Abstract
Background
The tumor microenvironment (TME) plays a crucial role in various aspects of breast cancer development and metastasis. Nevertheless, the expression, prognostic significance, and correlation with clinical features of SCARB2 in breast cancer, as well as the infiltrative characteristics of TME, remain largely unknown.
Methods
We analyzed the differential presentation of SCARB2 mRNA in breast cancer tissues and nontumorous breast tissues and prognosis by The Cancer Genome Atlas (TCGA) and the Genotype-Tissue Expression (GTEx) databases. Additionally, the Tumor Immunity Estimation Resource (TIMER) was taken to evaluate the correlation between SCARB2 mRNA presence and tumor-infiltrating immune cells and immune checkpoints in the TME in breast cancer. We performed multiple immunohistochemical staining to verify the SCARB2 protein expression in breast cancer tissues and its relationship to immune cells and checkpoints and clinicopathological features.
Results
We identified elevated SCARB2 expression in breast cancer tissues, and high SCARB2 protein presentation was associated with advanced clinical stage and unfavorable prognosis. In addition, enhanced SCARB2 protein presence was closely correlated with up-regulation CD66b+ neutrophils infiltration in tumor tissues (r = 0.210, P < 0.05) and CD68 + CD163+ M2 macrophages in the interstitium (r = 0.233, P < 0.05), as well as the immune checkpoints, including PD-1 (r = 0.314, P < 0.01) protein expression.
Conclusion
SCARB2 holds promise for predicting the clinical outcome of breast cancer patients and could serve as a potential therapeutic target.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
According to the latest data from the American Cancer Association (ACA), the incidence of breast cancer in women has been increasing annually, with a recent annual growth rate of approximately 0.5% [1]. This report also noted that by 2023, breast cancer is projected to account for 31% of the most commonly diagnosed cancers in women. Over time, cancer treatment has undergone obvious changes, shifting from single-surgical approaches to multidisciplinary treatments that encompass radiotherapy, chemotherapy, endocrine therapy, targeted therapy, and immunotherapy [2]. Immunotherapy has emerged as a promising area of research for breast cancer patients, offering potential improvements in survival and prognosis [3]. However, the effectiveness of immunotherapy is influenced by tumor cell heterogeneity and the immunosuppressive TME, resulting in lasting effects from immunotherapy being observed only in a small proportion of patients [4]. Therefore, there is a need to identify accurate biomarkers for assessing the efficacy of immunotherapy to further improve the survival of breast cancer patients.
The TME is a complex composition incorporating cancer-associated fibroblasts (CAFs), tumor-infiltrating immune cells (TIICs), angiogenic vascular cells (AVCs), and the extracellular matrix (ECM) [5,6,7]. Numerous studies have revealed a close connection between tumor development and the TME [6]. During tumor growth, tumor cells dynamically interact with TME components, contributing to cancer cell survival, local invasion, and metastatic spread [8]. TIICs, as prominent components of the TME, play a crucial role in tumor development, metastasis, and can serve as prognostic indicators and treatment response predictors in cancer patients [6, 9]. Elevated expression of TIICs and TIICs-related genes has been associated with poor survival in breast cancer patients [9]. Overexpression of immune checkpoint molecules such as programmed cell death receptor 1 (PD-1) and its ligand programmed cell death ligand 1 (PD-L1) and cytotoxic T-lymphocyte antigen 4 (CTLA-4) suppresses immune function and enables cancer cells to avoid being cleared by the immune system [10]. Therefore, the identification of biomarkers related to TME is vital for the treatment and prognosis of breast cancer.
SCARB2, a member of the scavenger receptor family B and also known as lysosomal integral membrane protein type 2 (LIMP2), is a lysosomal membrane glycoprotein encoded by the SCARB2 gene [11, 12]. However, the specific biological properties of SCARB2 in breast cancer, particularly its relationship with the TME, remain poorly studied.
In this research, we used bioinformatics techniques to assess the differential presentation of SCARB2 mRNA in breast cancer tissues compared to non-cancerous human breast tissues. The relationship between mRNA abundance and protein abundance is complex and nonlinear [13]. So we next evaluated the expression of SCARB2 protein and its pertinence to TIICs and immune checkpoints by multiple immunohistochemistry (mIHC) staining, to explore its immunological and prognostic value in breast cancer.
