Abstract
Being able to accurately quantify the hemodynamic response function (HRF) that links the blood oxygen level dependent functional magnetic resonance imaging (BOLD-fMRI) signal to the underlying neural activity is important both for elucidating neurovascular coupling mechanisms and improving the accuracy of fMRI-based functional connectivity analyses. In particular, HRF estimation using BOLD-fMRI is challenging particularly in the case of resting-state data, due to the absence of information about the underlying neuronal dynamics. To this end, using simultaneously recorded electroencephalography (EEG) and fMRI data is a promising approach, as EEG provides a more direct measure of neural activations. In the present work, we employ simultaneous EEG-fMRI to investigate the regional characteristics of the HRF using measurements acquired during resting conditions. We propose a novel methodological approach based on combining distributed EEG source space reconstruction, which improves the spatial resolution of HRF estimation and using block-structured linear and nonlinear models, which enables us to simultaneously obtain HRF estimates and the contribution of different EEG frequency bands. Our results suggest that the dynamics of the resting-state BOLD signal can be sufficiently described using linear models and that the contribution of each band is region specific. Specifically, it was found that sensory-motor cortices exhibit positive HRF shapes, whereas the lateral occipital cortex and areas in the parietal cortex, such as the inferior and superior parietal lobule exhibit negative HRF shapes. To validate the proposed method, we repeated the analysis using simultaneous EEG-fMRI measurements acquired during execution of a unimanual hand-grip task. Our results reveal significant associations between BOLD signal variations and electrophysiological power fluctuations in the ipsilateral primary motor cortex, particularly for the EEG beta band, in agreement with previous studies in the literature.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the last 30 years, blood oxygen level-dependent functional magnetic resonance imaging (BOLD-fMRI) (Ogawa et al. 1990) has become the leading imaging technique for studying brain function and its organization into brain networks in both health and disease. Although most fMRI studies use BOLD contrast imaging to determine the brain regions that are active, the BOLD fMRI signal is an indirect measure of neuronal activity through a series of complex events, which are collectively referred to as the hemodynamic response (Buxton et al. 2004). Therefore, accurate interpretation of fMRI data requires understanding of the underlying link between neuronal activity and the BOLD fMRI signal. To this end, using intracranial electrophysiology (Logothetis et al. 2001) has confirmed that BOLD fluctuations are associated with changes in neuronal activity, with stronger correlations reported between BOLD and local field potential (LFP) changes, as compared to spiking activity. However, the physiology of the BOLD signal and its exact association with oscillations within specific frequency bands of the LFP spectrum (< 30 Hz) is still poorly understood.
At the macroscopic scale, simultaneous EEG-fMRI is a commonly used non-invasive technique for the study of the relationship between electrophysiological activity, which is a more direct measure of the underlying neural activity, and the regional changes in the BOLD signal. This technique allows non-invasive recording of brain activity with both high spatial and high temporal resolution, overcoming some of the limitations associated with unimodal EEG or fMRI. Many different analysis methods have been proposed for EEG-fMRI data fusion (Abreu et al. 2018; Jorge et al. 2014). Typically, features extracted from raw EEG time-series are transformed using a static linear or nonlinear transformation and subsequently convolved with a hemodynamic response function (HRF) (Buxton et al. 2004) to explain BOLD signal fluctuations. The accuracy of these predictions depends on both a proper transformation of the EEG features, as well as the shape of the HRF (Sato et al. 2010; Rosa et al. 2010a, b). Symmetric EEG-fMRI data fusion techniques allowing examination of the dynamic interplay between the fMRI and EEG sources have been also suggested (Calhoun et al. 2006).
Two classes of algorithms for EEG feature extraction are typically found in the literature. The first class, which has been mainly employed in task-related studies, refers to the detection of large scale neural events, such as evoked or event-related potentials in response to motor, sensory or cognitive stimuli (Bénar et al. 2007; Fuglø et al. 2012; Nguyen and Cunnington 2014; Wirsich et al. 2014), as well as to epileptic discharges (Bagshaw et al. 2005; Bénar et al. 2002; Murta et al. 2016; Thornton et al. 2010).
The second class, which is the most widely used in the literature, refers to the decomposition of the EEG data into frequency bands of rhythmically sustained oscillations and extraction of the power profile of each band. Along these lines, early attempts to infer BOLD signal dynamics from features extracted from the EEG spectrum focused on the alpha band (8–12 Hz), particularly for the brain in the resting-state (de Munck et al. 2007; Goldman et al. 2002; Laufs et al. 2003, 2006). Similarly, standard frequency bands of the LFP spectrum, such as the delta (2–4 Hz) (de Munck et al. 2009), theta (5–7 Hz) (Scheeringa et al. 2008), beta (15–30 Hz) (Laufs et al. 2006), and gamma (30–80 Hz; Ebisch et al. 2005; Scheeringa et al. 2011, 2016) bands have also been used. However, focusing on specific EEG (or LFP) frequency bands while disregarding the information from others may result in less accurate BOLD signal predictions. Therefore, the importance of including multiple frequency bands in EEG-fMRI data fusion has been suggested (Bridwell et al. 2013; de Munck et al. 2009; Mantini et al. 2007; Tyvaert et al. 2008). Other studies pointed out the importance of using broadband EEG signal transformations, such as a linear combination of band-specific power values (Goense and Logothetis 2008), total power (Wan et al. 2006), and root mean square frequency (Kilner et al. 2005; Rosa et al. 2010a, b). Higher nonlinear or information theoretic transformations have been also suggested (Portnova et al. 2018).
Most of the aforementioned studies performed EEG-fMRI data fusion after imposing constraints that allowed the authors to restrict their attention to a certain number of EEG sensors or within specific frequency bands. More recently, a number of studies proposed using data-driven techniques, such as spectral blind source separation (sBSS) or multiway decomposition to detect information hidden in the structure of both EEG and fMRI, without imposing any prior constraints with regards to the spatial, spectral, or temporal dimensions of the data (Bridwell et al. 2013; Marecek et al. 2016). This approach yielded a set of paired EEG spatial-spectral and fMRI spatial–temporal atoms blindly derived from the data, where each pair of atoms was associated with a distinct source of underlying neuronal activity. The detected pairs of spatial-spectral and spatial–temporal patterns were subsequently used to model the coupling between the two modalities using finite impulse response (FIR) analysis. This method was shown to improve BOLD signal prediction compared to alternative fusion techniques using individual EEG frequency bands. While a finite number of active sources in the brain evoked during task execution might be a reasonable assumption, this may not be the case for the resting-state.
In this work, we extend previous work operating in the time–frequency domain by investigating linear and nonlinear dynamic interactions between EEG and BOLD-fMRI measurements acquired from 12 healthy subjects during resting experimental conditions. To this end, we performed distributed EEG source space reconstruction and employed linear and nonlinear (Hammerstein and Weiner-Hammerstein) block-structured models that have been extensively used for modeling of physiological systems (Westwick and Kearney 2003). This framework allowed us to obtain smooth and accurate HRF estimates directly from the data, as well as to investigate the contribution of the delta (2–4 Hz), theta (5–7 Hz), alpha (8–12 Hz) and beta (15–30 Hz) EEG frequency bands on the BOLD signal variance across the entire cerebral cortex in a high spatial resolution.
Our results suggest that during the resting-state all the examined EEG bands contribute to the fluctuations in the BOLD signal and that the contribution of each EEG band is region specific. They also suggest that increases in the power within lower EEG bands are followed by positive BOLD responses in the sensory-motor cortices. In contrast, increases in the alpha and beta power are followed by negative BOLD responses in the superior and inferior parietal lobule and lateral occipital cortices. Furthermore, increases in the beta band are followed by negative BOLD responses in most brain regions. To validate the proposed method for resting-state HRF estimation, we repeated the same analysis using EEG-fMRI measurements collected during execution of a unimanual hand-grip task. Our results suggest that BOLD signal variance in this case is mainly explained by EEG oscillations in the beta band in the ipsilateral primary motor cortex, in agreement with previous studies is the literature.
Methods
Experimental Methods
Twelve healthy volunteers (age range 20–29 years) participated in this study after giving a written informed consent in accordance with the McGill University Ethical Advisory Committee. All participants were right-handed according to the Edinburgh Handedness Inventory (EHI; Oldfield 1971): mean EHI score = 76.66 ± 14.74 (SD); EHI score range [44.44–100]. Measurements were recorded at the McConnel Brain Imaging center (BIC) of the Montreal Neurological Institute (MNI), at McGill University.
