Abstract
Analysis of the genetic diversity and structure of declining populations is critical as species and populations are increasingly fragmented globally. In the Great Basin Desert in particular, climate change, habitat alteration, and fragmentation threaten aquatic habitats and their endemic species. Tui chubs, including the Lahontan tui chub and Dixie Valley tui chub, (Siphateles bicolor ssp.) are native to the Walker, Carson, Truckee and Humboldt River drainages in the Great Basin Desert. Two populations, Walker Lake and Dixie Valley, are under threat from habitat alteration, increased salinity, small population sizes, and nonnative species. We used nine microsatellite markers to investigate the population genetic structure and diversity of these and nine other tui chub populations to provide information to managers for the conservation of both Walker Lake and Dixie Valley tui chubs. Genetic population structure reflects both historical and contemporary factors, such as connection with Pleistocene Lake Lahontan in addition to more recent habitat fragmentation. Dixie Valley was the most highly differentiated population (pairwise F ST = 0.098−0.217, p < 0.001), showed evidence of a past bottleneck, and had the lowest observed heterozygosity (Ho = 0.607). Walker Lake was not substantially differentiated from other Lahontan tui chub populations, including those located in different watersheds (pairwise F ST = 0.031−0.103, p < 0.001), and had the highest overall observed heterozygosity (Ho = 0.833). We recommend that managers continue to manage and monitor Dixie Valley as a distinct Management Unit, while continuing to maximize habitat size and quality to preserve overall genetic diversity, evolutionary potential, and ecological processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The field of conservation genetics operates on the premise that the genetic diversity of a population has consequences for its health and viability (Frankham 2002). Small populations in particular are at greater risk of extinction due to demographic (e.g. stochastic or catastrophic events Lande 1993) and genetic [inbreeding, increased genetic load, and reduced evolutionary potential, (Frankham 2002)] reasons, the combination of which can lead to what is known as an “extinction vortex”. Therefore managers are increasingly looking to genetic studies to quantify and monitor the genetic diversity of threatened populations as part of comprehensive management plans. Such genetic studies have the added benefit of helping managers prioritize conservation actions when resources are limited and populations are fragmented, such as those of aquatic species in arid regions.
The Great Basin desert, located primarily in Nevada, exhibits extreme aquatic endemism due to climatic and geological processes—Pleistocene lakes in the area receded around 10,000 ya allowing aquatic populations to speciate in isolation (Hubbs and Miller 1948; Hershler 1998; Smith et al. 2002). Many of these endemic species are vulnerable as a result of habitat alteration, water extraction, introduced species, population fragmentation, and climate change (Sada and Vineyard 2002). Sada and Vineyard (2002) reviewed the status of aquatic taxa in 2002 and found sixteen to be extinct, 78 declining, and only 28 to be somewhat stable, yet still facing threats. Efforts to preserve this threatened biodiversity may necessitate management actions that benefit from conservation genetics research (e.g. Frankham et al. 2002). Such actions can include, but are not limited to, founding refuge populations, habitat restoration, reintroductions and translocations, designating Evolutionarily Significant (Moritz 1994) or Management Units (Avise 2000), and genetic monitoring (e.g. Frankham et al. 2002).
The Lahontan tui chub (Siphateles bicolor obesa, S. b. pectinifer), and the closely related Dixie Valley tui chub (S. b. ssp.) are two Great Basin subspecies. Lahontan and Dixie Valley tui chubs are in the minnow family (Cyprinidae) and serve as a major prey item for native species such as migratory birds and the threatened native Lahontan cutthroat trout (Coffin and Cowan 1995). Tui chubs are typically abundant when found; archeological evidence suggests they were used as a food source for Native Americans (Raymond and Soble 1990). Like many desert fishes, tui chubs are adapted to harsh and variable environments due to their longevity (up to 35 years) and high fecundity. Tui chubs can mature at 2–3 years (Kucera 1978) and produce as many as ~70,000 eggs per year (Kimsey 1954). This life history enables them to survive unfavorable conditions and rapidly repopulate, much like the cui ui (Chasmistes cujus; Scoppetone et al. 1986) and native fishes in the Colorado River (e.g. Garrigan et al. 2002).
