Abstract
Sirtuin2 (SIRT2) is a deacetylase enzyme predominantly expressed in myelinating glia of the central nervous system (CNS). We have previously demonstrated that Sirt2 expression enhances oligodendrocyte (OL) differentiation and arborization in vitro, but the molecular targets of SIRT2 in OLs remain speculative. SIRT2 has been implicated in cholesterol biosynthesis by promoting the nuclear translocation of sterol regulatory element binding protein (SREBP)-2. We investigated this further in CNS myelination by examining the role of Sirt2 in cholesterol biosynthesis in vivo and in vitro employing Sirt2 −/− mice, primary OL cells and CG4-OL cells. Our results demonstrate that expression of cholesterol biosynthetic genes in the CNS white matter or cholesterol content in purified myelin fractions did not differ between Sirt2 −/− and age-matched wild-type mice. Cholesterol biosynthetic gene expression profiles and total cholesterol content were not altered in primary OLs from Sirt2 −/− mice and in CG4-OLs when Sirt2 was either down-regulated with RNAi or overexpressed. In addition, Sirt2 knockdown or overexpression in CG4-OLs had no effect on SREBP-2 nuclear translocation. Our results indicate that Sirt2 does not impact the expression of genes encoding enzymes involved in cholesterol biosynthesis, total cholesterol content, or nuclear translocation of SREBP-2 during OL differentiation and myelination.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the developing central nervous system (CNS), oligodendrocyte precursor cells (OPCs) originate from the ventricular/subventricular zone. OPCs proliferate and migrate throughout the cortical white matter, such as the corpus callosum (CC), and ultimately differentiate into mature, myelinating oligodendrocytes (OLs) (Baumann and Pham-Dinh 2001; Bergles and Richardson 2015; Nicolay et al. 2007). These differentiating OLs, which are rich in myelin lipids and structural proteins, extend their plasma membranes to wrap axons with compact, multilayered membranous sheath. Proper myelination is critical for rapid signal conduction and long-term axonal survival (Baumann and Pham-Dinh 2001; Simons and Nave 2015).
Myelin contains greater than 70% lipids in its dry weight with 25% cholesterol in the total lipid content (Norton and Poduslo 1973a; Saher and Simons 2010). In the brain, cholesterol biosynthesis increases during the peak of myelination (Dietschy and Turley 2004; Jurevics et al. 1997; Jurevics and Morell 1995; Muse et al. 2001), and cholesterol needed for the myelin biogenesis is primarily derived from in situ biosynthesis in differentiating OLs with little imported via circulation (Jurevics et al. 1997; Jurevics and Morell 1995). In addition to being a large structural component of myelin membranes, cholesterol may also facilitate transport and sorting of proteins to form compact myelin sheath (Saher and Simons 2010; Saher et al. 2005; Saher et al. 2009; Simons et al. 2000; Werner et al. 2013). Inactivation or mutation of enzymes involved in the cholesterol biosynthetic pathway in OLs, such as squalene synthase (SQS; Saher et al. 2005) and Hmg-CoA synthase 1 (HMGCS1; Mathews et al. 2014), perturbs myelin gene expression and axon ensheathment. Thus, the supply of cholesterol is a rate-limiting factor for proper CNS myelination (Mathews et al. 2014; Mathews and Appel 2016; Saher et al. 2005).
Two master regulatory transcription factors, sterol regulatory element binding protein (SREBP)-1 and SREBP-2, regulate lipid and cholesterol biosynthesis, respectively (Brown and Goldstein 1997; Eberlé et al. 2004). SREBP-2 is synthesized as a precursor (125 kDa) and is bound to SREBP cleavage activating protein (SCAP) in the endoplasmic reticulum. Under low sterol levels, the SREBP-SCAP complex is exported to the Golgi where the cytoplasmic N-terminal active domain of SREBP-2 is cleaved (Sakai et al. 1996; Wang et al. 1994; Yabe et al. 2002; Yang et al. 2002). Subsequently, this mature form (N-terminal domain) of SREBP-2 (68 kDa) translocates to the nucleus and binds to sterol response element (SRE) inducing genes encoding enzymes involved in cholesterol biosynthesis (Amemiya-Kudo et al. 2002; Eberlé et al. 2004; Horton et al. 2002, 2003). The activity of SREBP family of transcription factors is regulated by acetylation and deacetylation of lysine residues within the DNA binding domain, which modulates SREBPs activity and downstream target gene expression (Giandomenico et al. 2003; Ponugoti et al. 2010; Walker et al. 2010). However, relatively little is known about the regulatory factors governing SREBP-2 activity and cholesterol biosynthesis during OL differentiation and myelination in the developing CNS.
Sirtuin2 (SIRT2), an NAD+-dependent deacetylase enzyme has been implicated in cholesterol biosynthesis by facilitating the nuclear translocation of SREBP-2 (Luthi-Carter et al. 2010; Taylor et al. 2011). SIRT2 is primarily expressed in OLs and incorporated into the myelin sheath near paranodal loops (Li et al. 2007; Werner et al. 2007). Sirt2 expression promotes process formation and induces myelin gene expression during OL differentiation in vitro (Ji et al. 2011). Loss of Sirt2 in the peripheral nervous system (PNS) delays Schwann cell myelin formation via deacetylation of Par-3 (Beirowski et al. 2011), but its role in CNS myelination in vivo remains speculative. It was reported that pharmacological inhibition of SIRT2 or deacetylase mutation in Sirt2 impairs cholesterol biosynthesis by reducing nuclear translocation of SREBP-2 leading to downregulation of several key genes in cholesterol biosynthetic pathways in neuronal cultures (Luthi-Carter et al. 2010). In contrast, Bobrowska et al. (2012) reported no effect on nuclear translocation of SREBP-2 or cholesterol biosynthesis in brain tissue from one month-old Sirt2 −/− mice. As SIRT2 expression is upregulated dramatically during the peak of myelination (Li et al. 2007; Southwood et al. 2007; Werner et al. 2007; Zhu et al. 2012), we speculated that SREBP-2 may be a target of SIRT2 in CNS white matter when OLs are undergoing active differentiation and myelination.