Methods
Bioinformatics analysis
The differential expression of SCARB2 mRNA was analyzed by accessing tissue characteristics and clinical information from the TCGA (https://portal.gdc.cancer.gov/) and the GTEx (https://www.gtexportal.org) databases of breast cancer patients and healthy individuals. To explore the impact of SCARB2 mRNA on overall survival (OS) in breast cancer patients, we utilized GEPIA (http://gepia.cancer-pku.cn/), a web-based tool that provides interactive and customizable features [14]. Additionally, to assess the relationship between SCARB2 mRNA and the abundance of TIICs and immune checkpoints in breast cancer, we employed TIMER (https://cistrome.shinyapps.io/timer/), a publicly available resource that enables comprehensive analysis and visualization of TIICs [15].
Breast cancer patient samples and clinicopathological data
The breast cancer samples used in this study were obtained from the clinical biospecimen bank of Nantong University Hospital. The sample consisted of 124 breast cancer patients and 115 non-cancerous individuals who underwent breast cancer resection at Nantong University Hospital from 2002 to 2010. Samples were prepared using tissue microarray (TMA) and were expressed as 2-mm diameter cores. Patients who had received preoperative chemotherapy, radiotherapy, or immunotherapy were excluded from the sample. Clinical and pathological data, such as age, molecular subtype, tumor size, lymph node and distant metastases, and TNM stage were collected. This thesis was approved by the Human Ethics Committee of Nantong University Hospital (number: 2018-K020).
Fluorescence-based mIHC
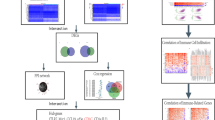
Due to the mismatch between mRNA and protein expression levels, we therefore performed mIHC experiments to explore the relationship between TIICs and SCARB2 protein presentation [16]. TMA sections were deparaffinized with xylene and then rehydrated. Antigen retrieval was performed by heating the slides in AR6 buffer (AR600, AKOYA) using a microwave. After cooling, the slides were incubated overnight at 4 °C with the primary antibody reagent. Following a half-hour re-warming, secondary antibodies were added, and staining was performed. Antigens were then repaired again through heat induction and cooling. Finally, the nuclei were stained using the DAPI Opal7 Color Technology Kit (NEL81001KT, USA) and blocked. The slides were scanned with the Vectra 3.0 automated quantitative pathology imaging system to obtain positive images of the biomarkers. The inForm®CellAnalysis software (V.2.6.0) was employed to differentiate and analyze areas of tumor cells and stroma cells. The images were then analyzed and scored, with a positive or negative threshold set for each cell. The percentage of cells in each region (0–100) was calculated and scored. The antibodies used in this inspection are listed as follows: anti-SCARB2 antibody (1:200, ab176317, Abcam); anti-CD8 antibody (1:1000, 85336s, Cell Signaling Technology); anti-CD3 antibody (1:1000, 85061s, Cell Signaling Technology); anti-CD4 antibody (1:100, NBP2-52663, Novus); anti-CD68 antibody (1:1000, 76437s, Cell Signaling Technology); anti-CD86 antibody (1:500, orb388891, Biorbyt); anti-CD163 antibody (1:500, 93498s, Cell Signaling Technology); anti-CD66b antibody (1:500, ARG66287, Ango. Biolaboratories); anti-CD20 antibody (1:1000, ab78237, Abcam); anti-LAMP3 antibody (1:1500, ab271053, Abcam); anti-cytokeratin antibody (1:4000, orb69073, Biorbyt).
Statistical analysis
The data were analyzed and processed using X-tile and SPSS (v.28.0). The differences in SCARB2 protein presentation between malignant and nonmalignant tissue samples were compared through the Student t-test. SCARB2 protein presence in breast cancer was divided into high and low presentation groups using X-tile software, and the correlations between SCARB2 protein expression and previously obtained clinicopathological parameters and TIICs and immune checkpoints were determined by Pearson’s Chi-squared test. Cox regression analysis was exploited to investigate factors associated with prognosis. A P value < 0.05 was considered statistically significant for all tests.
