Abstract
Microbiota are essential to normal immune development and there is growing recognition of its importance to human health and disease and deepening understanding of the complexity of host-microbe interactions in the human gut and other tissues. Commensal microbes not only can influence host immunity locally through impacts of bioactive microbial metabolites and direct interactions with epithelial cells and innate immune receptors but also can exert systemic immunomodulatory effects via impacts on host immune cells capable of trafficking beyond the gut. Emerging data suggest microbiota influence the development of multiple myeloma (MM), a malignancy of the immune system derived from immunoglobulin-producing bone marrow plasma cells, through the promotion of inflammation. Superior treatment outcomes for MM correlate with a higher abundance of commensal microbiota capable of influencing inflammatory responses through the production of butyrate. In patients with hematologic malignancies, higher levels of diversity of the gut microbiota correlate with superior outcomes after hematopoietic stem cell transplantation. Correlative data support the impact of commensal microbiota on survival, risk of infection, disease relapse, and graft-versus-host disease (GVHD) after transplant. In this review, we will discuss the current understanding of the role of host-microbe interactions and the inflammatory tumor microenvironment of multiple myeloma, discuss data describing the key role of microbiota in hematopoietic stem cell transplantation for treatment of hematologic malignancies, and highlight several possible concepts for interventions directed at the gut microbiota to influence treatment outcomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
1.1 Microbiota in human health and disease
The human body is home to complex communities of microorganisms collectively called the human microbiota or microbiome, which live in and on the body and interface with the host in the mucosal tissues, lungs, skin, mammary glands, and the gastrointestinal tract [1]. High-throughput sequencing enabled us to observe microbial communities [2] without the bias of artificial culture systems [3]. We have learned that the intestinal microbiota plays a vital role in the normal physiology of the host [4], facilitates energy harvest from the diet, and provides crucial signals for the development of the immune system. A recent report estimates the number of microbial cells is almost equal to that of the host, representing nearly 40 trillion cells [5]. For a particular individual, microbiota are transmitted vertically at birth and subsequently are shaped by diet and the microbiota of others in the same household, and can vary by age, geography, and genetics [6]. Environmental factors weigh more heavily than inherited features in determining the content of an individual’s microbiota [7]. Microbiota are relatively stable over time within the same individual. However, an individual’s microbiota can shift rapidly in response to dietary changes and exposure to antibiotics [8].
Detrimental changes in the microbiota affecting the diversity or community structure of the microbiota can negatively affect the host. Alpha diversity is a measure of the number of different microbes in a community and several states which negatively impact the microbiome lead to lower alpha diversity. Gut dysbiosis is a state of disruption in the content or structure of a host’s community of commensal microbiota (e.g., an overabundance of pathogenic microbes or loss of beneficial taxa). Conversely, a balanced gut microbial community can be described as being in a state of gut eubiosis [9]. Dysbiosis, loosely defined, has been associated with inflammatory bowel diseases [10,11,12], other autoimmune disorders [13], metabolic diseases such as diabetes [14,15,16], and cancer [17,18,19,20]. Microbiota have been directly linked to the pathogenesis of gastrointestinal cancers (colorectal [21], hepatocellular [22, 23], pancreatic [24]), and breast cancers as well as with hematologic malignancies such as lymphoma [25, 26] and multiple myeloma [27, 28].
In addition to impacting host immune defense against microbial pathogens, commensal microbiota also influence immune responses against malignancy [20]. Several recent studies have shown that microbiota can influence the response to cancer immunotherapy [19, 29,30,31,32]. In addition to anti-tumor responses, microbial dysbiosis has also been associated with immune checkpoint inhibitor-related colitis [33]. Microbiota can modulate the immune tumor microenvironment [34], drive anticancer immune responses [35], and mediate immune cell recovery after hematopoietic stem cell transplantation [36]. Emerging data suggest key relationships for microbiota in both the pathogenesis and response to cancer therapy. In this review, we will discuss the importance of host-microbe interactions in the biology and treatment of multiple myeloma and review the impact of microbiota on treatment outcomes after hematopoietic stem cell transplantation for hematologic cancers.
1.2 Microbiota and the development of the immune system
Host-microbe interactions are essential for the development of a functional immune system [37]. The critical role of microbiota in immune development has been established using germ-free (GF) animal models. GF mice are reared in isolators to prevent exposure to microbes and develop into adults without intestinal microbiota [38]. GF mice have multiple defects in mucosal immunity, including an absent mucous layer, reduced size and function of mesenteric lymph nodes and Peyer’s patches, and dysregulation of secreted immunoglobulin A (IgA) [39,40,41]. The absence of microbiota in GF mice leads to impaired tolerance mediated by fewer regulatory T cells (Tregs) in the gut-associated lymphoid tissue. Moreover, the Tregs present in GF animals have reduced functional suppressive capacity, and this functional impairment may impair tolerance and promote local and systemic inflammation [42]. These GF experiments show a key example by which microbiota might affect immune tolerance, promote inflammation, and potentially drive tumor cell growth in malignancies responsive to inflammation, such as multiple myeloma.
Commensal microbes provide necessary tonic stimulation to the host immune system and continuously challenge the immune system without eliciting an inflammatory response. Signaling from the commensal microbiota to the host is also thought to influence the activation threshold of innate immunity against viral pathogens [43].
Gnotobiotic studies in which defined microbiota are introduced into GF mice have allowed for interrogation of the role of specific host-microbe relationships in immune development [44]. For example, induction of Th17 cells by segmented filamentous bacteria has been demonstrated [45]. Th17 cells produce IL-17, a cytokine with conflicting roles in oncology. IL-17 has been shown to have anti-tumor effects [46] and tumor-promoting effects [47,48,49]. IL-17 has been shown to promote the progression of myeloma in the Vk*-Myc mouse model of multiple myeloma [27] and IL-17 producing lymphocytes also mediate lytic bone disease in myeloma [50].
1.3 Metabolites as mediators of beneficial host-microbe interactions: short-chain fatty acids
A core way the microbiome and host affect one another is through metabolic interchange mediated by small molecules produced by the host tissues and the microbiome. For example, SFCA (butyrate, acetate, lactate, and propionate) are created through anaerobic microbial metabolism and are waste products for the microbes’ anaerobic energy-generating mechanisms. Butyrate is an SCFA produced by microbial fermentation of dietary fibers and has been of particular interest due to its biological properties and observed correlation of increased abundance of butyrate producers with favorable health outcomes [51, 52]. Several extracellular receptors recognize butyrate, e.g. G-protein-coupled receptor (GPR) 41 (GPR41), also known as free fatty acid receptor 3 (FFAR3), GPR43 (FFAR3), and GPR109A [53, 54]. Butyrate is an agonist of intracellular peroxisome proliferator-activated receptor gamma (PPAR-γ) in colonic epithelial cells and lymphocytes. PPAR-γ activation by butyrate promotes apoptosis in normal lymphocytes, lymphoma [55], and malignant plasma cells in multiple myeloma [56]. Signaling induced by butyrate binding to its receptors leads to negative regulation of nuclear factor kappa B (NF-kB), a family of core transcription factors activated by a variety of canonical and non-canonical pathways which result in transcription of proinflammatory cytokines and which are central coordinators of innate and adaptive immune responses [57,58,59,60]. NF-kB signaling plays an important role in cancer development and disease progression in multiple myeloma [61]. Furthermore, SCFA impacts immunologic tolerance in the gut mucosa by promoting the differentiation of regulatory T cells (Treg) [62] and inhibiting immune effector cell activation through promoting TGF-β1 expression in epithelial cells [63]. Butyrate promotes host antibody responses [64], intestinal IgA secretion [65], and modulation of CD8 + T cell responses [66]. SCFA are not the only microbial metabolite with broad effects on the host. Indoles, sphingolipids, and others have all been implicated as functional mediators of host-microbe interaction in the gut.
1.4 Stimulation of host immune responses from the microbiota
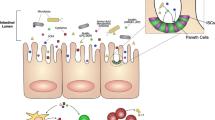
Host sensing of microbiota occurs through the binding of conserved microbial-associated molecular patterns (MAMPs) or pathogen-associated molecular patterns (PAMPs) with host pattern recognition receptors (PRRs) such as Toll-like receptors (TLR) and/or NOD-like receptors (NLR). Binding of MAMPs to PRRs can induce inflammatory signaling processes to defend the host from invasive pathogens [67]. Inflammasomes are large cytosolic multiprotein complexes expressed in monocytes, macrophages, granulocytes, dendritic cells, and epithelial cell osteoblasts, which are activated by a variety of microbial stimuli such PAMPs (e.g., lipopolysaccharide (LPS), flagellin) or host-derived danger-associated molecular patterns (DAMPs) through binding to TLR or NLR [68, 69]. Inflammasome activation mobilizes host immune responses through caspase-1 mediated proteolytic cleavage of inactive pro-cytokines pro-IL1-β and pro-IL-18 into their active forms and can induce a form of inflammatory cell death called pyroptosis [70]. IL-18 upregulates the production of antimicrobial peptides (AMPs) which are expressed by epithelial cells and mediate bacterial clearance [71], and AMP production is also promoted by SCFA [72]. IL-18 and IL-1β play an important role in host defense against microbial pathogens, but these cytokines may have detrimental effects in malignancy. For example, IL-18 induces IFN-γ [73] by T helper 1 (Th1) cells and natural killer (NK) cells, but may also promote malignancy as it is now recognized that tumor-promoting inflammation is a hallmark of cancer (discussed below) [74]. Systemic inflammation driven by inflammasome-mediated sensing of microbiota may drive inflammation-related cancers such as multiple myeloma [75, 76]. Impaired inflammasome activity due to mutations in NLR and reduced IL-18 production are each associated with dysbiosis, and both inflammasome dysfunction and activation by β-2-microglobulin have been linked to pathogenesis and progression of multiple myeloma [75,76,77]. Given the tumor-promoting nature of inflammasome-related cytokines IL-18 and IL-1 in multiple myeloma, it is possible certain microbiota may drive promotion of myeloma through activation of inflammasome (Fig. 1).
Impacts of gut microbiota on systemic inflammation and the bone marrow microenvironment of multiple myeloma. CTL = cytotoxic T lymphocyte, DC = dendritic cell, Eos = eosinophil, IL = interleukin, Glu = glutamate, HDACi = histone deacetylase inhibitor, MM = multiple myeloma, Mφ = macrophage. NK = natural killer cell, PC = plasma cell, SCFA = short-chain fatty acids, sIgA = secretory immunoglobulin A, Treg = regulatory T cell. Created with BioRender.com
This shaping of the immune response can, in turn, affect the microbial community. Two core mechanisms are AMPs and the production of secretory immunoglobulin A (IgA). IgA is produced by plasma cells mostly residing within Peyer’s patches within the gut and is secreted into the human gut lumen in large quantities (40–60 mg/kg/day of body weight) [78]. Secreted IgA binds to both commensal and pathogenic microbes and coats approximately 20–50% of the gut microbiota. Defective IgA production or secretion can lead to dysbiosis, bacterial invasion, and inflammatory disease [79, 80]. Inborn defects such as selective IgA deficiency or acquired hypogammaglobulinemia or secondary suppression of IgA production would also presumably impair the hosts’ ability to regulate its microbiota. Whether secretory IgA and microbiota bound by it are affected by the hypogammaglobulinemia caused by plasma cell dyscrasias is not known.