In the present study, we have used Sirt2 −/− mutant mice to investigate the role of SIRT2 in regulating cholesterol biosynthesis in the developing CNS white matter and primary OL cultures. Furthermore, we evaluated the impact of loss or gain of Sirt2 function in CG4-OLs on the expression of cholesterol biosynthetic genes, cholesterol content, and nuclear translocation of SREBP-2.
Materials and Methods
Animals
Sirt2 −/− mice (lacking exons 5, 6 and part of exon 7 of the Sirt2 gene) (Strain name: B6.129-Sirt2 tm1.1FWa/J; Stock number: 012772) and C57BL6/J mice were obtained from The Jackson Laboratory for breeding in-house at the University of Saskatchewan. Genotyping was performed using the following primers: Common—5′-GACTGGAAGTGATCAAAGCTC-3′, Wild-type—5′-CAGGGTCTCACGAGTCTCATG-3′, and Mutant—5′-TCAAATCTGGCCAGAACTTATG-3′. All animal studies were conducted in compliance with the University of Saskatchewan’s Animal Care Committee guidelines.
Primary OL Cell Culture
Primary OLs were isolated from C57BL/6 mice or Sirt2 −/− mice at post-natal day 1, as previously described (Chen et al. 2007; Niu et al. 2012; Thangaraj et al. 2017). Briefly, cortices from the neonatal pups were digested with accumax solution (Sigma®) and passed through a 70 μm nylon cell strainer. The filtered cell suspensions were maintained as mixed glial cells in medium containing Dulbecco’s modified Eagle medium/nutrient mixture F-12 (DMEM/F12), 10% fetal bovine serum (FBS), and 1% penicillin–streptomycin solution. After 7 days of culture, mixed glial cells were grown in OL growth medium (OGM) containing 10 ng/mL biotin, 5 μg/mL insulin, 50 μg/mL transferrin, 2 mM glutamine, 30 nM sodium selenite, 0.1% BSA, 10 nM hydrocortisone, 1% penicillin–streptomycin solution, and 30% B104-conditioned medium in DMEM/F12 to enrich OL progenitors. After 14 days of culture, primary OLs were shaken off (at 200 rpm; 37 °C) and isolated from the mixed glial cultures. Differentiation of purified primary OLs was induced by plating in OL differentiation medium (ODM) containing 10 ng/mL biotin, 5 μg/mL insulin, 50 μg/mL transferrin, 2 mM glutamine, 30 nM sodium selenite, 0.1% BSA, 10 nM hydrocortisone, 5 μg/ml N-acetyl-l-cysteine, 1% penicillin–streptomycin solution, and 1% FBS in DMEM/F12. In some experiments, primary OLs were differentiated using 1% delipidated FBS (DLFBS; cholesterol-depleted FBS) to avoid the interference of any external source of cholesterol from the media components. Primary OLs were harvested 6 days after differentiation for RNA extraction and cholesterol assay.
CG4-OL Cell Culture, Small Interference RNA, Overexpression Vector Construct, and Transfection
The CG4-OL cell line was derived from O-2A progenitors isolated from the rat forebrain (Louis et al. 1992). CG4-OLs were cultured on Poly-d-Lysine (Sigma)-coated tissue culture dishes in growth medium (GM) containing DMEM, 50 μg/mL transferrin, 5 μg/mL insulin, 9.8 ng/mL biotin, 50 ng/mL selenium, 1% antibiotic–antimycotic solution, and 30% B104 conditioned medium (Thangaraj et al., 2017; Ji et al. 2011; Louis et al. 1992; Wang et al. 2011). Differentiation was induced by changing GM to differentiation medium (DM) containing DMEM, 50 μg/mL transferrin, 5 μg/mL insulin, 9.8 ng/mL biotin, 50 ng/mL selenium, 1% antibiotic–antimycotic solution, and 2% FBS or DLFBS (Thangaraj et al. 2017; Ji et al. 2011; Wang et al. 2011).
Sirt2 siRNA with sense sequence 5′-AGGGAGCAUGCCAACAUAGAU-3′ commercially synthesized (Qiagen®) and pcDNA vector carrying full length Sirt2 insert have been described previously (Ji et al. 2011). Transfection was carried out with 40 nM of scramble control siRNA or Sirt2 siRNA and 3 μg of pcDNA control vector or pcDNA-Sirt2 vector on (day) d0, d2, d4 using 0.3% Lipofectamine 2000 (Invitrogen®) in serum-free/antibiotic-free medium. Cells were harvested for biochemical analyses after 6 days of differentiation.