Two tui chub populations in particular, Walker Lake and Dixie Valley, are currently managed and monitored by Nevada Department of Wildlife (NDOW). Lahontan tui chubs in Walker Lake, one of the largest in both habitat area and historical census size, are threatened by upstream water extraction, which has dramatically increased salinity in the lake (Russell 1885; Stockwell 1994; Lopes and Allender 2009). Between 1882 and 2010, mostly as a result of upstream water diversions, the lake level dropped more than 150 ft and total dissolved salts have increased to 19,200 mg/L (Lopes and Allender 2009; NDOW 2011). This increase in salinity has extirpated all other fish species in the lake, leaving Lahontan tui chubs as the last self-sustaining fish species in the lake (NDOW 2011). The Dixie Valley tui chub, on the other hand, is endemic to Dixie Valley and restricted to a few small geothermal and cold springs on the Fallon Naval Air Station. Dixie Valley tui chubs are considered by some to be an un-described subspecies (Moyle 1995). Previous allozyme research (May 1999) and phylogenetic analysis using Cytochrome B (Harris 2000) found them to be genetically differentiated from Lahontan tui chubs, although geographically proximate locations such as Topaz Lake and Little Soda Lake were not included in either analysis. Dixie Valley tui chub are listed as a category 2 species by the USFWS (USFWS 1985), and managed as a species of concern by NDOW. Threats include introduced species, habitat alteration, small population size, and current and proposed geothermal power plant operations that may alter spring hydrology (Kris Urqhart, NDOW, personal communication).
For conservation of both Dixie Valley and Walker Lake tui chub populations, it is important to understand levels of differentiation between these and other populations in the Lahontan basin. Many of these populations are fragmented, recipients of conspecific translocations, or of unknown origin. In the event of further declines or increased threats, refuge populations of Walker Lake or Dixie Valley tui chubs can be founded as a source of backup genetic material if future augmentation or reintroductions are deemed necessary. Alternatively, if Walker Lake or Dixie Valley is extirpated, managers may introduce fish from genetically similar populations for reintroduction or restoration. By estimating population structure, estimating effective population size (N e ), and assessing overall genetic diversity, managers can identify suitable populations for reintroductions and use the information for future genetic monitoring (Schwartz et al. 2007).
The objectives of this genetic study are to (1) use microsatellite data to assess population genetic structure of Lahontan and Dixie Valley tui chubs; and (2) assess genetic diversity, test for genetic bottlenecks, and estimate effective population size N e of eleven Lahontan and Dixie Valley tui chub populations, and (3) discuss the conservation implications of our findings.
Methods
Sample collection
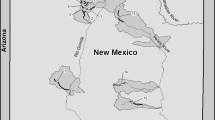
Between 10 and 50 individual Lahontan and Dixie Valley tui chub samples were collected from each of eleven locations: Topaz Lake, NV (TPZ), Spooner Lake, NV (SPN), Little Soda Lake, NV (LSL), Stillwater National Wildlife Refuge, NV (STW), Tahoe Keys, CA (TKS), Pyramid Lake, NV (PYR), East Fork Walker River, CA (EWR), South Fork Reservoir, NV (Humboldt River; SFH), Twin Lakes, CA (TWN), Walker Lake, NV (WLK), and Casey Pond, Dixie Valley, NV (DXV; Table 1; Fig. 1). These locations are in or are adjacent to the Walker, Carson, Humboldt, or Truckee River drainages. Each sample consists of a single 1 mm2 pelvic fin clip placed in a coin envelope and dried for storage. Whole genomic DNA was extracted using the Promega Wizard SV 96 Genomic DNA Purification System (Promega Corporation, Madison, WI).
Map of sample locations where Lahontan and Dixie Valley tui chubs were collected. Location codes are listed in Table 1. Gray shading indicates maximum area of Lake Lahontan during the late Pleistocene (Max Pleistocene). Stippled shading indicates other potential Pleistocene connections (Add. Pleistocene). Pleistocene Lake Lahontan layer taken from USGS (http://pubs.usgs.gov/mf/1999/mf-2323/)
Microsatellite genotyping
Samples were genotyped at five microsatellite loci from [Meredith and May (2002); Gbi-G13, Gbi-G38, Gbi-G39, Gbi-G79 and Gbi-G87)] and four from [Baerwald and May (2004); Cyp-G3, Cyp-G41, Cyp-G47, and Cyp-G48]. PCR reactions were conducted under conditions from (Chen 2013) and electrophoresed on an ABI 3730XL capillary electrophoresis instrument (Applied Biosystems, Carlsbad, California) after a 1:5 dilution with water. Peaks were scored using GeneMapper software (Applied Biosystems). We used the software MICRO-CHECKER 2.2.3 (Van Oosterhout et al. 2004) to detect and correct any unusual values in the data set and to look for significant homozygote excess that might indicate the presence of null alleles.
Population structure
Pairwise F ST values, a measure of the proportion of genetic diversity due to allele frequency differences among populations, were calculated with the software package Arlequin version 3.5 (Excoffier and Lischer 2010), using the option of exact tests of population differentiation. Significance of each pairwise F ST value was calculated in Arlequin using 5,000 bootstrap permutations. We used SPAGeDi v1.4 (Hardy and Vekemans 2002) to calculate R ST and its significance, and to test for significant difference between pairwise F ST and R ST values using 5,000 iterations of the allelic identity permutation test (Hardy et al. 2003). R ST is a measure of global variance in allele size, rather than allele identity, as in F ST. When R ST = F ST, one can assume that genetic differentiation is largely caused by genetic drift, rather than stepwise mutations (Hardy et al. 2003). We applied a sequential Bonferroni correction (α = 0.05) to comparisons between F ST and R ST to correct for multiple tests.