Results
SCARB2 mRNA expression and its relationship with prognosis
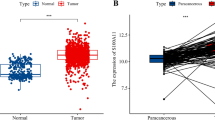
In this research, clinicopathological data of 1099 breast cancer patients were downloaded from the TCGA database and compared with 292 benign breast tissues from the GTEx database. The expression of SCARB2 mRNA was significantly higher in breast cancer tissues than in non-cancerous tissues (P < 0.001, Fig. 1A). To analyze the prognostic value of SCARB2 in breast cancer, we utilized the GEPIA database to assess the impact of SCARB2 mRNA expression on overall patient survival. High SCARB2 mRNA expression was strongly linked to lower OS among breast cancer patients (HR = 1.8, P = 0.006, Fig. 1B).
SCARB2 protein expression and its relevance to clinical characteristics and prognosis of breast cancer
The mIHC assay results showed a conspicuously higher representation of SCARB2 protein in breast cancer tissues compared to benign breast tissues (P < 0.01, Fig. 2A–C), which was consistent with the findings of SCARB2 mRNA presence. Based on the expression of the SCARB2 protein, we categorized the samples into high and low expression groups using a cut-off value of 23.42 (0–100). Subsequently, we analyzed the relationship between SCARB2 protein presence and clinicopathological features. The results revealed a visible association between SCARB2 and the TNM stage (P = 0.019, Table 1). Patients with TNM stages III-IV had relatively greater protein presence of SCARB2 in their breast cancer tissues than patients with stages I-II. Nevertheless, no palpable relevance was detected with age, molecular subtype, tumor size, lymph node metastasis, and distant metastasis (P > 0.05, Table 1). To further investigate the relation between SCARB2 protein expression, clinicopathological characteristics, and prognosis in breast cancer, Cox regression analysis was utilized. The results of univariate regression analysis demonstrated correlations between SCARB2 protein presentation, tumor size, lymph node metastasis, distant metastasis, TNM stage, and OS (Table 2). Multifactorial Cox survival analysis confirmed that SCARB2 can serve as an independent prognostic predictive biomarker for breast cancer (HR 2.893; 95% CI 1.107–7.556; P = 0.03), along with TNM stage as another independent prognostic factor (HR 5.805; 95% CI 2.567–13.125; P = 0.005) (Table 2). Kaplan–Meier curve analysis illustrated that breast cancer patients with high SCARB2 protein expression had poor outcomes (P < 0.05, Fig. 2D), suggesting that SCARB2 protein presence is a reliable predictor of prognosis in breast cancer patients.
Differential presentation of SCARB2 protein and CD66b in tissues. SCARB2 protein and CD66b expression in A breast cancer tissues and B non-cancerous tissues. CK: cytokeratin. Scale bar, 20 μm. C Differences in SCARB2 protein representation between breast cancer tissues and paracancerous tissues. D Patients with high SCARB2 expression showed conspicuously unfavorable survival than patients with low SCARB2 expression
Association of SCARB2 presentation with TIICs
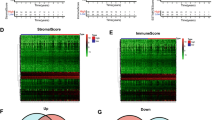
Considering the importance of the tumor immune microenvironment in breast cancer progression, it is crucial to explore the connection between SCARB2 and TIICs, which has not been addressed in this study. According to the TIMER database, SCARB2 mRNA expression was significantly relevant to the abundance of immune infiltration cells, including neutrophils (r = 0.192, P = 2.40e−09), macrophages (r = 0.331, P = 1.40e−26) and CD8+ T cells (r = 0.362, P = 1.63e−31) (Fig. 3A). However, no apparent correlation was observed with the abundance of B cells and CD4+ T cells (Fig. 3A). Furthermore, we mined the relationship between SCARB2 protein expression and the abundance of TIICs through mIHC experiments. The results revealed that high representation of SCARB2 protein exhibited a noticeable positive relativity with the abundance of CD66b+ neutrophils in tumor tissues (r = 0.210, P = 0.027) (Fig. 3B) and CD68+CD163+ M2 macrophages in the interstitium (r = 0.233, P = 0.014) (Fig. 3C). Although CD3+CD4+ T cells, CD3+CD8+ T cells, CD20+ B cells, and CD68+CD86+ macrophages were infiltrated in the TME of breast cancer (Fig. 3D), there was no marked pertinence were observed between SCARB2 and their abundance (P > 0.05, Fig. 3E, F).