RNA Isolation, Reverse Transcription-PCR (RT-PCR), and Quantitative Real-Time-PCR (qRT-PCR)
Total RNA was isolated from the CC, primary OLs, or CG4-OL cells using Ambion® total RNA miniprep kit (Invitrogen®) as per the manufacturer’s protocol. RNA concentration was determined using NanoVue UV/Visible spectrophotometer (GE Healthcare Life Sciences). First strand cDNA synthesis was performed with 250–500 ng of total RNA using High-Capacity cDNA Reverse Transcription kit (Applied Biosystems®) with random primers. Final concentration of total RNA for RT-PCR or qRT-PCR was 5–25 ng/μL. RT-PCR was carried out for Sirt2 in CG4-OLs transfected with Sirt2 siRNA and pcDNA-Sirt2 according to Ji et al. (2011). The following primer pairs were used: (i) Sirt2, forward 5′-TGAATGGCACCTACAGAGAC-3′ and reverse 5′-CAAAGGCATTATGGTAGGGC-3′; and (ii) β-Actin, forward 5′-ATTGTAACCAACTGGGACG-3′ and reverse 5′-TTGCCGATAGTGATGACCT-3′. The qRT-PCR was carried out using SYBR select master mix in ABI 7300 (Applied Biosystems®) with cDNA samples from CG4-OL cells, primary OLs, or CC of mouse brain. Expression levels of Srebp-2, Hmgcr, Sqs, Sqle, and Dhcr7 were quantified using the following primers pairs: (i) Srebp-2, forward 5′-TGCAGGTCAAAGTCTCTCCT-3′ and reverse 5′-GCAGGACTTGAAAGCTGGT-3′; (ii) Hmgcr, forward 5′-CTGGTCCTAGAGCTTTCTCG-3′ and reverse 5′-TGCTGTTCTGAGGAGAAGGA-3′; (iii) Sqs, forward 5′-GAAGATTCGGAAGGGGCAAG-3′ and reverse 5′-CTCAAGTACTGCCAGCTCAG-3′; (iv) Sqle, forward 5′-TAAGAAATGCGGGGATGTCA-3′ and reverse 5′-GAATATCTGAGAAGGCAGCG-3′; and (v) Dhcr7, forward 5′-TTTATGGCCATGTGACCAAC-3′ and reverse 5′-AACAGGTCCTTCTGATGGTT-3′. Relative quantification of transcripts was determined by threshold cycle differences (2−ΔΔCT) of target normalized to the endogenous control β-actin as described previously (Doucette et al. 2010; Thangaraj et al. 2017).
Immunocytochemistry
Immunocytochemistry was carried out as previously described in (Thangaraj et al. 2017; Wang et al. 2011). CG4-OL cells were cultured on coverslips at a density of 2 × 102 cells/mm2. Cells were fixed with 4% paraformaldehyde and rinsed twice with PBS for 10 min. Cells were blocked with 3% skim milk in phosphate-buffered saline (PBS) containing 0.1% Triton X-100, incubated with the anti-SREBP-2 (Abcam®) antibody overnight at 4 °C followed by incubation with Alexa Fluor® 594 (Molecular Probes®) secondary antibody for 1 h at room temperature. Coverslips were mounted with Prolong Gold anti-fade reagent containing DAPI (Invitrogen®) to visualize nuclei. Blinded cell counts were performed (Doucette et al. 2010; Thangaraj et al. 2017) using Image J software (NIH). Mean % nuclear SREBP-2 (SREBP-2/DAPI co-localization) was calculated by counting >1500 cells and expressed as a percentage of total cells (DAPI).
Cholesterol Assay
Cholesterol assay was carried out using Amplex Red Cholesterol Assay Kit (Invitrogen Molecular Probes®) according to manufacturer’s instructions. CG4-OLs and primary OLs were rinsed with 1× PBS and lysed with reaction buffer (Invitrogen®). Cholesterol assay in wild-type and Sirt2 −/− mice were carried out using purified myelin fractions. Total cholesterol values were normalized to total protein as estimated with the BCA Protein Assay (Pierce®).
Preparation of Purified Myelin Fraction
Myelin was isolated from the whole brain of wild-type and Sirt2 −/− mice as described previously (Larocca and Norton 2007; Norton and Poduslo 1973b). Briefly, brain was homogenized in ice-cold 0.3 M sucrose solution containing 20 mM Tris–HCl buffer (pH 7.45), 1 mM EDTA, 1 mM DTT, 100 μM phenylmethylsulfonyl fluoride (PMSF), and 10 μg/mL leupeptin. Brain homogenate was layered over 0.83 M sucrose and centrifuged in an ultracentrifuge at 75,000×g. Myelin membranes were recovered from the 0.3 and 0.83 M interface and further purified by two rounds of hypo-osmotic shock by resuspension in Tris–HCl buffer, followed by a second round of centrifugation in a sucrose gradient. Purified myelin was collected from the interface and resuspended in Tris–HCl buffer.
Statistical Analysis
All quantitative data were compared using student’s t test (Prism® Software Corporation) and are presented as mean ± standard error of the mean (SEM).
Results
Cholesterol Biosynthesis is not Altered in CNS White Matter of Sirt2 −/− Mice
To investigate the role of Sirt2 in regulating cholesterol biosynthesis in vivo, gene expression profiles of key cholesterogenic enzymes were characterized in Sirt2 −/− mice during the period of peak myelination (P15) and in adulthood (P60). Mice were genotyped using specific primers for wild-type (WT) and Sirt2 −/− mutant alleles (Fig. 1a). A complete loss of Sirt2 mRNA (Fig. 1b) and SIRT2 protein (Fig. 1c) was observed in the CC and purified myelin isolated from the whole brain, respectively, from Sirt2 −/− mice. Expression levels of Srebp-2, Hmgcr, Sqs, Sqle, and Dhcr7 were not altered in the CC of Sirt2 −/− mice compared to their age-matched WT controls at P15 (Fig. 2a) or P60 (Fig. 2b). In addition, total cholesterol content in the purified myelin isolated from the whole brain of Sirt2 −/− and WT mice did not differ significantly (Fig. 2c).