2 Microbiota, inflammation, and the pathogenesis of multiple myeloma
Multiple myeloma (MM) is a chronic and currently incurable malignancy of clonal plasma cells more common in older adults, with a median age of diagnosis in the United States (US) of 69 years [81]. MM represents 1% of new cancer cases in the US, and there are approximately 32,000 new diagnoses annually. All patients with myeloma evolve from a precursor state, monoclonal gammopathy of undetermined significance (MGUS), and smoldering multiple myeloma (SMM), which are prevalent in the population, affecting at least 3% of the general population over age 50. The incidence of myeloma increases with age. Although there are recurrent genomic events that play an important role in initiating and driving plasma cell neoplasms, similar events are found both in the malignant and premalignant states of the disease [82,83,84]. Chronic sterile inflammation associated with aging, also known as “inflammaging,” is correlated with cardiovascular disease [85] and the development of cancer [86]. IL-1 family cytokines are critical mediators of this type of inflammation [87]. IL-1 has been implicated in microbiota-induced inflammaging of hematopoietic stem cells in mice [88]. Clonal hematopoiesis (CH) driven by mutations in myeloid transcription factors such as DNMT3A and TET2 becomes more common with age and is associated with increased expression of pro-inflammatory cytokines and contributes to inflammaging [89]. Chronic inflammation associated with aging, CH, and the immune tumor microenvironment may play a role in driving malignant transformation in plasma cell neoplasia [90]. There is a great degree of unaccounted heterogeneity in the pathogenesis of myeloma and several recent reports suggest the microbiota may be implicated in this process, at least indirectly by modulating systemic inflammation. MM plasma cell growth and persistence are dependent upon the bone marrow tumor microenvironment which is both immunosuppressive and associated with inflammatory cytokines that promote malignant plasma cell growth, such as IL-6, IL-1β, IL-18, and IL-17 [91, 92].
Importantly, recent data suggest that microbiota-driven proinflammatory cytokine expression can influence the pathogenesis of MM (Fig. 1). The genetically modified immunocompetent Vκ*Myc mouse model in which C57BL/6 mice reliably develop spontaneous MM has provided an avenue to study the role of microbiota and inflammation in the pathogenesis of MM. Differences in the rate of progression of MM in this model in different geographic sites led to a subsequent investigation that demonstrated an association of certain microbiota with shorter time to progression in mice [27]. Calcinotto et al. demonstrated that higher abundance of Prevotella led to stimulation of T helper 17 (Th17) cell differentiation in the gut, with subsequent trafficking of these cells to the bone marrow tumor microenvironment where they release IL-17, which in turn both directly drives MM cell growth and stimulates eosinophils to produce IL-6 also driving MM progression [27]. In SMM, increased bone marrow IL-17 levels are associated with a higher risk of progression to active MM. IL-17 production driven by microbiota is well-established, particularly by segmented filamentous bacteria in the setting of inflammatory disorders [45], and butyrate has been shown to reduce IL-17 production in experimental colitis [93]. Methods to reduce inflammation may have therapeutic value in multiple myeloma, but as of yet, have had limited clinical utility other than corticosteroids. Although IL-6 has been recognized as a key paracrine and autocrine factor important for the growth and persistence of MM plasma cells, approaches to neutralize IL-6 have had limited success in clinical trials [94, 95]. A preliminary signal of efficacy from a phase 1 trial showed 2 of 13 patients with MM achieved complete responses (CR) after treatment with siltuximab [96]. Use of anti-IL-6 antibody siltuximab in combination with a triplet regimen (bortezomib, melphalan, and prednisone) increased the rate of very good partial responses (VGPR) over the triplet regimen alone [95]. However, this trial did not meet its primary endpoint of efficacy (CR rate) and came at the cost of an increased trend toward more infections with use of siltuximab [95]. A phase 2 trial of siltuximab in relapsed/refractory MM used alone or in combination with dexamethasone showed no objective clinical responses [94]. Use of the monoclonal antibody drug anakinra (anti-IL-1Rα) in smoldering MM patients attenuated C-reactive protein (CRP) levels in a subset of patients in whom a significantly longer interval to progression to active MM was noted [97]. These data suggest it may be important to modify the inflammatory microenvironment to influence the disease biology of myeloma, but that this approach may have more value prior to the onset of advanced or treatment-refractory disease [97].
There are preliminary data to suggest a potential correlation between certain microbial taxa and clinical outcomes to upfront treatment in multiple myeloma patients. Stool samples were collected to evaluate the gut microbiota in stool samples from MM patients who recently completed induction therapy with or without a stem cell transplant and a higher relative abundance of butyrate producers Faecalibacterium prausnitzii and Eubacterium hallii was noted in individuals with superior responses to therapy who were minimal residual disease (MRD) negative [98]. Follow-up data from this cohort showed an association of a diet with increased seafood and plant protein intake with higher abundance of butyrogenic bacteria and butyrate levels in stool samples and with sustained MRD negativity at 1 year after starting lenalidomide maintenance [99]. Associations of butyrate-producing bacteria with therapeutic efficacy in myeloma are preliminary and warrant further study to evaluate the impacts of diet and butyrate production on disease control, inflammation, and anti-tumor immunity in MM patients. Jian et al. compared microbiota in newly diagnosed myeloma patients at diagnosis (n = 19) and healthy gender and age-matched controls (n = 18), noting higher alpha diversity in MM patients, enrichment of nitrogen-recycling bacteria in the stool of MM patients, and lower abundance of SCFA-producing species in MM patients compared with healthy controls [100]. In this work, the authors noted that an increased relative abundance of Klebsiella spp. in newly diagnosed MM patients led to increased glutamate synthesis which promoted malignant plasma cell growth (Fig. 1).
Whether clinical approaches for modulation of the microbiota to reduce or impact inflammation, increase butyrate production, or eliminate possibly pathogenic microbiota would be feasible or efficacious as an approach has not yet been prospectively evaluated. Prophylactic antibiotics have been proposed as a method to reduce microbial-driven inflammation to impact the progression of MM. However, a higher rate of deaths from progressive MM in patients with newly diagnosed MM treated with 12 weeks of levofloxacin prophylaxis compared with placebo in the TEAMM study raises concern that microbiota injury caused by antibiotic prophylaxis may have unintended consequences and promote loss of immune control of myeloma [101,102,103]. Indeed, the importance of microbiota to responses to cancer immunotherapy and chemotherapy as demonstrated in other tumor types highlights that microbiota injury caused by antibiotic use could in theory adversely affect immune control of myeloma [30, 34, 104, 105]. The impact of other organisms beyond bacteria on myeloma pathogenesis and treatment outcomes such as commensal fungi or endogenous virii may also have relevance and may be assessed in future studies [106]. A plant-based dietary intervention in overweight and obese patients with the MM precursor conditions to reduce the risk of the progression to MM and evaluate its impacts on the microbiome is being evaluated in a prospective pilot phase clinical trial (NCT04920084).
3 Microbiota and stem cell transplantation for hematologic cancers
3.1 Allogeneic stem cell transplant
Evaluation of the microbiome and correlation with clinical outcomes was first noted in the field of stem cell transplantation. Before the development of immune checkpoint inhibitors, historically, the most common type of immunotherapy for cancer therapy has been hematopoietic stem cell transplantation for patients with hematologic malignancies, most commonly for leukemia and lymphoma and rarely in MM. Allogeneic hematopoietic stem cell transplantation (allo-HSCT) can be a potentially curative treatment for hematologic cancers. Prior to allo-HSCT, patients are treated with a conditioning regimen involving chemotherapy and/or radiation followed by an infusion of hematopoietic precursor cells from a donor who is matched for major histocompatibility complex antigens. Suppression of the recipient’s immune system allows for engraftment of the donor’s immune system to induce a graft-versus-tumor response to achieve durable immune control of the malignancy. A significant limitation of this approach is the potential for pathological recognition of the recipient tissues by the donor immune system, leading to graft-versus-host disease (GVHD), with potentially devastating consequences. Relapsed disease after allo-HSCT, toxic effects on organs, and risk of infection are significant sources of morbidity and mortality for patients receiving allo-HSCT. Microbiota have been shown to have powerful impacts on each of these outcomes and overall survival in hematopoietic stem cell transplantation [107,108,109,110,111,112,113,114,115,116,117,118].
3.2 Infection risk and antimicrobial prophylaxis: effects on the microbiome
Patients receiving allo-HSCT already have severely impacted gut microbiota with dramatically reduced alpha diversity at time of transplant. Iatrogenic dysbiosis, also referred to here as microbiota injury, worsens during the transplant course often when patients are exposed to antibiotics for treatment or prevention of infections [112, 118]. Patients treated with allo-HSCT are at high risk of bacterial infections during allo-HSCT due to a transient period of intense immunocompromise. Before transplantation, patients receive an intense conditioning regimen, which leads to neutropenia, injury to the oral and enteric mucosal lining, and an increased risk of bloodstream infections (BSI) [119,120,121]. Early in the history of transplant, approaches like gut decontamination with non-absorbable antibiotics and laminar airflow isolation were used to reduce the negative impact of microbial pathogens [122]. For years, efforts to reduce the risk of BSI from translocated oral and bowel microbiota in cancer patients have led to trials of numerous antibiotics, including neomycin and polymyxin, trimethoprim-sulfamethoxazole, and fluoroquinolones [123,124,125]. Several studies showed that antimicrobial prophylaxis led to a reduced rate of BSI in cancer patients without improving mortality rates [126,127,128]. A recent meta-analysis of 17 randomized trials of antimicrobial prophylaxis in 1,453 hematopoietic stem cell transplant recipients demonstrated prophylactic antibiotic use reduced incidence of febrile episodes, documented infection, and bacteremia but did not significantly affect all-cause mortality or infection-related mortality [129].
Using antibiotics to prevent or treat infections may have unintended consequences that may paradoxically increase the risk of secondary infections through multiple mechanisms, including suppression of myelopoiesis, immune, and hematopoietic reconstitution. Antibiotic prophylaxis may promote the emergence of resistant organisms and other infections such as Clostridium difficile (CDI) [130,131,132]. GF or antibiotic-treated mice are at increased risk of infection, which can be reversed through fecal microbiota transplantation [130, 131, 133]. Nutritional support from microbiota is a critical factor that impacts hematopoietic reconstitution after bone marrow transplantation in mice [134]. Depleting microbiota increases intestinal permeability [129], which may increase BSI risk.
There can be impacts globally on microbiota during HSCT that affect the risk of infection via effects on commensal microbial communities driven by specific organisms. Domination of the gut microbial community by a single organism (vancomycin-resistant enterococcus (VRE)) has been demonstrated to have a direct relationship to the risk of BSI in patients treated with allo-HSCT [108, 135], and can be precipitated by antibiotic use [115, 136]. During allo-HSCT, microbiota diversity is markedly reduced, and domination of the gut microbial community by certain bacterial species can predict the onset of bloodstream infection [108]. Patients who received allo-HSCT who developed E. coli or Klebsiella BSI were colonized with these organisms in the gut, supporting that the gut microbiota may also serve as a reservoir for BSI in this population [137]. A diverse microbiota can prevent hospital-acquired infection from VRE by inhibiting colonization of the individual by the organism [138, 139]. Certain commensal groups of anaerobic bacteria prevent infection from pathogenic microbes; depletion of specific microbiota capable of bile acid metabolism impacts susceptibility to CDI and can be reconstituted, restoring resistance to CDI [140]. During the first 6 months after allo-HSCT, 20–30% of patients develop a viral respiratory infection. Allo-HSCT recipient patients with higher abundances of butyrate-producing bacteria had a fivefold lower risk of viral lower respiratory tract infection in allo-HSCT [116].
Antibiotic exposure and microbiota injury may also affect outcomes after allo-HSCT through negative impacts on immune reconstitution [36]. Fecal microbial diversity at 3-month post-transplant was an independent predictor of CD4 T cell count after CD34-selected allo-HSCT, and higher relative abundance of microbes of the genus Staphylococcus was associated with impaired CD4 T cell recovery, suggesting an essential role of microbial diversity on immune recovery and potential negative impacts of specific microbiota on immune function [141].
