Abstract
Glacial relict plants are often endangered because extant populations can be small, geographically isolated and persist in suboptimal environments, leading to increased clonality and reduced genetic diversity putting their survival at further risk. This study examines how restriction to interglacial refugia has impacted the genetic diversity and structure of the threatened Tasmanian palaeoendemic, Pherosphaera hookeriana W. Archer bis. This species is a poorly dispersed, dioecious conifer that, having once been a major component of Last Glacial vegetation, is now limited to 30 known populations. Genetic diversity and structure were assessed using fifteen nuclear and nine chloroplast SSRs in 23 populations representing the species’ entire range. Changes in distribution and abundance from the Last Glacial to present were investigated by examining the fossil record, approximate Bayesian computation (ABC) and species distribution modelling. Despite fossil and ABC based evidence for a postglacial bottleneck, species-level genetic diversity (He = 0.56 and Ne = 2.86) exceeded that of some conifers with far wider distributions. Significant genetic structure (Fst = 0.127, Jost’s D = 0.203) was present, with most populations dominated by distinct nuclear SSR genetic clusters and having unique chloroplast haplotypes. Unexpectedly, clonality plays only a small role in population level regeneration. Genetic diversity has likely been maintained due to dioecy, persistence in multiple parts of its range and extant populations being directly descended from proximate glacial populations. Protecting populations from the mounting threat of fire will remain crucial for the in situ conservation of P. hookeriana.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Most of the Quaternary was far colder and drier than the present with glacial conditions reaching a climax during the Last Glacial Maximum (LGM; 26.5 to 19 kya; Clark et al. 2009). Severe environmental changes associated with the transition between glacials to wetter and warmer interglacials resulted in major shifts in the distribution and abundance of biota, particularly at mid to high latitudes. Fossil evidence show that during glacials, cold adapted plants dominated the vegetation, including in lowland areas (Birks and Willis 2008). Almost all such plants remain extant today but with ranges restricted to refugia in mountainous regions or at high latitudes (Gentili et al. 2015). Such species can be at risk of extinction due to the small number and size of surviving populations and geographic isolation resulting in enhanced inbreeding, loss of genetic diversity and vulnerability to stochastic events (Jadwiszczak et al. 2012). In some cases, limited capacity for sexual reproduction may result in increased dependence on regeneration by clonal growth (Migliore et al. 2013).
Studies of cold-adapted plants in interglacial refugia have shown that genetic diversity of even highly outcrossing wind pollinated plants can be impacted by small population sizes and geographic isolation. For example, the glacial relict conifers, Picea martínezii, P. chihuahuana, and Pinus torreyana all harbour markedly low genetic diversity due to late Quaternary genetic bottlenecks (Ledig et al. 2000; Jaramillo-Correa et al. 2006; Whittall et al. 2010). In the latter two species, remaining populations are effectively completely genetically isolated.
The Tasmanian paleoendemic conifer Pherosphaera hookeriana W. Archer bis (Podocarpaceae), is a classic example of a once widespread cold-adapted species now confined to interglacial refugia. During the Last Glacial, the island of Tasmania was at least 4 °C (Fletcher and Thomas 2010) and plausibly 6–8 °C colder than present (Mackintosh et al. 2006). While the drier eastern half of Tasmania was mostly ‘glacial arid’, the wetter western half remained densely vegetated with fossil evidence of an extensive alpine shrubland dominated by P. hookeriana, Asteraceae, Chenopodiaceae, and Poaceae occurring down to near sea level and forest species persisting in scattered small refugia (Macphail 1979; Colhoun 1985). At the onset of postglacial warming 15–11 kya, this alpine shrubland declined abruptly and was replaced by expanding rainforest and sclerophyll species (Colhoun et al. 1991, 1999). The contraction of P. hookeriana was part of a longer-term decline of the species beginning in the Middle Pleistocene (460,000 years ago to present) as is documented by the species declining pollen abundance over the length of a sea core 220 km southeast of Tasmania (De Deckker et al. 2019).
Today P. hookeriana is a minor element of the Tasmanian vegetation confined to montane areas of central and southern Tasmania (Fig. 1), where it occurs in montane conifer heath communities (Jackson 1972) usually associated with the fire protection afforded by water bodies or boggy areas (Minchin 1983). The species is listed as vulnerable to extinction under Tasmanian Government legislation with only 30 confirmed populations (defined as discrete patches of individuals at least 1 km apart) and an estimated 20,000 individual plants (Threatened Species Sect. 2016). Despite recent extensions to the species’ known range (Rudman and Schahinger 2016), it remains the rarest of the Tasmanian mesic conifers (see Figure S1 in the Supporting Information (S1)).
While better understanding of the species’ distribution has been vital for the conservation of P. hookeriana, crucial gaps in knowledge of the species’ biology remain including: (1) whether individual plants are actually genetically distinct given the rarity of observations of sexual reproduction (Threatened Species Section, 2016); (2) whether the species’ long term decline, compounded by a sudden contraction at the onset of Holocene warming, caused bottlenecking, inbreeding and low genetic diversity; and (3) whether P. hookeriana has maintained high genetic connectivity between its interglacial refugia populations. Worth et al. (2017) showed that two Tasmanian palaeoendemic conifers that have ranges overlapping that of P. hookeriana, Athrotaxis cupressoides and Diselma archeri, have low genetic differentiation across their ranges, indicating efficient gene flow via wind disseminated pollen. However, it remains uncertain whether the rarer and more fragmented P. hookeriana has similarly efficient gene flow. Indeed, despite their wind pollination mechanism, some conifers with patchy distributions and evidence of past bottlenecking harbour significant genetic structure over even relatively short geographic distances (< 100 km) (Tsumura 2006; Worth et al. 2018).
In this study we use a multi-disciplinary approach to address these questions. Firstly, we use range-wide sampling and nuclear and chloroplast simple sequence repeat (SSR) data sets to: 1) characterize contemporary levels of genetic diversity and structure and 2) assess levels of clonality and inbreeding. Nuclear SSRs (nSSRs) are bi-parentally inherited, exhibit high levels of polymorphism and have been widely used in studies of conservation genetics and phylogeography (Hodel et al. 2016). On the other hand, the chloroplast genome, which is paternally inherited in Podocarpaceae (Wilson and Owens 2003), provides information independent of the nuclear genome. High variation and lower effective population size make chloroplast SSRs (cpSSRs) an ideal marker to investigate range-wide genetic processes of conifers (Vendramin et al. 2008). Secondly, a fossil record of the species from the Late Pleistocene to present (compiled for the first time in this study) combined with investigation of past demographic changes using approximate Bayesian computation and species distribution modelling were used to explore past changes in the distribution and abundance of the species, specifically during the abrupt climatic changes at the Last Glacial-Holocene transition. The outcomes gained from this study will be essential to effectively prioritize management of the species in the face of rising threats from dry lightning ignited fires and a drying climate.