Sirt2 −/− mice show complete loss of Sirt2 mRNA and protein expression: (a) Genotyping of Sirt2 −/− mice was performed using primers specific for wild-type (WT; lane 2 and 4) and Sirt2 −/− mutants (lane 1 and 3) (see “Materials and Methods” section). The presence of amplicon at 538 bp (lane 2) and absence of amplicon at 700 bp (lane 1) confirms the WT phenotype, whereas the presence of amplicon at 700 bp (lane 3) and absence of amplicon at 538 bp (lane 4) confirms the Sirt2 mutant phenotype. (b) RT-PCR show the complete absence of Sirt2 mRNA in the corpus callosum (CC) of Sirt2 −/− mutants and (c) immunoblots of purified myelin isolated from the whole brain show complete absence of the SIRT2 protein
Cholesterol biosynthesis is not altered in CNS white matter of Sirt2 −/− mice. Quantification of mRNA expression of key cholesterol biosynthetic genes by qRT-PCR at P15 (a) and P60 (b) showed no difference between Sirt2 −/− mice and age-matched wild-type (WT) mice. The bar graphs represent the transcript levels of Srebp2, Hmgcr, Sqs, Sqle, and Dhcr7 normalized to the housekeeping gene β-actin and represented relative to WT controls (n = 6). Data represented as Mean ± SEM; unpaired t test; n.s. (c) Quantification of total cholesterol content from purified myelin fraction of the whole brain at P15 showed no difference between Sirt2 −/− and WT mice. Protein content in the purified myelin fraction was used to normalize total cholesterol (n = 4). Data represented as Mean ± SEM; unpaired t test; n.s
Sirt2 Does not Regulate the Cholesterol Biosynthesis Pathway in OLs
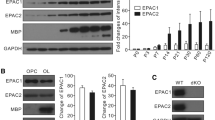
To directly evaluate the impact of Sirt2 on cholesterol biosynthesis in OLs, progenitor cells were isolated from the neonatal forebrain of Sirt2 −/− and WT mice followed by differentiation in serum-supplemented media for 6 days. The loss of Sirt2 −/− in primary OLs did not result in altered expression of Hmgcr or Sqle when cultured in FBS (Fig. 3a) or in DLFBS (Fig. 3c). Consistent with this, total cholesterol content did not differ between Sirt2 −/− and WT primary OL cultures (Fig. 3b, d).
Cholesterol biosynthesis is not altered in primary OLs isolated from Sirt2 −/− mice. Primary OLs isolated from WT and Sirt2 −/− mice were allowed to differentiate for 6 days. Quantification of mRNA expression levels of Hmgcr and Sqle by qRT-PCR showed no change in Sirt2 −/− primary OLs compared to WT primary OLs cultured with FBS (a) or DLFBS (c). qRT-PCR data were normalized to the housekeeping gene β-actin and represented relative to WT controls (n = 6). Mean ± SEM; unpaired t test; n.s. Quantification of total cholesterol content showed no difference between Sirt2 −/− and WT primary OLs cultured with FBS (b) or DLFBS (d). Protein content in the cell lysate was used to normalize total cholesterol (n = 6). Mean ± SEM; unpaired t test; n.s
Furthermore, Sirt2 was knocked down or overexpressed in CG4-OL cells during differentiation in media containing either FBS or DLFBS. In Sirt2-siRNA-treated CG4-OLs, expression of Sirt2 mRNA was significantly down-regulated by ~70% compared to control siRNA-treated cells (Fig. 4a, b). However, the expressions of Hmgcr, Sqs, and Sqle (Fig. 4c, e) and total cholesterol content (Fig. 4d, f) were not altered in Sirt2-siRNA-treated cells. In CG4-OLs transfected with pcDNA-Sirt2, expression of Sirt2 mRNA is significantly upregulated by ~500-fold (Fig. 5a, b). Again, neither the expression of Hmgcr, Sqs, and Sqle (Fig. 5c, e) nor total cholesterol content (Fig. 5d, f) was altered in cells overexpressing Sirt2.