Due to the concerns about potential harm from routine use of antibiotic prophylaxis, several groups, including the Centers for Disease Control and Prevention (CDC), Infectious Disease Society of America (IDSA), and the American Society of Transplantation and Cellular Therapy (ASTCT), recommend restraint against routine antibiotic prophylaxis [142]. Despite these recommendations, institutional practices vary in the use of antimicrobials to prevent infection [143, 144]. Antibiotic sparing approaches to reduce the risk of harm from infection while preserving the commensal microbiota are being considered at many transplant centers.
3.3 Graft-versus-host disease
Graft-versus-host-disease (GVHD) is a common complication of hematopoietic stem cell transplantation with potentially devastating consequences to the recipient, affecting 30–70% of allo-HSCT recipients [145,146,147]. Immune cells engrafting in the recipient recognize host tissues as foreign and lead to acute and or chronic immune injury by donor T cells to multiple recipient tissues, commonly gut, skin, oral mucosa, liver, and others [148]. GVHD commonly affects tissues that interface with microbiota, including skin, oral mucosa, gut, and liver. Experimental models of allo-HSCT in animals and human studies have demonstrated strong evidence that the gut microbiota affect the risk of GVHD and may directly contribute to the severity of GVHD. Enterococcus expansion in the gut is a common phenomenon observed in patients receiving allo-HSCT and has been linked to the onset of GVHD and reduced survival after transplant. Recent work showed that expansion of Enterococcus exacerbates the severity of GVHD in gnotobiotic mouse models of allo-HSCT, and directly linked promotion of Enterococcus growth to dietary lactose [149]. Dietary lactose depletion attenuated the severity of GVHD in mice, speaking to a possible mechanism through which microbiota might directly drive GVHD through impact of the diet [149].
Earlier animal studies of HCT showed superior survival of germ-free mice compared with conventional animals [150] and that GF mice were protected from GVHD [151, 152]. As GF mice are known to have aberrant immune development, this raises an important question: is reduced risk of GVHD observed in GF animal models of HCT due to absence of direct effects of microbiota or due to the dysfunction of an immune system that developed in a GF environment [153,154,155], or both?
There are a number of potential mechanisms by which microbiota might influence GVHD [156]. There may be impacts of microbiota on stimulation of host inflammation through host-sensing of MAMPs across epithelial tissues injured by conditioning chemotherapy or radiation [157].
Broad spectrum antibiotic use is associated with transplant-related mortality and GVHD-related mortality [112, 158]. A retrospective analysis of allo-HSCT patients who received the non-absorbable antibiotic rifaximin vs. prophylaxis with ciprofloxacin and metronidazole found those on rifaximin had lower rates of gastrointestinal GVHD-related treatment related mortality [159]. Antibiotic exposure in early peri-transplant period is associated with higher GVHD-associated treatment-related mortality [158]. Prolonged suppression of butyrate-producing microbiota is associated with a higher risk of acute GVHD and transplant-related mortality after allo-HSCT. Concentrations of human fecal SCFA butyrate and propionate may be decreased after HSCT and are likely influenced by antibiotic exposure and impact the incidence of GVHD [160]. In a retrospective case–control study, patients with higher systemic concentrations of microbial-derived SCFAs butyrate and propionate are associated with protection from GVHD [161]. It is important to note that although butyrate producers are associated with reduced rate of GVHD, in those with established GVHD, butyrate can have mixed effects. In GVHD, damaged crypts allow for butyrate to impair healing of the gut by inhibiting the regenerative function of intestinal stem cells [162]. Clinically, this manifests as an association of butyrate-generating microbes in the gut after the onset of severe GVHD with treatment-refractory GVHD [163]. This may suggest that a targeted antibiotic approach to inhibit butyrate production may have an initial benefit for steroid-refractory GVHD to allow gut healing, and perhaps could be followed with the restoration of microbiota using fecal microbiota transplantation. Associations of microbiota with GVHD in transplant have driven efforts to explore the modulation of the microbiota as a way to treat steroid-refractory GVHD and potentially prevent GVHD through preserving microbial community structure in the peri-transplant period.
3.4 Relapse
Collateral damage in the form of injury to the commensal microbiota caused by broad-spectrum antibiotic use not only impacts infection and GVHD but also affects an individual’s risk of disease relapse [114, 164]. Patients with hematologic malignancies already have marked alterations in their commensal microbiota at the time of transplant. Several groups have reported that adverse outcomes in hematopoietic cell transplant recipients are associated with microbiota injury caused in part by antibiotic exposure, manifested by expansions of potentially pathogenic microbiota and dramatic alterations in the diversity and content of microbes present in stool samples collected from these patients. Lower diversity in the intestinal microbiota is associated with worse mortality after allo-HSCT [110].
Concerning possible impacts of immune modulation of the host by microbiota, there are data to suggest important effects of the microbiota both on immune-mediated toxicity and immune control of the disease after allo-HSCT. Additionally, butyrogenic microbes may impact disease control in allo-HSCT, with patients with a higher relative abundance of Eubacterium limosum having a lower rate of disease relapse and longer overall survival after allo-HSCT [107]. In a large study evaluating microbiota and treatment outcomes after allo-HSCT, a higher relative abundance of Eubacterium limosum post-transplant was associated with a lower rate of disease relapse and increased overall survival, indicative of the role that butyrogenic microbes have in disease control after allo-HSCT [114].
In conclusion, it is becoming clear that microbiota are a key factor to consider in optimizing outcomes in allo-HSCT for hematologic cancers to minimize treatment-related mortality (TRM), prevent morbidity and mortality from infections, and reduce the incidence and severity of GVHD.
3.5 Autologous stem cell transplant
The use of autologous stem cells transplanted for reconstitution of bone marrow function after administration of high-dose chemotherapy allows for dose intensification for the treatment of hematologic malignancies. Autologous hematopoietic stem cell transplantation (auto-HSCT) using myeloablative high-dose melphalan conditioning is a standard approach used for the treatment of multiple myeloma which prolongs overall survival and progression-free survival compared with patients who do not receive this therapy [165, 166]. Comparable to the experience of patients receiving allo-HSCT, treatment outcomes from auto-HSCT correlate with changes in microbiota which may impact host immunity. Several groups have noted significant and reliable reductions in microbial diversity (microbiota injury) in lymphoma and myeloma patients treated with auto-HSCT during and after transplant [167, 168]. Additionally, lower peri-engraftment diversity in fecal samples of patients treated with auto-HSCT with lymphoma and multiple myeloma is associated with worse overall and progression-free survival [167]. It is unclear whether decreased microbial diversity may be a surrogate marker of other factors that could impact patient outcomes, or whether decreased microbial diversity is directly impacting patient outcomes. The exact mechanisms of how microbial diversity affects auto-HSCT outcomes also remain unclear. Although the mechanism of action of auto-HSCT has generally been thought to be related primarily to the cytotoxic effects of the conditioning regimen on tumor cells, there is evidence of immunological anti-tumor effects of auto-HSCT that may be impacted by microbiota injury. Auto-HSCT in a MM animal model leads to an augmentation of autologous T-cell-mediated control of MM [169], and in patients, reductions in regulatory T cell populations in the early post-transplant period may support antitumor immunity and use of immunotherapy for MM [170]. Melphalan has been shown to augment the effects of adoptive immunotherapy using CD4 + T cells [171]. These data further support possible beneficial effects of commensal microbes, including regeneration of the protective gut barrier and reduction of systemic inflammation, and raise questions about whether the microbiota may be a modifiable characteristic for patients receiving autologous-HSCT for MM.
4 The microbiome as prognostic marker and therapeutic target: future directions
As summarized above, the structure (e.g., alpha diversity), composition (specific microbes), and metabolic function of the gut microbiome (e.g., SCFA production) as estimated by techniques like 16S rRNA sequencing have been associated with outcomes for cancer patients, including MM and patients treated with stem cell transplantation. The microbiome relates to disease onset, progression, treatment response, and overall outcomes for people with hematologic malignancies, including MM. A remaining challenge is how to turn this insight into the improved determination of prognosis (e.g., the gut microbiome as a clinically relevant biomarker) and to use the microbiome to improve outcomes.
4.1 Limitations of microbiome analysis
Providing patients with a more accurate prognosis of their disease using personalized microbiome data is an enormous unrealized opportunity but is one currently limited by fundamental biological and technical challenges. The dominant technique for microbiome science has been 16S rRNA gene variable region amplicon sequencing. This technique is mature, cost-effective, and effective at being able to characterize metrics like alpha diversity and composition (i.e., a lack of typical butyrate-generating microbes), but 16S rRNA gene variable region sequencing is highly sensitive to specific technical details (which PCR primers are selected, PCR conditions, sequencing library preparation) as well as batch effects (even with all technical details the same, one can achieve different results from the same underlying community from different batches of reagents and/or sequencers used). This technical variability has limited the clinical applicability of 16S rRNA gene variable region amplicon sequencing. WGS is moderately less subject to technical variance (e.g., primer selection is less of a concern) but is complicated by increased cost, increased computational challenges, and reduced sensitivity for rarer organisms. The underlying person-to-person variance in the microbiome adds another layer of complexity. Targeted metabolomics may end up the most tractable technique. Measurement of SCFA levels (particularly butyrate) in the stool and serum may be a viable approach to improve prognostic accuracy for a patient.
4.2 The gut microbiome as a therapeutic target in MM and HCT
Beyond using the microbiome to improve our prediction of outcomes for MM and HCT patients, there is an opportunity to improve outcomes by directly affecting the microbiome. A combination of doing less harm to the gut microbiome and direct manipulations of the composition and metabolic output of the gut microbiome are all promising avenues based on our knowledge to date.
Doing less harm with careful antibiotic use and selection is a core opportunity to improve outcomes. In hematopoietic stem cell transplantation, judicious antibiotic use or antibiotic-sparing approaches to reduce microbiota injury may limit GVHD and infectious risk, and potentially reduce the risk of relapse. Although strategies that limit microbiota injury during treatment of patients with hematologic cancers undergoing treatment with immunotherapies could reduce harmful treatment outcomes, increased general risk of infections in this population makes some injury through broad-spectrum antibiotic use unavoidable; thus, methods to repair or replenish a healthy microbial community are needed.
Efforts to directly manipulate the microbiota through nutrition, prebiotic, or probiotic approaches may optimize the microbiome for improving treatment outcomes. Patients with MM with a diet that contains specific nutrients may impact clinical outcomes diets [172], suggesting a possible role for dietary intervention to impact the disease course. Reduction of inflammation through modulation of the microbiota by various methods may impact malignancies such as MM which are driven by inflammatory cytokine signaling. Additionally, hematopoietic stem cell transplantation remains an integral part of the treatment paradigm for patients with hematologic cancers. Transfer of healthy microbiota through fecal microbiota transplantation is an established therapy for the treatment of Clostridium difficile infection [173]. Fecal microbiota transplantation has been evaluated as a method to restore the diversity of the gut microbiota after injury associated with allo-HSCT [174, 175] and as a therapy for patients with steroid-resistant acute GVHD [176,177,178]. The potential risk of the transfer of drug-resistant pathogens is a complicating feature of fecal microbiota transplantation [179] and underscores the need for rigor in selecting and screening stool donors. Targeted transfer of engineered microbial communities is another approach aimed at reducing the risk of transmission of antibiotic-resistant pathogens in the preclinical stage of development. Multicenter studies show a strong negative effect of microbiota injury, and avenues to address this important problem may have significant benefits for patients with hematologic cancers.