To determine the optimal number of genetic clusters (K) and to assign individuals to specific genetic clusters, we used STRUCTURE 2.3.3 (Pritchard et al. 2000). We performed three independent runs of K = 1–11, each with a burn-in period of 100,000 and 1,000,000 MCMC repetitions. We assumed no prior information, admixture, and correlated allele frequencies (Falush et al. 2003). We used STRUCTURE HARVESTER (Earl and vonHoldt 2012) to implement the DeltaK method (Evanno et al. 2005) to determine the optimal K. The three STRUCTURE outputs for each K (total N outputs = 33) were compiled with the software CLUMPP (Jakobsson and Rosenberg 2007) using the Greedy K algorithm (described in Jakobsson and Rosenberg 2007). CLUMPP aligns multiple replicate analyses of the same data set and creates an infile for the software DISTRUCT (Rosenberg 2004). DISTRUCT was used to create a graphical representation of the mean STRUCTURE outputs for a chosen K. We used GENETIX (Belkhir et al. 2003) to perform a factorial correspondence analysis (FCA) to visually depict the genetic relationships between individuals and populations and confirm STRUCTURE results.
Genetic diversity
The scoring of private alleles, calculations of allelic frequencies, observed heterozygosity (Ho), expected heterozygosity (He) were calculated in GDA (Lewis and Zaykin 2001). Deviations from Hardy–Weinberg equilibrium (HWE) and detection of linkage disequilibrium (LD) were calculated using Genepop ver. 4.0 (Raymond and Rousset 1995). A sequential Bonferonni correction (α = 0.05) was applied to determine the significance of multiple tests in LD and HWE calculations (Rice 1989). We used the software program HP-Rare (Kalinowski 2005b) to calculate allelic richness (A r ), a measure of genetic diversity, and private allelic richness (A P ), a measure of genetic distinctiveness using rarefaction to correct for sample size (Kalinowski 2004). F IS values and their significance (α = 0.05) were calculated using the software program SPAGeDi v1.4 (Hardy and Vekemans 2002) with 5,000 permutations.
Population bottlenecks and effective population size
We used two methods to estimate genetic bottlenecks. First, to detect the probability of a more recent population bottleneck (between 0.8 and 4.0 N e generations ago) we used the software program Bottleneck 1.2.02 (Piry et al. 1999), which uses simulations to determine if the population of interest has an excess of heterozygosity greater than expected at mutation-drift equilibrium, an indication of a genetic bottleneck (Cornuet and Luikart 1996). For this test we conducted the Wilcoxan sign-rank test over 5,000 iterations for two mutation models: the stepwise mutation model (SMM) and the two-phase model (TPM; Di Rienzo et al. 1994). TPM parameters were 12 % variance in non-stepwise mutations, 95 % stepwise and 5 % non-stepwise mutations (Estoup and Cornuet 1999; Spencer et al. 2000). The most likely mutation model is likely somewhere between the two models (Di Rienzo et al. 1994). The second bottleneck test is the M-ratio test, which calculates the ratio “M” of number of alleles (k)/allele size range (r). Populations where M is smaller than expected are likely to have undergone a severe genetic bottleneck (Garza and Williamson 2001). M was calculated in M_p_val (http://swfsc.noaa.gov/textblock.aspx?Division=FED&id=3298), using the following parameters: proportion of one-step mutations (ps) = 0.9, average size of non-one-step mutations (deltag) = 3.5, and ⊖ = 10. The ⊖ parameter is calculated with the equation ⊖ = 4Neµ, where µ is = 5.0 × 10−4 mutations/locus/generation (Estoup and Angers 1998). To determine the sensitivity of M to changes in ⊖, we calculated M varying ⊖ from 0.1 to 20, corresponding with N e values of 50–10,000. M was compared to the critical M (M c) calculated in Mcrit (http://swfsc.noaa.gov/textblock.aspx?Division=FED&id=3298), where 95 % of 10,000 simulations have M > M c. M c parameters were ps = 0.9, deltag = 3.5, and ⊖ = 10. As with calculating M, we varied ⊖ from 0.1 to 20 to determine sensitivity of M c to changes in ⊖.