Materials and methods
The species
Pherosphaera hookeriana is a dioecious, multi-stemmed shrub, usually between 5 cm- 1 m high but (very rarely) a small tree up to 5 m tall. The species’ range is well understood particularly due to search efforts in the last decade except in the Southern Ranges (~ 60 km south of the nearest known population) where the species has not been recorded since 1898. The genus diverged from other podocarps approximately 115 million years ago (Biffin et al., 2011) with the only other species in the genus, P. fitzgeraldii (F.Muell.) Hook.f., being endangered and restricted to waterfall ledge habitat in the Blue Mountains, New South Wales. The lifespan of P. hookeriana is unknown but may be many hundred years (Minchin, 1983). The unwinged seeds are gravity dispersed (Kirkpatrick and Bridle 2013) although dispersal by water may occur for plants at the edge of lakes or rivers (Threatened Species Section, 2016). The species occurs from 650 m above sea level (masl) to 1200 masl usually as part of alpine coniferous heath. However, in some rare lower elevation sites, it persists within forests dominated by cool temperate rainforest and sclerophyll species.
Sampling and nSSR genotyping
A total of 599 samples consisting of branchlets from adult plants were collected from 23 populations spanning the species’ entire known range (Fig. 1 and Table S1(S2)). Whether the individuals collected were male or female was not determined because reproductive structures were mostly not present at the time of sampling. We aimed to sample at least 20 samples per population to accurately estimate genetic diversity and structure (Rodríguez-Peña et al. 2018). However, this was not possible in five of the 23 populations due to their small size and the need to avoid sampling related individuals. Thus, individual sampling was done at least 8 m apart except in the tiny, isolated population at Tahune Ridge, Frenchmans Cap, where all distinct individual clumps were sampled. Due to variation in sample size we tested whether sample size was correlated with four genetic indices (Ne, He, Ar and PAr). Significant relationships were found for Ne (P = 0.02) and Ar (P = 0.048) but not for He or PAr. However, these correlations were not significant when the population at Tahune Ridge was removed (P = 0.269 for Ne and P = 0.55 for Ar) (results not shown). This suggested that the variation in sample size has had little effect on genetic diversity values, except for Tahune Ridge.
DNA was extracted with the DNeasy 96 Plant Kit (Qiagen). Sixteen EST nSSR loci developed for P. hookeriana by Worth et al. (2018a) and for this study (Phero_4084: Forward primer 5`CGCCGTTAAGTGACACTCTG–3` and R primer 5`CTGTGAAAGGAGGGTGGGTA–3`) were used to genotype all samples. The PCR thermocycle and scoring followed Worth et al. (2018a). To check for genotyping accuracy, at least 39.8% of all samples were repeated for each of the four primer -pair multiplex sets. For information on the development of cpSSR loci for P. hookeriana and PCR conditions see S3.
Data integrity of nSSR loci
Allelic correlations among loci (i.e. linkage disequilibrium) and deviations from Hardy–Weinberg equilibrium were examined at the population level in Genepop v 4.2 (Raymond 1995) with sequential Bonferroni correction (P < 0.05). Null alleles were checked for using FreeNA (Chapuis and Estoup 2007). In the same program, unbiased estimates of Fst were also estimated (Chapuis and Estoup 2007). Because EST SSR loci are located in genes some loci may be under directional or balancing selection (Ellis and Burke 2007). The possibility that such selection of any EST-SSR loci could influence genetic parameters is low given that most genes do not experience positive selection (Tiffin and Hahn 2002) and EST-SSR loci have been found to display comparable genetic differentiation to genomic SSRs (Woodhead et al. 2005; Lind and Gailing 2013). To detect if any loci were potentially subject to selection the Fst outlier method, BayeScan v2.1 (Foll and Gaggiotti 2008), was used with each run repeated three times. The false discovery rate of BayeScan can be impacted by demographic history (e.g. island model, isolation by distance or expansion from refugia), therefore, we specified a more realistic prior odds of 1000 following the recommendation of Lotterhos and Whitlock (2014) and assessed each locus under Jeffreys’ scale of evidence (Jeffreys 1998). Existing genetic structuring can increase the number of false positives (Excoffier et al. 2009), therefore, the analyses were undertaken using genetic clusters detected in Bayesian Analysis of Population Structure (BAPS) version 6 (Corander et al. 2003) using all 16 nSSR loci, implementing ‘clustering of groups of individuals’ with a maximum K of 20 and ten independent runs. BAPS identified 15 clusters with probability of K = 15 being 0.994 versus the next best K = 16 with a probability of 0.005.
Tests for clonality and evaluation of range-wide genetic structure
The extent of clonality was investigated in Genalex 6.5 (Peakall and Smouse 2006) using the ‘find clones’ function and all 16 loci. If clonality was detected, one individual from each clone was retained for all further analyses.
To investigate range-wide genetic structure we used four population-based and two individual-based analyses. Firstly, for the nSSR data, population-level matrices of Fst, and Jost’s D (Jost 2008) were calculated using GenoDive 2.0b23 (Meirmans and Van Tienderen 2004) with the significance of pairwise population differentiation assessed using 999 permutations. Secondly, isolation by distance (IBD) at the nSSR level was tested for using a matrix of Fst/(1—Fst) and the logarithm of geographic distance (Rousset 1997) in Genalex 6.5 with 999 permutations. Thirdly, neighbour-joining trees were constructed in SplitsTree 4.14.4 (Huson and Bryant 2006) using a matrix of DA (Nei et al. 1983) for the nSSR data calculated in Populations 1.2.32 (Langella 2011) and Nei’s unbiased genetic distance calculated in GenAlEx 6.5 for the cpSSR data. Neighbour-joining trees are effective at displaying population structure under a range of evolutionary histories (Kalinowski 2009) while DA is able to obtain the correct tree topology under different SSR mutation models and demographic scenarios (Takezaki and Nei 1996). Lastly, BAPS was used to investigate genetic clustering for both nSSR and cpSSR datasets using the same method as described above. For individual based analyses of the nSSR dataset, we used the Bayesian clustering method of Structure ver. 2.3 (Pritchard et al. 2000) which assigns individuals into genetic clusters (K) based on allele frequencies. This analysis used the admixture model with sampling locations set as prior and correlated allele frequencies. The likelihood of K from 1 to 20 was estimated, with each analysis repeated 10 times. All runs involved 10,000 Markov chain Monte Carlo (MCMC) generations, after a burn-in period of 10,000 iterations. Structure Selector (Li and Liu 2018) was used to determine the best value of K by implementing the delta K method, which tends to identify the uppermost hierarchy of population structure (Evanno et al. 2005), and the estimates of Puechmaille (2016) (MedMedK, MedMeanK, MaxMedK and MaxMeanK) which have been shown to be accurately identify lower-level hierarchical structure (Puechmaille, 2016). However, because our results indicate an isolation by distance pattern (see Results) which can impact the results of Structure (Perez et al. 2018) we also undertook Discriminant Analysis of Principal Coordinates (DAPC) (Jombart et al. 2010). DAPC is a multivariate method that identifies and describes clusters of genetically related individuals using sequential K-means clustering (Jombart et al. 2010). The method has been shown to outperform Structure under a range of dispersal scenarios including when genetic structuring is clinal (Jombart et al. 2010). DAPC analyses were implemented in R using the ‘find.clusters’ command with max.n.clust = 20 and 50 PCs retained. To enable direct comparison with the results of Structure, DAPC plots of individual cluster assignment were drawn using the same number of inferred K from the Structure analysis. Both Structure and DAPC plots were drawn in Structure Plot v2 (Ramasamy et al. 2014).