Expression of cholesterol biosynthetic genes and total cholesterol content in CG4-OLs after knockdown of Sirt2. CG4-OLs were transfected with control siRNA or Sirt2 siRNA under differentiation conditions. (a, b) Transfection with Sirt2 siRNA down-regulated the expression of Sirt2 mRNA after 6 days. qRT-PCR data normalized to β-actin and represented relative to control siRNA (n = 3). Mean ± SEM; unpaired t test; ***p < 0.001. Knockdown of Sirt2 did not alter the expression of Hmgcr, Sqs, and Sqle mRNAs in CG4-OL cells differentiated with FBS (c) or DLFBS (e). qRT-PCR data were normalized to the housekeeping gene β-actin and represented relative to control siRNA (n = 3). Mean ± SEM; unpaired t test; n.s. Total cholesterol content did not change after knockdown of Sirt2 in CG4-OL cells differentiated with FBS (d) or DLFBS (f). Protein concentration in the cell lysate was used to normalize total cholesterol (n = 3). Mean ± SEM; unpaired t test; n.s
Expression of cholesterol biosynthetic genes and total cholesterol content in CG4-OLs overexpressing Sirt2. CG4-OLs were transfected with pcDNA or pcDNA-Sirt2 under differentiation conditions. (a, b) Overexpression of Sirt2 displays ~500-fold increase in the Sirt2 mRNA levels in CG4-OL cells after 6 days. qRT-PCR data normalized to β-actin and represented relative to pcDNA control vector (n = 3). Mean ± SEM; unpaired t test; **p < 0.01. Overexpression of Sirt2 did not alter the expression of Hmgcr, Sqs, and Sqle mRNAs in CG4-OL cells differentiated with FBS (c) or DLFBS (e). qRT-PCR data were normalized to the housekeeping gene β-actin and represented relative to pcDNA vector (n = 3). Mean ± SEM; unpaired t test; n.s. Total cholesterol content did not change after overexpression of Sirt2 in CG4-OL cells differentiated with FBS (d) or DLFBS (f). Protein content in the cell lysate was used to normalize total cholesterol (n = 3). Mean ± SEM; unpaired t test; n.s
Sirt2 Does not Impact Nuclear Translocation of SREBP-2
Nuclear translocation was assessed in CG4-OL cells transfected with Sirt2-siRNA or pcDNA-Sirt2 by immunostaining with SREBP-2 antibody (Fig. 6a–b, d–e). SREBP-2 was largely expressed in the perinuclear region of differentiating cells, presumably associated with the ER. Approximately 35–40% of cells also showed nuclear localization of SREBP-2. Knockdown (Fig. 6c) or overexpression (Fig. 6f) of Sirt2 resulted in no change in the subcellular distribution of SREBP-2 compared to their respective controls.
Sirt2 knockdown or Sirt2 overexpression in CG4-OL cells does not impact nuclear translocation of SREBP-2. Representative images of CG4-OLs cells transfected with the control siRNA (a) and Sirt2 siRNA (b) or pcDNA (d) and pcDNA-Sirt2 (e) at day 6 under differentiation conditions. Scale bar indicates 100 μm. Blinded counts (n = 3; >1500 cells) of SREBP-2 (red) expression in the nucleus (DAPI; blue) showed no difference in the subcellular localization of SREBP-2 in CG4-OL cells after Sirt2 downregulation (c) or Sirt2 overexpression (f). Mean ± SEM; unpaired t test, n.s
Discussion
Myelin is highly enriched in cholesterol, comprising ~80% of total brain cholesterol (Dietschy and Turley 2004; Muse et al. 2001). Cholesterol is thought to play a vital role in transport and sorting of the myelin proteins (Saher and Simons 2010; Saher et al. 2005, 2009; Simons et al. 2000; Werner et al. 2013). Cholesterol-enriched membrane domains in OPCs may also be important for efficient signal transduction to facilitate the differentiation (Mathews and Appel 2016). Defects in the cholesterol biosynthesis pathway can cause dysmyelination, reduced myelin gene expression, and impaired axon wrapping in both the CNS (Mathews et al. 2014; Mathews and Appel 2016; Saher et al. 2005) and PNS (Saher et al. 2009; Verheijen et al. 2009).
SREBP-2 is a key transcription factor that modulates cholesterol biosynthesis. SREBP-2 is synthesized in the endoplasmic reticulum, where it binds to SCAP. Low cellular sterol level induces SREBP-2 processing and translocation to the nucleus where it activates downstream target genes (Horton et al. 2002, 2003). Previous studies using SIRT2 specific inhibitors AGK2, AK-1, and AK-7 reported that the deacetylase activity of SIRT2 impacted sterol biosynthesis by regulating nuclear trafficking of SREBP-2 protein in neuronal cells (Luthi-Carter et al. 2010; Taylor et al. 2011). Overexpression of Sirt2 increased nuclear translocation of SREBP-2, while overexpression of deacetylase-deficient Sirt2 (Sirt2H150Y) or pharmacological inhibition of SIRT2 reduced the nuclear translocation of SREBP-2 (Luthi-Carter et al. 2010; Taylor et al. 2011). Our in silico analysis revealed the presence of two lysine residues at K310 and K329 in the DNA binding domain of SREBP-2 (Fig. 7a). These lysine residues are highly conserved across various SREBPs of which K283 and K303 in SREBP-1c have been shown to be deacetylated by SIRT1 (Ponugoti et al. 2010; Walker et al. 2010). Hence, it is possible that the two conserved lysine residues in SREBP-2 would be potential deacetylation sites for SIRT2.
Schematic diagram depicting the speculative role of Sirt2 in SREBP-2 nuclear trafficking and cholesterol biosynthesis. (a) In silico analysis revealed putative lysine acetylation sites in the DNA binding domain of SREBP-2 protein. Sequence alignment of the conserved lysine residues in mouse, rat, and human SREBP proteins with predicted acetylation lysine residues highlighted in red. (b) Acetylation/deacetylation regulates nuclear trafficking and activity of mature SREBP-2 (N-terminal fragment). In the nucleus, SREBP-2 binds to sterol response element (SRE) and activates the downstream genes encoding enzymes involved in cholesterol biosynthesis. Collectively, our finding shows no impact of Sirt2 expression on the cholesterol biosynthesis pathway indicating that SREBP-2 is not a primary target of SIRT2 during oligodendroglial differentiation
Loss of Sirt2 function in Schwann cells has been shown to delay myelin formation in PNS (Beirowski et al. 2011). Sirt2 expression is also upregulated during OL differentiation and in CNS myelination (Ji et al. 2011; Li et al. 2007; Thangaraj et al., 2017; Werner et al. 2007; Zhu et al. 2012). Our previous work demonstrates that Sirt2 expression promotes differentiation in CG4-OLs, including upregulation of myelin basic protein (MBP) expression and enhanced outgrowth of cellular processes (Ji et al. 2011). While α-tubulin has been identified as a primary target of SIRT2 in oligodendroglial cells in vitro (Li et al. 2007), this may not be the case in vivo (Bobrowska et al. 2012). Thus, we postulated that SREBP-2 may be a possible target of SIRT2 during OL differentiation and myelination. This would not only drive sterol synthesis for the formation of myelin membranes (Saher et al. 2005; Mathews et al. 2014) but may indirectly facilitate myelin gene expression (Mathews and Appel 2016).