To estimate the N e of individual sample locations, we used the corrected linkage disequilibrium method (Waples and Do 2008) implemented in NeEstimator (Do et al. 2014), using a Pcrit = 0.02 for populations where N > 25 samples and Pcrit = 0.03 where N ≤ 25 individuals. The assumptions of this method include random mating, isolation, selective neutrality of the markers used, no genetic structure within the population, and discrete generations. Do et al. (2014) performed simulations and found that the linkage disequilibrium method performed better than both the molecular coancestry method (Zhdanova and Pudovkin 2008) and the heterozygote excess method (Nomura 2008).
Results
One test in MICRO-CHECKER detected possible null alleles (p < 0.01) at Gbi-G87 in Twin Lakes. Out of 432 tests for HWE, 12 were significant after a sequential Bonferroni correction (p < 0.05). There is no consistent pattern across populations or loci for significant Hardy–Weinberg disequilibrium, so no loci were dropped. For LD in individual populations, after a sequential Bonferroni correction, there were 26 significant tests out of 550, but no significant LD tests across all loci and all populations. See Appendix I for allele frequencies in each sampling location.
Population structure
Computed pairwise F ST values ranged from 0.024 to 0.217 (Table 2). All F ST values were statistically significant (p < 0.001). The lowest pairwise F ST value was between Pyramid Lake and Tahoe Keys, which were once connected by the now dammed Truckee River. The highest pairwise F ST values were between Dixie Valley and other populations, and the lowest were between Walker Lake or Pyramid Lake and all other populations (Table 2). Pairwise R ST values ranged from −0.025 to 0.175. No R ST values were significantly greater than F ST values after a Bonferroni correction.
The optimal K-value for the STRUCTURE analysis based on the Evanno et al. (2005) method is K = 3, with additional substructure at K = 8 (see Fig. 2 for plot of DeltaK). When K = 3, the main cluster includes Topaz Lake, Tahoe Keys, Pyramid Lake, Twin Lakes, Walker Lake, Stillwater, East Walker River, and South Fork Humboldt. The second cluster includes Little Soda Lake and Dixie valley, and Spooner Lake forms its own cluster (Fig. 3). When K = 8, additional substructure is evident, with seven locations forming distinct clusters (Topaz Lake, Little Soda Lake, Spooner Lake, Stillwater, Dixie Valley, East Walker River, and South Fork Humboldt). The eighth genetic cluster is comprised of Tahoe Keys, Pyramid Lake, Twin Lakes and Walker Lake (Fig. 4). The FCA analysis supports the STRUCTURE analysis where K = 3; Walker Lake, Tahoe Keys, Twin Lakes, Pyramid Lake, Topaz Lake, Stillwater National Wildlife Refuge, East Walker River and South Fork Reservoir form an overlapping cluster. Spooner Lake forms a distinct cluster, and Little Soda Lake and Dixie Valley form an overlapping cluster (Fig. 5).
STRUCTURE output with K = 3 showing population substructure among 11 sample locations. Blue cluster consists of Topaz Lake, Tahoe Keys, Pyramid Lake, Twin Lakes, Walker Lake, Stillwater, East Walker River and South Fork Humboldt. Yellow cluster is Little Soda Lake and Dixie valley, and the orange cluster is Spooner Lake. (Color figure online)
STRUCTURE output with K = 8 showing population substructure among 11 sample locations. Topaz Lake, Little Soda Lake, Spooner Lake, Stillwater, Dixie Valley, East Walker River, and South Fork Humboldt form independent clusters, while Tahoe Keys, Pyramid Lake, Twin Lakes and Walker Lake form a cluster
STRUCTURE results, the FCA analysis, and low to moderate pairwise F ST values indicate that there is a single main overlapping cluster (hereafter Walker-Pyramid cluster) the core of which consists of Tahoe Keys, Pyramid Lake, Walker Lake, and Twin Lakes. East Walker River, Topaz, South Fork Humboldt, and Stillwater NWR are differentiated from this core when substructure is examined, but group with it when K = 3.
Population genetic diversity
The average number of alleles per locus (NA) ranged from 5.44 to 17.11 (Table 3). Ho values ranged from 0.607 to 0.833, and He values ranged from 0.623 to 0.831 (Table 3). Walker Lake had the most private alleles (N = 16), while several populations had only one private allele. A r and A P were calculated with a minimum number of genes N = 10, the smallest sample size included in the analysis; A r varied from 3.80 to 6.22. Walker Lake had the highest private allelic richness (A P = 0.730), and Twin Lakes had the lowest (A P = 0.160) (Table 3). F IS values ranged from −0.011 to 0.124, three of which were significant (p < 0.05).
Population bottlenecks and effective population size
Only Spooner Lake showed evidence of a population bottleneck using the Hk test (TPM, p = 0.082). The M-ratio test also showed evidence of a more severe bottleneck in Spooner Lake, Dixie Valley, and Tahoe Keys (Table 4). N e values ranged from 35.9 to ∞, and had wide confidence intervals, often with ∞ as an upper limit (Table 3).