Genetic diversity
For the nSSR dataset, the number of alleles (Na), number of effective alleles (Ne), observed (Ho) and expected heterozygosity (He) and F-statistics at both the locus and population level was assessed in GenAlEx 6.5. Rarefied values of allelic richness (Ar) and private allelic richness (PAr) were calculated in HpRare 1.1 (Kalinowski 2005) using a minimum sample size of nine. For the cpSSR dataset, locus and population level Na, Ne, haplotype diversity (h) and percentage of polymorphic loci were assessed in GenAlEx 6.5. Chloroplast SSR allele- and haplotype-based rarefied richness and private rarefied richness were calculated in ADZE 1.0 (Szpiech et al. 2008) with a minimum sample size of nine individuals. Chloroplast SSR haplotypes were determined in GenAlEx 6.5 and a median-joining haplotype network of all haplotypes was constructed in Network 5.0.1.1 (Bandelt et al. 1999) using maximum parsimony. Due to high levels of homoplasy of cpSSRs (Provan et al. 2001) a network was also constructed without singleton haplotypes.
Investigating past distributional change using the fossil record and species distribution modelling
The relatively low dispersal capability of P. hookeriana pollen suggests that the species fossil pollen record is informative about past distribution and abundance (Fletcher and Thomas 2010). Pollen records of P. hookeriana were searched for in the Indo-Pacific Pollen Database (Hope 2018), Colhoun and Shimeld (2012) and the Neotoma Paleoecology Database (Williams et al. 2018). Only dated pollen records were used and when calibrated ages were not available in the original publications, radiocarbon dates were calibrated using Calib rev7.1 (Reimer et al. 2013). For each core, the presence or absence of P. hookeriana and the maximum pollen frequencies within each of six time periods were recorded: pre-Last Glacial Maximum (126,000–26,500 years ago), Last Glacial Maximum (26,500–19,000), late Last Glacial (19,000–11,700), early Holocene (11,700–8,200), mid Holocene (8,200–4,200) and late Holocene (4,200–0). In addition, macrofossil records were also recorded based on literature searches and personal knowledge.
Species distribution models were developed following Worth et al. (2018b) using 19 bioclimatic (Hijmans et al. 2005) and four topographic variables to model the present and the Last Glacial Maximum distribution of suitable climate habitat for P. hookeriana. Species distribution models statistically associate the occurrence of a species across a landscape with environmental variables (usually climate) to model the fundamental niche in which the species can persist (Hutchinson 1957; Guisan et al. 2013). Species distribution models were estimated using the extendedForest package (Ellis et al. 2012) which implements the Random Forest algorithm (Breiman 2001) with conditional permutation variable importance (Strobl et al. 2008) to account for potential collinearity among predictor variables, using 142 occurrence records and a balanced number of randomly selected pseudo-absences. For details on the species distribution modelling methods used see S4.
Past demographic changes using ABC
The demographic simulation program fastsimcoal2 (Excoffier et al. 2013) was used to model the demographic history of individual P. hookeriana populations since the Last Glacial period. The sixteen populations were based on the results of BAPS analysis (see Results). Approximate Bayesian computation (ABC) is a flexible class of computation algorithm designed to perform complex model-based inferences, and can bypass exact likelihood calculations by using summary statistics and massive computer simulations (Cornuet et al. 2008; Csilléry et al. 2012). In this study, we assumed four alternative demographic models, Bottleneck, Constant, Decline and Expansion models (Figure S4 (S5)). The timing of the bottleneck at 15 kya was inferred from multiple fossil pollen-based studies documenting the species post-glacial decline while the expansion at 6 kya is based on increases in the species pollen abundance at mid-elevational range (900–1000 masl) at multiple sites (see Fossil data available on Dryad (https://doi.org/10.5061/dryad.g79cnp5mt)). For details on the ABC methods used see S5.
Results
Data integrity
The repeat genotyping found no genotyping errors. A total of 20 out of all 2552 population by locus-pair combinations were found to have significant allelic associations. Fourteen of these were due to a single locus combination (Phero_8380/Phero_12816) indicating that these loci are strong candidates for linkage, therefore, one locus of (Phero_12816) was removed from all analyses. No loci were found to have consistent significant deviations from HWE at the population level. In addition, no loci under potential selection were identified using BayeScan (Table S4 (S6)). For each of the 15 retained loci, the mean of the mean frequency of estimated null alleles per population was 2.2% and the mean across-loci Fst values estimated with and without null alleles were highly similar (0.126 versus 0.127) (Table S5 (S6)). These results indicated that the problem of null alleles was negligible for the 15 loci.
Clonality and range-wide genetic structure
Genotyping suggested low levels of clonality in this species. Of the 599 individuals sampled, 590 unique genotypes were observed and used for subsequent analysis. Three populations (1-JL, 4-SP and 7-GW1) had the same genotype in two individuals. The small and the isolated population 11-TR had three genotypes consisting of between 2–4 individuals (Table S1(S2)).