The connection between the presentation of SCARB2 and the degree of TIIC abundance. A SCARB2 mRNA presence was positively related to macrophages, neutrophils, CD8+ T cells, and NK cells, and not clearly connected with B cells and CD4+ T cells. B CD66b+ neutrophils were enriched in tumor areas with high expression of SCARB2 protein. C The high presence of SCARB2 protein in the stroma region showed a higher proportion of CD68+CD163+ M2 macrophages. Scale bar, 20 μm. CK: cytokeratin. D Penetration levels of TIICs in breast cancer. Correlation analysis of SCARB2 protein with TIICs in E tumor and F mesenchymal regions of breast cancer. *P < 0.05
Correlation of SCARB2 expression with immune checkpoints
We evaluated the connection between immune checkpoints and SCARB2 mRNA presentation. Breast cancer patients with increased SCARB2 mRNA expression had higher PD-Ll (CD 274) mRNA expression (r = 0.29, P = 9.13e−23, Fig. 4A). Meanwhile, PD-1 (PDCD1) and CTLA-4 did not show palpable differential representation (Fig. 4A). We further validated the relationship between SCARB2 protein presentation and immune checkpoints in breast cancer tissues. PD-1 protein expression was greater in the SCARB2 enhanced expression group (P = 0.0027, Fig. 4B). SCARB2 protein representation was positively correlated with PD-1 presence (r = 0.314, P < 0.01, Fig. 4C). The results were not fully consistent with mRNA expression level. We were able to observe PD-1 presentation in breast cancer (Fig. 4D).
A The relationship between SCARB2 mRNA expression with PD-1 (PDCD1), PD-L1 (CD274), and CTLA-4 (CTLA4) mRNA expression. B PD-1, PD-L1, and CTLA-4 protein presentation in high and low presentation motifs of SCARB2 protein. C The correlation of SCARB2 protein presence with immune checkpoints including PD-1, PD-L1 and CTLA-4 in breast cancer tissues. D The representative mIHC images of PD-1 expression in breast cancer tissues
Discussion
Breast cancer is one of the most prevalent and deadliest malignant tumors affecting women worldwide [17], significantly impacting their health and safety [18]. However, breast cancer treatment faces a critical challenge due to drug resistance [19], and early diagnosis, treatment monitoring, and prognostic evaluation play a vital role in the survival outcome of breast cancer patients [17]. Consequently, identifying specific biomarkers to predict treatment outcomes and prognosis has become a primary focus of our thesis.
Recent pan-cancer studies have reported variable SCARB2 presence in tumor and normal tissues depending on the cancer type [20]. In our investigation, we found that SCARB2 expression was relatively higher in breast cancer tissues. Further analysis of the relationship between SCARB2 presentation and OS in breast cancer patients indicated that SCARB2 over-representation in breast cancer was associated with dismal survival. Multivariate Cox survival analysis identified SCARB2 as an independent prognostic factor. Additionally, the exploration of SCARB2 and clinicopathological features showed a distinct connection between SCARB2 and the TNM stage. Based on these findings, we propose that SCARB2 may be involved in the disease progression of breast cancer and demonstrate its clinical value as a prognostic marker.
TME plays an essential role in breast cancer development [21], and TIICs are instrumental in disease progression and patient prognosis for various malignancies [22]. Among these TIICs, neutrophils, as the most abundant circulating leukocytes and immune and inflammatory cells infiltrating the TME [23, 24], play a pivotal role in tumor promotion [25] and are considered one of the most critical TIICs that promote cancer progression and metastasis [26]. In breast cancer patients, neutrophils usually infiltrate the carcinoma tissue [27].