Experimental Paradigm
The study was divided in two scans (Fig. S1). During the first scan (resting-state experiment—time of acquisition = 15 min 7 s), subjects were asked to perform no particular task other than to remain awake while looking at a white fixation cross displayed in a dark background. During the second scan (motor task experiment—time of acquisition = 14 min 14 s), subjects were asked to perform unimanual isometric right-hand grips to track a target as accurately as possible, while receiving visual feedback. At the beginning of each trial, an orange circle appeared on the screen and subjects had to adapt their force at 15% of their maximum voluntary contraction (MVC) to reach a white vertical block (low force level). This force was maintained at this level for 3 s. Subsequently, subjects had to progressively increase their force over a 3-s period following a white ramp to reach 30% of their MVC and to sustain their applied force at this level for another 3 s (high force level). A single trial lasted 11 s and was repeated 50 times. The inter-trial interval was randomly jittered between 3 and 5 s, during which subjects were able to rest their hand while looking on a white fixation cross. The MVC of each participant was obtained between the two scans, using the same hand gripper that was employed during the motor task.
Hand Grip Force Measurements
A non-magnetic hand clench dynamometer (Biopac Systems Inc, USA) was used to measure the subjects’ hand grip force strength during the motor paradigm. The dynamometer was connected to an MR compatible Biopac MP150 data acquisition system from which the signal was transferred to a computer.
EEG Data Acquisition
Scalp EEG signals were simultaneously acquired during fMRI scanning at 5 kHz using a 64 channel MR-compatible EEG system with ring Ag/AgCl electrodes (Brain Products GmbH, Germany). The electrodes were placed according to the 10/20 system and referenced to electrode FCz. The EEG data were synchronized with the MRI scanner clock via a synchronization device to improve the effectiveness of MRI artifact removal (see "EEG Data Preprocessing" section for details). Triggers indicating the beginning and end of each session, as well as the timing of each phase of the motor task during the motor task experiment were sent to both the Biopac and the EEG recording devices via a TriggerBox device (Brain Products GmbH, Germany). The electrodes were precisely localized using a 3-D electromagnetic digitizer (Polhemus Isotrack, USA).
BOLD Imaging
Whole-brain BOLD-fMRI volumes were acquired on a 3 T MRI scanner (Siemens MAGNETOM Prisma fit) with a standard T2*-weighted echo planar imaging (EPI) sequence using a 32-channel head coil for reception. EPI sequence parameters: TR/TE = 2120/30 ms (Repetition/Echo Time), Voxel size = 3 × 3 × 4 mm, 35 slices, Slice thickness = 4 mm, Field of view (FOV) = 192 mm, Flip angle = 90°, Acquisition matrix = 64 × 64 (RO × PE), Bandwidth = 2368 Hz/Px. Four hundred-twenty whole-brain volumes were acquired during the resting-state experiment and four hundred during the motor task experiment, respectively. A high-resolution T1-weighted MPRAGE structural image was also acquired to aid registration of the functional volumes to a common stereotactic space. MPRAGE sequence parameters: TI/TR/TE = 900/2300/2.32 ms (Inversion/Repetition/Echo Time), Flip angle = 8°, 0.9 mm isotropic voxels, 192 slices, Slice thickness = 0.9 mm, Field of view = 240 mm, Acquisition matrix = 256 × 256 (RO × PE), Bandwidth = 200 Hz/Px.
Data Preprocessing
EEG Data Preprocessing
EEG data acquired inside the scanner were corrected off-line for gradient and ballisto-cardiogram (BCG) artifacts using the BrainVision Analyser 2 software package (Brainproducts GmbH, Germany). The gradient artifact was removed via adaptive template subtraction (Allen et al. 2000). Gradient-free data were band-passed between 1 and 200 Hz, notch-filtered at 60, 120, and 180 Hz to remove power-line artifacts, and down-sampled to a 400 Hz sampling rate. Temporal independent component analysis (ICA) (Delorme and Makeig 2004) was performed on each subject separately. In each case, the number of components extracted was equal to the number of channels in the EEG data. The BCG-related component that accounted for most of the variance in the data was isolated and used to detect heartbeat events. Subsequently, BCG-related artifacts were removed via pulse artifact template subtraction, which was constructed using a moving average of EEG signal synchronized to the detected heartbeat events (Allen et al. 1998). Poorly connected electrodes were detected using visual inspection, as well as evaluation of their power spectrum, and interpolated using spherical interpolation (Delorme and Makeig 2004). Subsequently, a second temporal ICA was performed, and noisy components associated with non-neural sources, such as gradient and BCG residuals, ocular, or muscle artifacts were removed. The median of the retained ICA components was 21 (interquartile range (IQR) 13.5–23) for the resting-state data and 16 (IQR 13–20) for the motor task data, respectively. The remaining data were re-referenced to an average reference. After preprocessing, one subject was excluded from further analysis due to excessive noise that remained in the data.
MRI Data Pre-processing
Pre-processing of the BOLD images was carried out using the Oxford Centre for Functional Magnetic Resonance Imaging of the Brain Software Library (FMRIB, UK—FSL version 5.0.10) (Jenkinson et al. 2012). The following pre-processing steps were applied: brain extraction, high-pass temporal filtering (cutoff point = 90 s), spatial smoothing using a Gaussian kernel of 5 mm FWHM, volume realignment, and normalization to the MNI-152 template space, with resolution of 2 mm3. Spatial ICA was carried out for each subject using FSL’s MELODIC (Beckmann and Smith 2004) and spatial maps associated with head motion, cardiac pulsatility, susceptibility and other MRI-related artifacts with non-physiologically meaningful temporal waveforms were removed. MRI structural analysis and reconstruction of cortical surface models were performed with the FreeSurfer image analysis suite (version 5.3.0) (Fischl, 2012). 62 anatomical regions of interest (ROIs) were also defined in the native space of each individual according to the Mindboggle atlas using FreeSurfer (https://mindboggle.info) (Klein and Tourville 2012). The fMRI data were co-registered to the reconstructed EEG cortical source space (see "EEG Source Imaging" section for details) using volume-to-surface registration (Dickie et al. 2019).
Data Analysis
EEG Source Imaging
Our main aim was to model the dynamic interactions between individual EEG sources and BOLD-fMRI in high spatial resolution. To this end, we reconstructed the EEG source space for each subject using an extension of the linearly constrained minimum variance (LCMV) beamformer (Van Veen et al. 1997), which is implemented in Brainstorm (Tadel et al. 2011). Beamformers are adaptive linear spatial filters that isolate the contribution of a source located at a specific position of a 3D grid model of the cortical surface, while attenuating noise from all other locations yielding a 3D map of brain activity.
A set of 15,000 current dipoles distributed over the cortical surface was used. Source activity at each target location on the cortical surface was estimated as a linear combination of scalp field measurements, wherein the weights, as well as the orientation of the source dipoles were optimally estimated from the EEG data in the least-squares sense. A realistic head model for each subject was obtained using the subject’s individual cortical anatomy and precise electrode locations on the scalp. Lead fields were estimated using the symmetric boundary element method (BEM) (Gramfort et al. 2009). The relative conductivities assumed for estimation of the lead fields were 1 for the scalp, 0.0125 for the skull, and 1 for the brain.
Time–Frequency Analysis
EEG source waveforms were band-passed into the delta (2–4 Hz), theta (5–7 Hz), alpha (8–12 Hz) and beta (15–30 Hz) frequency bands and the complex analytic signal of each band was obtained via the Hilbert transform (Bruns 2004; Le Van Quyen et al. 2001). Band-pass filtering was performed using even-order linear phase FIR filters with zero-phase and zero-delay compensation implemented in Brainstorm. Subsequently, the instantaneous power time-series within each EEG band was calculated as the squared amplitude of the corresponding complex analytic signal. The EEG bandwidth was limited between 1 and 30 Hz, as above that frequency range MRI-related artifacts are more difficult to remove (Mullinger et al. 2008, 2011, 2014; Ryali et al. 2009), particularly for resting-state EEG, making the calculation of a signal of good quality more challenging. EEG instantaneous power time-series were down-sampled by averaging within the BOLD sampling interval yielding one value per fMRI volume. Representative band-specific EEG instantaneous power time-series from the left lateral occipital cortex superimposed with the corresponding BOLD time-series obtained from one representative subject during the motor task are shown in (Fig. S2) in the supplementary material.