We observed significant genetic differentiation across the species’ range with range-wide Fst = 0.127 (population pairwise values ranging between 0.006–0.324) and, Jost’s D = 0.203 (0.008–0.45). Apart from three populations in the Nive River catchment, all population pairwise comparisons were significantly different (P < 0.01) (Table S6 (S7)). The test for IBD showed a significant association (P < 0.001) (Figure S5 (S8)). Both the neighbour-joining tree and Structure, when K = 2 as supported by the delta K method (Figure S6 (S9)), identified the greatest genetic divergence as being between the most northern part of the range (Mersey Rv., Du Cane Range and the Nive River catchment) versus the south (Mt Field and Snowy Range) (Fig. 2a and 3). Populations in the upper Franklin River, Frenchmans Cap and King William Range and Mt Anne were genetically intermediate between these two major lineages (Fig. 2a). Results based on DAPC for K = 2 showed very similar results. Lower hierarchical genetic differentiation was evident in Structure using the estimates of Puechmaille (2016) whereby an optimal K = 15 was identified in three of four estimates (Figure S7 (S9)). This value was close to the number of genetic clusters inferred using BAPS (K = 16) (Fig. 3). Thus, except for genetically close populations along the Nive River and Mt Field, most populations were dominated by unique genetic clusters (Fig. 3). Notably, populations at relatively low elevations for the species, including 21-LST (895 masl), 19-HR (875 masl) and 12-BG (658 masl), were genetically diverged from the nearest higher elevation population. DAPC results for K = 15 showed similar results but there was evidence for more admixture within the northern and southern parts of the range compared to the Structure analysis (Fig. 3).
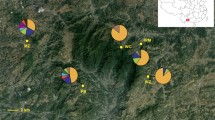
a Neighbour-joining tree based on Nei’s genetic distance (Da) calculated using 15 nuclear SSR loci. b Neighbour-joining tree of chloroplast SSR allelic variation at nine loci based on unbiased Nei’s genetic distance. The rarefied allelic richness of each population is indicated by the size of circles while colours of the circles correspond to the geographic regions. Population numbers and regions follow Table 1
Inference of population structure of Pherosphaera hookeriana using Structure and DAPC. The populations are ordered (from left to right) from north to south. Geographic regions and population numbers are indicated above while genetic clusters inferred by BAPS are indicated at the bottom. Asterisks indicate membership to a unique BAPS cluster while five populations in the Nive River catchment were inferred as belonging to BAPS cluster 1 and four in the Mt Field to BAPS cluster 2
Chloroplast SSRs showed significant genetic differentiation across the species range with PhiPT = 0.113 with similar north–south division to the nSSR data as shown by the neighbour joining tree (Fig. 2b). BAPS supported the existence of five genetic groups (Figure S8 (S10)). The northern and central part of the species range contained four geographically restricted groups: group 1 confined to the Nive River catchment; group 3, which apart from 4-SP, occurred in northern and central areas outside the Nive River catchment; group 4 in three disjunct populations (12-BG, 21-LST and 11-TR) all characterised by low chloroplast diversity (Table 2) and group 5 at population 8-LI. In contrast, the southern part of the species range (Mt Field, Snowy Range and Mt Anne) is dominated by a unique diverged group (group 2).
Genetic diversity
We observed 158 alleles across the 15 loci with a mean of 10.53 alleles per locus (see Table S7 (S11)). Observed heterozygosity values ranged between 0.45–0.65 with highest values at Mt Field and the Nive River catchment and the smallest values at 11-TR and 12-BG (Table 1). Population level Ar varied between 2.68 to 4.30 with high Ar values (> 4) observed across the species range (Table 1). The lowest Ar values were observed at a relatively low elevation population from Mt Field (19-HR), 11-TR and one site on the Nive River (5-GW2). Similar to Ar, populations with private alleles were distributed across the species range with highest values at 23-EP and 2-PA (5 private alleles each) and 22-LSU and 3-LM (4 each) (Table 1).
Overall inbreeding coefficient values ranged from -0.15 to 0.10 while the mean was close to zero (-0.001) indicating that there is no evidence for inbreeding at the species level. Twelve populations spread across the species range had negative values indicating an increased likelihood that alleles are not identical (Reichel et al. 2016) via a lack of inbreeding.
Genetic diversity at cpSSR loci
We observed 43 alleles in the nine cpSSR loci with the number of alleles per locus varying between 3–13 (Table S8 (S11)) and diversity (h) ranging from 0.15–0.39 (Table 2). Rarefied haploid allelic richness varied from 1.52 to 2.33 and showed high values across the species range (Table 2). Five populations contained private chloroplast alleles with three at 17-LDB and two at 21-LST (Table 2). Some non-SSR repeat type indels up to 25 bp in length at the G93 locus were highly geographically restricted including the two unique alleles at 21-LST and single unique alleles at Mt Field, upper Franklin River and King William Range. The combination of alleles at the nine loci resulted in 78 haplotypes of which 38 were singletons. The haplotype relationships were poorly resolved due to homoplasy with a mean of 2.51 independent origins for each allele inferred when using all 78 haplotypes (Figure S8 (S12)) and 1.63 when excluding singletons (result not shown). The population level frequency of unique haplotypes varied from 6.25% to 70% (mean = 24%) with highest values at 11-TR (70%), 12-BG (60%) and 21-LST (40%) (Table 2). Of all 78 haplotypes, only eight were found in over 10 individuals and, of these, five (haplotypes 23, 24, 33, 36, 54) were common across the species range while two were restricted to the north (haplotypes 10 and 60) and one (haplotype 1) mostly to the south. Rarefied haplotype richness varied between 2.9 in the 11-TR population to 7.95 at 18-WOM (mean = 5.71) with three other populations having values over seven (23-EP, 17-LDB and 6-TM) (Table 2).
The fossil record
The fossil record shows that the species was abundant in the West Coast lowlands, including the North West and South West coasts, during the pre-LGM with pollen percentage values of up to 42% at Darwin Crater (170 masl) and 39% at Henty Bridge (115 masl) (Fig. 4). The highest recorded abundance during this time was in central Tasmania at Clarence Lagoon (961 masl) with 58%. The absence of the species from the only two pre-LGM records in eastern Tasmania and in all records in later periods (except one Late Holocene site in the Tasmanian Midlands, which showed trace levels) suggests that P. hookeriana did not occur in the drier eastern half of Tasmania during the mid-late Pleistocene even during the cooler glacial period. LGM pollen sites are only available in the central West Coast region and at three sites within its current northern range. LGM pollen frequencies in the West Coast region were high reaching 38% at Governor Bog (180 masl) and 26% at both Crotty Dam (250 masl) and Henty Bridge while ranging between 9–14% in the northern part of its current range. There are no LGM records from southern Tasmania except for a single trace (1%) record from Ooze Lake (Fig. 4). High levels of P. hookeriana pollen were found widely in the many Late glacial pollen sites, both within the species’ current range and outside its current range on the West coast. In the early Holocene, pollen abundance declined dramatically with only trace levels found and almost all of these within or very near to the species current range. Sites in the West Coast area disappeared completely by 13 kya. In the mid-late Holocene, pollen abundance increased at higher altitude sites within, or near to, its northern and southern ranges.
