Abstract
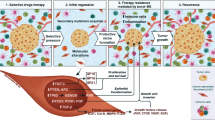
Cancer heterogeneity is a result of genetic mutations within the cancer cells. Their proliferation is not only driven by autocrine functions but also under the influence of cancer microenvironment, which consists of normal stromal cells such as infiltrating immune cells, cancer-associated fibroblasts, endothelial cells, pericytes, vascular and lymphatic channels. The relationship between cancer cells and cancer microenvironment is a critical one and we are just on the verge to understand it on a molecular level. Cancer microenvironment may serve as a selective force to modulate cancer cells to allow them to evolve into more aggressive clones with ability to invade the lymphatic or vascular channels to spread to regional lymph nodes and distant sites. It is important to understand these steps of cancer evolution within the cancer microenvironment towards invasion so that therapeutic strategies can be developed to control or stop these processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stanley P. Leong
Genetic mutation is the basis of cancer heterogeneity, which results in heterogeneous clones within the cancer population. Obviously, cancer grows within the cancer microenvironment consisting of “normal” stromal cells including infiltrating immune cells, cancer-associated fibroblasts, endothelial cells, pericytes, vascular and lymphatic channels [1]. The cancer microenvironment serves as a “natural selection” force, a term being borrowed from Darwin [2], over different cancer clones. Thus, within the cancer microenvironment, the process of cancer evolution is in play against the influence of the cancer microenvironment. In particular, the host immune system within the microenvironment may play an active role in its interaction with different clones within the cancer population resulting in the development of the “fittest clone” to proliferate and invade with acquisition of characteristics to spread to the lymph nodes and distant sites by lymphatic and vascular channels respectively. The genomic and molecular characteristics of cancer will enable us to appreciate genetic mutation relating to the spectrum of heterogeneity and molecular mechanisms that allow the cancer cells to invade and spread beyond the primary sites [3]. The relationship between cancer evolution and the cancer microenvironment is an important one.
In this review article, Isaac Witz, Orit Sagi-Assif and Sivan Izraely discuss the cross-talk between brain-metastasizing melanoma cells and the metastatic microenvironment. Again, using melanoma as a model, Jonathan Sleeman describes matrix-assisted autocrine signaling as a therapeutic target. In order to study cancer microenvironment, Brian Piening, Bernard A. Fox, and Carlo Bifulco use multiplex microscopy to analyze different cell populations within the cancer microenvironment. Cancer heterogeneity from genomic mutations is explained by Rachel Martini, Lisa Newman and Melissa Davis with respect to molecular, cellular, racial and environmental perspectives using breast cancer as a model system. Lauren Sanders, David Haussler, and Olena Vaske further analyze the genomic variations and mutations from the California Kids Cancer Comparison Project. Stanley Leong summarizes the concept and technique of CRISPR-Cas9 being extracted from the presentation of Christof Fellmann. Marlys Witte gives us the summary and future perspectives of the relationship between cancer evolution and its microenvironment.
The cross-talk between brain-metastasizing melanoma cells and the metastatic microenvironment
Isaac P. Witz, Orit Sagi-Assif and Sivan Izraely
A high proportion of melanoma, breast and lung cancer patients develop brain metastasis. These patients have poor survival outcomes and pose serious treatment challenges [4]. In view of the fact that neither genomic mutations nor epigenomic aberrations are significantly associated with the development of melanoma brain metastasis (MBM) [5], we hypothesized that the cellular and molecular brain microenvironment is involved in the formation of MBM.
The brain microenvironment contains unique cells such as neurons or astrocytes. These microenvironmental brain-specific cells confer upon the brain metastasizing cancer cells a different phenotype from that of cancer cells that metastasize to other organ sites [6, 7]. In order to develop unmet and critically needed novel treatment modalities for patients harboring brain metastasis, it is imperative to unravel the cellular and molecular mechanisms leading to such metastasis.
Aiming to identify and characterize signaling pathways that drive or inhibit MBM, we developed human to mouse xenograft MBM. Cutaneous and brain metastatic variants were generated from single human melanomas. Since each variant pair shared a common ancestry, we assume that transcriptomic, proteomic and other differences between these variants are linked to their different malignant phenotype. By identifying genes that are differentially expressed (up- or down-regulated) in MBM variants as compared to the matching cutaneous variants, we established a molecular signature of MBMs [8].
To identify pathways involved in the progression of cutaneous melanomas towards brain metastasis, we performed a proteomic profiling of four variant pairs of local and corresponding melanoma brain metastasis.
The expression level of several metastasis-associated proteins such as inflammatory cytokines, immune regulators, cell adhesion molecules and others, was higher in the four brain metastasis variants than in the corresponding cutaneous variants. Comparative analyses of eicosanoids and of multi parametric morphology yielded similar results. However, the four metastatic variants did not share any of the phenotypic molecular traits that characterize brain metastasis [9].
These results provided a strong indication for individually distinct patterns of metastasis-associated proteins. The results also highlighted the need for meta-analysis in the unraveling of the interactive complex pathways leading to melanoma brain metastasis.
Interactions of melanoma cells with skin, the microenvironment of the primary tumor as well as with the microenvironment of the brain drive or inhibit the progression of brain-metastasizing melanoma cells.
Below is an overview of studies on melanoma-intrinsic and brain microenvironmental drivers or inhibitors of brain metastasis employing the xenograft models described above. It should be stressed that the driver or inhibitor functions exerted by tumor-intrinsic or microenvironmental molecules refers to conditions described in the referenced studies. It is not unlikely that certain molecules may, under different circumstances, exhibit opposing functions to those described below [10].
Molecular drivers of melanoma progression
Several drivers of MBM were identified in our laboratory. The functional activity of two such drivers is described in some detail whereas others are mentioned in brief.
One of the genes whose expression was significantly higher in MBM variants than in matching cutaneous variants was Angiopoietin-like 4 (ANGPTL4). This gene whose expression is regulated by microenvironmental TGFβ1 [11] plays important but opposing roles in the progression of different cancers [12].
In vivo and functional in vitro studies employing the xenograft models described above indicated that ANGPTL4 promotes the malignancy phenotype of cutaneous melanomas [11]. In agreement with these results we found that ANGPTL4 expression is significantly higher in paired clinical specimens of melanoma metastases than in primary melanomas from the same patients.
Targeted migration of tumor cells to future metastatic cites is facilitated by the hijacking of chemokine receptors by metastasizing tumor cells thereby enabling chemotactic interactions with the corresponding ligands produced by and released from cells in the future metastatic microenvironment [13]. The expression of the chemokine receptor CCR4, another member of the molecular signature of MBM, was also higher in MBM variants than in matching cutaneous variants [14]. Similarly to ANGPTL4, the expression of CCR4 by melanoma cells was regulated by the brain microenvironment [15].
In view of the fact that MBM variants express higher levels of CCR4 than cutaneous variants and that the expression of CCR4 is significantly higher in paired clinical specimens of melanoma metastases than in samples of primary tumors from the same patients, we hypothesized that CCR4 ligands expressed in the brain interact with the CCR4-expressing melanoma cells thereby directing them to the brain.
The expression of the CCR4 ligands, CCL17 and CCL22 by brain endothelial cells, astrocytes and microglia is upregulated at the early stages of brain metastasis, preceding the infiltration of melanoma cells to the brain. In-vitro experiments indicated that CCL17 induced migration and transendothelial migration of melanoma cells. Melanoma cells over-expressing CCR4 generated a higher load of MBM than control cells. Blocking CCR4 with a small molecule CCR4 antagonist in-vivo, reduced MBM formation. All these results implicate CCR4 as a driver of melanoma brain metastasis [16]. Other molecular drivers of MBM characterized in our lab were: Extracellular cysteine protease inhibitor cystatin C (CysC) [17]. This protease inhibitor may either promote or confine tumor progression [18, 19]; IL-23 [20], a cytokine known to support growth of several types of tumors [21]; GM-CSF [22], a constituent of various anti-cancer immunotherapy trials [23]; Aldolase C (Izraely et al., in revision, Mol Oncol), a glycolytic enzyme with involvement in cancer progression [24]. In a collaborative study, the group of Dave Hoon demonstrated that GD3, a prominent cell surface ganglioside expressed by human cutaneous melanoma cells, plays a functional role in MBM formation [25].
Claudin-1 suppresses MBM
Claudin-1 (CLDN1) a tight junction protein, functions either as a tumor promoter or suppressor (or both). In some cancers, lower expression of CLDN1 is associated with cancer progression, while in others, loss of CLDN1 indicates restrained tumor progression [26].