Mathematical Methods
Block-Structured System Modeling
The dynamic interactions between EEG bands and BOLD were assessed using multiple-input single-output models. The nonlinear Hammerstein model (Fig. 1a) consists of the cascade connection of a static nonlinear map followed by a dynamic, linear time invariant (LTI) system. In the linear Hammerstein model (Fig. 1b), the static nonlinear map is substituted by a static linear map. The nonlinear Hammerstein-Wiener model (Fig. 1c) is an extension of the nonlinear Hammerstein model consisting of a second static nonlinearity that follows the output of the dynamic LTI system. The output nonlinearity of the Hammerstein-Wiener model allows modeling of nonlinear dynamic interactions between the input (source EEG frequency bands) and output data (BOLD-fMRI). These modular cascade models, which have been extensively used for modeling linear and nonlinear physiological systems (Westwick and Kearney 2003), are well suited for modeling the dynamics between EEG and BOLD-fMRI data as they provide estimates of the interactions between different EEG frequency bands and their effect on the BOLD signal, as well as the HRF without requiring a priori assumptions with regards to its shape.
a Multiple-input–single-output (MISO) nonlinear Hammerstein model consisting of a static (memoryless) nonlinearity N(⋅) followed by a linear time-invariant (LTI) system L. b Multiple-input–single-output (MISO) linear Hammerstein model consisting of a static linear map NL(⋅) followed by a LTI system L. c Multiple-input–single-output (MISO) Hammerstein-Wiener model consisting of a Hammerstein model followed by a static nonlinearity NL(⋅). Notation: n = 1,… denotes time. u1(n), …, uQ(n) denote the measured input (i.e. instantaneous power in Q different EEG bands) and y(n) the measured output (i.e. BOLD signal) of the different MISO systems. N(u(t)) and NL(u(t)) denote the static nonlinear and linear transformation of the Hammerstein and linear Hammerstein system input, respectively. yh(n) and F(yh(n)) denote the Hammerstein and Hammerstein-Weiner model output prediction, respectively. ε(n) denotes a noise process
Hammerstein Model Identification The MISO Hammerstein model (Fig. 1a) of the dynamic interactions between source EEG frequency bands and BOLD-fMRI consists of a nonlinear (polynomial) signal transformation of the EEG bands \(\text{N}\left(\cdot \right)\) in cascade with a LTI system \(\text{L}\left(\cdot \right)\), which is described in terms of an HRF \(\text{h}\left(\text{t}\right)\). The input–output relationship in discrete time is given by
where \(\text{y}\left(\text{n}\right)\) denotes the output (i.e. BOLD signal) and \(\mathbf{u}\left(\text{n}\right)\) the multivariate input (i.e. EEG frequency bands). The nonlinear block can be described by
where \({\text{g}}_{\text{i}}^{\left(\text{p}\right)}\left(\cdot \right)\) are polynomial terms of order \(\text{p}\) that allow representation of nonlinearities in the EEG bands, \({\text{a}}_{\text{I},\text{p}}\) are the unknown coefficients corresponding to the \(\text{p}\)-th polynomial term of the \(\text{i}\)-th EEG band, and Q denotes the total number of EEG bands. In this study, \(\text{Q}=4\) as the model input consists within four distinct source EEG frequency bands (see "Time-Frequency Analysis" section for details).
The MISO linear Hammerstein model is a special case of the Hammerstein model when \(\text{P}=1\). In this case, the output of the system is described as the convolution between a linear combination of the EEG frequency bands with the HRF. This model is consistent with the frequency response (FR) model that has been previously proposed in the neuroimaging literature (Goense and Logothetis 2008; Rosa et al. 2010a, b), which assumes that BOLD is best explained by a linear combination of synchronized activity within different EEG bands.
Hammerstein-Wiener Model Identification The MISO Hammerstein-Wiener (HW) model structure (Fig. 1b) consists of a nonlinear transformation \(\mathbf{F}\left(\cdot \right)\) in cascade with a Hammerstein system described by (1). The input–output relationship of the HW model in discrete time is given by
where \(\text{y}\left(\text{n}\right)\) denotes the system output (i.e. BOLD signal), and \({\text{y}}_{\text{H}}\left(\text{n}\right)\) the output of the preceding Hammerstein system. \({\text{F}}^{\left(\text{k}\right)}\left(\cdot \right)\) are polynomial terms of order \(\text{K}\) that allow representation of nonlinearities in the output of the preceding Hammerstein system, and \({\text{z}}_{\text{k}}\) is the coefficient of the \(\text{k}\)-th polynomial term.
The unknown polynomial coefficients \({\text{a}}_{\text{I},\text{p}}\) and \({\text{z}}_{\text{k}}\), and the unknown HRF of these block-structured models were estimated efficiently from the data using a function expansion technique (Marmarelis 1993), as described in Appendix.
Model Performance
Our goal was to compare models considering linear (linear Hammerstein) and nonlinear (Hammerstein) transformation of the power within different source EEG frequency bands, as well as linear and nonlinear dynamic behavior (Hammerstein-Wiener) that can be used to predict BOLD signal variations. To this end, we employed a threefold cross validation approach as follows: band-specific EEG and BOLD time-series were partitioned into three segments of equal length. Each segment was sequentially used as the validation set for assessing the performance of each model and the remaining two segments were used as the training set. For each segment, the parameters of the three models under consideration were estimated using the training set, and model performance was evaluated using the testing set in terms of the mean-squared prediction error (MSE). For the resting-state data, the validation and training set consisted of 280 and 140 data points, respectively. For the task data, the training and testing set consisted of 266 and 134 data points, respectively. In each case, the MSE was computed as
where \(\widehat{\text{y}}\left(\text{n}\right)\), and \(\text{y}\left(\text{n}\right)\) denote the predicted and measured BOLD, respectively. The average \(\text{MSE}\) value obtained across the three folds, which is typically referred to in the literature as the generalization error, was calculated and used for model comparisons. To prevent overfitting, particularly in the case of resting-state EEG-fMRI measurements where the SNR is considerably lower, the range for the total number \(\text{L}\) of spherical Laguerre functions used for modeling the impulse response of the linear filter \(\text{L}\left(.\right)\) and the range for \(\alpha\) were set to \(2<\text{L}\le 4\) and \(0.5<\alpha <1\), respectively (see Appendix for details). The range of these parameters was selected to ensure a reasonable complexity for the estimated hemodynamic models while providing a broad range of possible dynamics (fast/slow) (Fig. S12). The optimal value for the L and α parameters was determined based on model performance using a grid search.
Vertex-Wise Analysis
Contribution of Individual EEG Bands to BOLD Signal Variance
In each voxel, the contribution of individual source EEG frequency bands to the BOLD signal variance was evaluated in two steps. In the first step, the linear Hammerstein model, which is described by Eq. (1) for \(\text{P}=1\), was fitted to the full data set, and a BOLD prediction was obtained. In the second step, the linear Hammerstein model was refitted to a reduced data set from which the target EEG frequency band was excluded, and a BOLD prediction was obtained. Then the F-score was calculated using
where \({\text{SSE}}_{\text{F}}\) and \({\text{SSE}}_{\text{R}}\) are the residual sum of squares of the full and reduced model respectively. Likewise, \({\text{DFE}}_{\text{F}}\) and \({\text{DFE}}_{\text{R}}\) are the number of degrees of freedom for the full and reduced model, respectively. In each case, there are N –Q degrees of freedom, where N is the number of data points and Q is the number of regressors used in the model. The statistic \(\text{F}\) follows a \({\text{F}}_{\left({\text{DFE}}_{\text{R}}-{\text{DFE}}_{\text{F}},{\text{DFE}}_{\text{F}}\right)}\) distribution and a large value of \(\text{F}\) indicates that the target EEG band significantly contributes to BOLD signal variance while taking into consideration the difference in the number of regressors used in the full and reduced model, respectively.
Influence of Individual EEG Bands on HRF Scaling
The linear Hammerstein model described by Eq. (1) when \(\text{P}=1\) quantifies the interactions between EEG and fMRI as an HRF (impulse response of the LTI block) scaled by coefficients reflecting the relative contribution of each EEG band to BOLD signal variations (static linear MISO block). To investigate the influence of individual EEG bands on HRF scaling we proceeded in two steps (Fig. 2): First, we excited all inputs of the linear Hammerstein system at the same time using a Kronecker delta function \(\updelta \left(\text{n}\right)\) as input (Fig. 2a) to derive the system’s dynamic response to instantaneous changes in the power of all EEG bands (total HRF). The scaling of the total HRF was determined by the sum of the coefficients \(\mathbf{a}\) that define the static linear MISO block. Subsequently, we excited one input at a time (Fig. 2b). In each case, the scaling of the derived response (band-specific HRF), was determined by the coefficient \({\text{a}}_{\text{i}}\) corresponding to the \(\text{i}\)-th band.
Network representation of the multiple-input–single-output linear Hammerstein model used for quantifying the dynamical interactions between EEG and BOLD-fMRI. a The total HRF hT(n) is obtained by exciting all inputs of the linear Hammerstein system at the same time using a Kronecker delta function δ(n). The scaling of the total HRF is determined by the sum of all input coefficients ai, i = 1,…,Q. b A band-specific HRF hi(n) is obtained by exciting only the i-th input, which is associated with the i-th EEG frequency band. The scaling of the HRF in this case is determined by coefficient ai
To assess the contribution of individual EEG bands on the scaling of the total HRF in different brain regions, we compared the spatial maps of the HRF peak with the spatial maps of the band-specific HRF peak. The HRF peak describes the maximum instantaneous hemodynamic response to changes in neuronal activity.