Tumor cells produce a variety of potent neutrophil chemoattractants and activators, which in turn recruit circulating neutrophils to tumor tissues to form TANs, which are recruited to the tumor microenvironment and synthesize and secrete a large number of proteases to participate in the remodeling of the tumor microenvironment, leading to the accumulation of more inflammatory cells [28]. TANs can regulate tumor growth and progression in TME by secreting a variety of cytokines, including TGF-β2, IL-1β, TNF-α, IL-12, G-CSF, vascular endothelial growth factor (VEGF), CC-family chemokines, CXC-family chemokines [29]. During tumor proliferation, TANs play a role in synthesizing and secreting related proteases. Neutrophil elastase (NE) is one of the most important ones [30]. High levels of NE can promote the proliferation of breast cancer cells, and after silencing the NE gene in mice, the growth rate of tumors was significantly slowed down and the size of tumors was significantly reduced [31]. In most clinical studies, a large number of infiltrating TANs is considered to be one of the indicators of poor prognosis in patients with various malignant tumors [32]. Removing the infiltration of TANs in tumor tissues and suppressing the inflammatory state of the body is expected to be an effective way to inhibit tumor progression [33].
Scavenger receptors can promote cancer progression by influencing the function of tumor-associated immune cells [20], making them potential diagnostic and prognostic biomarkers, as well as therapeutic targets for cancer [34, 35]. As a member of the scavenger receptors, SCARB2 is a selective receptor for the enterovirus EV71, coxsackievirus, and β-glucosylceramide cerebrosidase, and is involved in immunization, viral or bacterial infections, and a variety of other biological processes [36]. SCARB2 presentation was greater in glioma cell lines than in normal glial cells [37]. Meanwhile, SCARB2 plays a vital role in innate immune cells, and SCARB2 was highly expressed in peripheral blood plasma cell-like dendritic cells [38]. Furthermore, SCARB2 exhibits distinct properties that can impact the immune microenvironment in patients with hepatocellular carcinoma and is involved in altered pathways in renal cell carcinoma, including PI3K-Akt, Foxo, endocytosis, MAPK, tight junction, and cytokine-cytokine-receptor interaction pathways [39]. In conclusion, there is increasing evidence that SCARB2 is involved in the immune response and plays a key role.
Our analysis suggested an apparent association between SCARB2 and TME in breast cancer. Increased SCARB2 content in breast cancer tissues corresponded to elevated TAN levels. These findings highlight SCARB2 as a potential prognostic biomarker allied to TANs, with promising implications for breast cancer clinical outcome and treatment.
PD-1 attaches to PD-L1, and the interaction between the two inhibits T lymphocyte proliferation, survival and effector functions, and transmits negative co-stimulatory signals that limit T cell activation [40, 41]. Overexpression and binding of PD-1 and PD-L1 to cancer cells or TME ligands leads to T cell attenuation, thus allowing tumor cells to avoid immune-mediated destruction [42]. Immunotherapy, which utilizes immune checkpoint inhibitors targeting PD-1 and PD-L1 thereby improving the prognosis and survival of cancer patients, has been shown to be effective in breast cancer patients [41, 43]. Herein, SCARB2 protein expression was obviously and positively correlated with PD1 in breast cancer tissues. SCARB2 may serve as a new immune-related biomarker to provide a new rationale for immunotherapy in breast cancer.
It is crucial to acknowledge the limitations of our study, which was conducted at a single center rather than a multicenter and utilized a retrospective rather than a prospective design. The use of a single center may limit the generalizability of our findings to a broader population. Additionally, the retrospective nature of our study design may introduce biases in data collection, analysis, and interpretation. Finally, further cytological experiments are necessary to validate the impact of SCARB2 on prognosis by elucidating its role in regulating the mechanisms of TME in breast cancer. These experiments will provide a deeper understanding of the underlying mechanisms and help translate our findings into clinical applications.
In conclusion, our research elucidated that SCARB2 presence is upregulated in breast cancer and strongly correlated with unfavorable survival. Additionally, the infiltration of neutrophils in the TME may play a conspicuous role in breast cancer progression and patient outcomes. SCARB2 may be a new target for prognostic and immunotherapeutic treatments of breast cancer.