An under-addressed challenge with any microbiome manipulation is the complexity and contextuality of host-microbe interactions. Even for the relatively well-characterized effects and mechanisms by which butyrate produced by microbes affects cancer outcomes, we have discovered some conflicting effects (i.e., reduced risk of developing GVHD, but perhaps increased treatment-refractory GVHD with butyrate). Embracing and acknowledging the contextual and complex effects of the microbiome during MM treatment will be key to the successful development of novel microbiome-oriented therapies.
5 Conclusions
In summary, the microbiota are essential to immune function and several studies have shown a potential role for the microbiota in the pathogenesis of hematologic malignancies such as multiple myeloma through impacts on inflammatory signaling pathways and host metabolism. Microbial metabolites can impact immunity and influence outcomes in patients with myeloma and hematologic malignancies. In patients, increased microbial diversity in allogeneic and autologous stem cell transplantation is associated with superior outcomes. The evaluation of potential strategies to impact the microbiota to improve outcomes for patients with hematologic malignancies is underway.
References
Human Microbiome Project Consortium. (2012). Structure, function and diversity of the healthy human microbiome. Nature, 486(7402), 207–214. https://doi.org/10.1038/nature11234
Ley, R. E., Hamady, M., Lozupone, C., Turnbaugh, P. J., Ramey, R. R., Bircher, J. S., … Gordon, J. I. (2008). Evolution of mammals and their gut microbes. Science, 320(5883), 1647–1651. https://doi.org/10.1126/science.1155725
Bik, E. M. (2016). The hoops, hopes, and hypes of human microbiome research. The Yale journal of biology and medicine, 89(3), 363–373. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/27698620. Accessed 3 Jan 2022
Belkaid, Y., & Hand, T. W. (2014). Role of the microbiota in immunity and inflammation. Cell, 157(1), 121–141. https://doi.org/10.1016/j.cell.2014.03.011
Sender, R., Fuchs, S., & Milo, R. (2016). Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell, 164(3), 337–340. https://doi.org/10.1016/j.cell.2016.01.013
Yatsunenko, T., Rey, F. E., Manary, M. J., Trehan, I., Dominguez-Bello, M. G., Contreras, M., … Gordon, J. I. (2012). Human gut microbiome viewed across age and geography. Nature, 486(7402), 222–227. https://doi.org/10.1038/nature11053
Rothschild, D., Weissbrod, O., Barkan, E., Kurilshikov, A., Korem, T., Zeevi, D., … Segal, E. (2018). Environment dominates over host genetics in shaping human gut microbiota. Nature, 555(7695), 210–215. https://doi.org/10.1038/nature25973
David, L. A., Maurice, C. F., Carmody, R. N., Gootenberg, D. B., Button, J. E., Wolfe, B. E., … Turnbaugh, P. J. (2014). Diet rapidly and reproducibly alters the human gut microbiome. Nature, 505(7484), 559–563. https://doi.org/10.1038/nature12820
Iebba, V., Totino, V., Gagliardi, A., Santangelo, F., Cacciotti, F., Trancassini, M., … Schippa, S. (2016). Eubiosis and dysbiosis: The two sides of the microbiota. The new microbiologica, 39(1), 1–12. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/26922981. Accessed 3 Jan 2022
Kostic, A. D., Xavier, R. J., & Gevers, D. (2014). The microbiome in inflammatory bowel disease: Current status and the future ahead. Gastroenterology, 146(6), 1489–1499. https://doi.org/10.1053/j.gastro.2014.02.009
Dalal, S. R., & Chang, E. B. (2014). The microbial basis of inflammatory bowel diseases. The Journal of clinical investigation, 124(10), 4190–4196. https://doi.org/10.1172/JCI72330
Lloyd-Price, J., Arze, C., Ananthakrishnan, A. N., Schirmer, M., Avila-Pacheco, J., Poon, T. W., … Huttenhower, C. (2019). Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature, 569(7758), 655–662. https://doi.org/10.1038/s41586-019-1237-9
Dehner, C., Fine, R., & Kriegel, M. A. (2019). The microbiome in systemic autoimmune disease: Mechanistic insights from recent studies. Current opinion in rheumatology, 31(2), 201–207. https://doi.org/10.1097/BOR.0000000000000574
Paun, A., Yau, C., & Danska, J. S. (2017). The influence of the microbiome on type 1 diabetes. The Journal of Immunology. Retrieved from https://www.jimmunol.org/content/198/2/590.short. Accessed 3 Jan 2022
Udayappan, S., Manneras-Holm, L., Chaplin-Scott, A., Belzer, C., Herrema, H., Dallinga-Thie, G. M., … Nieuwdorp, M. (2016). Oral treatment with Eubacterium hallii improves insulin sensitivity in db/db mice. NPJ Biofilms Microbiomes, 2, 16009. https://doi.org/10.1038/npjbiofilms.2016.9
Wu, H., Esteve, E., Tremaroli, V., Khan, M. T., Caesar, R., Mannerås-Holm, L., … Bäckhed, F. (2017). Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nature medicine, 23(7), 850–858. https://doi.org/10.1038/nm.4345
Elinav, E., Garrett, W. S., Trinchieri, G., & Wargo, J. (2019). The cancer microbiome. Nature reviews. Cancer, 19(7), 371–376. https://doi.org/10.1038/s41568-019-0155-3
Xavier, J. B., Young, V. B., Skufca, J., Ginty, F., Testerman, T., Pearson, A. T., … Wargo, J. A. (2020). The cancer microbiome: Distinguishing direct and indirect effects requires a systemic view. Trends in cancer research, 6(3), 192–204. https://doi.org/10.1016/j.trecan.2020.01.004
Routy, B., Gopalakrishnan, V., Daillère, R., Zitvogel, L., Wargo, J. A., & Kroemer, G. (2018). The gut microbiota influences anticancer immunosurveillance and general health. Nature reviews. Clinical oncology, 15(6), 382–396. https://doi.org/10.1038/s41571-018-0006-2
Gopalakrishnan, V., Helmink, B. A., Spencer, C. N., Reuben, A., & Wargo, J. A. (2018). The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell, 33(4), 570–580. https://doi.org/10.1016/j.ccell.2018.03.015
Yu, A. I., & Chen, G. Y. (2021). The gut microbiome and colorectal cancer. In J. Sun (Ed.), Inflammation, Infection, and Microbiome in Cancers: Evidence, Mechanisms, and Implications (pp. 63–96). Cham: Springer International Publishing. https://doi.org/10.1007/978-3-030-67951-4_3
Ren, Z., Li, A., Jiang, J., Zhou, L., Yu, Z., Lu, H., … Zheng, S. (2019). Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut, 68(6), 1014–1023. https://doi.org/10.1136/gutjnl-2017-315084
Roderburg, C., & Luedde, T. (2014). The role of the gut microbiome in the development and progression of liver cirrhosis and hepatocellular carcinoma. Gut microbes, 5(4), 441–445. https://doi.org/10.4161/gmic.29599
Pushalkar, S., Hundeyin, M., Daley, D., Zambirinis, C. P., Kurz, E., Mishra, A., … Miller, G. (2018). The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer discovery, 8(4), 403–416. https://doi.org/10.1158/2159-8290.CD-17-1134
Portlock, C. S., Hamlin, P. A., Gerecitano, J. F., Noy, A., Palomba, M. L., Walkley, J., … Markowitz, A. J. (2015). A positive prospective trial of antibiotic therapy in advanced stage, non-bulky indolent lymphoma. Tumor Microenviron Ther, 2(1), 14–18. https://doi.org/10.1515/tumor-2015-0001
Wotherspoon, A. C., Doglioni, C., Diss, T. C., Pan, L., Moschini, A., de Boni, M., & Isaacson, P. G. (1993). Regression of primary low-grade B-cell gastric lymphoma of mucosa-associated lymphoid tissue type after eradication of Helicobacter pylori. The Lancet, 342(8871), 575–577. https://doi.org/10.1016/0140-6736(93)91409-f
Calcinotto, A., Brevi, A., Chesi, M., Ferrarese, R., Garcia Perez, L., Grioni, M., … Bellone, M. (2018). Microbiota-driven interleukin-17-producing cells and eosinophils synergize to accelerate multiple myeloma progression. Nature communications, 9(1), 4832. https://doi.org/10.1038/s41467-018-07305-8
Jian, X., Zhu, Y., Ouyang, J., Lei, Q., Xia, J., Zhang, J., … Zhou, W. (2019). Alterations of gut microbiome accelerate multiple myeloma progression by increasing the relative abundances of nitrogen recycling bacteria. Blood, 134(Supplement_1), 688–688. https://doi.org/10.1182/blood-2019-125051
Frankel, A. E., Coughlin, L. A., Kim, J., Froehlich, T. W., Xie, Y., Frenkel, E. P., & Koh, A. Y. (2017). Metagenomic shotgun sequencing and unbiased metabolomic profiling identify specific human gut microbiota and metabolites associated with immune checkpoint therapy efficacy in melanoma patients. Neoplasia , 19(10), 848–855. Retrieved from https://www.sciencedirect.com/science/article/pii/S1476558617302385. Accessed 3 Jan 2022
Gopalakrishnan, V., Spencer, C. N., Nezi, L., Reuben, A., Andrews, M. C., Karpinets, T. V., … Wargo, J. A. (2018). Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science, 359(6371), 97–103. https://doi.org/10.1126/science.aan4236
Chaput, N., Lepage, P., Coutzac, C., Soularue, E., Le Roux, K., Monot, C., … Carbonnel, F. (2017). Baseline gut microbiota predicts clinical response and colitis in metastatic melanoma patients treated with ipilimumab. Annals of oncology: official journal of the European Society for Medical Oncology / ESMO, 28(6), 1368–1379. https://doi.org/10.1093/annonc/mdx108
Vetizou, M., Pitt, J. M., Daillere, R., Lepage, P., Waldschmitt, N., Flament, C., … Zitvogel, L. (2015). Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science, 350(6264), 1079–1084. https://doi.org/10.1126/science.aad1329
Dubin, K., Callahan, M. K., Ren, B., Khanin, R., Viale, A., Ling, L., … Wolchok, J. D. (2016). Intestinal microbiome analyses identify melanoma patients at risk for checkpoint-blockade-induced colitis. Nature communications, 7, 10391. https://doi.org/10.1038/ncomms10391
Iida, N., Dzutsev, A., Stewart, C. A., Smith, L., Bouladoux, N., Weingarten, R. A., … Goldszmid, R. S. (2013). Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science, 342(6161), 967–970. https://doi.org/10.1126/science.1240527
Paulos, C. M., Wrzesinski, C., Kaiser, A., Hinrichs, C. S., Chieppa, M., Cassard, L., … Restifo, N. P. (2007). Microbial translocation augments the function of adoptively transferred self/tumor-specific CD8+ T cells via TLR4 signaling. The Journal of clinical investigation, 117(8), 2197–2204. https://doi.org/10.1172/JCI32205
Schluter, J., Peled, J. U., Taylor, B. P., Markey, K. A., Smith, M., Taur, Y., … Xavier, J. B. (2020). The gut microbiota is associated with immune cell dynamics in humans. Nature, 588(7837), 303–307. https://doi.org/10.1038/s41586-020-2971-8