Statistics
Model comparisons were performed using MSE values obtained for all the 62 Mindboggle atlas ROIs, which were derived in the native space of each individual using FreeSurfer. The selected ROIs are illustrated in Fig. S11 in the Supplementary material. Group-level statistical comparisons were carried out using linear models including the averaged MSE across ROIs within each subject as outcome variable and the block-structured model type as predictor variable. The MSE values were averaged across ROIs to account for spatial correlation within subjects. The optimal model for explaining the dynamic relation between source EEG frequency bands and fMRI was determined as the most parsimonious model exhibiting the smallest generalization error. Comparison of the MSE values indicated that the linear Hammerstein model is sufficient for describing the dynamic relation between different source EEG bands and BOLD for both experimental conditions (Fig. 3). Furthermore, to assess the performance of our approach, we compared the HRF estimates obtained using the proposed function expansion technique to three other HRF estimation methods that have been previously used in the neuroimaging literature for HRF estimation: (i) non-parametric, direct impulse response estimation with stable spline regularization (Chen et al. 2012; Lu et al. 2006; Sato et al. 2010), (ii) function expansions using the canonical HRF along with its time and dispersion derivatives (Friston et al. 1998), and (iii) function expansions using the first two components derived by applying principal component analysis (PCA) on an extended set of gamma density functions (Hossein-Zadeh et al. 2003). In the latter case, the extended set of gamma density functions was constructed by varying the peak (τ) and dispersion (σ) parameters of the gamma density functionFootnote 1 as follows: 0.2 ≤ σ ≤ 0.4 and 1 ≤ τ ≤ 14. For the purpose of this comparison, we employed the hand grip force time-series. Statistical comparisons of the MSE values achieved by each method were performed using linear models including MSE as the outcome variable and estimation method as the predictor variable. Voxel-wise statistical comparisons were performed at the group level using one-sample t-tests. The resulting statistical parametric maps were corrected for multiple comparisons using false discovery rate (FDR).
Boxplots of mean square error (MSE) values between measured versus predicted BOLD in large structurally defined ROIs from all subjects. BOLD predictions were obtained using the block-structured Hammerstein-Wiener (HW), Hammerstein (H), and linear Hammerstein (LH) models, and the instantaneous power time series within the delta (2–4 Hz), theta (5–7 Hz), alpha (8–12 Hz) and beta (15–30 Hz) bands. Statistical comparisons between the MSE values obtained from each model were performed using linear models including averaged MSE values across ROIs within each subject as outcome variable and block-structured model type as predictor variable. During resting conditions, the MSE obtained from the H and LH were significantly lower compared to the WH (WH vs H: B = − 0.07, t(30) = − 3.27, p = 0.003; WH vs LH: B = − 0.1, t(30) = − 4.355, p < 0.001). No significant differences were detected between the H and LH models. Also, no significant differences were detected during the task. The results collectively suggest that increasing model complexity does not significantly improve model performance and that the LH model is adequate to describe the dynamics between EEG and BOLD during both experimental conditions
Results
Model Comparisons
The Hammerstein-Wiener, Hammerstein, and linear Hammerstein block-structured models were compared in terms of their mean square prediction error (MSE) obtained in large structurally defined ROIs according to the Mindboggle atlas. Boxplots of the averaged MSE values across ROIs within subjects are shown in Fig. 3 for each model and experimental condition. During resting conditions, the MSE values achieved by the linear Hammerstein model were significantly lower compared to the Hammerstein-Wiener model (B = − 0.1, t(30) = − 4.355, p < 0.001), but not when compared to the standard Hammerstein model. No significant differences were detected during the task. These results suggest that increasing model complexity does not improve its performance. They also suggest that the BOLD signal can be sufficiently described as the convolution between a linear combination of the power profile within different frequency bands and a HRF, which can be estimated from the data using the functional expansion technique along with the spherical Laguerre basis (see "Block-Structured System Modeling" section for details). Representative HRF estimates obtained using the proposed method, as well as three other methods previously used in the literature (see "Statistics" section) are illustrated in Fig. S3a. Figure S3b shows boxplots of the cross-validation MSE values achieved by each method. The MSE achieved by the spherical Laguerre model was lower compared to all other methods within each subject, and statistically lower compared to the direct impulse response method (p < 0.01).
Contribution of Individual EEG Bands to BOLD Signal Variance
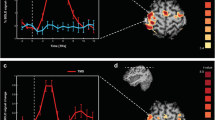
Figure 4 illustrates the contribution of individual frequency bands of EEG current sources to the resting-state BOLD signal variance. EEG source space reconstruction was performed using distributed source imaging, whereby dipolar current sources were estimated along the cortical surface in high spatial resolution (see "EEG Source Imaging" section for details). BOLD signal predictions were obtained using the linear Hammerstein model and the custom spherical Laguerre HRF. The results suggest significant contributions from all EEG frequency bands (\(\text{p}<0.0001\), FDR corrected for multiple comparisons), and for each band the significant contributions were found to be region-specific. Specifically, lower EEG bands, such as the delta and theta frequency bands, exhibited significant contribution to the BOLD signal in the primary motor and somatosensory cortices (Fig. 4—upper row). On the other hand, higher EEG bands, such as the alpha and beta frequency bands, exhibited significant contribution to the BOLD signal in visual-related areas in the occipital cortex (Fig. 4—bottom row). To validate the proposed method, we repeated the same analysis using the data obtained during the hand grip task (Fig. S4). Using the handgrip force time-series, we observed significant BOLD signal predictions in the left primary motor cortex, which was used as a gold standard (Fig. S4d). As expected, similar results were obtained using the source space EEG frequency bands (Fig. S4c). In this case, larger contributions to the BOLD signal variance in this area were observed from the beta frequency band (15–30 Hz).
Contribution of individual frequency bands of distributed EEG sources to BOLD signal variance during resting-state conditions. The analysis was performed using custom voxel-specific HRFs, which were estimated directly from the data using the linear Hammerstein model and spherical Laguerre basis functions. Group-level one-sample t-statistical maps were obtained for each frequency band separately (\(\text{p}<0.0001\), FDR corrected for multiple comparisons). The delta (2–4 Hz) and theta (4–8 Hz) frequency bands contributed significantly to BOLD signal variance in the primary motor and somatosensory cortices. The alpha (8–12 Hz) and beta (15–30 Hz) frequency bands contributed significantly to BOLD signal variance in the occipital cortex
Group average HRF estimates obtained in functionally defined ROIs in which EEG explained a large fraction of BOLD variance during resting conditions are shown in (Fig. 5). These ROIs included the right primary motor and lateral occipital cortices (Fig. 4). Representative BOLD signal predictions obtained from one subject for the right lateral occipital cortex is also shown in the same Figure. These results suggest that the linear Hammerstein model can be used to obtain reliable estimates of the HRF as well as BOLD signal predictions from the EEG even during the resting state, where SNR is particularly low. Similar results were also obtained during the motor task for the left primary motor and superior parietal lobule cortices (Fig. S5). It should be noted similar results were also obtained when using EEG instantaneous power time-series at a higher sampling rate (256 Hz). Therefore, using down-sampled EEG instantaneous power time-series did not affect the accuracy of the BOLD signal predictions (Fig. S10).
Group average normalized HRF curves obtained in the right primary motor and right lateral occipital cortices under resting conditions. The red curve corresponds to the mean HRF curve across all subjects. The blue shaded area corresponds to the standard error. The ROIs were functionally defined based on regions where EEG explained a large fraction of the variance in the BOLD signal (Fig. 4). A representative BOLD prediction in the right occipital cortex obtained from one subject is shown in the lower panel. The same plot superimposed with the instantaneous power of individual EEG bands is shown in (Fig. S9) the supplementary material
Influence of Individual EEG Bands on HRF Scaling
To investigate the regional variability of the total HRF in high spatial resolution, we excited all inputs of the estimated linear Hammerstein model at each voxel at the same time using one Kronecker delta function for each input. The derived dynamic response was determined from both the shape of the HRF provided by the impulse response of the LTI block, as well as the total scaling coefficient provided by the sum of the \(\mathbf{a}\) coefficients that define the static linear MISO block (see "Mathematical methods" section for details). Average maps of total HRF peak values obtained during resting conditions across all subjects are shown in (Fig. 6). The results suggest that areas in the attention cortical network, such as the dorsal lateral prefrontal and inferior parietal lobule cortices, as well as areas in the default mode network, such as the medial prefrontal and precuneus cortices exhibit a negative response to abrupt instantaneous increases in the source EEG power. On the other hand, areas in the primary sensory, primary motor, medial occipital, insular, and auditory cortices exhibit a positive hemodynamic response. During the motor task (Fig. S6), the vast majority of brain areas spanning the cortical surface exhibited a negative hemodynamic response to abrupt instantaneous increases of the source EEG power for all frequency bands. The largest negative responses were observed in the superior parietal lobule and lateral occipital cortices. On the other hand, areas in the primary somatosensory, primary motor and medial occipital cortices exhibited a positive hemodynamic response.