Employing the melanoma xenograft models described above, we found that the expression of CLDN1 was lower in the brain‐metastasizing variants than in cutaneous variants from the same melanoma. In order to establish the function, if any, of CLDN1 downregulation/loss in melanoma brain metastasis, we transduced melanoma brain metastatic cells expressing low levels of CLDN1 with CLDN1 cDNA [27]. CLDN1‐overexpression eliminated the formation of micrometastasis in the brain. In sharp contrast, the ability of the CLDN1‐overexpresing cells to form lung micrometastasis was not impaired. The differential effect of CLDN1 overexpression on brain and lung metastasis is due to a differential expression of CLDN1 by brain and lung endothelial cells; the former cells expressed significantly higher levels of this tight junction component than the lung endothelial cells. The CLDN1 overexpressing melanoma cells adhered firmly to the brain endothelial cells via a hemophilic CLDN1–CLDN1 interaction [28], which blocked melanoma cell penetration into the brain. In view of the fact that lung endothelium expresses very low levels of CLDN1, such a hemophilic interaction does not occur allowing metastasizing melanoma to colonize the lungs.
Astrocytes, microglial and brain endothelial cells take part in the colonization and maintenance of human MBM
Astrocytes play important but contradictory roles in the homeostasis of the central nervous system. On the one hand they contribute to neuroprotection and on the other hand they may exacerbate neurological diseases [29].
Activated astrocytes are involved in the formation of MBM [30]. As reported above factors released from microenvironmental brain cells shape the malignant behavior of MBM [15]. Reciprocal interactions between astrocytes and melanoma cells were then identified as the source of some of these factors. Astrocyte‐derived factors up‐regulated the secretion of MMP2 from MBM whereas the melanoma cells up‐regulated the expression of the pro‐inflammatory cytokine IL‐23 in microglia. This cytokine enhanced melanoma cell invasion in vitro implying its function in MBM formation [20].
Microglia cells are the main immune cells of the brain [31]. Microglia and brain metastasizing melanoma cells are engaged in a dialogue which modifies gene expression patterns and cell signaling in, and cytokine secretion from both interacting cell types [32]. Brain metastasizing melanoma cells prompted significant morphological changes in microglia cells, enhanced their proliferation and migration and induced MMP-2 activation. Reciprocally, microglia cells generated phenotypic changes in melanoma cells and amplified their malignant phenotype. Specifically, microglia cells increased proliferation, migration and ability to penetrate the brain endothelium, and augmented MMP-2 activity of melanoma cells. Microglia also supported 3D spheroid formation by melanoma cells.
Cystatin C (CysC) (see above) is involved in melanoma-microglia interactions. Melanoma and microglia reciprocally upregulated CysC secretion from both cells. In vitro and in vivo experiments led to the conclusion that secreted CysC promotes melanoma brain metastasis [17].
Putting together the work on the bidirectional interactions between melanoma and microglia cells, indicates that microglia contributes to melanoma brain metastasis formation. The endothelium of the brain functioning as the blood brain barrier plays multiple (sometimes contradictory) roles in the generation of brain metastasis [10].
Interactions between neural progenitor cells and the vasculature form a neurovascular niche. Injured neurovascular niches, induced, for example by stroke, elicit repair processes that regenerate the neurovascular niche. These repair processes are mediated by cytokines and other growth factors [33].
A collaborative study with the Carmichael group at UCLA indicated that factors that promote melanoma brain metastasis and those mediating brain tissue repair share similar cellular processes. We identified a hitherto undescribed function of the stroke-induced regenerative neurovascular niche, predominantly endothelial cells, in shaping the brain metastatic microenvironment and in promoting melanoma brain metastasis [34].
Conclusion
Studying interactions between brain-metastasizing melanoma cells and their metastatic microenvironment reveals an intricate signaling web. Its comprehension is a prerequisite for the development of novel anti-metastasis therapy modalities.
Matrix-assisted autocrine signaling in melanoma as a therapeutic target
Jonathan Sleeman
During tumorigenesis, the progressive development of the tumor stroma serves to support the survival and growth of cancer cells [35]. Upon metastatic spread, cancer cells become divorced from this supportive stroma. At secondary sites, disseminated tumor cells (DTCs) encounter a new stromal microenvironment that determines whether they die, remain dormant or grow as metastases [36]. Microenvironments that support the survival and outgrowth of DTCs at secondary sites are termed metastatic niches [37]. In addition to cellular components such as myeloid-derived suppressor cells and cancer-associated fibroblasts, the constitution and conformation of the extracellular matrix (ECM) has emerged as an important component of metastatic niches [38]. Accordingly, the formation of metastatic niches is often associated with extensive remodelling of the ECM, including deposition of a range of structural and matricellular ECM components, crosslinking of ECM proteins by enzymes such as lysyl oxidases, and ECM degradation by a number of proteolytic enzymes such as MMPs [39].
Although particular characteristics of the ECM within metastatic niches have been associated with the initiation of metastases by DTCs, the mechanisms through which the ECM impacts on metastasis initiation are poorly understood. Enhanced matrix stiffness via ECM crosslinking and collagen deposition that is associated with metastatic niches results in mechanotransduction in tumor cells, which stimulates integrin signalling [40], the induction of EMT [41] and migration [42]. In the context of metastasis initiation, this may be relevant for suppressing dormancy [43]. Matricellular proteins such as tenascin C and periostin that are associated with metastatic niches have been implicated in regulating metastasis-initiating Wnt and Notch signalling [44, 45]. Laminin- or fibronectin-rich ECM can tether tumor-derived exosomes that bear appropriate integrin receptors, allowing them to fuse with and regulate organ-specific resident cells that contribute to developing metastatic niches, thereby shaping the organotropism of metastasis [46]. Despite these insights, more work is required to understand how the ECM can regulate metastasis initiation.
Malignant melanoma (MM) is the most lethal form of skin cancer. Cutaneous metastasis is a frequent and early event during the progression of MM, and represents the first site of metastasis formation for more than half of all MM patients [47]. In a recently published study [48] we used subcutaneous injection of melanoma cells together with specific ECM components as a model to investigate the initiation of cutaneous metastases. We found that a number of ECM components strongly increase the efficiency of cutaneous metastasis initiation. To understand how 3D ECM environments impact on gene expression that is associated with metastasis initiation, we compared the transcriptional profiles of melanoma cells growing in 2D and in 3D ECM environments. These data and subsequent validation revealed that a number of 3D ECM environments including Matrigel and collagen strongly induced Id1 and Id3 expression in melanoma cells.
Inhibitor of DNA binding 1 and 3 (Id1 and Id3) are transcriptional regulators whose expression is regulated by bone morphogenetic protien (BMP) and TGF-β signalling [49, 50]. They act as dominant negative inhibitors of basic helix-loop-helix (bHLH) transcription factors, by heterodimerising with them and preventing them from binding to DNA [51]. Expression of Id1 and Id3 has been implicated in tumor initiation and metastatic growth [52,53,54]. Accordingly, their expression correlates with poor prognosis for many types of cancer [55], including melanoma [56]. Loss of Id3 results in an impaired B-cell proliferation that can be rescued by ectopic overexpression of Id1 [57], indicating functional redundancy between Id1 and Id3.
To demonstrate a role for Id1 and Id3 expression in the initiation of cutaneous melanoma, we used CRISPR-Cas9 to disrupt Id1 and Id3 expression in two independent melanoma models. The concept and technique of CRISPR-Cas 9 are described below in this review article. Genetic ablation of Id1 and Id3 expression suppressed melanoma cell outgrowth and invasiveness in 3D ECM, and inhibited melanoma initiation and growth in vivo. Mechanistically, we found the physical properties of 3D matrix environments promote autocrine BMP signalling. Specifically, we used fluorescence correlation spectroscopy to demonstrate that specific 3D ECM microenvironments inhibit the diffusion of endogenously-produced BMPs. This serves to increase pericellular BMP concentrations, and thereby fosters autocrine BMP signalling. We have termed this mechanism matrix-assisted autocrine signalling (Fig. 1).