Data availability
The data used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Siegel RL, Miller KD, Wagle NS et al (2023) Cancer statistics, 2023. CA Cancer J Clin 73(1):17–48
Agostinetto E, Gligorov J, Piccart M (2022) Systemic therapy for early-stage breast cancer: learning from the past to build the future. Nat Rev Clin Oncol 19(12):763–774
Zhou Z, Hu Y, Wu Y et al (2022) The immunosuppressive tumor microenvironment in hepatocellular carcinoma-current situation and outlook. Mol Immunol 151:218–230
Zheng H, Li M, Wu L et al (2023) Progress in the application of hydrogels in immunotherapy of gastrointestinal tumors. Drug Deliv 30(1):2161670
Hanahan D, Coussens LM (2012) Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21(3):309–322
Mu G, Zhang W, Huang J et al (2022) Research status of tumor-associated fibroblasts regulating immune cells. Zhongguo Fei Ai Za Zhi 25(3):207–213
Liu Y, Li C, Lu Y et al (2022) Tumor microenvironment-mediated immune tolerance in development and treatment of gastric cancer. Front Immunol 13:1016817
Sepúlveda F, Mayorga-Lobos C, Guzmán K et al (2023) EV-miRNA-mediated intercellular communication in the breast tumor microenvironment. Int J Mol Sci 24(17):13085. https://doi.org/10.3390/ijms241713085
Li HX, Wang SQ, Lian ZX et al (2022) Relationship between tumor infiltrating immune cells and tumor metastasis and its prognostic value in cancer. Cells 12(1):64
Xie J, Huang H, Li X et al (2023) The role of traditional chinese medicine in cancer immunotherapy: current status and future directions. Am J Chin Med 51:1627–1651
Gonzalez A, Valeiras M, Sidransky E et al (2014) Lysosomal integral membrane protein-2: a new player in lysosome-related pathology. Mol Genet Metab 111(2):84–91
He M, Liu Z, Tang B et al (2015) Advances in research of SCARB2 functions and related disorders. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 32(5):723–727
Upadhya SR, Ryan CJ (2022) Experimental reproducibility limits the correlation between mRNA and protein abundances in tumor proteomic profiles. Cell Rep Methods 2(9):100288
Tang Z, Li C, Kang B et al (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45(W1):W98-w102
Li T, Fan J, Wang B et al (2017) TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res 77(21):e108–e110
Li J, Zhang Y, Yang C et al (2020) Discrepant mRNA and protein expression in immune cells. Curr Genomics 21(8):560–563
Li M, Zhao Y, Li H et al (2023) Application value of circulating LncRNA in diagnosis, treatment, and prognosis of breast cancer. Funct Integr Genomics 23(1):61
Liang Z, Liu L, Zhou Y et al (2023) Research progress on bioactive metal complexes against ER-positive advanced breast cancer. J Med Chem 66(4):2235–2256
Cen Y, Chen L, Liu Z et al (2023) Novel roles of RNA-binding proteins in drug resistance of breast cancer: from molecular biology to targeting therapeutics. Cell Death Discov 9(1):52
Niculae AM, Dobre M, Herlea V et al (2022) Lipid handling protein gene expression in colorectal cancer: CD36 and targeting miRNAs. Life (Basel) 12(12):2127
Chen X, Zhang J, Lei X et al (2023) CD1C is associated with breast cancer prognosis and immune infiltrates. BMC Cancer 23(1):129
Su Y, Zhang T, Lu J et al (2023) Identification and validation of the prognostic panel in clear cell renal cell carcinoma based on resting mast cells for prediction of distant metastasis and immunotherapy response. Cells 12(1):180
Lu C, Liu Y, Ali NM et al (2022) The role of innate immune cells in the tumor microenvironment and research progress in anti-tumor therapy. Front Immunol 13:1039260
Shang B, Cui H, Xie R et al (2023) Neutrophil extracellular traps primed intercellular communication in cancer progression as a promising therapeutic target. Biomark Res 11(1):24
Mantovani A, Cassatella MA, Costantini C et al (2011) Neutrophils in the activation and regulation of innate and adaptive immunity. Nat Rev Immunol 11(8):519–531