Group-level averaged maps of total HRF peak obtained by exciting all inputs of the linear Hammerstein model estimated at each voxel at the same time, using one Kronecker delta function (resting conditions). The total HRF was determined by both the HRF shape provided by the impulse response of the LTI block, as well as the total scaling coefficient provided by the sum of the a coefficients that define the static linear MISO block of the linear Hammerstein model. Areas in the attention cortical network, such as the dorsal-lateral prefrontal and inferior parietal lobule cortices, as well as areas in the default mode network, such as the medial prefrontal and precuneus cortices, exhibited a negative hemodynamic response to abrupt instantaneous increases in the resting-state EEG power. On the other hand, areas in the primary somatosensory, primary motor, medial occipital, insular, and auditory cortices exhibited a positive hemodynamic response. mPFC medial prefrontal cortex, PCC precuneus cortex, S1 primary sensory cortex, M1 primary motor cortex, PMC premotor cortex, SPL superior parietal lobule, IPL inferior parietal lobule
Figure 7 shows group-level average band-specific HRF peak maps obtained under the resting-state condition for the delta (2–4 Hz), theta (5–7 Hz), alpha (8–12 Hz) and beta (15–30 Hz) frequency bands. Band-specific HRF peak maps were obtained by exciting one input of the linear Hammerstein model estimated at each voxel at a time, using a Kronecker delta function. The obtained band-specific HRF associated with the i-th input was determined by both the HRF shape provided by the impulse response of the LTI block, as well as the coefficient \({\text{a}}_{\text{i}}\), which reflects the relative contribution of the i-th input to the BOLD signal. The alpha and beta bands exhibited widespread negative responses. The largest negative responses for the alpha band were observed in the occipital cortex, whereas for the beta band in areas involved in the cortical attention network, such as the dorsal lateral prefrontal cortex. On the other hand, the delta and theta frequency bands exhibited strong positive responses in the motor, somatosensory, superior parietal lobule, auditory and insular cortices. Also, the medial occipital cortex exhibited negative responses for the alpha and beta bands, and strong positive responses for the delta and theta frequency bands. During the motor task (Fig. S7), the alpha and beta frequency bands exhibited strong negative responses in visual related areas, such as the lateral occipital and superior parietal lobule cortices. On the other hand, the delta and theta frequency bands exhibited strong positive responses in the primary motor and somatosensory cortices. All frequency bands exhibited a positive hemodynamic response in the medial occipital cortex, with the strongest responses being observed for the delta and beta frequency bands.
Group-level average maps of band-specific HRF peak values obtained during resting-state by exciting one input of the linear Hammerstein model estimated at each voxel at a time, using a Kronecker delta function. Each input was associated with a different frequency band, and the relative contribution of the i-th input to BOLD signal variance was quantified in terms of the coefficient ai. In each case, the band-specific HRF was determined by both the HRF shape provided by the impulse response of the LTI block, as well as the coefficient ai of the associated i-th input. The alpha and beta frequency bands exhibited widespread negative hemodynamic responses spanning in multiple cortical regions. For the alpha band, the largest negative responses were observed in the lateral occipital cortex, and for the beta band in areas in the attention cortical network, such as the dorsal lateral prefrontal and inferior parietal lobule cortices. Moreover, the delta and theta frequency bands exhibited strong positive responses in areas in the primary somatosensory, motor, insular, and auditory cortices, as well as in visual-related areas, such as the lateral occipital and superior parietal lobule cortices
Discussion
In the present work, we investigated in detail the dynamic interactions between changes in neuronal activity and the BOLD signal measured with simultaneous EEG-fMRI under resting-state conditions. To quantify these interactions in high spatial resolution, we reconstructed the EEG source space along the cortical surface using distributed source space analysis, in contrast to similar previous studies, which performed this investigation using EEG sensor level measurements (de Munck et al. 2009, 2007; Laufs et al. 2003, 2006; Mantini et al. 2007; Portnova et al. 2018; Rosa et al. 2010a, b; Sclocco et al. 2014). Source space reconstruction allows the spatial information present in the multi-channel EEG to be better exploited, providing more information regarding the local neuronal input within a given cortical area. The dynamic interactions between EEG and BOLD were investigated using block-structured linear and nonlinear models that describe the BOLD signal as the convolution between a static linear (linear Hammerstein) or nonlinear (standard Hammerstein) polynomial transformation of the EEG power within different frequency bands with a hemodynamic response function. We also investigated the possible presence of dynamic nonlinearities in the BOLD signal using the Hammerstein-Wiener model. These nonlinearities may result from suppression and increased latency of present BOLD responses that are incurred by preceding changes in the source EEG power (Friston et al. 2000).
The degree and coefficients of the polynomial transformation preceding (Hammerstein structure) and following (Hammerstein-Wiener structure) the linear hemodynamic system (Fig. 1), as well as the shape of the unknown HRF curve were determined from the data using partial least squares regression (PLSR). PLSR was employed to account for the high collinearity between the instantaneous power of different frequency bands, as it provides unbiased estimates of the unknown model parameters. Moreover, the unknown HRF curves estimated in both large ROIs and individual voxels were estimated efficiently from the data using function expansions in terms of the spherical Laguerre basis functions. The use of an orthonormal basis reduces the number of required free parameters in the model and allows parameter estimation using least-squares regression, which leads to increased estimation accuracy in the presence of noise even from short experimental data-records (Marmarelis 2004).
To validate the proposed method used for the analysis of resting-state EEG-fMRI data, we repeated the same procedure for the analysis of data acquired during a hand grip task that elicits a well-described neuronal response in the ipsilateral primary motor cortex (Xifra-Porxas et al. 2019; Sclocco et al. 2014). The comparison of the BOLD variance explained by the different EEG frequency bands using sensor versus source space analysis (Fig. S4a, b), as well as using the canonical, double gamma versus a custom HRF (Fig. S4b, c) revealed increased detection sensitivity and region specificity of brain activation when source space analysis and a custom HRF were employed. Similar activation patterns were obtained using the handgrip force time-series (Fig. S4d), which reflect the dynamics of neural activation in the ipsilateral primary motor cortex in response to the handgrip task. The similarity between the activation maps obtained using the handgrip force and power in the beta band during the task suggests the potential of the proposed method for obtaining reliable BOLD predictions and HRF estimates even from resting-state data, where there is no explicit task.
Model comparisons revealed that the convolution between a linear combination of the power profile of different EEG frequency bands with a hemodynamic response function is sufficient to describe the dynamics observed between fluctuations in the power of different frequency bands and the BOLD signal. Using this linear model, we showed that the contribution of different frequency bands to the BOLD signal variance strongly depends on brain region and experimental condition. Our results suggest that the proposed method yields robust HRF estimates even during resting conditions, despite the lower SNR associated with them. This has important implications particularly in the context of resting-state functional connectivity, as accurate HRF estimates are important for removing the hemodynamic blurring that is inherent in the fMRI time series, resulting in more accurate functional connectivity maps (Rangaprakash et al. 2018; Wu et al. 2013).
The Proposed Linear Model for the Dynamic Interactions Between Source Space EEG and BOLD-fMRI
The model comparisons shown in Fig. 3 suggest that the linear Hammerstein model achieves smaller mean squared error (\(\text{MSE}\)) values compared to the standard Hammerstein and Hammerstein-Wiener model, for both experimental conditions. In each case the \(\text{MSE}\) values were obtained using a threefold cross-validation approach, which was implemented to assess model performance (see "Model performance" section for details). In this context, the linear Hammerstein model was found to yield the optimal balance between predictive accuracy and parsimony, which suggests that it can sufficiently describe the dynamics between source EEG and BOLD-fMRI without overfitting.
The linear Hammerstein model assumes that the BOLD signal is best explained by a linear combination of activity within different frequency bands in agreement with the frequency response (FR) modelFootnote 2 previously used to predict BOLD activity from intra-cortical LFP recordings in alert behaving monkeys (Goense and Logothetis 2008). The main difference between the FR and the linear Hammerstein hemodynamic model proposed herein is that the latter employs a custom HRF to describe the dynamic interactions between source EEG power and BOLD-fMRI, which is estimated directly from the experimental data. This provides additional flexibility in modeling the dynamic relation between changes in neuronal activity and BOLD as compared to the FR model. Also, it allows for the investigation of the regional variability of the HRF in high spatial resolution.