In the present study, we found no impact of Sirt2 expression on cholesterol biosynthesis in CNS white matter in vivo and OL cultures in vitro (Fig. 7b). Although Sirt2 −/− mice showed a complete absence of Sirt2 mRNA and SIRT2 protein in white matter tracts, no difference in the expression level of Srebp2 or four of the key genes in the biosynthesis pathway was observed (Figs. 1 and 2). Furthermore, there was no difference in the cholesterol content of purified myelin from Sirt2 −/− and WT mice (Fig. 2). Although OLs are thought to be primarily responsible for the production of sterol required for myelin formation, some of the cholesterol may be derived from extracellular sources to maintain the integrity of myelin sheath (Saher et al. 2005; Verheijen et al. 2009). Thus, it is possible that in Sirt2 −/− mice OLs might be dependent on cholesterol coming from an external source for myelin synthesis in vivo. To negate this caveat, primary OL and CG4-OL cultures were differentiated in vitro to assess the role of SIRT2 in cholesterol biosynthesis. In mammalian cells, the presence of excess sterol induces feedback regulation by promoting the retention of SREBP-SCAP complex in the endoplasmic reticulum and inhibiting SREBP processing (Wang et al. 1994; Yang et al. 2002). To control the presence of cholesterol in the media, CG4-OLs and primary OLs were cultured in media containing normal serum (FBS) or delipidated serum (DLFBS). The loss of Sirt2 in primary OLs (Fig. 3), as well as knockdown of Sirt2 or overexpression of Sirt2 in CG4-OLs (Figs. 4 and 5), did not alter the expression of cholesterol biosynthetic genes and total cholesterol content.
The findings presented here are in agreement with Bobrowska et al. (2012) who did not find a change in the gene expression profile of enzymes involved in cholesterol biosynthesis in the cortex, hippocampus, or brain stem of Sirt2 −/− mice. Unlike neuronal cells (Luthi-Carter et al. 2010; Taylor et al. 2011), knockdown or overexpression of Sirt2 did not impact nuclear translocation of SREBP-2 in OLs (Fig. 6). This discrepancy could be due to cell type-specific differences in the expression pattern, as Sirt2 is more abundantly expressed by OLs than other cell types in the CNS (Zhang et al. 2014). Moreover, the predominate isoform in OLs is SIRT2.2 (Thangaraj et al. 2017; Zhu et al. 2012), whereas in neurons it is SIRT2.1 (Luthi-Carter et al. 2010) which may result in selective deacetylation targets. In summary, we have presented several lines of evidence (Fig. 7b) to conclude that Sirt2 does not impact cholesterol biosynthesis during OL differentiation and myelination in the developing CNS white matter.
References
Amemiya-Kudo M, Shimano H, Hasty AH, Yahagi N, Yoshikawa T, Matsuzaka T, Okazaki H, Tamura Y, Iizuka Y, Ohashi K, Osuga J, Harada K, Gotoda T, Sato R, Kimura S, Ishibashi S, Yamada N (2002) Transcriptional activities of nuclear SREBP-1a, -1c, and -2 to different target promoters of lipogenic and cholesterogenic genes. J Lipid Res 43:1220–1235
Baumann N, Pham-Dinh D (2001) Biology of oligodendrocyte and myelin in the mammalian central nervous system. Physiol Rev 81:871–927
Beirowski B, Gustin J, Armour SM, Yamamoto H, Viader A, North BJ, Michán S, Baloh RH, Golden JP, Schmidt RE, Sinclair DA, Auwerx J, Milbrandt J (2011) Sir-two-homolog 2 (Sirt2) modulates peripheral myelination through polarity protein Par-3/atypical protein kinase C (aPKC) signaling. Proc Natl Acad Sci 108:E952–E961
Bergles DE, Richardson WD (2015) Oligodendrocyte development and plasticity. Cold Spring Harb Perspect Biol 8:a020453
Bobrowska A, Donmez G, Weiss A, Guarente L, Bates G (2012) SIRT2 ablation has no effect on tubulin acetylation in brain, cholesterol biosynthesis or the progression of Huntington’s disease phenotypes in vivo. PLoS ONE 7:e34805
Brown MS, Goldstein JL (1997) The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell 89:331–340
Chen Y, Balasubramaniyan V, Peng J, Hurlock EC, Tallquist M, Li J, Lu QR (2007) Isolation and culture of rat and mouse oligodendrocyte precursor cells. Nat Protoc 2:1044–1051
Dietschy JM, Turley SD (2004) Cholesterol metabolism in the central nervous system during early development and in the mature animal. J Lipid Res 45:1375–1397
Doucette JR, Jiao R, Nazarali AJ (2010) Age-related and cuprizone-induced changes in myelin and transcription factor gene expression and in oligodendrocyte cell densities in the rostral corpus callosum of mice. Cell Mol Neurobiol 30:607–629
Eberlé D, Hegarty B, Bossard P, Ferré P, Foufelle F (2004) SREBP transcription factors: master regulators of lipid homeostasis. Biochimie 86:839–848