Matrix-assisted BMP autocrine signaling. Tumor cells produce low levels of BMPs that are insufficient to activate the cognate receptors on their surface under normal diffusion conditions. When the tumor cells are placed within particular 3D ECM microenvironments, the diffusion of the endogenously produced BMPs is significantly reduced, leading to pericellular accumulation of BMP proteins and subsequent receptor activation. Downstream signaling then induces expression of Id1 and Id3, fostering tumor initiation and metastasis
To leverage these findings therapeutically we synthesized and screened a custom chemical library. Thereby we identified a novel coumarin-like substance class that inhibits Id1 and Id3 expression. In proof of principle experiments, one of the most promising of these compounds was investigated further. Similar to the impact of genetic ablation of Id1 and ID3, the compound exerted a strong inhibitory effect on the outgrowth and invasiveness of melanoma cells grown in 3D ECM, and potently suppressed the initiation and growth of cutaneous metastasis in experimental animals. These data suggest that Id1 and Id3 represent promising therapeutic targets for melanoma, and identify new chemical inhibitors of Id1 and Id3 that can serve as lead compounds for drug development. More broadly, our findings suggest that the physical properties of a particular matrix environment can regulate metastasis formation through matrix-assisted autocrine signalling, and may be operative in other types of cancer as well. The mechanism of matrix-assisted autocrine signalling may additionally be relevant not only for BMP signalling, but also for other signalling molecules. Preconditions for this mechanism are that the signalling molecule is produced at low levels endogenously by cancer cells, that the cancer cells also express the cognate receptor, and that the diffusion of the signalling molecule is reduced by the 3D matrix environment.
Utility of multiplex microscopy to study the cancer microenvironment
Brian Piening, Bernard A. Fox and Carlo B. Bifulco
With the adoption of checkpoint inhibitors (CPI), immunoncology (IO) is rapidly transforming outcomes and therapeutic approaches in a variety of cancers, leading to durable responses in a significant subset of patients with previously-incurable disease, such as metastatic melanoma or metastatic non-small cell lung cancers [58,59,60]. These effects, driven by the inhibition of the PD1/PD-L1 axis, are believed to be ultimately mediated by the activation, in the context of the tumor microenvironment, of the cytotoxic effects of previously exhausted T-effector cells [61]. The clinical efficacy of CPI validates retrospectively a series of observations made in the pathology literature, going back all the way to the early 1990s, describing a strong correlation in multiple tumor types between the density of T cells present in the intratumoral and peritumoral microenvironment and outcomes. This body of work is best exemplified by the pioneering work of Jerome Galon, demonstrating that the quantification via digital pathology of the combined CD3+ and CD8+ T-cell density at the tumor margin and intratumor compartments (Immunoscore) predicts outcomes independently and significantly better than conventional locoregional pathologic stage in colon cancer [62]. These findings have been confirmed in a large multinational cohort of colon cancer patients [63], and beyond staging, are now being actively explored as predictors of response to adjuvant chemotherapy in stage III colon cancer, where emerging evidence suggests that the benefits from systemic chemotherapy are predicated on a functional immune system as assessed by a high Immunoscore [64]. Notwithstanding the available strong evidence supporting the importance of density and localization of T-cells in cancer outcomes and the requirement of effector T-cell tumor killing for IO efficacy, these findings have not yet been incorporated in the prediction of response to CPI, which is instead currently clinically either based on tumor intrinsic features, such as the tumor mutational burden [65] or microsatellite high status [66], or on a simple quantification of the expression of PD-L1, one of the components of the PD1/PD-L1 axis [67]. While these biomarkers have led to improved response rates in a variety of tumors, there are still a significant subset of biomarker-positive tumors that exhibit little to no response to CPI, a phenomenon likely reflective of the lack of comprehensive tumor microenvironment characterization with these current strategies. These paradigms may be soon changing as there are now multiple novel multiplexed immunohistochemistry/immunofluorescence (mIHC/IF) platforms that enable the co-visualization of large numbers of biomarkers in a spatial and morphological context, allowing for an accurate characterization of immune cell distributions in the tumor microenvironment and, beyond that, of their complex spatial relationships [68]. The current generation of mIHC/IF platforms solutions are based on a variety of technologies, including, among others, multiplexed tyramide fluorescent immunohistochemistry [69], repetitive cycling stain-stripping workflows [70], mass spectrometry-based detection of elemental isotopes [70,71,72] and oligoprobes hybridization based solutions [73], with the latter potentially enabling the concurrent multiplexed detection and spatial resolution of thousands of biomarkers. The emergent potential for the clinical utility of these novel technologies, here defined as the applicability of mIHC/IF platforms to empower clinical decision making processes, was first suggested in head and neck non-HPV-positive squamous SCC by the finding that the density of immunosuppressive populations, assessed by a tyramide mIHC/IF platform and identified by the expression of PD-L1 and FOXP3 in macrophages and T-cells, had, when colocalizing within a 30 micron radius of an effector CD8+ cells, a detrimental effect on the overall survival of these patients [74, 75]. Several research groups have now extended the application of these technologies beyond prognosis into the prediction of response to PD-1/PD-L1 checkpoint blockade. Their results, summarized in a recent review and meta-analysis, demonstrate that mIHC/IF is an independent predictor of response to CPI and both outperforms and synergizes with PD-L1 IHC, TMB and mRNA based signatures [66]. Despite these successes, a number of challenges mostly related to the inherent complexity of these novel technologies are currently slowing adoption into the clinic. Depending on the platform, multiplexing itself requires careful planning in order to avoid steric hindrances, secondary to the deposition of either chromogenic/fluorescence substrates or for antibody complexes as well as the loss of antigenicity due to harsh chemical treatments [68]. Careful attempts to address these multiplexing signal quantification reproducibility aspects with quantitative controls, such as exemplified in the T-cell activation marker proficiency panel (TAMPP) study, relying on a comparison of flow cytometric and multiplexed imaging technologies, are currently underway and may contribute to increase the robustness of in-situ antigen quantification approaches. Another existing challenge is the lack of a standardized strategy to guide the selection of regions of interest (ROI) for downstream high-resolution image analysis, a process that is often reliant, with a few exceptions such as the Immunoscore, on a manual and subjective human operator-driven process, potentially resulting in poor reproducibility. Possible strategies being explored to address these challenges take advantage of existing H&E images, as these can guide field selection, via co-registration of the IF and H&E digital slides, facilitating ROI selection, or fully or semiautomated selection tiling based procedures driven by image analysis derived features such as the density of immune cells in specific image tiles. The application of machine learning and specifically deep learning-based methodologies in histopathology is currently a significant area of focus, and such approaches have the potential to rapidly automate image analysis and classification (Fig. 2). There is also the potential to use H&Es to automate cell type recognition by leveraging data generated via mIF/IHC and the application deep convolutional neural networks (CNN). This process relies on the tagging of cell types based on mIHC/IF labels, and uses the generated labels to train a CNN to enable cell type recognition based on H&E features only. The results have the potential to provide an intrinsic quality control to multiplexed images, by anchoring them to observable features that are independent of complex experimental procedures and are embedded in an easily interpretable morphological background. In addition, this approach could enable an assessment of the tumor microenvironment that is scalable, as H&Es are routinely used in clinical diagnostic pathology around the world, with the availability of millions of historical slides, correlated to clinical outcomes and easily digitizable thorough high throughput FDA-approved scanning platforms. In summary, the development of multiple novel multiplexed imaging technologies as well as supporting algorithms for detection, segmentation and/or classification is expected to transform the assessment of the tumor microenvironment both in research and clinically, offering the opportunity to rationally select patients for CPI-based and other immunologic therapies, ultimately leading to better response rates and outcomes.
Breast cancer heterogeneity: molecular and cellular mechanisms are further complicated by racial and genetic diversity
Rachel Martini, Lisa Newman and Melissa Davis
Breast cancer is a collection of phenotypically distinct diseases, typically defined by hormone receptor status, or more recently gene signature profiling, which define intrinsic molecular tumor subtypes [76,77,78]. Disparities in breast cancer (BC) mortality emerged in a similar timeframe as the advent of targeted treatment therapies for hormone receptor positive (HR+) breast tumors [79], where we observed a distinct increase of mortality among Black or African American (AA) women compared to White or European American (EA) women, and an overall excess of 40% mortality among AA women is still observed today [80, 81]. Retrospectively, we can attribute divergence in mortality as an unmasking of this diversity in tumor phenotypes, where increase in AA mortality corresponds to disproportionately higher prevalence of HR− or Triple-negative BC (TNBC) tumors among AA women [79, 80, 82, 83]. HR− or TNBC tumors, which lack targeted treatments, are more molecularly aggressive, and have worst survival outcomes of all BC subtypes [80, 81, 84].