Sengupta S, Hein LE, Parent CA (2021) The recruitment of neutrophils to the tumor microenvironment is regulated by multiple mediators. Front Immunol 12:734188
Sheng Y, Peng W, Huang Y et al (2023) Tumor-activated neutrophils promote metastasis in breast cancer via the G-CSF-RLN2-MMP-9 axis. J Leukocyte Biol 113:383–399
Kim J, Bae JS (2016) Tumor-associated macrophages and neutrophils in tumor microenvironment. Mediators Inflamm 2016:6058147
Tian S, Chu Y, Hu J et al (2022) Tumour-associated neutrophils secrete AGR2 to promote colorectal cancer metastasis via its receptor CD98hc-xCT. Gut 71(12):2489–2501
Garratt LW, Breuer O, Schofield CJ et al (2021) Changes in airway inflammation with pseudomonas eradication in early cystic fibrosis. J Cyst Fibros 20(6):941–948
Caruso JA, Akli S, Pageon L et al (2015) The serine protease inhibitor elafin maintains normal growth control by opposing the mitogenic effects of neutrophil elastase. Oncogene 34(27):3556–3567
Wang X, Li X, Wu Y et al (2023) The prognostic significance of tumor-associated neutrophils and circulating neutrophils in glioblastoma (WHO CNS5 classification). BMC Cancer 23(1):20
Zheng D, Lin Y, Yu Y et al (2016) The value of preoperative neutrophil to lymphocyte ratio in indicating lymph node metastasis in patients with resectable T2 stage gastric adenocarcinoma. Clin Lab 62(4):659–665
Wang J, Li Y (2019) CD36 tango in cancer: signaling pathways and functions. Theranostics 9(17):4893–4908
Yu X, Guo C, Fisher PB et al (2015) Scavenger receptors: emerging roles in cancer biology and immunology. Adv Cancer Res 128:309–364
Perry JSA, Russler-Germain EV, Zhou YW et al (2018) Transfer of cell-surface antigens by scavenger receptor CD36 promotes thymic regulatory T cell receptor repertoire development and allo-tolerance. Immunity 48(6):1271
Zhang X, Wang H, Sun Y et al (2020) Enterovirus A71 oncolysis of malignant gliomas. Mol Ther 28(6):1533–1546
Guo H, Zhang J, Zhang X et al (2015) SCARB2/LIMP-2 regulates IFN production of plasmacytoid dendritic cells by mediating endosomal translocation of TLR9 and nuclear translocation of IRF7. J Immunol 194(10):4737–4749
Patten DA, Wilkinson AL, O’Keeffe A et al (2022) Scavenger receptors: novel roles in the pathogenesis of liver inflammation and cancer. Semin Liver Dis 42(1):61–76
Hatic H, Sampat D, Goyal G (2021) Immune checkpoint inhibitors in lymphoma: challenges and opportunities. Ann Transl Med 9(12):1037
Marin-Acevedo JA, Kimbrough EO, Lou Y (2021) Next generation of immune checkpoint inhibitors and beyond. J Hematol Oncol 14(1):45
Marin-Acevedo JA, Dholaria B, Soyano AE et al (2018) Next generation of immune checkpoint therapy in cancer: new developments and challenges. J Hematol Oncol 11(1):39
Cejuela M, Vethencourt A, Pernas S (2022) Immune checkpoint inhibitors and novel immunotherapy approaches for breast cancer. Curr Oncol Rep 24(12):1801–1819
Funding
The current study was supported by Jiangsu Province Capability Improvement Project through Science, Technology and Education (ZDXK202234), Jiangsu Provincial Research Hospital (YJXYY202204), China, and Postgraduate Research & Practice Innovation Program of Jiangsu Province (SJCX23_1796). We sincerely appreciate all the study participants.
Author information
Authors and Affiliations
Contributions
Dan Zhang, Jun Fang and Jiali Shan contributed equally to this work.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest for this article.
Ethical approval
The Human Research Ethics Committee of the Affiliated Hospital of Nantong University approved the research method (2018-K020).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, D., Fang, J., Shan, J. et al. SCARB2 associates with tumor-infiltrating neutrophils and predicts poor prognosis in breast cancer. Breast Cancer Res Treat 207, 15–24 (2024). https://doi.org/10.1007/s10549-024-07401-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-024-07401-y