Discussion
In this study we analyzed the population genetic structure and diversity of 11 tui chub populations to measure the genetic distinctiveness of two populations of concern: Walker Lake and Dixie Valley. This analysis provides genetic data in the event of future monitoring, and can be used to recommend source populations for reintroduction in the event of extirpation of the Walker Lake or Dixie Valley populations of tui chubs.
Genetic structure
Both contemporary and historical factors, as well as relative habitat size and isolation have contributed to the population genetic structure of Lahontan-Dixie tui chubs today. All of the populations analyzed here were once connected by Lake Lahontan. At its last highstand 14,500–13,000 ya, Lake Lahontan covered much of Northwestern Nevada and collected waters from the Truckee, Carson, Walker, Humboldt, Susan and Quinn rivers (Benson 1991). At this time, the fish populations in these watersheds were connected, allowing gene flow. When Lake Lahontan receded, gene flow was restricted to connected inundated areas. Though Lake Lahontan has receded since ~13,000 ya, flood events may have reconnected populations more recently, allowing gene flow and reducing differentiation. This has been observed in the golden perch (Macquaria ambigua) in Australia (Faulks et al. 2010). As indicated by significantly greater F ST values versus R ST values, genetic drift rather than mutation may be the main cause of existing population differentiation (Hardy et al. 2003). It should be noted that Kalinowski (2005a) suggests that if the true F ST of a population is greater than 0.05, fewer than 20 individuals may be used, though when the F ST is very low (0.01), fewer than 20 may not be not sufficient for accurate estimation (Table 1).
Surprisingly, even when substructure is examined, Pyramid Lake and Walker Lake cluster together despite being separated by the Carson River drainage. Both Walker and Pyramid Lakes have the lowest pairwise F ST values between them and other populations, and both have among the greatest observed heterozygosity levels. One explanation for these results is the large sizes of both Pyramid (~49,000 ha) and Walker Lake (~13,000 ha) that likely supported very large population sizes over time since the Lakes were connected. Pyramid Lake and Walker Lake are the main remnants of Pleistocene Lake Lahontan, and it is possible their large population sizes resulted in less genetic drift. This would have allowed retention of a greater proportion of historical neutral genetic diversity from Lake Lahontan, resulting in both higher genetic diversity and lower differentiation between Pyramid and Walker Lakes today.
Dixie Valley is the most differentiated population of those sampled based on pairwise F ST values and structure analysis. This finding, in combination with previous genetic research (May 1999; Harris 2000) and the unique and isolated habitat in Dixie Valley (Garside and Schilling 1979), supports continued efforts to conserve Dixie Valley as a Management Unit. Little Soda Lake and Spooner Lake are also genetically distinct. Both Dixie Valley and Little Soda Lake are small locations (<200 m across). Dixie Valley is thought to be a natural population, while Little Soda Lake was stocked from an unknown source in 1931 (Kim Tisdale, NDOW, personal communication). Spooner Lake, created in 1927, is a relatively small impoundment (0.4 ha) thought to be a natural (unstocked) population. Though tui chubs are currently abundant in Spooner Lake and Little Soda Lake (Kim Tisdale, NDOW, personal communication), isolation, founder effect, genetic drift, and/or bottlenecks may have led to differentiation of this population.
Genetic diversity
The mean heterozygosity value of sampled tui chub populations is higher than the mean of most freshwater fishes (0.54, DeWoody and Avis 2000), though sample sizes from some locations were lower than recommended. Hale et al. (2012) recommends at least 25–30 individuals to accurately estimate allele frequencies using microsatellites, and for five of the populations, 25–30 individuals were not available (Table 1). Only Spooner Lake and Dixie Valley presented evidence of genetic bottlenecks, though it is possible that the sample sizes of some populations or the markers used do not provide enough power to detect bottlenecks (Peery et al. 2012). The significant positive F IS values in Little Soda Lake, Twin Lakes, and Pyramid Lake suggest increased inbreeding, though the sample size in Twin Lakes was too small to accurately calculate a F IS value. Our N e estimates with infinite confidence intervals are due to small sample sizes relative to true N e (Waples and Do 2010). Many of these populations, such as Walker and Pyramid Lakes, are thought to have census sizes in the hundreds of thousands (Kris Urqhart, NDOW, personal communication). A sample of only 30–50 individuals from such locations may not be enough to get a precise estimate, even if true N e is as low as ~1,000 (Waples and Do 2010). Given the overall wide confidence intervals and small sample sizes, we recommend N e be used in the context of additional genetic analysis and monitoring when considering management actions.