Honda, K., & Littman, D. R. (2016). The microbiota in adaptive immune homeostasis and disease. Nature, 535(7610), 75–84. https://doi.org/10.1038/nature18848
Al-Asmakh, M., & Zadjali, F. (2015). Use of germ-free animal models in microbiota-related research. Journal of microbiology and biotechnology, 25(10), 1583–1588. https://doi.org/10.4014/jmb.1501.01039
Johansson, M. E. V., Jakobsson, H. E., Holmén-Larsson, J., Schütte, A., Ermund, A., Rodríguez-Piñeiro, A. M., … Hansson, G. C. (2015). Normalization of host intestinal mucus layers requires long-term microbial colonization. Cell Host & Microbe. https://doi.org/10.1016/j.chom.2015.10.007
Hapfelmeier, S., Lawson, M. A. E., Slack, E., Kirundi, J. K., Stoel, M., Heikenwalder, M., … Macpherson, A. J. (2010). Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science, 328(5986), 1705–1709. https://doi.org/10.1126/science.1188454
Spiljar, M., Merkler, D., & Trajkovski, M. (2017). The immune system bridges the gut microbiota with systemic energy homeostasis: Focus on TLRs, mucosal barrier, and SCFAs. Frontiers in immunology, 8, 1353. https://doi.org/10.3389/fimmu.2017.01353
Ostman, S., Rask, C., Wold, A. E., Hultkrantz, S., & Telemo, E. (2006). Impaired regulatory T cell function in germ-free mice. European journal of immunology, 36(9), 2336–2346. https://doi.org/10.1002/eji.200535244
Abt, M. C., Osborne, L. C., Monticelli, L. A., Doering, T. A., Alenghat, T., Sonnenberg, G. F., … Artis, D. (2012). Commensal bacteria calibrate the activation threshold of innate antiviral immunity. Immunity, 37(1), 158–170. https://doi.org/10.1016/j.immuni.2012.04.011
Goodman, A. L., Kallstrom, G., Faith, J. J., Reyes, A., Moore, A., Dantas, G., & Gordon, J. I. (2011). Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proceedings of the National Academy of Sciences of the United States of America, 108(15), 6252–6257. https://doi.org/10.1073/pnas.1102938108
Ivanov, I. I., Atarashi, K., Manel, N., Brodie, E. L., Shima, T., Karaoz, U., … Littman, D. R. (2009). Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell, 139(3), 485–498. https://doi.org/10.1016/j.cell.2009.09.033
Muranski, P., & Restifo, N. P. (2013). Essentials of Th17 cell commitment and plasticity. Blood, 121(13), 2402–2414. https://doi.org/10.1182/blood-2012-09-378653
McAllister, F., Bailey, J. M., Alsina, J., Nirschl, C. J., Sharma, R., Fan, H., … Leach, S. D. (2014). Oncogenic Kras activates a hematopoietic-to-epithelial IL-17 signaling axis in preinvasive pancreatic neoplasia. Cancer cell, 25(5), 621–637. https://doi.org/10.1016/j.ccr.2014.03.014
Tartour, E., Fossiez, F., Joyeux, I., Galinha, A., Gey, A., Claret, E., … Sautès-Fridman, C. (1999). Interleukin 17, a T-cell-derived cytokine, promotes tumorigenicity of human cervical tumors in nude mice. Cancer research, 59(15), 3698–3704. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/10446984. Accessed 3 Jan 2022
Wang, L., Yi, T., Kortylewski, M., Pardoll, D. M., Zeng, D., & Yu, H. (2009). IL-17 can promote tumor growth through an IL-6-Stat3 signaling pathway. The Journal of experimental medicine, 206(7), 1457–1464. jem.20090207 [pii] https://doi.org/10.1084/jem.20090207
Noonan, K., Marchionni, L., Anderson, J., Pardoll, D., Roodman, G. D., & Borrello, I. (2010). A novel role of IL-17–producing lymphocytes in mediating lytic bone disease in multiple myeloma. Blood, 116(18), 3554–3563. https://doi.org/10.1182/blood-2010-05-283895
Baxter, N. T., Schmidt, A. W., Venkataraman, A., Kim, K. S., Waldron, C., Schmidt, T. M., & Blaser, M. J. (2019). Dynamics of human gut microbiota and short-chain fatty acids in response to dietary interventions with three fermentable fibers. mBio, 10(1). https://doi.org/10.1128/mBio.02566-18
Davis, J. A., Collier, F., Mohebbi, M., Pasco, J. A., Shivappa, N., Hébert, J. R., … Loughman, A. (2021). The associations of butyrate-producing bacteria of the gut microbiome with diet quality and muscle health. Gut Microbiome, 2(e2), 1–36. https://doi.org/10.1017/gmb.2021.2
Thangaraju, M., Cresci, G. A., Liu, K., Ananth, S., Gnanaprakasam, J. P., Browning, D. D., … Ganapathy, V. (2009). GPR109A is a G-protein-coupled receptor for the bacterial fermentation product butyrate and functions as a tumor suppressor in colon. Cancer research, 69(7), 2826–2832. https://doi.org/10.1158/0008-5472.CAN-08-4466
Wu, J., Zhou, Z., Hu, Y., & Dong, S. (2012). Butyrate-induced GPR41 activation inhibits histone acetylation and cell growth. Yi chuan xue bao [Journal of genetics and genomics], 39(8), 375–384. https://doi.org/10.1016/j.jgg.2012.05.008
Padilla, J., Leung, E., & Phipps, R. P. (2002). Human B lymphocytes and B lymphomas express PPAR-γ and are killed by PPAR-γ agonists. Clinical immunology, 103(1), 22–33. https://doi.org/10.1006/clim.2001.5181
Garcia-Bates, T. M., Bernstein, S. H., & Phipps, R. P. (2008). Peroxisome proliferator-activated receptor γ overexpression suppresses growth and induces apoptosis in human multiple myeloma cells. Clinical Cancer Research. https://doi.org/10.1158/1078-0432.ccr-08-0457
Molitor, J. A., Walker, W. H., Doerre, S., Ballard, D. W., & Greene, W. C. (1990). NF-kappa B: A family of inducible and differentially expressed enhancer-binding proteins in human T cells. Proceedings of the National Academy of Sciences of the United States of America, 87(24), 10028–10032. https://doi.org/10.1073/pnas.87.24.10028
Guilloteau, P., Martin, L., Eeckhaut, V., Ducatelle, R., Zabielski, R., & Van Immerseel, F. (2010). From the gut to the peripheral tissues: The multiple effects of butyrate. Nutrition research reviews, 23(2), 366–384. https://doi.org/10.1017/S0954422410000247
Segain, J.-P. (2000). Butyrate inhibits inflammatory responses through NFkappa B inhibition: Implications for Crohn’s disease. Gut. https://doi.org/10.1136/gut.47.3.397
Lee, C., Kim, B. G., Kim, J. H., Chun, J., Im, J. P., & Kim, J. S. (2017). Sodium butyrate inhibits the NF-kappa B signaling pathway and histone deacetylation, and attenuates experimental colitis in an IL-10 independent manner. International immunopharmacology, 51, 47–56. https://doi.org/10.1016/j.intimp.2017.07.023
Karin, M. (2006). Nuclear factor-κB in cancer development and progression. Nature. https://doi.org/10.1038/nature04870
Furusawa, Y., Obata, Y., Fukuda, S., Endo, T. A., Nakato, G., Takahashi, D., … Ohno, H. (2013). Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature, 504(7480), 446–450. https://doi.org/10.1038/nature12721
Martin-Gallausiaux, C., Béguet-Crespel, F., Marinelli, L., Jamet, A., Ledue, F., Blottière, H. M., & Lapaque, N. (2018). Butyrate produced by gut commensal bacteria activates TGF-beta1 expression through the transcription factor SP1 in human intestinal epithelial cells. Scientific reports, 8(1), 9742. https://doi.org/10.1038/s41598-018-28048-y
Kim, M., Qie, Y., Park, J., & Kim, C. H. (2016). Gut microbial metabolites fuel host antibody responses. Cell host & microbe, 20(2), 202–214. https://doi.org/10.1016/j.chom.2016.07.001
Wu, W., Sun, M., Chen, F., Cao, A. T., Liu, H., Zhao, Y., … Cong, Y. (2017). Microbiota metabolite short-chain fatty acid acetate promotes intestinal IgA response to microbiota which is mediated by GPR43. Mucosal immunology, 10(4), 946–956. https://doi.org/10.1038/mi.2016.114
Bachem, A., Makhlouf, C., Binger, K. J., de Souza, D. P., Tull, D., Hochheiser, K., … Bedoui, S. (2019). Microbiota-derived short-chain fatty acids promote the memory potential of antigen-activated CD8+ T cells. Immunity, 51(2), 285-297.e5. https://doi.org/10.1016/j.immuni.2019.06.002
Kawasaki, T., & Kawai, T. (2014). Toll-like receptor signaling pathways. Frontiers in immunology, 5, 461. https://doi.org/10.3389/fimmu.2014.00461
Broz, P., & Dixit, V. M. (2016). Inflammasomes: Mechanism of assembly, regulation and signalling. Nature reviews. Immunology, 16(7), 407–420. https://doi.org/10.1038/nri.2016.58
Chassaing, B., Ley, R. E., & Gewirtz, A. T. (2014). Intestinal epithelial cell toll-like receptor 5 regulates the intestinal microbiota to prevent low-grade inflammation and metabolic syndrome in mice. Gastroenterology, 147(6), 1363–77.e17. https://doi.org/10.1053/j.gastro.2014.08.033
Chen, G. Y. (2017). Regulation of the gut microbiome by inflammasomes. Free radical biology & medicine, 105, 35–40. https://doi.org/10.1016/j.freeradbiomed.2016.11.011
Levy, M., Thaiss, C. A., Zeevi, D., Dohnalová, L., Zilberman-Schapira, G., Mahdi, J. A., … Elinav, E. (2015). Microbiota-modulated metabolites shape the intestinal microenvironment by regulating NLRP6 inflammasome signaling. Cell, 163(6), 1428–1443. https://doi.org/10.1016/j.cell.2015.10.048
Zhao, Y., Chen, F., Wu, W., Sun, M., Bilotta, A. J., Yao, S., … Cong, Y. (2018). GPR43 mediates microbiota metabolite SCFA regulation of antimicrobial peptide expression in intestinal epithelial cells via activation of mTOR and STAT3. Mucosal Immunology. https://doi.org/10.1038/mi.2017.118
Novick, D., Kim, S., Kaplanski, G., & Dinarello, C. A. (2013). Interleukin-18, more than a Th1 cytokine. Seminars in immunology, 25(6), 439–448. https://doi.org/10.1016/j.smim.2013.10.014
Hanahan, D., & Weinberg, R. A. (2011). Hallmarks of cancer: The next generation. Cell, 144(5), 646–674. https://doi.org/10.1016/j.cell.2011.02.013
Hofbauer, D., Mougiakakos, D., Broggini, L., Zaiss, M., Büttner-Herold, M., Bach, C., … Bruns, H. (2021). β2-microglobulin triggers NLRP3 inflammasome activation in tumor-associated macrophages to promote multiple myeloma progression. Immunity. https://doi.org/10.1016/j.immuni.2021.07.002
Li, Y., Li, N., Yan, Z., Li, H., Chen, L., Zhang, Z., … Li, Z. (2016). Dysregulation of the NLRP3 inflammasome complex and related cytokines in patients with multiple myeloma. Hematology , 21(3), 144–151. https://doi.org/10.1179/1607845415Y.0000000029
Zhao, X., Hua, M., Yan, S., Yu, J., Han, F., Zhong, C., … Ma, D. (2018). The genetic polymorphisms of NLRP3 inflammasome associated with T helper cells in patients with multiple myeloma. Journal of immunology research, 2018, 7569809. https://doi.org/10.1155/2018/7569809
Macpherson, A. J., McCoy, K. D., Johansen, F. E., & Brandtzaeg, P. (2007). The immune geography of IgA induction and function. Mucosal Immunology, 2008(1), 11–22. PMID: 19079156.