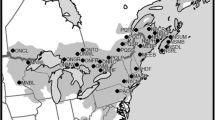
The fossil pollen record of Pherosphaera hookeriana mapped in six time periods: pre-Last Glacial Maximum (126,000–26,500 years ago), Last Glacial Maximum (26,500–19,000), late Last Glacial (19,000–11,700), early Holocene (11,700–8,200), mid Holocene (8,200–4,200) and late Holocene (4,200–0). Pale green circles represent the presence of fossil pollen of the species in the relevant time period; the radius of the circle is proportional the pollen abundance, while macrofossil records are indicated by stars. Small red x’s represent available pollen cores where Pherosphaera hookeriana was not recorded for the relevant time period. The current range of P. hookeriana is encircled by blue dashed lines. The hatched area in the LGM panel shows the extent of glaciers during the LGM (Barrows et al. 2002)
Species distribution modelling
The PCA reduced the climate variation across the distribution of P. hookeriana into three orthogonal axes that captured 96% of the variance in the climate dataset and were characterised by precipitation during the driest quarter, mean temperature of the wettest quarter, maximum temperature of the warmest period, and temperature and precipitation seasonality (Figure S2(S4)). The Random Forest (RF) model showed that variation in climate and topography could predict the contemporary suitable habitat for P. hookeriana (pseudo-\({\mathrm{r}}_{\mathrm{test}}^{2}\) = 0.91, AUC = 0.99), with precipitation seasonality and temperature seasonality being the most important predictors (Figure S3(S4)). A total of 814 points (93%) were predicted within suitable habitat (i.e. predict probability ≥ 0.9) with the remaining point records predicted with probabilities ranging between 0.73 and 0.89 except for one point with a predicted probability of 0.57. While the RF model predicted 100% of the area within the convex hull around all occurrence records (334 km2), it did predict an additional 2,583 km2 of suitable habitat (i.e. predicted probability ≥ 0.73) (Fig. 5). Most of this potential habitat was in central Tasmania with a small pocket located in NE Tasmania.
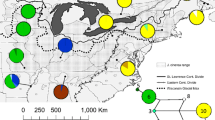
Predicted distribution of suitable habitat for Pherosphaera hookeriana under contemporary (1960–1990) and Last Glacial Maximum (22 kya; LGM) climates. The predictions for the LGM are means of three global circulation models. The convex hull around 814 occurrence records is shown by small black polygons. Black circles in the LGM represent the locations of P. hookeriana pollen in the fossil record, where the size is proportional to the frequency of pollen detected
Hindcasting the RF model to the LGM supported the persistence of suitable habitat for P. hookeriana in western and central Tasmania, including where the species is no longer present in the West Coast (Fig. 5). LGM fossil records had suitable habitat probabilities ranging from 0.14–0.57. The area of suitable habitat for P. hookeriana was found to have contracted by 62.2% since the LGM to the present.
Demographic changes inferred by ABC
ABC analysis supported all BAPS based populations as having undergone a post-Glacial bottleneck with posterior probability values exceeding 0.8 in all populations except for Lake Skinner Track (0.631), The Parthenon (0.785) and Lake Undine (0.770) (Table 3). For these populations, the second most likely demographic model was decline while no support was evident for either constant population size or expansion in any population.
Discussion
This study has revealed a markedly high genetic structure and diversity of Pherosphaera hookeriana, with clonality having a minor role. Unlike some other relictual conifer species, the species retains relatively high levels of genetic diversity. This is in spite of a long-term decrease in abundance since the Middle Pleistocene and fossil and ABC based evidence for an abrupt decline following the Last Glacial that involved the complete extirpation of a stronghold for the species to the west of its modern range (Fig. 4). Apart from some small, isolated populations, we observed high and evenly distributed genetic diversity across the species’ range. Strong geographic structuring of genetic diversity at both the nuclear and chloroplast level indicates that despite the species’ narrow range and wind pollination, past and contemporary gene flow has been insufficient to counter the effects of population geographic isolation. This contrasts with some other glacial relict conifer populations such as Pinus sylvestris in Spain where effective wind pollination has enabled genetic connectivity between isolated populations (Robledo‐Arnuncio et al. 2005). Major genetic differentiation between populations in the northern, central and southern part of the range as evidenced by both marker types indicate the persistence of populations in these regions over the Last Glacial to present. Lower hierarchical genetic differentiation and geographic restriction of nuclear and cpSSR alleles was also evident. Thus, outside of Mt Field and the Nive River catchment, most populations were dominated by distinct genetic clusters including geographically proximal populations occurring at different elevations. This, may be due local processes involving establishment via distinct colonization events and poor dispersal.
Maintenance of high diversity in interglacial refugia
The high level of genetic diversity of P. hookeriana is well demonstrated by comparison to other population genetic studies of conifers using EST SSRs and cpSSR markers. Thus, the species has higher levels of EST SSR alleles per locus and number of effective alleles and similar levels of heterozygosity as some other species including those with much wider distributions and/ or lower endangerment category (Table S9 (S13)). Similarly, P. hookeriana has comparable, or in some cases much higher, cpSSR diversity compared to other conifers, irrespective of whether the cpSSR primers have been developed specifically for the target species (Breidenbach et al. 2019; Wu et al. 2020) or using universal Pinaceae cpSSR primers (see Table 3 of Vendramin et al. (1999)). Overall, this comparison shows that the current rarity and Quaternary decline of P. hookeriana has not resulted in the species having markedly low genetic diversity versus other conifers and is in fact higher than some other palaeoendemic conifers like Glyptostrobus pensilis (Wu et al. 2020) and Sciadopitys verticillata (Worth et al. 2014).
