populations. Am Nat 106(949):283–292
Ochoa A, Storey JD (2021) Estimating FST and kinship for arbitrary population structures. PLoS Genet 17(1):e1009241. https://doi.org/10.1371/journal.pgen.1009241
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20(2):289–290. https://doi.org/10.1093/bioinformatics/btg412
Parks Canada Agency (2022a) Jasper National Park Caribou Program Progress Report 2017–2020. Jasper National Park, Parks Canada Agency
Parks Canada Agency (2022b) Proposal for consultation: conservation breeding strategy to rebuild small caribou populationpopulations in Jasper National Park. Jasper National Park of Canada, Parks Canada Agency
Pelletier F, Réale D, Watters J, Boakes EH, Garant D (2009) Value of captive populations for quantitative genetics research. Trends Ecol Evol 24(5):263–270. https://doi.org/10.1016/j.tree.2008.11.013
Pembleton LW, Cogan NO, Forster JW (2013) St AMPP: An R package for calculation of genetic differentiation and structure of mixed-ploidy level populations. Mol Ecol Resour 13(5):946–952. https://doi.org/10.1111/1755-0998.12129
Pimm SL, Jenkins CN, Abell R, Brooks TM, Gittleman JL, Joppa LN et al (2014) The biodiversity of species and their rates of extinction, distribution, and protection. Science 344(6187):1246752. https://doi.org/10.1126/science.1246752
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38(8):904–909. https://doi.org/10.1038/ng1847
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575. https://doi.org/10.1086/519795
Ralls K, Sunnucks P, Lacy RC, Frankham R (2020) Genetic rescue: a critique of the evidence supports maximizing genetic diversity rather than minimizing the introduction of putatively harmful genetic variation. Biol Cons 251:108784
Robert A (2009) Captive breeding genetics and reintroduction success. Biol Cons 142(12):2915–2922. https://doi.org/10.1016/j.biocon.2009.07.016
Russello MA, Jensen EL (2018) Ex situ wildlife conservation in the age of population genomics. Population genomics: wildlife. Springer, Cham, pp 473–492
Salafsky N, Irvine R, Boshoven J, Lucas J, Prior K, Bisaillon JF et al (2022) A practical approach to assessing existing evidence for specific conservation strategies. Conserv Sci Pract 4(4):e12654. https://doi.org/10.1111/csp2.12654
Schweizer RM, Vonholdt BM, Harrigan R, Knowles JC, Musiani M, Coltman D et al (2016) Genetic subdivision and candidate genes under selection in North American grey wolves. Mol Ecol 25(1):380–402. https://doi.org/10.1111/mec.13364
Seddon PJ, Armstrong DP, Maloney RF (2007) Developing the science of reintroduction biology. Conserv Biol 21(2):303–312. https://doi.org/10.1111/j.1523-1739.2006.00627.x
Seddon PJ, Griffiths CJ, Soorae PS, Armstrong DP (2014) Reversing defaunation: restoring species in a changing world. Science 345(6195):406–412. https://doi.org/10.1126/science.1251818
Serrouya R, Paetkau D, McLellan BN, Boutin S, Campbell M, Jenkins DA (2012) Population size and major valleys explain microsatellite variation better than taxonomic units for caribou in western Canada. Mol Ecol 21(11):2588–2601. https://doi.org/10.1111/j.1365-294X.2012.05570.x
Serrouya R, Seip DR, Hervieux D, McLellan BN, McNay RS, Steenweg R et al (2019) Saving endangered species using adaptive management. Proc Natl Acad Sci USA 116(13):6181–6186. https://doi.org/10.1073/pnas.1816923116
Serrouya R, Bollefer K, Cook R, Gilbert S, Gill R, Legebokow C, MacBeth B, Schwantje H, Thacker C (2021) Final Report for Revelstoke Caribou Rearing in the Wild. Nelson: British Columbia Ministry of Forests, Lands, Natural Resource Operations and Rural Development
Slatkin M (1987) Gene flow and the geographic structure of natural populations. Science 236(4803):787–792. https://doi.org/10.1126/science.3576198
Solmundson K, Bowman J, Wilson PJ, Taylor RS, Horn RL, Keobouasone S, Manseau M (2020) Genomic islands of heterozygosity maintained across caribou populations despite inbreeding. bioRxiv. https://doi.org/10.1101/2020.12.29.424772
Soorae PS (ed) (2018) Global reintroduction perspectives, 2018: case studies from around the globe. IUCN-International Union for Conservation of Nature and Natural Resources, Gland
Taylor RS, Manseau M, Horn RL, Keobouasone S, Golding GB, Wilson PJ (2020) The role of introgression and ecotypic parallelism in delineating intraspecific conservation units. Mol Ecol 29(15):2793–2809. https://doi.org/10.1111/mec.15522
Taylor RS, Manseau M, Klütsch CF, Polfus JL, Steedman A, Hervieux D et al (2021) Population dynamics of caribou shaped by glacial cycles before the Last Glacial Maximum. Mol Ecol 30(23):6121–6143. https://doi.org/10.1111/mec.16166
Taylor RS, Manseau M, Redquest B, Keobouasone S, Gagné P, Martineau C, Wilson PJ (2022) Whole genome sequences from non-invasively collected caribou faecal samples. Conserv Genet Resour 14(1):53–68. https://doi.org/10.1007/s12686-021-01235-2