Chiba, T., Honjo, T., & Fagarasan, S. (2004). Aberrant expansion of segmented filamentous bacteria in IgA-deficient gut. Proceedings of the. Retrieved from https://www.pnas.org/content/101/7/1981.short. Accessed 3 Jan 2022
Suzuki, K., & Nakajima, A. (2014). New aspects of IgA synthesis in the gut. International immunology, 26(9), 489–494. https://doi.org/10.1093/intimm/dxu059
SEER, National Cancer Institute. (n.d.). SEER cancer stat facts: Myeloma. National Cancer Institute. Bethesda, MD. SEER Cancer Stat Facts: Myeloma. Retrieved September 19, 2020, from https://seer.cancer.gov/statfacts/html/mulmy.html. Accessed 3 Jan 2022
Bolli, N., Maura, F., Minvielle, S., Gloznik, D., Szalat, R., Fullam, A., … Munshi, N. (2018). Genomic patterns of progression in smoldering multiple myeloma. Nature communications, 9(1), 3363. https://doi.org/10.1038/s41467-018-05058-y
Oben, B., Froyen, G., Maclachlan, K. H., Leongamornlert, D., Abascal, F., Zheng-Lin, B., … Maura, F. (2021). Whole-genome sequencing reveals progressive versus stable myeloma precursor conditions as two distinct entities. Nature communications, 12(1), 1861. https://doi.org/10.1038/s41467-021-22140-0
Maura, F., Bolli, N., Angelopoulos, N., Dawson, K. J., Leongamornlert, D., Martincorena, I., … Campbell, P. J. (2019). Genomic landscape and chronological reconstruction of driver events in multiple myeloma. Nature communications, 10(1), 3835. https://doi.org/10.1038/s41467-019-11680-1
Ferrucci, L., & Fabbri, E. (2018). Inflammageing: Chronic inflammation in ageing, cardiovascular disease, and frailty. Nature reviews. Cardiology, 15(9), 505–522. https://doi.org/10.1038/s41569-018-0064-2
Leonardi, G. C., Accardi, G., Monastero, R., Nicoletti, F., & Libra, M. (2018). Ageing: From inflammation to cancer. Immunity & ageing: I & A, 15, 1. https://doi.org/10.1186/s12979-017-0112-5
Garlanda, C., Dinarello, C. A., & Mantovani, A. (2013). The interleukin-1 family: Back to the future. Immunity, 39(6), 1003–1018. https://doi.org/10.1016/j.immuni.2013.11.010
Kovtonyuk, L. V., Caiado, F., Garcia-Martin, S., Manz, E.-M., Helbling, P., Takizawa, H., … Manz, M. G. (2022). IL-1 mediates microbiome-induced inflammaging of hematopoietic stem cells in mice. Blood, 139(1), 44–58. https://doi.org/10.1182/blood.2021011570
Jaiswal, S., & Ebert, B. L. (2019). Clonal hematopoiesis in human aging and disease. Science, 366(6465). https://doi.org/10.1126/science.aan4673
Zheng, M. M., Zhang, Z., Bemis, K., Belch, A. R., Pilarski, L. M., Shively, J. E., & Kirshner, J. (2013). The systemic cytokine environment is permanently altered in multiple myeloma. PLoS ONE, 8(3), e58504. https://doi.org/10.1371/journal.pone.0058504
Guillerey, C., Nakamura, K., Vuckovic, S., Hill, G. R., & Smyth, M. J. (2016). Immune responses in multiple myeloma: Role of the natural immune surveillance and potential of immunotherapies. Cellular and molecular life sciences: CMLS, 73(8), 1569–1589. https://doi.org/10.1007/s00018-016-2135-z. Accessed 3 Jan 2022
Musolino, C., Allegra, A., Innao, V., Allegra, A. G., Pioggia, G., & Gangemi, S. (2017). Inflammatory and anti-inflammatory equilibrium, proliferative and antiproliferative balance: The role of cytokines in multiple myeloma. Mediators of inflammation, 2017, 1852517. https://doi.org/10.1155/2017/1852517
Chen, L., Sun, M., Wu, W., Yang, W., Huang, X., Xiao, Y., … Cong, Y. (2019). Microbiota metabolite butyrate differentially regulates Th1 and Th17 cells’ differentiation and function in induction of colitis. Inflammatory bowel diseases, 25(9), 1450–1461. https://doi.org/10.1093/ibd/izz046
Voorhees, P. M., Manges, R. F., Sonneveld, P., Jagannath, S., Somlo, G., Krishnan, A., … Thomas, S. K. (2013). A phase 2 multicentre study of siltuximab, an anti-interleukin-6 monoclonal antibody, in patients with relapsed or refractory multiple myeloma. British journal of haematology, 161(3), 357–366. https://doi.org/10.1111/bjh.12266
San-Miguel, J., Bladé, J., Shpilberg, O., Grosicki, S., Maloisel, F., Min, C.-K., … Others. (2014). Phase 2 randomized study of bortezomib-melphalan-prednisone with or without siltuximab (anti--IL-6) in multiple myeloma. Blood, The Journal of the American Society of Hematology, 123(26), 4136–4142. Retrieved from https://ashpublications.org/blood/article-abstract/123/26/4136/32785
Kurzrock, R., Voorhees, P. M., Casper, C., Furman, R. R., Fayad, L., Lonial, S., … van Rhee, F. (2013). A phase I, open-label study of siltuximab, an anti-IL-6 monoclonal antibody, in patients with B-cell non-Hodgkin lymphoma, multiple myeloma, or Castleman disease. Clinical cancer research: an official journal of the American Association for Cancer Research, 19(13), 3659–3670. https://doi.org/10.1158/1078-0432.CCR-12-3349
Lust, J. A., Lacy, M. Q., Zeldenrust, S. R., Witzig, T. E., Moon-Tasson, L. L., Dinarello, C. A., & Donovan, K. A. (2016). Reduction in C-reactive protein indicates successful targeting of the IL-1/IL-6 axis resulting in improved survival in early stage multiple myeloma. American journal of hematology, 91(6), 571–574. https://doi.org/10.1002/ajh.24352
Pianko, M. J., Devlin, S. M., Littmann, E. R., Chansakul, A., Mastey, D., Salcedo, M., … Lesokhin, A. M. (2019). Minimal residual disease negativity in multiple myeloma is associated with intestinal microbiota composition. Blood advances, 3(13), 2040–2044. https://doi.org/10.1182/bloodadvances.2019032276
Shah, U., Derkach, A., Adintori, P., Cross, J., Maclachlan, K., Mailankody, S., … Lesokhin, A. (2021). P-042: Sustained minimal residual disease negativity in Multiple Myeloma is impacted positively by stool butyrate and healthier plant forward diets. Clinical lymphoma, myeloma & leukemia, 21, S61. https://doi.org/10.1016/s2152-2650(21)02176-5
Jian, X., Zhu, Y., Ouyang, J., Wang, Y., Lei, Q., Xia, J., … Zhou, W. (2020). Alterations of gut microbiome accelerate multiple myeloma progression by increasing the relative abundances of nitrogen-recycling bacteria. Microbiome, 8(1), 74. https://doi.org/10.1186/s40168-020-00854-5
Drayson, M. T., Bowcock, S., Planche, T., Iqbal, G., Pratt, G., Yong, K., … TEAMM Trial Management Group and Trial Investigators. (2019). Levofloxacin prophylaxis in patients with newly diagnosed myeloma (TEAMM): a multicentre, double-blind, placebo-controlled, randomised, phase 3 trial. The lancet oncology, 20(12), 1760–1772. https://doi.org/10.1016/S1470-2045(19)30506-6
Albrich, W. C., & Boggian, K. (2020, February). Levofloxacin prophylaxis in patients with myeloma. The lancet oncology. https://doi.org/10.1016/S1470-2045(20)30008-5
Teh, B. W., Harrison, S. J., Worth, L. J., Thursky, K. A., & Slavin, M. A. (2020, February). Levofloxacin prophylaxis in patients with myeloma. The lancet oncology. https://doi.org/10.1016/S1470-2045(19)30824-1
Viaud, S., Saccheri, F., Mignot, G., Yamazaki, T., Daillere, R., Hannani, D., … Zitvogel, L. (2013). The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science, 342(6161), 971–976. https://doi.org/10.1126/science.1240537
Matson, V., Fessler, J., Bao, R., Chongsuwat, T., Zha, Y., Alegre, M.-L., … Gajewski, T. F. (2018). The commensal microbiome is associated with anti–PD-1 efficacy in metastatic melanoma patients. Science. https://doi.org/10.1126/science.aao3290
El Jurdi, N., Filali-Mouhim, A., Salem, I., Retuerto, M., Dambrosio, N. M., Baer, L., … de Lima, M. (2019). Gastrointestinal microbiome and mycobiome changes during autologous transplantation for multiple myeloma: Results of a prospective pilot study. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 25(8), 1511–1519. https://doi.org/10.1016/j.bbmt.2019.04.007
Holler, E., Butzhammer, P., Schmid, K., Hundsrucker, C., Koestler, J., Peter, K., … Oefner, P. J. (2014). Metagenomic analysis of the stool microbiome in patients receiving allogeneic stem cell transplantation: Loss of diversity is associated with use of systemic antibiotics and more pronounced in gastrointestinal graft-versus-host disease. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 20(5), 640–645. https://doi.org/10.1016/j.bbmt.2014.01.030
Taur, Y., Xavier, J. B., Lipuma, L., Ubeda, C., Goldberg, J., Gobourne, A., … Pamer, E. G. (2012). Intestinal domination and the risk of bacteremia in patients undergoing allogeneic hematopoietic stem cell transplantation. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America, 55(7), 905–914. https://doi.org/10.1093/cid/cis580
Golob, J. L., Pergam, S. A., Srinivasan, S., Fiedler, T. L., Liu, C., Garcia, K., … Fredricks, D. N. (2017). Stool microbiota at neutrophil recovery is predictive for severe acute graft vs host disease after hematopoietic cell transplantation. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America, 65(12), 1984–1991. https://doi.org/10.1093/cid/cix699
Taur, Y., Jenq, R. R., Perales, M. A., Littmann, E. R., Morjaria, S., Ling, L., … Pamer, E. G. (2014). The effects of intestinal tract bacterial diversity on mortality following allogeneic hematopoietic stem cell transplantation. Blood, 124(7), 1174–1182. https://doi.org/10.1182/blood-2014-02-554725
Jenq, R. R., Taur, Y., Devlin, S. M., Ponce, D. M., Goldberg, J. D., Ahr, K. F., … van den Brink, M. R. (2015). Intestinal Blautia is associated with reduced death from graft-versus-host disease. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 21(8), 1373–1383. https://doi.org/10.1016/j.bbmt.2015.04.016
Shono, Y., Docampo, M. D., Peled, J. U., Perobelli, S. M., Velardi, E., Tsai, J. J., … Jenq, R. R. (2016). Increased GVHD-related mortality with broad-spectrum antibiotic use after allogeneic hematopoietic stem cell transplantation in human patients and mice. Science translational medicine, 8(339), 339ra71. https://doi.org/10.1126/scitranslmed.aaf2311
Hidaka, D., Hayase, E., Shiratori, S., Hasegawa, Y., Ishio, T., Tateno, T., … Teshima, T. (2018). The association between the incidence of intestinal graft-vs-host disease and antibiotic use after allogeneic hematopoietic stem cell transplantation. Clinical transplantation, 32(9), e13361. https://doi.org/10.1111/ctr.13361
Peled, J. U., Devlin, S. M., Staffas, A., Lumish, M., Khanin, R., Littmann, E. R., … van den Brink, M. R. M. (2017). Intestinal microbiota and relapse after hematopoietic-cell transplantation. Journal of clinical oncology: official journal of the American Society of Clinical Oncology, 35(15), 1650–1659. https://doi.org/10.1200/JCO.2016.70.3348