Both Walker and Pyramid Lakes are expected to have higher genetic diversity relative to other populations due to their larger population sizes and terminal locations. Large populations experience less genetic drift than small populations. In addition, terminal or downstream locations like Pyramid and Walker Lakes will accrue genetic diversity from their upstream populations, a pattern observed in Gambusia holbrooki (Hernandez-Martich and Smith 1997). Life history also plays a role in the strength of genetic drift. Populations of longer-lived organisms often contain multiple generations that act as a reservoir of diversity, making it more difficult to detect increasing genetic drift associated with declining populations (e.g. ornate box turtles, Kuo and Janzen 2004). Similar patterns have been observed in other inland fish species of concern, such as the copper redhorse (Moxostoma hubbsi) which has an estimated census size of 500 and low recruitment, yet has both high genetic diversity and N e (Lippe et al. 2006). Another example is the endangered razorback sucker (Xyrauchen taxanus), a species that has declined steeply over the last few decades with little evidence of successful recruitment. Most surviving razorback suckers are large, old adults, and the population has maintained high genetic variation (Dowling et al. 1996a, b; Garrigan et al. 2002; Dowling et al. 2005). Garrigan et al. (2002) suggests that this is due to the long generation time and the historically large and geographically wide range of the razorback sucker. As recently as the mid-20th century, there was probably a large population of razorback suckers in the Lower Colorado River (Hedrick 2004). Therefore, any decline in census number has not yet significantly reduced the genetic diversity of the remaining population, though if declines continue genetic diversity will be lost. Similarly, if the tui chub population in Walker Lake continues to experience low recruitment, it will ultimately lose genetic variation.
Conservation implications
Our analysis has different implications for the conservation of tui chub populations in Walker Lake and Dixie Valley. First, Dixie Valley is genetically differentiated from other tui chub populations, and has lowest genetic diversity, lowest estimated N e , and evidence of a genetic bottleneck. Small, isolated populations such as Dixie Valley are at increased risk of inbreeding depression and extinction by stochastic events (e.g. Gilpin and Soulé 1986; Caughley 1994; Frankham et al. 2002). Given these findings and known threats, we recommend continuing to manage Dixie Valley tui chub as a separate Management Unit (Avise 2000). We also recommend genetic monitoring, and maximizing habitat area and quality through control of emergent vegetation, managing grazing, and control of introduced species. If further declines of Dixie Valley tui chub occur, managers may consider founding one or more refuge populations of Dixie Valley tui chub. If founding refuge population(s) is deemed necessary, managers should follow recommendations of population size requirements that retain and sustain both short and long term evolutionary potential of both the founding and the refuge population (i.e., 50/500 rule; for discussion and clarification of the “rule” see Jamieson and Allendorf (2012), but also consider Lynch and Lande (1998), and Franklin and Frankham (1998)).
Walker Lake on the other hand, is not substantially differentiated from other sampled Lahontan tui chub populations, including those that are in other watersheds. However Walker Lake is a valuable population, with the greatest overall genetic diversity and an important ecological role as the last self-sustaining native fish in the Lake. The recovery of the threatened Lahontan cutthroat trout is dependent upon a healthy self-sustaining Walker Lake tui chub population. By preserving overall genetic diversity managers also preserve evolutionary potential, allowing populations to continue to adapt to changing environments, fill ecological roles, and conserve evolutionary processes.
References
Avise JC (2000) Phylogeography. The history and formation of species. Harvard University Press, Cambridge, MA
Baerwald MR, May B (2004) Characterization of microsatellite loci for five members of the minnow family Cyprinidae found in the Sacramento-San Joaquin Delta and its tributaries. Mol Ecol Notes 4:385–390
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (2003) GENETIX version 4.04, logiciel sous WindowsTM pour la genetique des populations. Laboratoire Genome, Populations, Interactions: CNRS UMR. 5000, Universite de Montpellier II, Montpellier, France
Benson LV (1991) Timing of the last highstand of Lake Lahontan. J Paleolimnol 5:115–126
Caughley G (1994) Directions in conservation biology. J Anim Ecol 63(2):215–244
Chen YZ (2013) Genetic characterization and management of the endangered Mohave tui chub. Conserv Genet 14:11–20
Coffin PD, Cowan WF (1995) Lahontan cutthroat trout (Oncorhynchus clarki henshawi) recovery plan. U. S. Fish and Wildlife Service, Portland