Although explaining the factors underlying the genetic diversity of species is generally difficult (Leffler et al. 2012), several processes may account for the current high diversity of P. hookeriana. Importantly, clonality was found to be of minor importance to regeneration despite a lack of observations of sexual reproduction in the field. This contrasts with the co-occurring palaeoendemic conifer, Athrotaxis cupressoides, where using the same spaced sampling strategy, up to half the individuals in some populations were clonal (Worth et al. 2017). Given that P. hookeriana forms distinct clumps consisting of many separate stems, finer scale sampling within populations may reveal more clonality, but the low number of clones identified at the sampling distance used here show that sexual reproduction dominates the species population demography. The high genetic diversity must imply that the species has either maintained large effective population sizes since the Last Glacial despite the abrupt decline in the species apparent in the fossil record and supported by ABC or the genetic diversity of the species has been able to recover. Likely both factors have played an important role. Some mechanisms for maintaining genetic diversity include the presence of distinct northern, central and southern gene pools. This population subdivision increases the species effective population size (Wright 1943) and therefore the ability to retain genetic diversity. This effect has been inferred in another glacial relict conifer Picea omorika (Aleksić and Geburek 2014) which also displays high genetic diversity (Table S9 (S13)). Also, the species distribution models and fossil data suggest that P. hookeriana was present within its current range during the LGM (Fig. 5) including above the glacial tree line (Astorga 2016). This means that any postglacial migration likely occurred over short distances involving vertical migration from local glacial populations reducing the impact of genetic diversity loss during migration (Hewitt 1996). During postglacial expansion, allelic diversity may in fact have been augmented by new mutations which have a greater potential to become fixed in expanding populations (Burton and Travis 2008; McInerny et al. 2009). Lastly, the dioecy of P. hookeriana may help to maintain genetic diversity due to obligate outcrossing (Vandepitte et al. 2010), which is supported by the low values of inbreeding. Dioecy, along with the species long lived nature, may also reduce meta-population turnover which has been shown by both genetic theory and observations to reduce genetic diversity (Obbard et al. 2006). Further study is required to examine the role of dioecy in the genetic diversity of the species including any impact of skewed sex ratios in small populations (Vandepitte et al. 2010).
Conservation implications
This study has important implications for conserving P. hookeriana in an increasingly drier and fire-prone landscape. Firstly, the fact that the species occurs in only 11.45% of its modelled suitable climatic range confirms observation that the species is often absent from areas of apparently suitable habitat (Elliott 1948). This suggests that severe restrictions in mobility constrain the species range (Macphail et al. 2014) most likely related to its poor seed dispersal and the possible impact of past fires and possibly drought. In fact, fires have been active within the past and present range of P. hookeriana and have been impacting fire-intolerant palaeoendemic conifers from the Late Pleistocene (Jordan et al. 1991; Colhoun et al. 1993). The large area of unoccupied habitat suggests that there is wide scope for establishing insurance populations, especially in locations near waterbodies or in boggy areas. While some of the existing populations occur in areas that are predicted to be refugia for P. hookeriana and other palaeoendemic species into the future (Mokany et al. 2017) even minor spatial disconnection between future and current suitable habitats may require assisted migration for the survival of some populations. Secondly, this study shows that priorities for conserving P. hookeriana based on genetic data differ from those of other Tasmanian palaeoendemic species for which there have been detailed population genetic studies. Ensuring the in situ survival of long-unburnt high genetic diversity stands is of highest priority in Athrotaxis cupressoides and Diselma archeri, due to low geographic structuring of genetic variation and the deleterious impact of past fires on genetic diversity in these species (Worth et al. 2017). Genetic diversity in P. hookeriana, however, has been little impacted by post-European colonization possibly because the species range already occupies extreme fire refugia (Holz et al. 2020) and, unlike A. cupressoides and D. archeri there is little evidence of post-European range range-loss. Thus, conservation actions should instead prioritize protecting all existing populations, which are nearly all genetically distinct from one another, and, where possible, to source seed material for rehabilitation programs (for example, in the event of fire) from the nearest populations. Lastly, genetic bottlenecks apparent in the most isolated population at Tahune Ridge show that extreme reductions in population size could result in long-term loss of diversity which cannot be easily recovered via gene flow from outside populations.
Conclusions
Despite long-term decline and the severe decrease in abundance at the end of the Last Glacial, the species maintains higher levels of genetic diversity than many more widespread conifers. The high genetic diversity bodes well for the future conservation of the species providing extant populations can be protected from the threat of fire.
Data availability
The draft whole chloroplast genome of P. hookeriana, nuclear and chloroplast SSR datasets in GenAlEx format and the fossil data are available in Dryad (https://doi.org/10.5061/dryad.g79cnp5mt) and from the corresponding author upon request. Auxiliary data (i.e. species distribution data and climate layers) can be accessed from published work and websites referenced in the Material and Methods section.
Code availability
Not applicable.
References
Aleksić JM, Geburek T (2014) Quaternary population dynamics of an endemic conifer, Picea omorika, and their conservation implications. Conserv Genet 15:87–107
Astorga GA (2016) Patterns of vegetation in south-central Tasmania: a view based on plant macrofossils.
Bandelt H-J, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Barrows TT, Stone JO, Fifield LK, Cresswell RG (2002) The timing of the Last Glacial Maximum in Australia. Quat Sci Rev 21:159–173
Birks HJB, Willis KJ (2008) Alpines, trees, and refugia in Europe. Plant Ecol Divers 1:147–160
Breidenbach N, Gailing O, Krutovsky K V (2019) Assignment of frost tolerant coast redwood trees of unknown origin to populations within their natural range using nuclear and chloroplast microsatellite genetic markers. bioRxiv Doi:https://doi.org/10.1101/732834
Breiman L (2001) Random forests. Mach Learn 45:5–32