Ubeda, C., Taur, Y., Jenq, R. R., Equinda, M. J., Son, T., Samstein, M., … Pamer, E. G. (2010). Vancomycin-resistant Enterococcus domination of intestinal microbiota is enabled by antibiotic treatment in mice and precedes bloodstream invasion in humans. The Journal of clinical investigation, 120(12), 4332–4341. https://doi.org/10.1172/JCI43918
Haak, B. W., Littmann, E. R., Chaubard, J.-L., Pickard, A. J., Fontana, E., Adhi, F., … Taur, Y. (2018). Impact of gut colonization with butyrate-producing microbiota on respiratory viral infection following allo-HCT. Blood, 131(26), 2978–2986. https://doi.org/10.1182/blood-2018-01-828996
Harris, B., Morjaria, S. M., Littmann, E. R., Geyer, A. I., Stover, D. E., Barker, J. N., … Pamer, E. G. (2016). Gut microbiota predict pulmonary infiltrates after allogeneic hematopoietic cell transplantation. American journal of respiratory and critical care medicine, 194(4), 450–463. https://doi.org/10.1164/rccm.201507-1491OC
Peled, J. U., Gomes, A. L. C., Devlin, S. M., Littmann, E. R., Taur, Y., Sung, A. D., … van den Brink, M. R. M. (2020). Microbiota as predictor of mortality in allogeneic hematopoietic-cell transplantation. The New England journal of medicine, 382(9), 822–834. https://doi.org/10.1056/NEJMoa1900623
Stoma, I., Karpov, I., Milanovich, N., Uss, A., & Iskrov, I. (2016). Risk factors for mortality in patients with bloodstream infections during the pre-engraftment period after hematopoietic stem cell transplantation. Blood research, 51(2), 102–106. https://doi.org/10.5045/br.2016.51.2.102
Trecarichi, E. M., Pagano, L., Candoni, A., Pastore, D., Cattaneo, C., Fanci, R., … HeMABIS Registry—SEIFEM Group, Italy. (2015). Current epidemiology and antimicrobial resistance data for bacterial bloodstream infections in patients with hematologic malignancies: An Italian multicentre prospective survey. Clinical microbiology and infection: the official publication of the European Society of Clinical Microbiology and Infectious Diseases, 21(4), 337–343. https://doi.org/10.1016/j.cmi.2014.11.022
Averbuch, D., Tridello, G., Hoek, J., Mikulska, M., Akan, H., Yaňez San Segundo, L., … Others. (2017). Antimicrobial resistance in Gram-negative rods causing bacteremia in hematopoietic stem cell transplant recipients: Intercontinental prospective study of the Infectious Diseases Working Party of the European Bone Marrow Transplantation Group. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America, 65(11), 1819–1828. Retrieved from https://academic.oup.com/cid/article-abstract/65/11/1819/4036247. Accessed 3 Jan 2022
Storb, R., Prentice, R. L., Buckner, C. D., Clift, R. A., Appelbaum, F., Deeg, J., … Thomas, E. D. (1983). Graft-versus-host disease and survival in patients with aplastic anemia treated by marrow grafts from HLA-identical siblings. Beneficial effect of a protective environment. The New England journal of medicine, 308(6), 302–307. https://doi.org/10.1056/NEJM198302103080602
Cruciani, M., Rampazzo, R., Malena, M., Lazzarini, L., Todeschini, G., Messori, A., & Concia, E. (1996). Prophylaxis with fluoroquinolones for bacterial infections in neutropenic patients: A meta-analysis. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America, 23(4), 795–805. https://doi.org/10.1093/clinids/23.4.795
Engels, E. A., Lau, J., & Barza, M. (1998). Efficacy of quinolone prophylaxis in neutropenic cancer patients: A meta-analysis. Journal of clinical oncology: Official journal of the American Society of Clinical Oncology, 16(3), 1179–1187. https://doi.org/10.1200/JCO.1998.16.3.1179
Nagatomo, A., Watanabe, K., Kunikane, H., Okamoto, H., & Kunitoh, H. (1998). A randomized controled trial of sulfamethoxazole/trimethoprim plus norfloxacin versus sulfamethoxazole/trimethoprim alone for the prophylaxis of bacterial infection during chemotherapy for lung cancer. Lung cancer , 19(2), 121–125. Retrieved from https://www.sciencedirect.com/science/article/pii/S0169500297000871?casa_token=rueotu1UrxQAAAAA:GFf46QwLznRrllvqNVW7dqcS0VnozNfQ2jHNdJsIj9oTJp7SCK4hBG7QCAOzcVcG79gIzxoy. Accessed 3 Jan 2022
Bucaneve, G., Micozzi, A., Menichetti, F., Martino, P., Stella Dionisi, M., Martinelli, G., … Del Favero, A. (2005). Levofloxacin to prevent bacterial infection in patients with cancer and neutropenia. New England Journal of Medicine. https://doi.org/10.1056/nejmoa044097
Cullen, M., Steven, N., Billingham, L., Gaunt, C., Hastings, M., Simmonds, P., … Stanley, A. (2005). Antibacterial prophylaxis after chemotherapy for solid tumors and lymphomas. New England Journal of Medicine. https://doi.org/10.1056/nejmoa050078
Gafter-Gvili, A., Fraser, A., Paul, M., & Leibovici, L. (2005). Meta-analysis: Antibiotic prophylaxis reduces mortality in neutropenic patients. Annals of Internal Medicine, 142(12 Pt 1), 979–995. https://doi.org/10.7326/0003-4819-142-12_part_1-200506210-00008
Kimura, S.-I., Akahoshi, Y., Nakano, H., Ugai, T., Wada, H., Yamasaki, R., … Kanda, Y. (2014). Antibiotic prophylaxis in hematopoietic stem cell transplantation. A meta-analysis of randomized controlled trials. The Journal of infection, 69(1), 13–25. https://doi.org/10.1016/j.jinf.2014.02.013
Khosravi, A., Yáñez, A., Price, J. G., Chow, A., Merad, M., Goodridge, H. S., & Mazmanian, S. K. (2014). Gut microbiota promote hematopoiesis to control bacterial infection. Cell host & microbe, 15(3), 374–381. https://doi.org/10.1016/j.chom.2014.02.006
Josefsdottir, K. S., Baldridge, M. T., Kadmon, C. S., & King, K. Y. (2017). Antibiotics impair murine hematopoiesis by depleting the intestinal microbiota. Blood, 129(6), 729–739. https://doi.org/10.1182/blood-2016-03-708594
Alonso, C. D., Treadway, S. B., Hanna, D. B., Huff, C. A., Neofytos, D., Carroll, K. C., & Marr, K. A. (2012). Epidemiology and outcomes of Clostridium difficile infections in hematopoietic stem cell transplant recipients. Clinical infectious diseases: An official publication of the Infectious Diseases Society of America, 54(8), 1053–1063. https://doi.org/10.1093/cid/cir1035
Yan, H., Baldridge, M. T., & King, K. Y. (2018). Hematopoiesis and the bacterial microbiome. Blood, 132(6), 559–564. https://doi.org/10.1182/blood-2018-02-832519
Staffas, A., Burgos da Silva, M., Slingerland, A. E., Lazrak, A., Bare, C. J., Holman, C. D., … van den Brink, M. R. M. (2018). Nutritional support from the intestinal microbiota improves hematopoietic reconstitution after bone marrow transplantation in mice. Cell host & microbe, 23(4), 447-457.e4. https://doi.org/10.1016/j.chom.2018.03.002
Stoma, I., Littmann, E. R., Peled, J. U., Giralt, S., van den Brink, M. R. M., Pamer, E. G., & Taur, Y. (2021). Compositional flux within the intestinal microbiota and risk for bloodstream infection with gram-negative bacteria. Clinical infectious diseases: An official publication of the Infectious Diseases Society of America, 73(11), e4627–e4635. https://doi.org/10.1093/cid/ciaa068
Donskey, C. J., Chowdhry, T. K., Hecker, M. T., Hoyen, C. K., Hanrahan, J. A., Hujer, A. M., … Rice, L. B. (2000). Effect of antibiotic therapy on the density of vancomycin-resistant enterococci in the stool of colonized patients. The New England journal of medicine, 343(26), 1925–1932. https://doi.org/10.1056/NEJM200012283432604
Tamburini, F. B., Andermann, T. M., Tkachenko, E., Senchyna, F., Banaei, N., & Bhatt, A. S. (2018). Precision identification of diverse bloodstream pathogens in the gut microbiome. Nature medicine, 24(12), 1809–1814. https://doi.org/10.1038/s41591-018-0202-8
Caballero, S., Kim, S., Carter, R. A., Leiner, I. M., Sušac, B., Miller, L., … Pamer, E. G. (2017). Cooperating commensals restore colonization resistance to vancomycin-resistant Enterococcus faecium. Cell host & microbe, 21(5), 592-602.e4. https://doi.org/10.1016/j.chom.2017.04.002
Gilmore, M. S., Clewell, D. B., Ike, Y., & Shankar, N. (Eds.). (2014). Enterococci: From commensals to leading causes of drug resistant infection. Boston: Massachusetts Eye and Ear Infirmary. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/24649510. Accessed 3 Jan 2022
Buffie, C. G., Bucci, V., Stein, R. R., McKenney, P. T., Ling, L., Gobourne, A., … Pamer, E. G. (2015). Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature, 517(7533), 205–208. https://doi.org/10.1038/nature13828
Miltiadous, O., Waters, N. R., Andrlova, H., Dai, A., Nguyen, C. L., Burgos da Silva, M., … van den Brink, M. R. M. (2022). Early intestinal microbial features are associated with CD4 T cell recovery after allogeneic hematopoietic transplant. Blood. https://doi.org/10.1182/blood.2021014255
Centers for Disease Control and Prevention, Infectious Disease Society of America, & American Society of Blood and Marrow Transplantation. (2000). Guidelines for preventing opportunistic infections among hematopoietic stem cell transplant recipients. MMWR. Recommendations and reports: Morbidity and mortality weekly report. Recommendations and reports / Centers for Disease Control, 49(RR-10), 1–125, CE1–7. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/11718124. Accessed 3 Jan 2022
Maakaron, J. E., Liscynesky, C., Boghdadly, Z. E., Huang, Y., Agyeman, A., Brammer, J., … Jaglowski, S. M. (2020). Fluoroquinolone prophylaxis in autologous stem cell transplantation: Worthy of a second look. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 26(8), e198–e201. https://doi.org/10.1016/j.bbmt.2020.03.027
Rashidi, A., Wangjam, T., Bhatt, A. S., Weisdorf, D. J., Holtan, S. G., & Investigators, B. M. T. C. T. N. (2018). Antibiotic practice patterns in hematopoietic cell transplantation: A survey of blood and marrow transplant clinical trials network centers. American journal of hematology, 93(11), E348–E350. https://doi.org/10.1002/ajh.25236
Arai, S., Arora, M., Wang, T., Spellman, S. R., He, W., Couriel, D. R., … Pavletic, S. Z. (2015). Increasing incidence of chronic graft-versus-host disease in allogeneic transplantation: A report from the center for international blood and marrow transplant research. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 21(2), 266–274. https://doi.org/10.1016/j.bbmt.2014.10.021
Jacobsohn, D. A., & Vogelsang, G. B. (2007). Acute graft versus host disease. Orphanet journal of rare diseases, 2, 35. https://doi.org/10.1186/1750-1172-2-35