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Di Rienzo A, Peterson A, Garza J, Valdes A, Slatkin M, Freimer N (1994) Mutational processes of simple-sequence repeat loci in human populations. PNAS 91:3166–3170
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovendon JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective size (N e ) from genetic data. Mol Ecol Resour 14:209–214
Dowling T, Minckley W, Marsh P (1996a) Mitochondrial DNA diversity within and among populations of razorback sucker (Xyrauchen taxanus) as determined by restriction endonuclease analysis. Copeia 1996:542–550
Dowling T, Minckley W, Marsh P, Goldstein E (1996b) Mitochondrial DNA diversity in the endangered razorback sucker (Xyrauchen texanus): analysis of hatchery stocks and implications for captive propagation. Conserv Biol 10:120–127
Dowling T, Marsh P, Kelsen A, Tibbets C (2005) Genetic monitoring of wild and repatriated populations of endangered razorback sucker (Xyrauchen texanus, Catostomidae, Teleostei) in Lake Mohave, Arizona-Nevada. Mol Ecol 14:123–136
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Estoup A, Angers B (1998) Microsatellites and minisatellites for molecular ecology: theoretical and empirical considerations. In: Carvlho GR (ed) Advances in molecular ecology. NATO Science Series, IOS Press, Amsterdam, pp 55–86
Estoup A, Cornuet JM (1999) Microsatellite evolution: inferences from population data. In: Goldstein DB, Schlöterrer C (eds) Microsatellites: evolution and applications. Oxford University Press, Oxford, pp 49–65
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetic analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotyped data: linked loci and correlated allele frequences. Genetics 164(4):1567–1587
Faulks LK, Gilligan DM, Beheregaray LB (2010) Islands of water in a sea of dry land: hydrological regime predicts genetic diversity and dispersal in a widespread fish from Australia’s arid zone, the golden perch (Macquaria ambigua). Mol Ecol 19:4723–4737
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Franklin IR, Frankham R (1998) How large must populations be to retain evolutionary potential? Anim Conserv 1:69–73
Garrigan D, Marsh PC, Dowling TE (2002) Long-term effective population size of three endangered Colorado Fishes. Anim Conserv 5:95–102
Garside LJ, Schilling JH (1979) Thermal waters of Nevada: Nevada bureau of mines and geology. Bulletin 91:163
Garza JK, Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10(2):305–318
Gilpin ME, Soulé ME (1986) Minimum viable populations: processes of species extinction. In: Soule ME (ed) Conservation biology: the science of scarcity and diversity. Sinauer and Associates, Sunderland, pp 19–34
Hale ML, Burg TM, Steeves TE (2012) Sampling for microsatellite-based population genetic studies: 25 to 30 individuals per population is enough to accurately estimate allele frequencies. PLoS One 7(9):e45170
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hardy OJ, Charbonnel N, Freville H, Heuertz M (2003) Microsatellite allele sizes: a simple test to assess their significance on genetic differentiation. Genetics 163:1467–1482
Harris PM (2000) Systematic studies of the genus Siphateles (Ostariophysi: Cyprinidae) from western North America. PhD dissertation, Oregon State University
Hedrick PC (2004) Recent developments in conservation genetics. Forest Ecol Manag 197:3–19
Hernandez-Martich JD, Smith MA (1997) Downstream gene flow and genetic structure of Gambusia holbrooki (eastern mosquitofish). Heredity 79:295–301
Hershler R (1998) A systematic review of hydrobiid snails (Gastropoda: Risooidea) of the Great Basin, Western United States, Part I: Genus Pyrgulopsis. Veliger 41:1–132
Hubbs CL, Miller RR (1948) The zoological evidence: correlation between fish distribution and hydrographic history in the desert basins of Western United States. In: The Great Basin, with emphasis on glacial and postglacial times. Bulletin of the University of Utah, vol 38. pp 17–166
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Jamieson IG, Allendorf FW (2012) How does the 50/500 rule apply to MVPs? Trends Ecol Evol 27:578–584
Kalinowski ST (2004) Counting alleles with rarefaction: private alleles and hierarchical sampling designs. Conserv Genet 5:539–543
Kalinowski ST (2005a) Do polymorphic loci require large sample sizes to estimate genetic distances? Heredity 94:33–36
Kalinowski ST (2005b) HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5:187–189
Kimsey JB (1954) The life history of the tui chub Siphateles bicolor (Girard) from Eagle Lake, California. Calif Fish Game 40:395–410
Kucera PA (1978) Reproductive biology of the tui chub, Gila bicolor, in Pyramid Lake, Nevada. Great Basin Nat 38(2):203–207
Kuo C, Janzen F (2004) Genetic effects of a persistent bottleneck on a natural population of ornate box turtles (Terrapene ornata). Conserv Genet 5:425–437
Lande R (1993) Risks of population extinction from demographic and environmental stochasticity and random catastrophes. Am Nat 142(6):911–927