In contrast to other linear hemodynamic models which assume a different HRF shape for each EEG frequency band (Bridwell et al. 2013; de Munck et al. 2009), the linear Hammerstein model employs a unique HRF curve shape for all EEG bands, which is estimated directly from the data. We hypothesized that the dynamics of the physiological mechanism that relates changes in neuronal activity to changes in cerebral blood flow do not depend on the specific frequency of the underlying neural oscillations. Instead, the dynamics of the hemodynamic response to changes in the EEG power, which determine the HRF curve shape, are an intrinsic property of the local cerebral vasculature that is related to elastance and compliance. On the other hand, the relative contribution of each EEG band to BOLD signal variance is reflected on the scaling coefficient \({\text{a}}_{\text{i}}\) of the HRF that is associated with each band. Hence, a large positive scaling coefficient corresponds to a frequency band that is positively correlated with the BOLD signal and explains a large portion of its variance. Likewise, a large negative coefficient corresponds to a frequency band that is negatively correlated with the BOLD signal. In contrast, a small positive (negative) scaling coefficient corresponds to a frequency band that is weakly positively (negatively) correlated with the BOLD signal.
A model that has been extensively used in the literature for modelling the dynamic interactions between neuronal activity and BOLD during task execution (Murta et al. 2015; Rosa et al. 2010a, b; Rosa et al. 2011; Sclocco et al. 2014), as well as during EEG epileptic activity (Leite et al. 2013) is the so called Heuristic model proposed by (Kilner et al. 2005). This model uses the root mean square frequency of the normalized power spectrum to define a nonlinear signal transformation of the EEG power that is used to predict changes in the BOLD signal. The power spectrum employed by this model is normalized with the total average power of the EEG (area under the power spectral density) at each time instant. Hence, direct comparison between the Heuristic and the linear Hammerstein model employed in this work is not straightforward, as the later uses an absolute power spectrum. However, the statistical comparisons shown in (Fig. 3) suggest that the linear Hammerstein model would be superior to the root mean square frequency model using an absolute power spectrum (unnormalized Heuristic model), as the latter can be adequately described with a standard Hammerstein model. Moreover, (Rosa et al. 2010a, b) performed a comparison between the normalized FR and Heuristic models, which revealed no significant differences. Considering the additional flexibility provided by the custom HRF in the linear Hammerstein model, which is estimated directly from the data as compared to the FR model, we speculate that the normalized linear Hammerstein model can explain a larger fraction of BOLD variance compared to the Heuristic model (Kilner et al. 2005). However, this remains to be investigated in a future study.
BOLD Signal Variance Explained by the Individual Frequency Bands
The comparison of the resting-state BOLD variance explained by the different frequency bands using EEG source space analysis and the linear Hammerstein model revealed significant contributions from all frequency bands, which are region specific (Fig. 4). Oscillations in the alpha band explained significant BOLD signal variance in visual-related areas. This finding agrees with previous studies in the literature which investigated the electrophysiology corelates of the BOLD signal during resting-state with eyes closed (de Munck et al. 2007; Laufs et al. 2003, 2006; Mantini et al. 2007), suggesting the important role of these regions in the generation of the alpha rhythm even during resting-state with eyes open. Furthermore, we also observed significant contributions from the beta band, which could be related to changes in the brain state associated with vigilance and alertness that occur during eyes open as compared to eyes closed (Falahpour et al. 2018; Chang et al. 2016). On the other hand, significant contributions from the delta and theta bands were detected in the primary motor cortex. We believe that it is less likely that activation in these areas is solely due to motion-related artifacts as proposed in (Jansen et al. 2012), since (i) we have employed stringent methods to remove motion-related artefacts from both the EEG and fMRI data, and (ii) the neuronally plausible patterns of activation predicted by motion-related EEG artifacts shown in (Jansen et al. 2012) do not include the primary motor cortices.
During the handgrip task, the source EEG beta frequency band was found to significantly contribute to the BOLD signal in the ipsilateral primary somatosensory and motor cortices (Fig. S4c). This finding is in agreement with previous similar studies that showed strong correlations between beta EEG oscillations and BOLD-fMRI in the same brain regions during motor tasks (Ohara et al. 2001; Ritter et al. 2009; Sclocco et al. 2014). Also, our results suggest significant contributions from the beta band in the occipital and the superior parietal lobule cortices, which become activated in response to the visual feedback that the subjects received during task execution. The superior parietal lobule is a polymodal association area integrating motor, somesthetic and visual information.
Influence of Individual EEG Bands on HRF Scaling
The average maps of the total HRF peak obtained during the resting-state condition across subjects (Fig. 6) show that the hemodynamic response in the occipital, parietal and frontal cortices is mainly negative. In accordance with the results of previous studies (de Munck et al. 2007; Goldman et al. 2002; Laufs et al. 2003, 2006; Moosmann et al. 2003) these negative hemodynamic responses were found to be associated with instantaneous increases in the alpha band (Fig. 7). Occipital BOLD deactivation was discussed in (Goldman et al. 2002) as a result of alpha synchronization and idling. It has been also linked to changes in vigilance (Moosmann et al. 2003). In addition to contributions of the alpha band in negative resting-state BOLD responses, we also observed contributions of the beta frequency band (Fig. 7). Specifically, our results revealed negative responses in almost all regions spanning the cerebral cortex for both the alpha and beta frequency bands in agreement with a previous study by (Mantini et al. 2007), which showed negative correlations between the power profile of these bands and the BOLD signal in the default mode, dorsal attention, visual, motor and auditory networks.
During the motor task, our results revealed large negative HRF peak values in the lateral occipital and superior parietal lobule cortices for the alpha and beta bands (Figs. S4, S7). Negative HRF peak values were also observed areas within the left primary motor and somatosensory cortices for the beta band. These findings are consistent with desynchronization in the alpha and beta bands observed in young adults during a handgrip task using MEG (van Wijk et al. 2012; Xifra-Porxas et al. 2019) and EEG (Erbil and Ungan, 2007). Alpha and beta band desynchronizations are associated with decreases in the instantaneous EEG power and increases in the BOLD signal, which result in negative hemodynamic responses.
Areas in the somatosensory and motor cortices exhibited large positive values during both the resting-state condition (Fig. 6) and the motor task (Fig. S6). Similar patterns were also observed in the average HRF peak maps obtained for the delta and theta frequency bands for both experimental conditions (Figs. 7, S7). These findings suggest that the positive hemodynamic responses observed in these areas are more strongly associated with activity in lower frequency bands. Moreover, during resting conditions, the medial occipital cortex exhibited positive responses for the delta and theta bands, while the alpha and beta bands exhibited negative responses. During the motor task, the same area exhibited a positive HRF peak values in all frequency bands, with the strongest responses being observed for the delta and beta bands. Interestingly, this increase in the HRF peak values observed for the beta band in the medial occipital cortex between the task versus the resting condition is in agreement with the Heuristic model (Kilner et al. 2005), which states that the shifts in the EEG spectral profile towards higher frequencies that observed during neuronal activation are related to increases in the rate of energy dissipation and the BOLD-fMRI signal. However, it should be also noted that since the medial occipital cortex is more densely vascularized compared to other brain regions (Bernier et al. 2018), it is plausible that the large BOLD signal increases observed in this region during the task are related to artificial BOLD signal amplification resulting from its dense vascular network and proximity to large draining veins compared to other brain regions.
Limitations
The present study set out to investigate the link between changes in the level of neuronal activity as these manifests in narrow frequency bands of the LFP spectrum with the corresponding changes in the BOLD signal, using simultaneous EEG-fMRI. The cohort size (n = 12) used to achieve this was relatively small compared to recent EEG-fMRI studies in the literature. However, as the main purpose of this work was to demonstrate a method for describing the relationship between EEG and BOLD-fMRI, the available data are sufficient. Furthermore, a large body of animal studies has pointed to the gamma band (30–80 Hz) exhibiting the highest correlations with fluctuations in the BOLD signal (Goense and Logothetis 2008; Logothetis et al. 2001; Magri et al. 2012; Shmuel and Leopold 2008; Scheeringa et al. 2011; Uji et al. 2018). In the present study, however, the gamma band was excluded from the analysis, as we were not able to sufficiently remove MRI-related artifacts, such as RF gradient, ballisto-cardiogram, and helium pump artifacts within this frequency band. Of note, the proposed method for modeling the dynamic relation between source EEG and BOLD-fMRI presented herein can be readily applied to any number of EEG frequency bands, including the gamma band, in cases where the non-neural related artefacts can be successfully removed. Future work performed using gradient-free multimodal imaging techniques, such as simultaneous EEG-FNIRS would help overcome these limitations, but at the cost of a reduced spatial resolution.