From a global perspective, African nations have some of the lowest incidence rates of BC among all nations yet suffer from highest BC mortality rates. The International Center for the Study of Breast Cancer Subtypes (ICSBCS) investigates BC disparities worldwide. The disparate outcomes observed in Africa are partially due to initial disease presentation occurring at later stages and limited resource access to standard treatment options [85]. ICSBCS has participated in capacity-building to increase HR testing and we find that African nations also have among the highest frequencies of TNBC, which is intrinsically associated with poor prognosis and increased mortality. West African and AA women report the highest frequencies of TNBC, compared to relatively lower prevalence of TNBC disease among East African and CA women [86,87,88]. While tracking the social and demographic history of ethnically diverse groups, our Oncologic Anthropology work allows us to characterize the genetic network differences among genetically distinct African populations. Intriguingly, their migration within and out of Africa correlates with the distribution of TNBC incidence and frequency. As ICSBCS also prospectively recruits newly diagnosed BC patients across our international sites, we are able to investigate the biological factors driving disparate outcomes. Using germline DNA testing, we have shown that quantified west African ancestry significantly increases risk for TNBC disease worldwide [79]. An individual’s genetic ancestry composition is determined through analysis of ancestral informative markers [89,90,91], or single nucleotide variants that are population private.
The genetic ancestry of African Americans in the US today is largely represented by European admixture with a majority of commonly shared African ancestry. The shared African ancestry is historically derived from ancestors who were enslaved and forced into the Americas over hundreds of years through the Trans-Atlantic Slave Trade (TST) [92, 93]. The activity of the TST had a profound impact across the African diaspora, which has health implications to this day. In the case of BC, that impact includes a distribution of risk alleles for aggressive tumor subtypes (i.e. TNBC) that are more prevalent in AA women as well as other women of African descent, globally. We sought to determine if shared African ancestry was a key to these observations. Our previous differential gene expression profile studies identified unique immune response in tumors among AA patients [94, 95], compared to EA patients. Intriguingly, this corresponds to other human evolutionary genetics studies that indicate distinct immune responses in infectious diseases, specifically the association of stronger immune response signals for cytokine production and inflammatory response, among individuals with West African ancestry [96]. We have identified an enrichment of immune response in AA, specifically within TNBC [97], which has vast implications in disease outcome and treatment response.
In our study of TNBC gene expression profiles, we compared between AA and EA women, using both self-reported race (SRR) and quantified genetic ancestry to determine differentially expressed genes [98] (Fig. 3). We found that the quantified ancestry approach yields genes with a more robust difference in gene expression. Our gene network analysis implicated known canonical cancer pathways (such as EGFR, TP52 and NFkB) as well as an enrichment of genes specifically related to immune response. This West African ancestry associated immune signature is strongly replicated in our preliminary analysis of a Pan-African dataset (not yet published), where increased signals observed among Ghanaian and AA women in the dataset appear to be decreased in Ethiopian and EA women. This indicates the potential of an evolutionarily enriched immune response to impact tumor biology, directed through shared West African ancestry among these women.
Quantified genetic ancestry reveals a more robust differentially expressed gene signature than self-reported race among TNBC cases. Proportional ancestry estimates are shown as a heatmap heading across all patient columns. Darker blue indicates higher ancestry within the genetic supergroups defined as European (EUR), East Asian (EAS), American Native (AMR), South Asian (SAS) and African (AFR). a Self-reported race (SRR) was used as a categorical variable to determine differentially expressed genes (DEGs) between European Americans (EA) and African Americans (AA). Over 1000 DEGs were identified using this approach and are shown in the unsupervised hierarchical cluster. b Quantified African ancestry was used as a continuous variable to determine DEGs associated with African ancestry. Approximately 150 DEGs were identified using this approach and are shown in the unsupervised hierarchical cluster, clearly filtering most differential gene patterns
Population-private mutations that have arisen in ancient populations in response to disease burden at different points through our history could be underlying the unique immune responses we have observed. Specifically, one such mutation is the Duffy-null allele, which arose across Sub Saharan Africa, and was fixed in populations in this region as it provided immunity towards certain malaria parasites [99], removing a route of entry for the pathogen into red blood cells [100]. While this mutation arose hundreds of years ago, it remains at almost 100% frequency across Sub Saharan Africa, and AAs are carriers of this allele in present day [101]. Global frequency of the Duffy-null allele follows similar global distribution as TNBC prevalence and BC mortality. We have found that this allele is significantly associated with TNBC risk among AA, after controlling for age and west African ancestry in a nested case-series analysis in our African ancestry enriched cohort [87].
The function of DARC and the Duffy-null allele in TNBC outcomes is still emerging. We know that Duffy-null is a promoter region variant of the Duffy Antigen Receptor for Chemokines/Atypical Chemokine Receptor 1 gene (DARC/ACKR1) and removes its expression from red blood cells (RBCs) specifically [100]. DARC/ACKR1 functions as an atypical chemokine receptor and is able to bind both structural classes of pro-inflammatory chemokines (i.e. CXCL and CCL) [102, 103]. Its primary function on RBCs is to modulate levels of chemokines in circulation through sequestering chemokines for degradation, returning homeostatic levels, thereby limiting duration of inflammation [103]. The known endothelial expression of DARC/ACKR1 functions in chemokine transcytosis, presenting chemokines to rolling leukocytes in circulation to aid in immune cell recruitment and diapedesis [103,104,105].
In the BC context, we were the first to show DARC/ACKR1 expression on tumor epithelial cells, its co-localization with pro-inflammatory chemokine ligands and corresponding increase of immune cells associated with higher DARC/ACKR1 levels [106]. In TCGA data, we found that AA and EA patients have broad variation of DARC/ACKR1 gene expression in BC tumors, AA had the highest prevalence of tumors with low DARC/ACKR1 expression. High DARC/ACKR1 expression was found to be significantly associated with better overall and relapse-free BC survival outcomes among all BC subtypes. We also observed that the Duffy-null allele regulates the availability of CCL2 in circulation, revealing that newly diagnosed BC patients who were homozygous for the Duffy-null allele showed significantly lower levels of CCL2 in circulation compared to heterozygotes or non-carriers of this mutation. This suggests that inflammation from tumors may be dampened in AA’s who carry the Duffy Null mutation.
To specifically investigate the association of DARC/ACKR1 expression with immune cell response, we used CIBERSORT RNAseq deconvolution methods and reported significant positive correlation between DARC/ACKR1 tumor expression and tumor-associated leukocyte abundance [106]. While bulk signals of immune cell infiltration are compelling, we are missing spatial acuity with the deconvolution method. Currently, we are utilizing imaging mass cytometry (IMC) methods to phenotypically define and quantify immune response. As we characterize the differences in spatial distribution of immune cell types, we will utilize the DARC/ACKR1 immuno-tumor phenotype in prognostics. Preliminary findings shown that DARC/ACKR1 positive tumors have more immune cell infiltration into the solid tumor space, compared to DARC/ACKR1 low tumors having more stromal compartmentalization of immune cells.
In summary, BC is a heterogeneous disease, where both tumor characteristics and patient race/ethnicity play an important role in prognostic outcomes of a BC diagnosis. With the expansion of our genomic toolkit, we have been able to identify potential biological drivers of BC disparities, with the goal to identify actionable targets for therapeutic development. These findings will have global impact, especially in the case of TNBC, where the African diaspora has increased burden and targeted treatments are not currently available. We have seen immune response biomarker differences between race/ethnic groups, and these signals appear to be driven by shared west African ancestry, as a consequence of evolutionary adaptation. Specifically, we have highlighted the Duffy-null allele, and further work done around the DARC/ACKR1 gene, as this chemokine receptor plays a significant role in immune cell recruitment and is also associated with West African ancestry and increased TNBC disease risk. As we advance in the field of precision medicine, it is imperative that through study conception and design, we plan for inclusion of diverse patient populations to fully characterize tumor heterogeneity across subtypes and racial/ethnic groups, to identify targets for treatment that are effective across all BC patients.
The California Kids Cancer Comparison Project
Lauren M. Sanders, David Haussler and Olena M. Vaske
Demonstrated clinical utility of large genomic datasets for individual childhood cancer patients
Worldwide genomic data sharing has come to the forefront of scientific focus as large consortia such as The Cancer Genome Atlas and the Human Brain Atlas have revealed the molecular underpinnings of human physiology and disease. Nowhere is the need for genomic data sharing more apparent than in rare and understudied diseases, including many childhood cancers and germline disorders. Because no one institution generates enough data from these rare diseases to fully understand them, it is vitally important to share these genomic data and make them public to the scientific community.