Giandomenico V, Simonsson M, Grönroos E, Ericsson J (2003) Coactivator-dependent acetylation stabilizes members of the SREBP family of transcription factors. Mol Cell Biol 23:2587–2599
Horton JD, Goldstein JL, Brown MS (2002) SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest 109:1125–1131
Horton JD, Shah NA, Warrington JA, Anderson NN, Park SW, Brown MS, Goldstein JL (2003) Combined analysis of oligonucleotide microarray data from transgenic and knockout mice identifies direct SREBP target genes. Proc Natl Acad Sci 100:12027–12032
Ji S, Doucette JR, Nazarali AJ (2011) Sirt2 is a novel in vivo downstream target of Nkx2.2 and enhances oligodendroglial cell differentiation. J Mol Cell Biol 3:351–359
Jurevics H, Morell P (1995) Cholesterol for synthesis of myelin Is made locally, not imported into brain. J Neurochem 64:895–901
Jurevics HA, Kidwai FZ, Morell P (1997) Sources of cholesterol during development of the rat fetus and fetal organs. J Lipid Res 38:723–733
Larocca JN, Norton WT (2007) Isolation of myelin. Curr Protoc Cell Biol Chapter 3:Unit3.25. doi:10.1002/0471143030.cb0325s33
Li W, Zhang B, Tang J, Cao Q, Wu Y, Wu C, Guo J, Ling E, Liang F (2007) Sirtuin 2, a mammalian homolog of yeast silent information regulator-2 longevity regulator, is an oligodendroglial protein that decelerates cell differentiation through deacetylating α-tubulin. J Neurosci 27:2606–2616
Louis JC, Magal E, Muir D, Manthorpe M, Varon S (1992) CG-4, A new bipotential glial cell line from rat brain, is capable of differentiating in vitro into either mature oligodendrocytes or type-2 astrocytes. J Neurosci Res 31:193–204
Luthi-Carter R, Taylor DM, Pallos J, Lambert E, Amore A, Parker A, Moffitt H, Smith DL, Runne H, Gokce O, Kuhn A, Xiang Z, Maxwell MM, Reeves SA, Bates GP, Neri C, Thompson LM, Marsh JL, Kazantsev AG (2010) SIRT2 inhibition achieves neuroprotection by decreasing sterol biosynthesis. Proc Natl Acad Sci 107:7927–7932
Mathews ES, Appel B (2016) Cholesterol biosynthesis supports myelin gene expression and axon ensheathment through modulation of P13K/Akt/mTor signaling. J Neurosci 36:7628–7639
Mathews ES, Mawdsley DJ, Walker M, Hines JH, Pozzoli M, Appel B (2014) Mutation of 3-hydroxy-3-methylglutaryl CoA synthase I reveals requirements for isoprenoid and cholesterol synthesis in oligodendrocyte migration arrest, axon wrapping, and myelin gene expression. J Neurosci 34:3402–3412
Muse ED, Jurevics H, Toews AD, Matsushima GK, Morell P (2001) Parameters related to lipid metabolism as markers of myelination in mouse brain. J Neurochem 76:77–86
Nicolay DJ, Doucette JR, Nazarali AJ (2007) Transcriptional control of oligodendrogenesis. Glia 55:1287–1299
Niu J, Wang L, Liu S, Li C, Kong J, Shen H, Xiao L (2012) An efficient and economical culture approach for the enrichment of purified oligodendrocyte progenitor cells. J Neurosci Methods 209:241–249
Norton WT, Poduslo SE (1973a) Myelination in rat brain: changes in myelin composition during brain maturation. J Neurochem 21:759–773
Norton WT, Poduslo SE (1973b) Myelination in rat brain: method of myelin isolation. J Neurochem 21:749–757
Ponugoti B, Kim D, Xiao Z, Smith Z, Miao J, Zang M, Wu S, Chiang C, Veenstra TD, Kemper JK (2010) SIRT1 deacetylates and inhibits SREBP-1C activity in regulation of hepatic lipid metabolism. J Biol Chem 285:33959–33970
Saher G, Simons M (2010) Cholesterol and myelin biogenesis. Subcell Biochem 51:489–508
Saher G, Brugger B, Lappe-Siefke C, Mobius W, Tozawa R, Wehr MC, Wieland F, Ishibashi S, Nave K (2005) High cholesterol level is essential for myelin membrane growth. Nat Neurosci 8:468–475
Saher G, Quintes S, Möbius W, Wehr MC, Krämer-Albers EM, Brügger B, Nave KA (2009) Cholesterol regulates the endoplasmic reticulum exit of the major membrane protein P0 required for peripheral myelin compaction. J Neurosci 29(19):6094–6104
Sakai J, Duncan EA, Rawson RB, Hua X, Brown MS, Goldstein JL (1996) Sterol-regulated release of SREBP-2 from cell membranes requires two sequential cleavages, one within a transmembrane segment. Cell 85:1037–1046
Simons M, Nave K (2015) Oligodendrocytes: myelination and axonal support. Cold Spring Harb Perspect Biol 8:a020479
Simons M, Krämer E, Thiele C, Stoffel W, Trotter J (2000) Assembly of myelin by association of proteolipid protein with cholesterol- and galactosylceramide-rich membrane domains. J Cell Biol 151:143–154
Southwood CM, Peppi M, Dryden S, Tainsky MA, Gow A (2007) Microtubule deacetylases, SirT2 and HDAC6, in the nervous system. Neurochem Res 32(2):187–195