Burton OJ, Travis JMJ (2008) Landscape structure and boundary effects determine the fate of mutations occurring during range expansions. Heredity 101:329–340
Chapuis M-P, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24:621–631
Clark PU, Dyke AS, Shakun JD et al (2009) The Last Glacial Maximum. Science 325:710–714
Colhoun EA (1985) Pre-last glaciation maximum vegetation history at Henty Bridge, western Tasmania. New Phytol 100:681–690
Colhoun EA, Shimeld PW (2012) Late-Quaternary vegetation history of Tasmania from pollen records. Terra Australis. ANU E Press, Canberra, Australia, pp 297–328
Colhoun EA, van de Geer G, Fitzsimons SJ (1991) Late glacial and Holocene vegetation history at Governor Bog, King Valley, western Tasmania, Australia. J Quat Sci 6:55–66. https://doi.org/10.1002/jqs.3390060107
Colhoun EA, Benger SN, Fitzsimons SJ et al (1993) Quaternary Organic Deposit from Newton Creek Valley, Western Tasmania. Aust Geogr Stud 31:26–38. https://doi.org/10.1111/j.1467-8470.1993.tb00648.x
Colhoun EA, Pola JS, Barton CE, Heijnis H (1999) Late Pleistocene vegetation and climate history of Lake Selina, western Tasmania. Quat Int 57:5–23
Corander J, Waldmann P, Sillanpää MJ (2003) Bayesian analysis of genetic differentiation between populations. Genetics 163:367–374
Cornuet J-M, Santos F, Beaumont MA et al (2008) Inferring population history with DIY ABC: a user-friendly approach to approximate Bayesian computation. Bioinformatics 24:2713–2719
Csilléry K, François O, Blum MGB (2012) abc: an R package for approximate Bayesian computation (ABC). Methods Ecol Evol 3:475–479
De Deckker P, van der Kaars S, Macphail MK, Hope GS (2019) Land-sea correlations in the Australian region: 460 ka of changes recorded in a deep-sea core offshore Tasmania. Part 1: the pollen record. Aust J Earth Sci 66:1–15
Elliott CG (1948) Studies of the life histories and morphology of Tasmanian conifers. University of Tasmania
Ellis JR, Burke JM (2007) EST-SSRs as a resource for population genetic analyses. Heredity 99:125–132
Ellis N, Smith SJ, Pitcher CR (2012) Gradient forests: calculating importance gradients on physical predictors. Ecology 93:156–168
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Hofer T, Foll M (2009) Detecting loci under selection in a hierarchically structured population. Heredity 103:285–298
Excoffier L, Dupanloup I, Huerta-Sánchez E et al (2013) Robust demographic inference from genomic and SNP data. PLoS Genet 9:e1003905
Fletcher M-S, Thomas I (2010) A quantitative Late Quaternary temperature reconstruction from western Tasmania, Australia. Quat Sci Rev 29:2351–2361
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180:977–993
Gentili R, Bacchetta G, Fenu G et al (2015) From cold to warm-stage refugia for boreo-alpine plants in southern European and Mediterranean mountains: the last chance to survive or an opportunity for speciation? Biodiversity 16:247–261
Guisan A, Tingley R, Baumgartner JB et al (2013) Predicting species distributions for conservation decisions. Ecol Lett 16:1424–1435
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58:247–276
Hijmans RJ, Cameron SE, Parra JL et al (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978
Hodel RGJ, Segovia-Salcedo MC, Landis JB et al (2016) The report of my death was an exaggeration: a review for researchers using microsatellites in the 21st century. Appl Plant Sci 4:1600025
Holz A, Wood SW, Ward C et al (2020) Population collapse and retreat to fire refugia of the Tasmanian endemic conifer Athrotaxis selaginoides following the transition from Aboriginal to European fire management. Glob Chang Biol 26:3108–3121
Hope G (2018) Indo-Pacific Pollen Database.
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Hutchinson GE (1957) Concluding remarks: population studies, animal ecology and demography. In: Cold Spring Harbor Symposia on Quantitative Biology. pp 22–415
Jackson WD (1972) Vegetation of the central plateau. In: Papers and Proceedings of the Royal Society of Tasmania. pp 61–86
Jadwiszczak KA, Drzymulska D, Banaszek A, Jadwiszczak P (2012) Population history, genetic variation and conservation status of the endangered birch species Betula nana L. in Poland. Silva Fenn 46:465–477
Jaramillo-Correa JP, Beaulieu J, Ledig FT, Bousquet J (2006) Decoupled mitochondrial and chloroplast DNA population structure reveals Holocene collapse and population isolation in a threatened Mexican-endemic conifer. Mol Ecol 15:2787–2800. https://doi.org/10.1111/j.1365-294X.2006.02974.x
Jeffreys H (1998) The Theory of Probability. OUP Oxford
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94. https://doi.org/10.1186/1471-2156-11-94
Jordan GJ, Carpenter RJ, Hill RS (1991) Late Pleistocene vegetation and climate near Melaleuca Inlet, south-western Tasmania. Aust J Bot 39:315–333
Jost LOU (2008) GST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Kalinowski ST (2005) hp-rare 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189
Kalinowski ST (2009) How well do evolutionary trees describe genetic relationships among populations? Heredity 102:506–513
Kirkpatrick JB, Bridle KL (2013) Natural and cultural histories of fire differ between Tasmanian and mainland Australian alpine vegetation. Aust J Bot 61:465–474
Langella O (2011) Populations 1.2. 32 CNRS UPR9034.
Ledig FT, Bermejo-Velázquez B, Hodgskiss PD et al (2000) The mating system and genic diversity in Martinez spruce, an extremely rare endemic of Mexico’s Sierra Madre Oriental: an example of facultative selfing and survival in interglacial refugia. Can J For Res 30:1156–1164
Leffler EM, Bullaughey K, Matute DR et al (2012) Revisiting an old riddle: what determines genetic diversity levels within species? PLoS Biol 10:e1001388
Li Y, Liu J (2018) StructureSelector: A web-based software to select and visualize the optimal number of clusters using multiple methods. Mol Ecol Resour 18:176–177
Lind JF, Gailing O (2013) Genetic structure of Quercus rubra L. and Quercus ellipsoidalis EJ Hill populations at gene-based EST-SSR and nuclear SSR markers. Tree Genet genomes 9:707–722
Lotterhos KE, Whitlock MC (2014) Evaluation of demographic history and neutral parameterization on the performance of FST outlier tests. Mol Ecol 23:2178–2192
Mackintosh AN, Barrows TT, Colhoun EA, Fifield LK (2006) Exposure dating and glacial reconstruction at Mt. Field, Tasmania, Australia, identifies MIS 3 and MIS 2 glacial advances and climatic variability. J Quat Sci 21:363–376
Macphail MK (1979) Vegetation and climates in southern Tasmania since the last glaciation. Quat Res 11:306–341. https://doi.org/10.1016/0033-5894(79)90078-4
Macphail MK, Sharples C, Bowman D et al (2014) Coastal erosion reveals a potentially unique Oligocene and possible periglacial sequence at present-day sea level in Port Davey, remote South-West Tasmania. Pap Proc R Soc Tasmania 148:43–59
McInerny GJ, Turner JRG, Wong HY et al (2009) How range shifts induced by climate change affect neutral evolution. Proc R Soc B Biol Sci 276:1527–1534
Meirmans PG, Van Tienderen PH (2004) GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794