The study of childhood cancer provides a key example of the necessity of leveraging combined, publicly available genomic datasets into clinically translatable findings. Unlike adult cancers, childhood cancers are thought to arise from epigenetic or developmental aberrations during embryogenesis or early childhood [107]. As a result, pediatric cancers harbor lower overall DNA mutation rates than adult cancers, and recurrent genomic aberrations tend to be epigenetic in nature and impossible to target therapeutically [107, 108].
Nevertheless, despite the marked lack of upstream activating DNA mutations, pediatric cancer cell growth is driven by activated oncogenic pathways as in adult cancers. Targetable oncogenic gene and pathway activity can be detected through RNA sequencing (RNA-Seq) of cancer biopsy or resection samples [109, 110]. Due to the relative nature of RNA-Seq data, each individual sample must be compared against a background cohort to detect unusually highly expressed genes and pathways. For adult cancers, such comparative analysis often uses a background of normal tissue samples, such as the Genotype-Tissue Expression database (GTEx) [111]. However, the early developmental origin of most pediatric cancers makes it difficult to identify or source the appropriate normal tissue. For example, the ETV6-RUNX1 fusion positive subtype of acute lymphoblastic leukemia is thought to arise from B-cell progenitors during embryonic hematopoiesis, but the developmental stage and cancer cell of origin are as yet unknown [112].
To address this problem and maximize the clinical utility of childhood cancer genomic datasets, the Treehouse Childhood Cancer Initiative at UC Santa Cruz has generated a publicly available cancer compendium of RNA-Seq data from over 12,000 adult and pediatric tumors and 144 different tumor types [113]. The compendium contains 44 independent RNA-Seq datasets, all of which have been processed uniformly using the UCSC TOIL RNA-Seq pipeline to eliminate technical artifacts [114]. Included are the data from The Cancer Genome Atlas and the Therapeutically Applicable Research to Generate Effective Treatments program.
Treehouse has also developed a method to compare individual samples to a background cohort. This method, Treehouse Comparative Analysis of RNA Expression (Treehouse CARE), identifies genes with outlier expression in a single tumor sample as compared to a background cohort [109]. These genes are then used to identify significantly enriched gene sets with an existing drug or therapeutic targeting them. The analysis is performed against two different background cohorts: the “pan-cancer” analysis compares a child’s cancer to the entire Treehouse cancer compendium. The second analysis, “pan-disease”, compares the tumor to a subset of highly correlated tumors or tumors of the same disease type. The pan-disease analysis also aids in molecular subtyping of rare tumors by identifying most similar tumors through gene expression correlation. Treehouse partners with clinical sites and genomics trials to provide a report of CARE findings, including the outlier genes and enriched pathways, and the clinical characteristics of the tumors with highest correlation to the focus sample.
The development of this approach and the ability to compare within thousands of RNA-Seq cancer samples has led to the discovery of clinically actionable insights that would have been otherwise impossible to attain. A multi-center study of 144 childhood or young adult patients with relapsed, refractory or rare cancers found that the Treehouse CARE method identified actionable overexpressed genes or pathways in 68.8% of cases [109]. In 36.5% of cases, the actionable target was only identifiable through RNA analysis, and was not present in DNA variant analysis.
In a case of a child with relapsed sarcoma of the central nervous system, metastatic to the lungs, comparative RNA-Seq analysis revealed overexpression of JAK1 as compared to all cancers (pan-cancer analysis), and other sarcomas or other lung cancers (pan-disease analysis) [110]. This patient’s tumors had no actionable DNA variants, so the physician treated with Ruxolitinib, a JAK inhibitor. Within a week of Ruxolitinib initiation, the patient had dramatic improvement in energy level and resolution of several symptoms. Although most of his lung lesions remained stable, one lesion progressed after 5 months so Ruxilitinib was stopped and focal radiation was administered. Within 2 months of Ruxolitinib discontinuation, the other lung lesions progressed, and the patient’s family requested that Ruxolitinib be restarted for quality of life. The patient again showed dramatic improvement and a prolonged period of stable disease until dose reduction was required because of myelosuppression, and the patient passed away.
In a case of a young patient originally diagnosed with immature teratoma, comparative RNA-Seq analysis revealed that the top six most correlated tumors were all glioma. After additional histopathological analysis, the patient’s diagnosis was refined to gliomatosis peritonei, a rare tumor type involving mature glial tissue in the peritoneum which is difficult to identify via histopathological analysis alone [115].
These findings demonstrate the clinically translational value of sharing and combining genomic data for precision medicine trials. In the context of rare and difficult to treat childhood cancers, comparative RNA-Seq analysis using thousands of samples from various sources has aided in identifying therapeutics and refining diagnoses.
Application of CRISPR-Cas9 system in editing the genetic code
Stanley P Leong
This section of the review article has been extracted from the presentation by Christof Fellmann at the 2019 8th International Cancer Metastasis Congress in San Francisco.
The 2020 Nobel Prize for Chemistry was awarded to Emmanuelle Charpentier and Jennifer Duodna “for the development of a method for genome editing,” the CRISPR/Cas9 (Clustered Regularly-Interspaced Short Palindromic Repeats) genetic scissors [116]. This important topic was presented by Christof Fellmann, a colleague of Jennifer Duodna at the 2019 8th Cancer Metastasis Congress in San Francisco [117]. He was not able to write this section and this write-up has been extracted from his talk, Emerging Roles of CRISPR-Ca in Precision Oncology (available at: https://www.vumedi.com/video/emerging-roles-of-crispr-cas-in-precision-oncology/) with his permission. The mechanism of CRISPR-Cas in editing the genetic code has been discovered by Charpentier and Duodna based on the adaptive immunity in bacteria and archaea against plasmids and viral infections [118,119,120,121]. CRISPR-Cas being associated with adaptive immune systems are found in roughly 50% of bacteria and 90% of archaea [122].
Like the vertebrate adaptive immunity, CRISPR immunity functions similarly by generating memory of previous infections to launch a rapid and effective response during reinfection. Cas9 from S. pyogenes in particular has proven enormously useful for genome editing. The original Cas9, a two-component system can be rendered into one system by fusing the CRISPR RNA (crRNA) and tracer RNA (tracrRNA) into a single guide RNA (sgRNA) (Fig. 4), thus, allowing ease for genome editing by cutting the specific DNA segment and incorporation of donor DNA (genetic engineering), transcriptional control, RNA targeting, and imaging [123, 124]. CRISPR-Cas9 has been used in different cell types and organisms including mice and monkeys to primary human T cells and stem cells in addition to plants, bacteria, and fungi [123, 124].
Courtesy of Professor Jennifer Duodna. Molecular model of Cias9 protein and fused RNA onto the DNA based on crystallographic structures. The Cas9 protein is bound to the fused RNA as a guide to the targeted sequence of the DNA. Once the RNA is bound to the DNA at the specific targeted segment, the Cas9 protein cuts precisely the targeted segment of the DNA with resultant repair of the cut-off segment
With the rapid expansion of personalized and reference genomics, CRISPR-Cas genetic editing tools have opened unlimited genetic manipulation in different biological systems, including human cells [119, 125,126,127,128]. It enables editing, inhibition or activation of genes, it will allow more fundamental understanding of biology and create disease models to develop therapeutic strategies. It can be applied to drug target discovery, toxicology and diagnostics. It can help to develop more effective cellular therapies with novel therapeutics [129]. CRISPR may be applied in understanding cancer and metastasis to allow: (1) precision edits at single-gene level; (2) large scale in vitro and in vivo studies of functional genomics; (3) Screens for knockout, inhibition, activation and methylation and (4) growing toolkit of new editing modalities. Also, from the diagnostic and treatment points of view, CRISPR may be used to develop: (1) CAS12a, Cas13a-based detection kits, (2) cell based therapies, such as CAR T cells and (3) somatic in vivo editing for Mendelian diseases and cancer-related genes [130]. Indeed, CRISPR-Cas is a revolutionary molecular technique which can edit the genetic code of DNA and can be considered as a molecular scalpel for precision medicine. Thus, beyond the realm of cancer evolution within the cancer microenvironment, CRISPR-Cas system has opened new vistas with the ability of editing specific genes to modify cancer evolution and the cancer microenvironment with the goal to halt cancer evolution and progression, thus, controlling and eventually stopping the cancer growth and metastasis.