In the present study we employed source space reconstruction to investigate the dynamic interactions between different frequency bands of individual current sources and BOLD-fMRI. Source space reconstruction was performed using linearly constrained minimum variance beamformers. Our results (Figs. S4, 4) suggested that source space analysis improved BOLD signal prediction for both task-based and resting-state experimental conditions. They also suggested that under each condition, different frequency bands may explain more BOLD signal variance relative to others depending on brain region. However, we note that HRF estimation using EEG-fMRI may be affected by the localization error of the underlying neuronal activation associated with each individual modality. However, evaluation of this error is challenging and remains an open question for future investigation. Furthermore, the EEG bands might be localized with different errors since different EEG sensors might be affected in a different way from various sources of noise characterized by distinct frequency content. For example, it is well known that eyeblink and BCG artefacts mainly affect frontal sensors (Marino et al. 2018), whereas muscle artifacts affect more temporal sensors (Muthukumaraswamy 2013). In this study, although gradient and BCG artefact removal was performed on a channel-by-channel basis, it is likely that the levels of noise that remained after preprocessing might be different for each sensor, which might result in different localization error for each band.
Lastly, accurate estimation of the HRF shape requires matching the power-spectrum of the experimental task to the power spectrum of the HRF (Wager and Nichols 2003). During the resting-state, which was the main condition of interest in this work, it is reasonable to assume that that the underlying spontaneous neuronal oscillations excited most of the frequencies within the HRF bandwidth (< 0.2 Hz). During the motor task, on the other hand, the power spectrum of the experimental task was restricted in the 0.05–0.08 Hz range, which may have affected the HRF shape estimates to some extent.
Conclusion
We developed a novel methodological approach using linear and nonlinear block-structured models to investigate the dynamic interactions between distributed dipolar current sources and changes in BOLD-fMRI signal evaluated using simultaneous EEG-fMRI. We applied the proposed method to data collected during resting-state conditions with eyes open, as well as data collected during a handgrip task. Our results suggest that these interactions can be sufficiently described using a linear Hammerstein model, which describes the BOLD signal as the convolution between a linear combination of the power profile of individual frequency bands with a data-driven HRF. Using this model, we rigorously investigated the regional variability of the HRF during both experimental conditions. Our results reveal that the regional characteristics of the HRF depend on both brain region, as well as on specific frequency bands under each experimental condition. Moreover, during the motor task, the proposed method was shown to yield similar results to those obtained when using the subjects’ hand grip force. This suggests the potential of the proposed method for obtaining reliable BOLD predictions and HRF estimates even from resting-state data, where there is no explicit task and SNR is lower. The proposed method can be readily applied to studying resting-state functional connectivity, as accurate resting-state HRF estimates are important for removing the hemodynamic blurring, which is inherent in the fMRI data.
Data Availability
The data collected for the purpose of this work are currently being curated and will be made available upon reasonable request (pending approval from our funding agencies and research institution).
Code Availability
The code that supports the findings of this study can be downloaded from the GitHub repository https://github.com/prokopis.
Notes
We considered the gamma density function given by \(\text{h}\left(\text{t};\uptau ,\upsigma \right)=\left\{\begin{array}{c}\text{exp}\left\{-\text{t}\sqrt{\upsigma \bullet\uptau } \right\}{\left(\frac{\text{e}\bullet\uptau }{\uptau }\right)}^{\sqrt{\uptau /\upsigma }}, \tau >0\\ 0, \tau <0\end{array}\right.\)where and control the location of the peak and width (dispersion), respectively (Hossein-Zadeh et al. 2003).
References
Abreu R, Leal A, Figueiredo P (2018) EEG-informed fMRI: a review of data analysis methods. Front Hum Neurosci 12:1–23. https://doi.org/10.3389/fnhum.2018.00029
Allen PJ, Polizzi G, Krakow K, Fish DR, Lemieux L (1998) Identification of EEG events in the MR scanner: the problem of pulse artifact and a method for its subtraction. Neuroimage 8:229–239. https://doi.org/10.1006/nimg.1998.0361
Allen PJ, Josephs O, Turner R (2000) A method for removing imaging artifact from continuous EEG recorded during functional MRI. Neuroimage 12:230–239. https://doi.org/10.1006/nimg.2000.0599
Bagshaw AP, Hawco C, Bénar CG, Kobayashi E, Aghakhani Y, Dubeau F, Pike GB, Gotman J (2005) Analysis of the EEG-fMRI response to prolonged bursts of interictal epileptiform activity. Neuroimage 24:1099–1112. https://doi.org/10.1016/j.neuroimage.2004.10.010
Beckmann CF, Smith SM (2004) Probabilistic independent component analysis for functional magnetic resonance imaging. IEEE Trans Med Imaging 23:137–152. https://doi.org/10.1109/TMI.2003.822821
Bénar CG, Gross DW, Wang Y, Petre V, Pike B, Dubeau F, Gotman J (2002) The BOLD response to interictal epileptiform discharges. Neuroimage 17:1182–1192. https://doi.org/10.1006/nimg.2002.1164
Bénar CG, Schön D, Grimault S, Nazarian B, Burle B, Roth M, Badier JM, Marquis P, Liegeois-Chauvel C, Anton JL (2007) Single-trial analysis of oddball event-related potentials in simultaneous EEG-fMRI. Hum Brain Mapp 28:602–613. https://doi.org/10.1002/hbm.20289
Bernier M, Cunnane SC, Whittingstall K (2018) The morphology of the human cerebrovascular system. Hum Brain Mapp 39:4962–4975. https://doi.org/10.1002/hbm.24337
Bridwell DA, Wu L, Eichele T, Calhoun VD (2013) The spatiospectral characterization of brain networks: fusing concurrent EEG spectra and fMRI maps. Neuroimage 69:101–111. https://doi.org/10.1016/j.neuroimage.2012.12.024
Bruns A (2004) Fourier-, Hilbert-and wavelet-based signal analysis: are they really different approaches? J Neurosci Methods 137:321–332
Buxton RB, Uludaǧ K, Dubowitz DJ, Liu TT (2004) Modeling the hemodynamic response to brain activation. Neuroimage 23:220–233. https://doi.org/10.1016/j.neuroimage.2004.07.013
Calhoun VD, Adali T, Pearlson GD, Kiehl KA (2006) Neuronal chronometry of target detection: fusion of hemodynamic and event-related potential data. Neuroimage 30(2):544–553
Chang C, Leopold DA, Schölvinck ML, Mandelkow H, Picchioni D, Liu X, Frank QY, Turchi JN, Duyn JH (2016) Tracking brain arousal fluctuations with fMRI. Proc Natl Acad Sci USA 113:4518–4523
Chen T, Ohlsson H, Ljung L (2012) On the estimation of transfer functions, regularizations and Gaussian processes—revisited. Automatica 48:1525–1535
de Munck JC, Gonçalves SI, Huijboom L, Kuijer JPA, Pouwels PJW, Heethaar RM, Lopes da Silva FH, Goncalves SI, Huijboom L, Kuijer JPA, Pouwels PJW, Heethaar RM, Lopes da Silva FH, Gonçalves SI, Huijboom L, Kuijer JPA, Pouwels PJW, Heethaar RM, Lopes da Silva FH (2007) The hemodynamic response of the alpha rhythm: an EEG/fMRI study. Neuroimage 35:1142–1151. https://doi.org/10.1016/j.neuroimage.2007.01.022
de Munck JC, Gonçalves SI, Mammoliti R, Heethaar RM, Lopes da Silva FH (2009) Interactions between different EEG frequency bands and their effect on alpha-fMRI correlations. Neuroimage 47:69–76. https://doi.org/10.1016/j.neuroimage.2009.04.029
Delorme A, Makeig S (2004) EEGLAB: an open source toolbox for analysis of single-trial EEG dynamics including independent component analysis. J Neurosci Methods 134:9–21. https://doi.org/10.1016/j.jneumeth.2003.10.009