Migliore J, Baumel A, Juin M et al (2013) Surviving in mountain climate refugia: new insights from the genetic diversity and structure of the relict shrub Myrtus nivellei (Myrtaceae) in the Sahara Desert. PLoS ONE 8:e73795
Minchin PR (1983) A comparative evaluation of techniques for ecological ordination using simulated vegetation data and an integrated ordination-classification analysis of the alpine and subalpine plant communitites of the Mt. University of Tasmania, Field Plateau, Tasmania
Mokany K, Jordan GJ, Harwood TD et al (2017) Past, present and future refugia for Tasmania’s palaeoendemic flora. J Biogeogr 44:1537–1546
Nei M, Tajima F, Tateno Y (1983) Accuracy of estimated phylogenetic trees from molecular data. J Mol Evol 19:153–170
Obbard DJ, Harris SA, Pannell JR (2006) Sexual systems and population genetic structure in an annual plant: testing the metapopulation model. Am Nat 167:354–366
Peakall ROD, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Resour 6:288–295
Perez MF, Franco FF, Bombonato JR et al (2018) Assessing population structure in the face of isolation by distance: Are we neglecting the problem? Divers Distrib 24:1883–1889
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1111/j.1471-8286.2007.01758.x
Provan J, Powell W, Hollingsworth PM (2001) Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol 16:142–147. https://doi.org/10.1016/S0169-5347(00)02097-8
Puechmaille SJ (2016) The program structure does not reliably recover the correct population structure when sampling is uneven: subsampling and new estimators alleviate the problem. Mol Ecol Resour 16:608–627
Ramasamy RK, Ramasamy S, Bindroo BB, Naik VG (2014) STRUCTURE PLOT: a program for drawing elegant STRUCTURE bar plots in user friendly interface. Springerplus 3:1–3
Raymond M (1995) GENEPOP (version 1.2.): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Reichel K, Masson J-P, Malrieu F et al (2016) Rare sex or out of reach equilibrium? The dynamics of F IS in partially clonal organisms. BMC Genet 17:76
Reimer PJ, Bard E, Bayliss A et al (2013) IntCal13 and Marine13 radiocarbon age calibration curves 0–50,000 years cal BP. Radiocarbon 55:1869–1887
Robledo-Arnuncio JJ, Collada C, Alia R, Gil L (2005) Genetic structure of montane isolates of Pinus sylvestris L. in a Mediterranean refugial area. J Biogeogr 32:595–605
Rodríguez-Peña RA, Johnson RL, Johnson LA et al (2018) Investigating the genetic diversity and differentiation patterns in the Penstemon scariosus species complex under different sample sizes using AFLPs and SSRs. Conserv Genet 19:1335–1348
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rudman T, Schahinger R (2016) Pherosphaera hookeriana extension surveys: 2016 TWWHA Program Report: June 2016.
Strobl C, Boulesteix A-L, Kneib T et al (2008) Conditional variable importance for random forests. BMC Bioinformatics 9:307
Szpiech ZA, Jakobsson M, Rosenberg NA (2008) ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24:2498–2504
Takezaki N, Nei M (1996) Genetic distances and reconstruction of phylogenetic trees from microsatellite DNA. Genetics 144:389–399
Threatened Species Section (2016) Listing Statement for Pherosphaera hookeriana (mount mawson pine). Department of Primary Industries, Parks, Water and Environment, Tasmania
Tiffin P, Hahn MW (2002) Coding sequence divergence between two closely related plant species: Arabidopsis thaliana and Brassica rapa ssp. pekinensis. J Mol Evol 54:746–753
Tsumura Y (2006) The phylogeographic structure of Japanese coniferous species as revealed by genetic markers. Taxon 55:53–66. https://doi.org/10.2307/25065528
Vandepitte K, Honnay O, De Meyer T et al (2010) Patterns of sex ratio variation and genetic diversity in the dioecious forest perennial Mercurialis perennis. Plant Ecol 206:105–114
Vendramin GG, Degen B, Petit RJ et al (1999) High level of variation at Abies alba chloroplast microsatellite loci in Europe. Mol Ecol 8:1117–1126
Vendramin GG, Fady B, González-Martínez SC et al (2008) Genetically depauperate but widespread: the case of an emblematic Mediterranean pine. Evol Int J Org Evol 62:680–688
Whittall JB, Syring J, Parks M et al (2010) Finding a (pine) needle in a haystack: chloroplast genome sequence divergence in rare and widespread pines. Mol Ecol 19:100–114
Williams JW, Grimm EC, Blois JL et al (2018) The Neotoma Paleoecology Database, a multiproxy, international, community-curated data resource. Quat Res 89:156–177
Wilson VR, Owens JN (2003) Cytoplasmic inheritance in Podocarpus totara (Podocarpaceae). Acta Hortic 615:171–172
Woodhead M, Russell J, Squirrell J et al (2005) Comparative analysis of population genetic structure in Athyrium distentifolium (Pteridophyta) using AFLPs and SSRs from anonymous and transcribed gene regions. Mol Ecol 14:1681–1695
Worth JRP, Yokogawa M, Perez-Figueroa A et al (2014) Conflict in outcomes for conservation based on population genetic diversity and genetic divergence approaches: a case study in the Japanese relictual conifer Sciadopitys verticillata (Sciadopityaceae). Conserv Genet 15:1243–1257. https://doi.org/10.1007/s10592-014-0615-y
Worth JRP, Jordan GJ, Marthick JR et al (2017) Fire is a major driver of patterns of genetic diversity in two co-occurring Tasmanian palaeoendemic conifers. J Biogeogr 44:1254–1267
Worth JRP, Sakaguchi S, Harrison PA et al (2018) Pleistocene divergence of two disjunct conifers in the eastern Australian temperate zone. Biol J Lin Soc 125:459–474
Wright S (1943) Isolation by distance. Genetics 28:114
Wu X, Ruhsam M, Wen Y et al (2020) The last primary forests of the Tertiary relict Glyptostrobus pensilis contain the highest genetic diversity. Forestry 93:359–375
Acknowledgements
We would like to thank Matilda Brown, Pierre Feutry, Richard Pickup, Terry Reid, Laura van Galen, Mary Williams and Raymond Worth for their effort in collecting samples and the Department of Primary Industries, Parks, Water and Environment, Tasmanian Government, for providing collection permits (TFL16005 and TFL17332). We also thank to Andry Sculthorpe of the Tasmanian Aboriginal Centre for organising access to trawtha makuminya, and the Tasmanian Land Conservancy for access to Skullbone Plains. P.A.H. would like to acknowledge the support by the Australian Research Council Industrial Transformation Training Centre for Forest Value (IC150100004).
Funding
This work was funded by Forestry and Forest Products Research Institute, Tsukuba, Japan (grant no. 201430).
Author information
Authors and Affiliations
Contributions
JRPW, JRM and GJJ conceived the original idea; JRPW and JRM collected the data; JRPW, SS, and PAH analysed the data; and JRPW, JRM, PAH and GJJ wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest.
Ethical Approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Animal Research
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Worth, J.R.P., Marthick, J.R., Harrison, P.A. et al. The palaeoendemic conifer Pherosphaera hookeriana (Podocarpaceae) exhibits high genetic diversity despite Quaternary range contraction and post glacial bottlenecking. Conserv Genet 22, 307–321 (2021). https://doi.org/10.1007/s10592-021-01338-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-021-01338-1