Summary and future perspectives
Marlys Witte
New ideas replace the old but the old comes back again—epitomizes the growing emphasis on the critical cancer microenvironment, its interplay with the genomics of the cancer cell, and the evolution of the metastatic process, i.e. whether and even where the cancer will spread. Historically, for several centuries before Virchow’s demonstration that cancer involved aberrant cells, the term “cancer” was almost synonymous with “lymph”, i.e. a disease of the tissues and tissue fluid [131]. It is only relatively recently that there has been an increased recognition of the influence of surrounding non-cancer cells and their products along with alterations in the stracellular matrix in determining the fate of the cancer cell and even the genes expressed, particularly those related to epithelial-mesenchymal transition in a reversion to a migrating invading embryonic phenotype. The evolving integration of cancer cell genetics and epigenetics (environment within and outside the cancer cell) mirrors the larger recognition over the last several decades of the importance of epigenetics/environment on the expression of genes (heredity) more generally.
Moreover, understanding of lymphatic “systemomics” has simultaneously evolved, encompassing not only lymphatic vessels and “lymph” permeating the tissues and circulating in collecting channels through lymph nodes to return to the bloodstream, but also as a route of entry of harmful substances (e.g., carcinogens, cancer-causing and protective microbes), as a transport system for abnormal cells and their products to disseminate, and as the center of the immune network of resident and circulating “immunocytes” [132].
The communications in this symposium session illuminate the multifaceted aspects of the evolution of thinking about cancer, lymphatic systemomics, and potential therapeutic implications for modulating the influence of the cancer microenvironment on the fate of both the cancer cell and ultimately, the host.
Abbreviations
- MBM:
-
Brain metastasis
- ANGPTL4:
-
Angiopoietin-like 4
- CysC:
-
Cystatin C
- CLDN1:
-
Claudin-1
- DTCs:
-
Disseminated tumor cells
- ECM:
-
Extracellular matrix
- MM:
-
Malignant melanoma
- bHLH:
-
Helix-loop-helix
- CPI:
-
Checkpoint inhibitors
- IO:
-
Immunoncology
- TAMPP:
-
T-cell activation marker proficiency panel
- ROI:
-
Regions of interest
- CNN:
-
Convolutional neural networks
- BC:
-
Breast cancer
- AA:
-
African American
- EA:
-
White or European American
- TNBC:
-
Triple-negative BC
- ICSBCS:
-
International Center for the Study of Breast Cancer Subtypes
- TST:
-
Trans-Atlantic Slave Trade
- DARC/ACKR1 :
-
Duffy Antigen Receptor for Chemokines/Atypical Chemokine Receptor 1 gene
- IMC:
-
Imaging mass cytometry
- RNA-Seq:
-
RNA sequencing
- GTEx:
-
Genotype-Tissue Expression database
- Treehouse CARE:
-
Treehouse Comparative Analysis of RNA Expression
- CRISPR/Cas9:
-
Clustered Regularly-Interspaced Short Palindromic Repeats
References
Hanahan D, Coussens LM (2012) Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21(3):309–322
Darwin C (1859) On the origin of species by means of natural selection, or preservation of favoured races in the struggle for life. John Murray, London
Leong SP, Aktipis A, Maley C (2018) Cancer initiation and progression within the cancer microenvironment. Clin Exp Metastasis 35(5–6):361–367
Boire A et al (2020) Brain metastasis. Nat Rev Cancer 20(1):4–11
Orozco JIJ et al (2018) Epigenetic profiling for the molecular classification of metastatic brain tumors. Nat Commun 9(1):4627
Valiente M et al (2018) The evolving landscape of brain metastasis. Trends Cancer 4(3):176–196
Winkler F (2015) The brain metastatic niche. J Mol Med (Berl) 93(11):1213–1220
Izraely S, Klein A, Sagi-Assif O, Meshel T, Tsarfaty G, Hoon DSB, Witz IP (2010) The metastatic microenvironment: brain-residing melanoma metastasis and dormant micrometastasis. Int J Cancer 130(1–2):107–114. https://doi.org/10.1016/j.imlet.2009.12.003
Neuditschko B et al (2020) The challenge of classifying metastatic cell properties by molecular profiling exemplified with cutaneous melanoma cells and their cerebral metastasis from patient derived mouse xenografts. Mol Cell Proteomics 19(3):478–489
Izraely S, Witz IP (2021) Site-specific metastasis: a cooperation between cancer cells and the metastatic microenvironment. Int J Cancer 148(6):1308–1322. https://doi.org/10.1002/ijc.33247
Izraely S et al (2017) ANGPTL4 promotes the progression of cutaneous melanoma to brain metastasis. Oncotarget 8(44):75778–75796
Tan MJ et al (2012) Emerging roles of angiopoietin-like 4 in human cancer. Mol Cancer Res 10(6):677–688
Zlotnik A, Burkhardt AM, Homey B (2011) Homeostatic chemokine receptors and organ-specific metastasis. Nat Rev Immunol 11(9):597–606
Izraely S, Klein A, Sagi-Assif O, Meshel T, Tsarfaty G, Hoon DSB, Witz IP (2010) Chemokine–chemokine receptor axes in melanoma brain metastasis. Immunol Lett 130(1–2):107–114. https://doi.org/10.1016/j.imlet.2009.12.003
Klein A et al (2012) The metastatic microenvironment: brain-derived soluble factors alter the malignant phenotype of cutaneous and brain-metastasizing melanoma cells. Int J Cancer 131(11):2509–2518
Klein A et al (2017) CCR4 is a determinant of melanoma brain metastasis. Oncotarget 8(19):31079–31091
Moshe A et al (2018) Cystatin C takes part in melanoma-microglia cross-talk: possible implications for brain metastasis. Clin Exp Metastasis 35(5–6):369–378
Kopitz C et al (2005) Reduction of experimental human fibrosarcoma lung metastasis in mice by adenovirus-mediated cystatin C overexpression in the host. Cancer Res 65(19):8608–8612
Huh CG et al (1999) Decreased metastatic spread in mice homozygous for a null allele of the cystatin C protease inhibitor gene. Mol Pathol 52(6):332–340
Klein A et al (2015) Astrocytes facilitate melanoma brain metastasis via secretion of IL-23. J Pathol 236(1):116–127
Langowski JL et al (2006) IL-23 promotes tumour incidence and growth. Nature 442(7101):461–465
Moshe A, Izraely S, Sagi-Assif O, Malka S, Ben-Menachem S, Meshel T, Pasmanik-Chor M, Hoon DSB, Witz IP, (2020) Inter-tumor heterogeneity—melanomas respond differently to GM-CSF-mediated activation. Cells 9(7):1683. https://doi.org/10.3390/cells9071683
Hoeller C et al (2016) Systematic review of the use of granulocyte-macrophage colony-stimulating factor in patients with advanced melanoma. Cancer Immunol Immunother 65(9):1015–1034
Chang YC et al (2018) Roles of aldolase family genes in human cancers and diseases. Trends Endocrinol Metab 29(8):549–559
Ramos RI et al (2020) Upregulation of cell surface GD3 ganglioside phenotype is associated with human melanoma brain metastasis. Mol Oncol 14(8):1760–1778
Bhat AA et al (2020) Claudin-1, a double-edged sword in cancer. Int J Mol Sci 21(2):569
Izraely S et al (2015) The metastatic microenvironment: Claudin-1 suppresses the malignant phenotype of melanoma brain metastasis. Int J Cancer 136(6):1296–1307
Stebbing J, Filipovic A, Giamas G (2013) Claudin-1 as a promoter of EMT in hepatocellular carcinoma. Oncogene 32(41):4871–4872
Almad A, Maragakis NJ (2018) A stocked toolbox for understanding the role of astrocytes in disease. Nat Rev Neurol 14(6):351–362
Fidler IJ et al (2010) The brain microenvironment and cancer metastasis. Mol Cells 30(2):93–98
Eyo UB, Wu LJ (2019) Microglia: lifelong patrolling immune cells of the brain. Prog Neurobiol 179:101614
Izraely S et al (2019) The metastatic microenvironment: melanoma-microglia cross-talk promotes the malignant phenotype of melanoma cells. Int J Cancer 144(4):802–817
Ohab JJ et al (2006) A neurovascular niche for neurogenesis after stroke. J Neurosci 26(50):13007–13016
Prakash R et al (2019) Regeneration enhances metastasis: a novel role for neurovascular signaling in promoting melanoma brain metastasis. Front Neurosci 13:297
Sleeman JP et al (2012) Concepts of metastasis in flux: the stromal progression model. Semin Cancer Biol 22(3):174–186
Sleeman JP (2012) The metastatic niche and stromal progression. Cancer Metastasis Rev 31(3–4):429–440