Storb, R., Gyurkocza, B., Storer, B. E., Maloney, D. G., Sorror, M. L., Mielcarek, M., … Sandmaier, B. M. (2013). Allogeneic hematopoietic cell transplantation following minimal intensity conditioning: Predicting acute graft-versus-host disease and graft-versus-tumor effects. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 19(5), 792–798. https://doi.org/10.1016/j.bbmt.2013.02.006
Sun, Y., Tawara, I., Toubai, T., & Reddy, P. (2007). Pathophysiology of acute graft-versus-host disease: Recent advances. Translational research: The journal of laboratory and clinical medicine, 150(4), 197–214. https://doi.org/10.1016/j.trsl.2007.06.003
Stein-Thoeringer, C. K., Nichols, K. B., Lazrak, A., Docampo, M. D., Slingerland, A. E., Slingerland, J. B., … van den Brink, M. R. M. (2019). Lactose drives Enterococcus expansion to promote graft-versus-host disease. Science (New York, N.Y.), 366(6469), 1143–1149. https://doi.org/10.1126/science.aax3760
Connell, M. S., & Wilson, R. (1965). The treatment of x-irradiated germfree CFW and C3H mice with isologous and homologous bone marrow. Life sciences, 4, 721–729. https://doi.org/10.1016/0024-3205(65)90011-1
Jones, J. M., Wilson, R., & Bealmear, P. M. (1971). Mortality and gross pathology of secondary disease in germfree mouse radiation chimeras. Radiation research, 45(3), 577–588. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/4396814. Accessed 3 Jan 2022
Bealmear, P. M., Mirand, E. A., & Holtermann, O. A. (1983). Modification of graft-vs-host disease following bone marrow transplantation in germfree mice. Progress in clinical and biological research, 132C, 409–421. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/6227014. Accessed 3 Jan 2022
Bauer, H., Horowitz, R. E., Levenson, S. M., & Popper, H. (1963). The response of the lymphatic tissue to the microbial flora. Studies on germfree mice. The American journal of pathology, 42, 471–483. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/13966929. Accessed 3 Jan 2022
Hooper, L. V., Littman, D. R., & Macpherson, A. J. (2012). Interactions between the microbiota and the immune system. Science, 336(6086), 1268–1273. https://doi.org/10.1126/science.1223490
Round, J. L., & Mazmanian, S. K. (2009). The gut microbiota shapes intestinal immune responses during health and disease. Nature reviews. Immunology, 9(5), 313–323. https://doi.org/10.1038/nri2515
Fredricks, D. N. (2019). The gut microbiota and graft-versus-host disease. The Journal of clinical investigation, 129(5), 1808–1817. https://doi.org/10.1172/JCI125797
Gerbitz, A., Schultz, M., Wilke, A., Linde, H.-J., Schölmerich, J., Andreesen, R., & Holler, E. (2004). Probiotic effects on experimental graft-versus-host disease: Let them eat yogurt. Blood, 103(11), 4365–4367. https://doi.org/10.1182/blood-2003-11-3769
Weber, D., Jenq, R. R., Peled, J. U., Taur, Y., Hiergeist, A., Koestler, J., … Holler, E. (2017). Microbiota disruption induced by early use of broad-spectrum antibiotics is an independent risk factor of outcome after allogeneic stem cell transplantation. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 23(5), 845–852. https://doi.org/10.1016/j.bbmt.2017.02.006
Weber, D., Oefner, P. J., Dettmer, K., Hiergeist, A., Koestler, J., Gessner, A., … Holler, E. (2016). Rifaximin preserves intestinal microbiota balance in patients undergoing allogeneic stem cell transplantation. Bone marrow transplantation, 51(8), 1087–1092. https://doi.org/10.1038/bmt.2016.66
Romick-Rosendale, L. E., Haslam, D. B., Lane, A., Denson, L., Lake, K., Wilkey, A., … Davies, S. M. (2018). Antibiotic exposure and reduced short chain fatty acid production after hematopoietic stem cell transplant. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation, 24(12), 2418–2424. https://doi.org/10.1016/j.bbmt.2018.07.030
Markey, K. A., Schluter, J., Gomes, A. L. C., Littmann, E. R., Pickard, A. J., Taylor, B. P., … van den Brink, M. R. M. (2020). The microbe-derived short-chain fatty acids butyrate and propionate are associated with protection from chronic GVHD. Blood, 136(1), 130–136. https://doi.org/10.1182/blood.2019003369
Kaiko, G. E., Ryu, S. H., Koues, O. I., Collins, P. L., Solnica-Krezel, L., Pearce, E. J., … Stappenbeck, T. S. (2016). The colonic crypt protects stem cells from microbiota-derived metabolites. Cell, 167(4), 1137. https://doi.org/10.1016/j.cell.2016.10.034
Golob, J. L., DeMeules, M. M., Loeffelholz, T., Quinn, Z. Z., Dame, M. K., Silvestri, S. S., … Fredricks, D. N. (2019). Butyrogenic bacteria after acute graft-versus-host disease (GVHD) are associated with the development of steroid-refractory GVHD. Blood Advances. https://doi.org/10.1182/bloodadvances.2019000362
Weber, D., Hiergeist, A., Weber, M., Dettmer, K., Wolff, D., Hahn, J., … Holler, E. (2019). Detrimental effect of broad-spectrum antibiotics on intestinal microbiome diversity in patients after allogeneic stem cell transplantation: Lack of commensal sparing antibiotics. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America, 68(8), 1303–1310. Retrieved from https://academic.oup.com/cid/article-abstract/68/8/1303/5075542. Accessed 3 Jan 2022
Child, J. A., Morgan, G. J., Davies, F. E., Owen, R. G., Bell, S. E., Hawkins, K., … Medical Research Council Adult Leukaemia Working Party. (2003). High-dose chemotherapy with hematopoietic stem-cell rescue for multiple myeloma. The New England journal of medicine, 348(19), 1875–1883. https://doi.org/10.1056/NEJMoa022340
Attal, M., Lauwers-Cances, V., Hulin, C., Leleu, X., Caillot, D., Escoffre, M., … IFM 2009 Study. (2017). Lenalidomide, bortezomib, and dexamethasone with transplantation for myeloma. The New England journal of medicine, 376(14), 1311–1320. https://doi.org/10.1056/NEJMoa1611750
Khan, N., Peled, J. U., Gomes, A. L. C., Devlin, S. M., Rondon-Clavo, C., Clurman, A., … van den Brink, M. R. M. (2018). Loss of microbiota diversity after autologous stem cell transplant is comparable to injury in allogeneic stem cell transplant. Blood, 132(Supplement 1), 608–608. https://doi.org/10.1182/blood-2018-99-110857
El Jurdi, N., Filali-Mouhim, A., & Salem, I. (2019). … in relative abundance, diversity and richness of the oral and gastrointestinal microbiome during autologous transplantation for multiple myeloma: Results of a …. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation. Retrieved from https://www.tctjournal.org/article/S1083-8791(18)31768-3/abstract. Accessed 3 Jan 2022
Vuckovic, S., Minnie, S. A., Smith, D., Gartlan, K. H., Watkins, T. S., Markey, K. A., … Hill, G. R. (2019). Bone marrow transplantation generates T cell-dependent control of myeloma in mice. The Journal of clinical investigation, 129(1), 106–121. https://doi.org/10.1172/JCI98888
Chung, D. J., Pronschinske, K. B., Shyer, J. A., Sharma, S., Leung, S., Curran, S. A., … Young, J. W. (2016). T-cell exhaustion in multiple myeloma relapse after autotransplant: Optimal timing of immunotherapy. Cancer Immunol Res, 4(1), 61–71. https://doi.org/10.1158/2326-6066.CIR-15-0055
Lu, X., Ding, Z.-C., Cao, Y., Liu, C., Habtetsion, T., Yu, M., … Zhou, G. (2015). Alkylating agent melphalan augments the efficacy of adoptive immunotherapy using tumor-specific CD4+ T cells. Journal of immunology , 194(4), 2011–2021. https://doi.org/10.4049/jimmunol.1401894
Lee, D. H., Fung, T. T., Tabung, F. K., Marinac, C. R., Devore, E. E., Rosner, B. A., … Birmann, B. M. (2020). Prediagnosis dietary pattern and survival in patients with multiple myeloma. International journal of cancer. Journal international du cancer, 147(7), 1823–1830. https://doi.org/10.1002/ijc.32928
Gough, E., Shaikh, H., & Manges, A. R. (2011). Systematic review of intestinal microbiota transplantation (fecal bacteriotherapy) for recurrent Clostridium difficile infection. Clinical infectious diseases: An official publication of the Infectious Diseases Society of America, 53(10), 994–1002. https://doi.org/10.1093/cid/cir632
DeFilipp, Z., Peled, J. U., Li, S., Mahabamunuge, J., Dagher, Z., Slingerland, A. E., … Chen, Y. B. (2018). Third-party fecal microbiota transplantation following allo-HCT reconstitutes microbiome diversity. Blood Adv, 2(7), 745–753. https://doi.org/10.1182/bloodadvances.2018017731
Taur, Y., Coyte, K., Schluter, J., Robilotti, E., Figueroa, C., Gjonbalaj, M., … Xavier, J. B. (2018). Reconstitution of the gut microbiota of antibiotic-treated patients by autologous fecal microbiota transplant. Science translational medicine. https://doi.org/10.1126/scitranslmed.aap9489
Kakihana, K., Fujioka, Y., Suda, W., Najima, Y., Kuwata, G., Sasajima, S., … Ohashi, K. (2016). Fecal microbiota transplantation for patients with steroid-resistant acute graft-versus-host disease of the gut. Blood, 128(16), 2083–2088. https://doi.org/10.1182/blood-2016-05-717652
van Lier, Y. F., Davids, M., Haverkate, N. J. E., de Groot, P. F., Donker, M. L., Meijer, E., … Hazenberg, M. D. (2020). Donor fecal microbiota transplantation ameliorates intestinal graft-versus-host disease in allogeneic hematopoietic cell transplant recipients. Science translational medicine, 12(556). https://doi.org/10.1126/scitranslmed.aaz8926
Spindelboeck, W., Schulz, E., Uhl, B., Kashofer, K., Aigelsreiter, A., Zinke-Cerwenka, W., … Neumeister, P. (2017). Repeated fecal microbiota transplantations attenuate diarrhea and lead to sustained changes in the fecal microbiota in acute, refractory gastrointestinal graft-versus-host-disease. Haematologica, 102(5), e210–e213. https://doi.org/10.3324/haematol.2016.154351
DeFilipp, Z., Bloom, P. P., Torres Soto, M., Mansour, M. K., Sater, M. R. A., Huntley, M. H., … Hohmann, E. L. (2019). Drug-resistant E. coli bacteremia transmitted by fecal microbiota transplant. The New England journal of medicine, 381(21), 2043–2050. https://doi.org/10.1056/NEJMoa1910437
Funding
This work was supported by the National Institutes of Health grants (National Institute of Aging: P30-AG024824, M.J.P.), (National Cancer Institute: P30-CA046592, M.J.P.), (National Center for Advancing Translational Sciences: UL1-TR00457, M.J.P.), and the American Society for Transplantation and Cellular Therapy (ASTCT) New Investigator Award (J.L.G.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pianko, M.J., Golob, J.L. Host-microbe interactions and outcomes in multiple myeloma and hematopoietic stem cell transplantation. Cancer Metastasis Rev 41, 367–382 (2022). https://doi.org/10.1007/s10555-022-10033-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-022-10033-7