Taylor DM, Balabadra U, Xiang Z, Woodman B, Meade S, Amore A, Maxwell MM, Reeves S, Bates GP, Luthi-Carter R, Lowden PAS, Kazantsev AG (2011) A brain-permeable small molecule reduces neuronal cholesterol by inhibiting activity of sirtuin 2 deacetylase. ACS Chem Biol 6:540–546
Thangaraj MP, Furber KL, Gan JK, Ji S, Sobchishin L, Doucette JR, Nazarali AJ (2017) RNA binding protein Quaking stabilizes Sirt2 mRNA during oligodendroglial differentiation. J Biol Chem 292:5166–5182
Verheijen MHG, Camargo N, Verdier V, Nadra K, de Preux Charles A, Médard J, Luoma A, Crowther M, Inouye H, Shimano H, Chen S, Brouwers JF, Helms JB, Feltri ML, Wrabetz L, Kirschner D, Chrast R, Smit AB (2009) SCAP is required for timely and proper myelin membrane synthesis. Proc Natl Acad Sci 106:21383–21388
Walker AK, Yang F, Jiang K, Ji J, Watts JL, Purushotham A, Boss O, Hirsch ML, Ribich S, Smith JJ, Israelian K, Westphal CH, Rodgers JT, Shioda T, Elson SL, Mulligan P, Najafi-Shoushtari H, Black JC, Thakur JK, Kadyk LC, Whetstine JR, Mostoslavsky R, Puigserver P, Li X, Dyson NJ, Hart AC, Näär AM (2010) Conserved role of SIRT1 orthologs in fasting-dependent inhibition of the lipid/cholesterol regulator SREBP. Genes Dev 24:1403–1417
Wang X, Sato R, Brown MS, Hua X, Goldstein JL (1994) SREBP-1, a membrane-bound transcription factor released by sterol-regulated proteolysis. Cell 77:53–62
Wang M, Doucette JR, Nazarali AJ (2011) Conditional Tet-regulated over-expression of Hoxa2 in CG4 cells increases their proliferation and delays their differentiation into oligodendrocyte-like cells expressing myelin basic protein. Cell Mol Neurobiol 31:875–886
Werner HB, Kuhlmann K, Shen S, Uecker M, Schardt A, Dimova K, Orfaniotou F, Dhaunchak A, Brinkmann BG, Möbius W, Guarente L, Casaccia-Bonnefil P, Jahn O, Nave K (2007) Proteolipid protein Is required for transport of sirtuin 2 into CNS myelin. J Neurosci 27:7717–7730
Werner HB, Krämer-Albers E, Strenzke N, Saher G, Tenzer S, Ohno-Iwashita Y, De Monasterio-Schrader P, Möbius W, Moser T, Griffiths IR, Nave K (2013) A critical role for the cholesterol-associated proteolipids PLP and M6B in myelination of the central nervous system. Glia 61:567–586
Yabe D, Xia Z, Adams CM, Rawson RB (2002) Three mutations in sterol-sensing domain of SCAP block interaction with insig and render SREBP cleavage insensitive to sterols. Proc Natl Acad Sci 99:16672–16677
Yang T, Espenshade PJ, Wright ME, Yabe D, Gong Y, Aebersold R, Goldstein JL, Brown MS (2002) Crucial step in cholesterol homeostasis: sterols promote binding of SCAP to INSIG-1, a membrane protein that facilitates retention of SREBPs in ER. Cell 110:489–500
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, Keeffe S, Phatnani HP, Guarnieri P, Caneda C, Ruderisch N, Deng S, Liddelow SA, Zhang C, Daneman R, Maniatis T, Barres BA, Wu JQ (2014) An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci 34:11929–11947
Zhu H, Zhao L, Wang E, Dimova N, Liu G, Feng Y, Cambi F (2012) The QKI-PLP pathway controls SIRT2 abundance in CNS myelin. Glia 60:69–82
Acknowledgements
This work was supported by an operating grant from Canadian Institutes of Health Research (CIHR) and the Saskatchewan Health Research Foundation (SHRF) (to A. J. N and J. R. D). A. J. N acknowledges the support from the Natural Sciences and Engineering Research Council of Canada (NSERC). M. P. T has been a recipient of a Dean’s Graduate Scholarship, University of Saskatchewan and is a current recipient of an endMS Doctoral Studentship award from the Multiple Sclerosis Society of Canada (MSSC). K. L. F is the recipient of an endMS Postdoctoral Fellowship award from MSSC.
Author information
Authors and Affiliations
Contributions
A. J. N and J. R. D conceived and coordinated the study. M. P. T designed, performed experiments, analyzed data, and drafted the manuscript. A. J. N revised the manuscript for interpretation of data and critical content, and approved the final version for submission. K. L. F contributed to writing and editing of the manuscript. L. S assisted with the analysis of data in Fig. 6. S. J assisted in experimental design.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Ethical Approval
The research was approved by the University of Saskatchewan’s Animal Research Ethics Board and adhered to the Canadian Council on Animal Care guidelines for humane animal use.
Rights and permissions
About this article
Cite this article
Thangaraj, M.P., Furber, K.L., Sobchishin, L. et al. Does Sirt2 Regulate Cholesterol Biosynthesis During Oligodendroglial Differentiation In Vitro and In Vivo?. Cell Mol Neurobiol 38, 329–340 (2018). https://doi.org/10.1007/s10571-017-0537-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-017-0537-6