Lewis PO, Zaykin D (2001) Genetic data analysis: computer program for the analysis of allelic data. Version 1.0 (d16c) Free program distributed by the authors over the internet from http://lewis.eeb.uconn.edu/lewishome/software.html
Lippe C, Dumont P, Bernatchez L (2006) High genetic diversity and no inbreeding in the endangered copper redhorse, Moxostoma hubbsi (Catostomidae: Pisces): the positive sides of long generation time. Mol Ecol 15:1769–1780
Lopes TJ, Allender KK (2009) Water budgets of the Walker River basin and Walker Lake, California and Nevada: U.S. Geological Survey Scientific Investigations. Report 2009–5157:44p
Lynch M, Lande R (1998) The critical effective size for a genetically secure population. Anim Conserv 1:70–72
May B (1999) Genetic purity and subspecific status of the Dixie Valley tui chub. Report to the Department of the Navy, N68711-98-LT-80018
Meredith EP, May B (2002) Microsatellite loci in the Lahontan tui chub, Gila bicolor obesa, and their utilization in other chub species. Mol Ecol Notes 2:156–158
Moritz C (1994) Defining ‘Evolutionarily significant units’ for conservation. Trends Ecol Evol 9(10):373–375
Moyle PB, Yoshiyama RM, Williams JE, Wikramanayake ED (1995) Fish species of special concern in California. Final Report for Contract N. 2128IF to California Department of Fish and Game
Nevada Department of Wildlife (2011) Walker Lake Fishery Improvement Program. Final Report for FWS Cooperative Agreement No. 84240-6-J, 52 pp
Nomura T (2008) Estimation of effective number of breeders from molecular ancestry of single cohort sample. Evol Appl 1(3):462–474
Peery MZ, Kirby R, Reid BN, Stoelting R, Doucet-Beer E, Robinson S, Vasquez-Carillo C, Pauli JN, Palsboll PJ (2012) Reliability of genetic bottleneck tests for detecting recent population declines. Mol Ecol 21:3403–3418
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. Heredity 90:502–503
Pritchard JK, Stephens M, Donnelly PJ (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Raymond AW, Sobel E (1990) The use of tui chub as food by Indians of the Western Great Basin. J Cailf Great Basin Anthropol 12:2–18
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43(1):223–225
Rosenberg NA (2004) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Russell IC (1885) Geological History of Lake Lahontan, A Quaternary Lake of Northwestern Nevada. Monograph XI, Geological Survey, U.S. Department of the Interior, Government Printing Office, Washington, D. C
Sada DW, Vineyard GL (2002) Anthropogenic changes in biogeography of Great Basin Aquatic Biota. Smithson Contrib Earth Sci 33:277–293
Scoppettone GG, Coleman M, Wedemeyer GA (1986) Life history and status of the endangered cui-ui of Pyramid Lake, Nevada. U.S Fish and wildlife service. Fish Wildl Res 1:1–23
Schwartz MK, Luikart G, Waples RS (2007) Genetic monitoring as a promising tool for conservation and management. TREE 22(1):25–33
Smith GR, Dowling TE, Gobalet KW, Lugaski T, Shiazawa D, Evans RP (2002) Biogeography and timing of evolutionary events amont Great Basin fishes. In: R Hershler, DB Madsen, Curry DR (eds) Great Basin aquatic systems history, smithsonian contributions to earth sciences, vol 33. pp 175–234
Spencer CC, Neigel JE, Leberg PL (2000) Experimental evaluation of the usefulness of microsatellite DNA for detecting demographic bottlenecks. Mol Ecol 9:1517–1528
Stockwell CA (1994) The biology of Walker Lake, The University Report. Department of Biology, University of Nevada, Reno
U.S. Fish and Wildlife Service (1985) Endangered and threatened wildlife and plants: review of vertebrate wildlife; Notice of review. Federal Register 50 CFR Part 17
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotype errors in microsatellite data. Mol Ecol Notes 4:535–538
Waples RS, C Do (2008) LDNe: a program for estimating effective population size from data on linkage disequilibrium. Mol Ecol Resour 8:753–756
Waples RS, Do C (2010) Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl 3(3):244–262
Zhdanova OL, Pudovkin AI (2008) Nb_HetEx: a program to estimate the effective number of breeders. J Hered 99(6):694–695
Acknowledgments
The authors would like to thank Kathleen Fisch, Mariah Meek, Ben Sacks, Karrigan Bork, Molly Stephens, and three anonymous reviewers for valuable comments. We would also like to thank NDOW biologists Kim Tisdale, Karie Wright, and Kris Urqhart for samples, insight, and a greater understanding of tui chub populations. Funding for this project was provided by Nevada Department of Wildlife, Task order 84240-9-J002; CESU agreement 81332-5-G004.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
See Table 5.
Rights and permissions
About this article
Cite this article
Finger, A.J., May, B. Conservation genetics of a desert fish species: the Lahontan tui chub (Siphateles bicolor ssp.). Conserv Genet 16, 743–758 (2015). https://doi.org/10.1007/s10592-015-0697-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-015-0697-1