Tokarska M, Marshall T, Kowalczyk R, Wójcik JM, Pertoldi C, Kristensen TN et al (2009) Effectiveness of microsatellite and SNP markers for parentage and identity analysis in species with low genetic diversity: the case of European bison. Heredity 103(4):326–332. https://doi.org/10.1038/hdy.2009.73
Van Oort H, McLellan BN, Serrouya R (2011) Fragmentation, dispersal and metapopulation function in remnant populations of endangered mountain caribou. Anim Conserv 14(3):215–224. https://doi.org/10.1111/j.1469-1795.2010.00423.x
Vors LS, Boyce MS (2009) Global declines of caribou and reindeer. Glob Change Biol 15(11):2626–2633. https://doi.org/10.1111/j.1365-2486.2009.01974.x
Vucetich JA, Waite TA (2000) Is one migrant per generation sufficient for the genetic management of fluctuating populations? Anim Conserv 3(3):261–266. https://doi.org/10.1111/j.1469-1795.2000.tb00111.x
Wang J (2004) Application of the one-migrant-per-generation rule to conservation and management. Conserv Biol 18(2):332–343. https://doi.org/10.1111/j.1523-1739.2004.00440.x
Weckworth BV, Musiani M, McDevitt AD, Hebblewhite M, Mariani S (2012) Reconstruction of caribou evolutionary history in Western North America and its implications for conservation. Mol Ecol 21(14):3610–3624. https://doi.org/10.1111/j.1365-294X.2012.05621.x
Weeks AR, Sgro CM, Young AG, Frankham R, Mitchell NJ, Miller KA et al (2011) Assessing the benefits and risks of translocations in changing environments: a genetic perspective. Evolut Appl 4(6):709–725
Williams SE, Hoffman EA (2009) Minimizing genetic adaptation in captive breeding programs: a review. Biol Cons 142(11):2388–2400. https://doi.org/10.1016/j.biocon.2009.05.034
Williams SH, Steenweg R, Hegel T, Russell M, Hervieux D, Hebblewhite M (2021) Habitat loss on seasonal migratory range imperils an endangered ungulate. Ecol Solut Evid 2(1):e12039
Witzenberger KA, Hochkirch A (2011) Ex situ conservation genetics: a review of molecular studies on the genetic consequences of captive breeding programmes for endangered animal species. Biodivers Conserv 20(9):1843–1861. https://doi.org/10.1007/s10531-011-0074-4
Wright S (1931) Evolution in Mendelian populations. Genetics 16(2):97. https://doi.org/10.1093/genetics/16.2.97
Wright BR, Hogg CJ, McLennan EA, Belov K, Grueber CE (2021) Assessing evolutionary processes over time in a conservation breeding program: a combined approach using molecular data, simulations and pedigree analysis. Biodivers Conserv 30(4):1011–1029. https://doi.org/10.1007/s10531-021-02128-4
Yannic G, Pellissier L, Ortego J, Lecomte N, Couturier S, Cuyler C et al (2014) Genetic diversity in caribou linked to past and future climate change. Nat Clim Chang 4(2):132–137. https://doi.org/10.1038/nclimate2074
Acknowledgements
We would like to thank the Aseniwuche Winewak Caribou Patrol, Bevan Ernst, Leo DeGroot, Mike Klaczek, and Aaron Reid for their efforts for caribou monitoring and conservation management. We also thank the Italian PNRR project Spoke 4.4, Scenarios of Area-based conservation planning and management, for supporting MM’s line of research.
Funding
This work was supported by the Alberta Conservation Association, Alberta Innovates, Alberta Upstream Petroleum Research Fund, Canadian Association of Petroleum Producers, Canada's Oil Sands Innovation Alliance, Conoco-Phillips, Environment and Climate Change Canada's Canadian Wildlife Service, Exxon, fRI Research, Governments of Alberta and British Columbia, Natural Sciences and Engineering Research Council of Canada, Parks Canada Agency, and Teck Resources.
Author information
Authors and Affiliations
Contributions
Maria Cavedon designed the study (with Marco Musiani and Lalenia Neufeld), conducted lab analyses (with Anita Michalak), analyzed genomic data, and drafted the manuscript (with Marco Musiani, Anita Michalak, and Jocelyn Poissant). Laura Finnegan, Dave Hervieux, Agnes Pelletier, Jean L. Polfus, Helen Schwantje, Geoff Skinner, Robin Steenweg, and Caeley Thacker gathered genetic and/or biological samples. All authors critically reviewed drafts and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethics Approval
Research was conducted under research permits of Governments of British Columbia and Alberta, Parks Canada and University of Calgary. Approval was granted by the University of Calgary Animal Care Committee Studies AC16-0195 and AC20-0110.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cavedon, M., Neufeld, L., Finnegan, L. et al. Genomics of founders for conservation breeding: the Jasper caribou case. Conserv Genet 24, 855–867 (2023). https://doi.org/10.1007/s10592-023-01540-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-023-01540-3