Bethan Psaila DL (2009) The metastatic niche: adapting the foreign soil. Nat Rev Cancer 9:285–292
Celia-Terrassa T, Kang Y (2018) Metastatic niche functions and therapeutic opportunities. Nat Cell Biol 20(8):868–877
Hoye AM, Erler JT (2016) Structural ECM components in the premetastatic and metastatic niche. Am J Physiol Cell Physiol 310(11):C955–C967
Levental KR et al (2009) Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139(5):891–906
Wei SC, Fattet L, Tsai JH, Guo Y, Pai VH, Majeski HE, Chen AC, Sah RL, Taylor SS, Engler AJ, Yang J (2015) Matrix stiffness drives epithelial–mesenchymal transition and tumour metastasis through a TWIST1–G3BP2 mechanotransduction pathway. Nat Cell Biol 17(5):678–688. https://doi.org/10.1038/ncb3157
Zaman MH, Trapani LM, Sieminski AL, MacKellar D, Gong H, Kamm RD, Wells A, Lauffenburger DA, Matsudaira P (2006) Migration of tumor cells in 3D matrices is governed by matrix stiffness along with cell-matrix adhesion and proteolysis. Proc Natl Acad Sci USA 103(29):10889–10894. https://doi.org/10.1073/pnas.0604460103
Barkan D, Green JE, Chambers AF (2010) Extracellular matrix: a gatekeeper in the transition from dormancy to metastatic growth. Eur J Cancer 46(7):1181–1188
Malanchi I et al (2011) Interactions between cancer stem cells and their niche govern metastatic colonization. Nature 481(7379):85–89
Oskarsson T et al (2011) Breast cancer cells produce tenascin C as a metastatic niche component to colonize the lungs. Nat Med 17(7):867–874
Hoshino A et al (2015) Tumour exosome integrins determine organotropic metastasis. Nature 527(7578):329–335
Savoia P et al (2009) Skin metastases of malignant melanoma: a clinical and prognostic survey. Melanoma Res 19(5):321–326
Sedlmeier G, Al-Rawi V, Buchert J, Yserentant K, Rothley M, Steshina A, Gräßle S, Wu RL, Hurrle T, Richer W, Decraene C, Thiele W, Utikal J, Abuillan W, Tanaka M, Herten DP, Hill CS, Garvalov BK, Jung N, Bräse S, Sleeman JP (2020) Id1 and Id3 are regulated through matrix-assisted autocrine BMP signaling and represent therapeutic targets in melanoma. Adv Ther 4(2):2000065. https://doi.org/10.1002/adtp.202000065
Hollnagel A et al (1999) Id genes are direct targets of bone morphogenetic protein induction in embryonic stem cells. J Biol Chem 274(28):19838–19845
Liang YY, Brunicardi FC, Lin X (2009) Smad3 mediates immediate early induction of Id1 by TGF-beta. Cell Res 19(1):140–148
Sun X-H, Copeland NG, Jenkins NA, Baltimore D (1991) Id proteins Idl and Id2 selectively inhibit DNA binding by one class of helix-loop-helix proteins. Mol Cell Biol 11(11):5603–5611. https://doi.org/10.1128/MCB.11.11.5603
Gupta GP, Perk J, Acharyya S, de Candia P, Mittal V, Todorova-Manova K, Brogi E, Gerald WL, Benezra R, Massague J (2007) ID genes mediate tumor reinitiation during breast cancer lung metastasis. Proc Natl Acad Sci USA 104(49):19506–19511. https://doi.org/10.1073/pnas.0709185104
Lai X et al (2014) Inhibitor of DNA-binding protein 1 knockdown arrests the growth of colorectal cancer cells and suppresses hepatic metastasis in vivo. Oncol Rep 32(1):79–88
O’Brien CA et al (2012) ID1 and ID3 regulate the self-renewal capacity of human colon cancer-initiating cells through p21. Cancer Cell 21(6):777–792
Roschger C, Cabrele C (2017) The Id-protein family in developmental and cancer-associated pathways. Cell Commun Signal 15(1):7
Straume O, Akslen LA (2005) Strong expression of ID1 protein is associated with decreased survival, increased expression of ephrin-A1/EPHA2, and reduced thrombospondin-1 in malignant melanoma. Br J Cancer 93(8):933–938
Pan L, Sato S, Frederick JP, Sun X-H, Zhuang Y (1999) Impaired immune responses and B-cell proliferation in mice lacking the Id3 gene. Mol Cell Biol 19(9):5969–5980. https://doi.org/10.1128/MCB.19.9.5969
Darvin P et al (2018) Immune checkpoint inhibitors: recent progress and potential biomarkers. Exp Mol Med 50(12):1–11
Wolchok JD et al (2017) Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med 377(14):1345–1356
Garon EB et al (2015) Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 372(21):2018–2028
Wei SC et al (2017) Distinct cellular mechanisms underlie anti-CTLA-4 and anti-PD-1 checkpoint blockade. Cell 170(6):1120–1133 e17
Galon J et al (2006) Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313:1960–1964
Pagès F et al (2018) International validation of the consensus Immunoscore for the classification of colon cancer: a prognostic and accuracy study. Lancet 391(10135):2128–2139
Mlecnik B et al (2020) Multicenter International Society for immunotherapy of cancer study of the consensus immunoscore for the prediction of survival and response to chemotherapy in stage III colon cancer. J Clin Oncol 38(31):3638–3651
Samstein RM et al (2019) Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet 51(2):202–206
Li K et al (2020) Microsatellite instability: a review of what the oncologist should know. Cancer Cell Int 20:16
Diggs LP, Hsueh EC (2017) Utility of PD-L1 immunohistochemistry assays for predicting PD-1/PD-L1 inhibitor response. Biomark Res 5:12
Taube JM, Akturk G, Angelo M, et al (2020) The Society for Immunotherapy of Cancer statement on best practices for multiplex immunohistochemistry (IHC) and immunofluorescence (IF) staining and validation. J Immunother Cancer 8:e000155. https://doi.org/10.1136/jitc-2019-000155
Hoyt C et al (2019) Abstract LB-318: Multi-institutional TSA-amplified Multiplexed Immunofluorescence Reproducibility Evaluation (MITRE study): reproducibility assessment of an automated multiplexed immunofluorescence slide staining, imaging, and analysis workflow. Cancer Res 79:LB-LB-318
Pirici D et al (2009) Antibody elution method for multiple immunohistochemistry on primary antibodies raised in the same species and of the same subtype. J Histochem Cytochem 57(6):567–575
Giesen C et al (2014) Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat Methods 11(4):417–425
Toki MI et al (2017) Proof of the quantitative potential of immunofluorescence by mass spectrometry. Lab Investig 97(3):329–334
Goltsev Y et al (2018) Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174(4):968–981 e15
Feng Z, Seliger B, Bernard AF (2017) Multiparametric immune profiling in HPV-oral squamous cell cancer. JCI Insight 2(14):e93652. https://doi.org/10.1172/jci.insight.93652
Hum L et al (2021) Cumulative suppressive index as a predictor of relapse free survival and overall survival in Human Papilloma Virus-negative oral squamous cell carcinomas with negative resection margins. Head Neck 43(2):568–576
Perou CM et al (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752
Sorlie T et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98(19):10869–10874
Parker JS et al (2009) Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 27(8):1160–1167
Newman LA (2016) Parsing the etiology of breast cancer disparities. J Clin Oncol 34(9):1013–1014
DeSantis CE et al (2019) Breast cancer statistics, 2019. CA Cancer J Clin 69(6):438–451
DeSantis CE et al (2016) Breast cancer statistics, 2015: convergence of incidence rates between black and white women. CA Cancer J Clin 66(1):31–42
Allott EH et al (2018) Frequency of breast cancer subtypes among African American women in the AMBER consortium. Breast Cancer Res 20(1):12
Ihemelandu CU et al (2007) Molecular breast cancer subtypes in premenopausal and postmenopausal African-American women: age-specific prevalence and survival. J Surg Res 143(1):109–118
Costa RLB, Gradishar WJ (2017) Triple-negative breast cancer: current practice and future directions. J Oncol Pract 13(5):301–303
Torre LA et al (2017) Global cancer in women: burden and trends. Cancer Epidemiol Biomark Prev 26(4):444–457
Jemal A, Fedewa SA (2012) Is the prevalence of ER-negative breast cancer in the US higher among Africa-born than US-born black women? Breast Cancer Res Treat 135(3):867–873
Newman LA et al (2019) Hereditary susceptibility for triple negative breast cancer associated with Western Sub-Saharan African ancestry: results from an International Surgical Breast Cancer Collaborative. Ann Surg 270(3):484–492
Jiagge E et al (2016) Comparative analysis of breast cancer phenotypes in African American, White American, and West versus East African patients: correlation between African ancestry and triple-negative breast cancer. Ann Surg Oncol 23(12):3843–3849
Al-Alem U et al (2014) Association of genetic ancestry with breast cancer in ethnically diverse women from Chicago. PLoS One 9(11):e112916
Kosoy R et al (2009) Ancestry informative marker sets for determining continental origin and admixture proportions in common populations in America. Hum Mutat 30(1):69–78
Nassir R et al (2009) An ancestry informative marker set for determining continental origin: validation and extension using human genome diversity panels. BMC Genet 10:39
Mersha TB, Abebe T (2015) Self-reported race/ethnicity in the age of genomic research: its potential impact on understanding health disparities. Hum Genomics 9:1
Newman LA, Carpten J (2018) Integrating the genetics of race and ethnicity into cancer research: trailing Jane and John Q. Public. JAMA Surg 153(4):299–300
Martin DN et al (2009) Differences in the tumor microenvironment between African-American and European-American breast cancer patients. PLoS One 4(2):e4531
Kim G et al (2020) The contribution of race to breast tumor microenvironment composition and disease progression. Front Oncol 10:1022
Nedelec Y et al (2016) Genetic ancestry and natural selection drive population differences in immune responses to pathogens. Cell 167(3):657–669 e21
Davis M et al (2018) AR negative triple negative or “quadruple negative” breast cancers in African American women have an enriched basal and immune signature. PLoS One 13(6):e0196909
Davis M, Martini R, Newman L, Elemento O, White J, Verma A, Datta I, Adrianto I, Chen Y, Gardner K, Kim HG, Colomb WD, Eltoum IE, Frost AR, Grizzle WE, Sboner A, Manne U, Yates C (2020) Identification of distinct heterogenic subtypes and molecular signatures associated with African ancestry in triple negative breast cancer using quantified genetic ancestry models in admixed race populations. Cancers (Basel) 12(5):1220. https://doi.org/10.3390/cancers12051220
Livingstone FB (1984) The Duffy blood groups, vivax malaria, and malaria selection in human populations: a review. Hum Biol 56(3):413–425
Tournamille C et al (1995) Disruption of a GATA motif in the Duffy gene promoter abolishes erythroid gene expression in Duffy-negative individuals. Nat Genet 10(2):224–228
Howes RE et al (2011) The global distribution of the Duffy blood group. Nat Commun 2:266
Lu ZH et al (1995) The promiscuous chemokine binding profile of the Duffy antigen/receptor for chemokines is primarily localized to sequences in the amino-terminal domain. J Biol Chem 270(44):26239–26245
Nibbs RJ, Graham GJ (2013) Immune regulation by atypical chemokine receptors. Nat Rev Immunol 13(11):815–829
Peiper SC et al (1995) The Duffy antigen/receptor for chemokines (DARC) is expressed in endothelial cells of Duffy negative individuals who lack the erythrocyte receptor. J Exp Med 181(4):1311–1317
Pruenster M et al (2009) The Duffy antigen receptor for chemokines transports chemokines and supports their promigratory activity. Nat Immunol 10(1):101–108
Jenkins BD et al (2019) Atypical chemokine receptor 1 (DARC/ACKR1) in breast tumors is associated with survival, circulating chemokines, tumor-infiltrating immune cells, and African ancestry. Cancer Epidemiol Biomark Prev 28(4):690–700
Filbin M, Monje M (2019) Developmental origins and emerging therapeutic opportunities for childhood cancer. Nat Med 25(3):367–376
Grobner SN et al (2018) The landscape of genomic alterations across childhood cancers. Nature 555(7696):321–327
Vaske OM et al (2019) Comparative tumor RNA sequencing analysis for difficult-to-treat pediatric and young adult patients with cancer. JAMA Netw Open 2(10):e1913968
Newton Y, Rassekh SR, Deyell RJ, Shen Y, Jones MR, Dunham C, Yip S, Leelakumari S, Zhu J, McColl D, Swatloski T, Salama SR, Ng T, Hendson G, Lee AF, Ma Y, Moore R, Mungall AJ, Haussler D, Stuart JM, Jantzen C, Laskin J, Jones SJM, Marra MA, Morozova O (2018) Comparative RNA-sequencing analysis benefits a pediatric patient with relapsed cancer. JCO Precis Oncol 2:1–16
Consortium GT et al (2017) Genetic effects on gene expression across human tissues. Nature 550(7675):204–213
Ford AM et al (2009) The TEL-AML1 leukemia fusion gene dysregulates the TGF-beta pathway in early B lineage progenitor cells. J Clin Investig 119(4):826–836
Treehouse Public Data. https://treehousegenomics.soe.ucsc.edu/public-data/
Vivian J et al (2017) Toil enables reproducible, open source, big biomedical data analyses. Nat Biotechnol 35(4):314–316
Liang L et al (2015) Gliomatosis peritonei: a clinicopathologic and immunohistochemical study of 21 cases. Mod Pathol 28(12):1613–1620
Nevelius E (2020) Press release: the Nobel Prize in Chemistry 2020, in Genetic scissors: a tool for rewriting the code of life. The Royal Swedish Academy of Sciences, Stockholm
Fellmann C (2019) Emerging roles of Crispr-cas9 in precision-oncology. In: 8th International Cancer Metastasis Congress. San Francisco
Labrie SJ, Samson JE, Moineau S (2010) Bacteriophage resistance mechanisms. Nat Rev Microbiol 8(5):317–327
Jinek M et al (2012) A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–820
Channel TNPY (2020) Nobel Lecture: Emmanuelle Charpentier, Nobel Prize in Chemistry 2020. Youtube. https://www.youtube.com/watch?v=3POrtQEpV2s
Channel TNPY (2020) Nobel Lecture: Jennifer Doudna, Nobel Prize in Chemistry 2020. Youtube.com. https://www.youtube.com/watch?v=KSrSIErIxMQ
Makarova KS et al (2015) An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol 13(11):722–736
Jiang W, Marraffini LA (2015) CRISPR-Cas: new tools for genetic manipulations from bacterial immunity systems. Annu Rev Microbiol 69:209–228
Sternberg SH et al (2015) Conformational control of DNA target cleavage by CRISPR-Cas9. Nature 527(7576):110–113
Mali P et al (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Jinek M et al (2013) RNA-programmed genome editing in human cells. Elife 2:e00471
Cong Le et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–822
Cho SW et al (2013) Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31(3):230–232
Fellmann C et al (2017) Cornerstones of CRISPR-Cas in drug discovery and therapy. Nat Rev Drug Discov 16(2):89–100
McBride DA et al (2019) Applications of molecular engineering in T-cell-based immunotherapies. Wiley Interdiscip Rev Nanomed Nanobiotechnol 11(5):e1557
Triolo VA (1965) Nineteenth century foundations of cancer research advances in tumor pathology, nomenclature, and theories of oncogenesis. Cancer Res 25:75–106
Dellinger MT, Witte MH (2018) Lymphangiogenesis, lymphatic systemomics, and cancer: context, advances and unanswered questions. Clin Exp Metastasis 35(5–6):419–424
Acknowledgements
Isaac P. Witz, Orit Sagi-Assif and Sivan Izraely thank their colleagues Shlomit Ben-Menachem and Tsipi Meshel, and the Laboratory of Tumor Microenvironment & Metastasis Research for their valuable help and support.
Funding
The research conducted by the authors is supported by the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (Needham, MA, USA).
Author information
Authors and Affiliations
Contributions
Equal contribution regarding data/information collection, analysis and writing among the listed authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
Not applicable for this review article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Presented at the 8th International Cancer Metastasis Congress in San Francisco, CA, USA from October 25–27, 2019 (http://www.cancermetastasis.org). To be published in an upcoming Special Issue of Clinical and Experimental Metastasis: Novel Frontiers in Cancer Metastasis.
Rights and permissions
About this article
Cite this article
Leong, S.P., Witz, I.P., Sagi-Assif, O. et al. Cancer microenvironment and genomics: evolution in process. Clin Exp Metastasis 39, 85–99 (2022). https://doi.org/10.1007/s10585-021-10097-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-021-10097-9