Dickie EW, Anticevic A, Smith DE, Coalson TS, Manogaran M, Calarco N, Viviano JD, Glasser MF, Van Essen DC, Voineskos AN (2019) Ciftify: a framework for surface-based analysis of legacy MR acquisitions. Neuroimage 197:818–826. https://doi.org/10.1016/j.neuroimage.2019.04.078
die Jong S (1993) SIMPLS: an alternative approach squares regression to partial least. Chemom Intell Lab Syst 18:251–263. https://doi.org/10.1016/0169-7439(93)85002-X
Ebisch B, Schmidt KE, Niessing M, Singer W, Galuske RAW, Niessing J (2005) Hemodynamic signals correlate tightly with synchronized gamma oscillations. Science 309:948–951
Erbil N, Ungan P (2007) Changes in the alpha and beta amplitudes of the central EEG during the onset, continuation, and offset of long-duration repetitive hand movements. Brain Res 1169:44–56. https://doi.org/10.1016/J.BRAINRES.2007.07.014
Falahpour M, Chang C, Wong CW, Liu TT (2018) Template-based prediction of vigilance fluctuations in resting-state fMRI. Neuroimage 174:317–327
Fischl B (2012) FreeSurfer. Neuroimage. https://doi.org/10.1016/j.neuroimage.2012.01.021
Friston KJ, Fletcher P, Josephs O, Holmes ANDREW, Rugg MD, Turner R (1998) Event-related fMRI: characterizing differential responses. Neuroimage 7(1):30–40
Friston KJ, Mechelli A, Turner R, Price CJ (2000) Nonlinear responses in fMRI: the balloon model, Volterra kernels, and other hemodynamics. Neuroimage 12:466–477. https://doi.org/10.1006/nimg.2000.0630
Fuglø D, Pedersen H, Rostrup E, Hansen AE, Larsson HBW (2012) Correlation between single-trial visual evoked potentials and the blood oxygenation level dependent response in simultaneously recorded electroencephalography-functional magnetic resonance imaging. Magn Reson Med 68:252–260. https://doi.org/10.1002/mrm.23227
Goense JBM, Logothetis NK (2008) Neurophysiology of the BOLD fMRI signal in awake monkeys. Curr Biol 18:631–640. https://doi.org/10.1016/j.cub.2008.03.054
Goldman RI, Stern JM, Engel J, Cohen MS (2002) Simultaneous EEG and fMRI of the alpha rhythm. NeuroReport 13:2487–2492. https://doi.org/10.1097/01.wnr.0000047685.08940.d0
Gómez JC, Baeyens E (2004) Identification of block-oriented nonlinear systems using orthonormal bases. J Process Control 14:685–697. https://doi.org/10.1016/j.jprocont.2003.09.010
Gramfort A, Papadopoulo T, Olivi E, Clerc M (2009) OpenMEEG: opensource software for quasistatic bioelectromagnetics. Biomed Eng Online 8:1. https://doi.org/10.1186/1475-925X-8-1
Heuberger PSC, Van den Hof PMJJ, Wahlberg B (2005) Modeling and identification with rational orthogonal basis functions. Springer, Berlin
Hossein-Zadeh GA, Ardekani BA, Soltanian-Zadeh H (2003) A signal subspace approach for modeling the hemodynamic response function in fMRI. Magn Reson Imaging 21:835–843. https://doi.org/10.1016/S0730-725X(03)00180-2
Jansen M, White TP, Mullinger KJ, Liddle EB, Gowland PA, Francis ST, Bowtell R, Liddle PF (2012) Motion-related artefacts in EEG predict neuronally plausible patterns of activation in fMRI data. Neuroimage 59:261–270. https://doi.org/10.1016/J.NEUROIMAGE.2011.06.094
Jenkinson M, Beckmann CF, Behrens TEJ, Woolrich MW, Smith SM (2012) Fsl. Neuroimage 62:782–790. https://doi.org/10.1016/j.neuroimage.2011.09.015
Jorge J, Van der Zwaag W, Figueiredo P (2014) EEG-fMRI integration for the study of human brain function. Neuroimage 102:24–34. https://doi.org/10.1016/j.neuroimage.2013.05.114
Kilner JM, Mattout J, Henson R, Friston KJ (2005) Hemodynamic correlates of EEG: a heuristic. Neuroimage 28:280–286. https://doi.org/10.1016/j.neuroimage.2005.06.008
Klein A, Tourville J (2012) 101 labeled brain images and a consistent human cortical labeling protocol. Front Neurosci 6:171. https://doi.org/10.3389/fnins.2012.00171
Laufs H, Kleinschmidt A, Beyerle A, Eger E, Salek-Haddadi A, Preibisch C, Krakow K (2003) EEG-correlated fMRI of human alpha activity. Neuroimage 19:1463–1476. https://doi.org/10.1016/S1053-8119(03)00286-6
Laufs H, Holt JL, Elfont R, Krams M, Paul JS, Krakow K, Kleinschmidt A (2006) Where the BOLD signal goes when alpha EEG leaves. Neuroimage 31:1408–1418. https://doi.org/10.1016/j.neuroimage.2006.02.002
Leistedt B, McEwen JD (2012) Exact wavelets on the ball. IEEE Trans Signal Process 60:6257–6269. https://doi.org/10.1109/TSP.2012.2215030
Leite M, Leal A, Figueiredo P (2013) Transfer function between EEG and BOLD signals of epileptic activity. Front Neurol. https://doi.org/10.3389/fneur.2013.00001
Le Van Quyen M, Foucher J, Lachaux JP, Rodriguez E, Lutz A, Martinerie J, Varela FJ (2001) Comparison of Hilbert transform and wavelet methods for the analysis of neuronal synchrony. J Neurosci Methods 111:83–98
Logothetis NK, Pauls J, Augath M, Trinath T, Oeltermann A (2001) Neurophysiological investigation of the basis of the fMRI signal. Nature 412:150–157. https://doi.org/10.1038/35084005
Lu Y, Bagshaw AP, Grova C, Kobayashi E, Dubeau F, Gotman J (2006) Using voxel-specific hemodynamic response function in EEG-fMRI data analysis. Neuroimage 32:238–247
Magri C, Schridde U, Murayama Y, Panzeri S, Logothetis NK (2012) The amplitude and timing of the BOLD signal reflects the relationship between local field potential power at different frequencies. J NeuroSci 32:1395–1407. https://doi.org/10.1523/JNEUROSCI.3985-11.2012
Mantini D, Perrucci MG, Del Gratta C, Romani GL, Corbetta M (2007) Electrophysiological signatures of resting state networks in the human brain. Proc Natl Acad Sci USA 104:13170–13175. https://doi.org/10.1073/pnas.0700668104
Marecek R, Lamos M, Mikl M, Barton M, Fajkus J, Rektor I, Brazdil M (2016) What can be found in scalp EEG spectrum beyond common frequency bands. EEG-fMRI study. J Neural Eng. https://doi.org/10.1088/1741-2560/13/4/046026
Marino M, Liu Q, Koudelka V, Porcaro C, Hlinka J, Wenderoth N, Mantini D (2018) Adaptive optimal basis set for BCG artifact removal in simultaneous EEG-fMRI. Sci Rep 8:1–11. https://doi.org/10.1038/s41598-018-27187-6
Marmarelis VZ (1993) Identification of nonlinear biological systems using Laguerre expansions of kernels. Ann Biomed Eng 21:573–589
Marmarelis VZ (2004) Nonlinear dynamic modeling of physiological systems. Wiley, New York. https://doi.org/10.1002/9780471679370
Moosmann M, Ritter P, Krastel I, Brink A, Thees S, Blankenburg F, Taskin B, Obrig H, Villringer A (2003) Correlates of alpha rhythm in functional magnetic resonance imaging and near infrared spectroscopy. Neuroimage 20:145–158. https://doi.org/10.1016/S1053-8119(03)00344-6
Mullinger KJ, Morgan PS, Bowtell RW (2008) Improved artifact correction for combined electroencephalography/functional MRI by means of synchronization and use of vectorcardiogram recordings. J Magn Reson Imaging 27:607–616. https://doi.org/10.1002/jmri.21277
Mullinger KJ, Yan WX, Bowtell R (2011) Reducing the gradient artefact in simultaneous EEG-fMRI by adjusting the subject’s axial position. Neuroimage 54:1942–1950. https://doi.org/10.1016/j.neuroimage.2010.09.079
Mullinger KJ, Chowdhury MEH, Bowtell R (2014) Investigating the effect of modifying the EEG cap lead configuration on the gradient artifact in simultaneous EEG-fMRI. Front Neurosci 8:1–10. https://doi.org/10.3389/fnins.2014.00226
Murta T, Leite M, Carmichael DW, Figueiredo P, Lemieux L (2015) Electrophysiological correlates of the BOLD signal for EEG-informed fMRI. Hum Brain Mapp 36:391–414. https://doi.org/10.1002/hbm.22623
Murta T, Hu L, Tierney TM, Chaudhary UJ, Walker MC, Carmichael DW, Figueiredo P, Lemieux L (2016) A study of the electro-haemodynamic coupling using simultaneously acquired intracranial EEG and fMRI data in humans. Neuroimage 142:371–380. https://doi.org/10.1016/j.neuroimage.2016.08.001
Muthukumaraswamy SD (2013) High-frequency brain activity and muscle artifacts in MEG/EEG: a review and recommendations. Front Hum Neurosci 7:138. https://doi.org/10.3389/fnhum.2013.00138
Nguyen VT, Cunnington R (2014) The superior temporal sulcus and the N170 during face processing: single trial analysis of concurrent EEG-fMRI. Neuroimage 86:492–502. https://doi.org/10.1016/j.neuroimage.2013.10.047
Ogawa S, Lee TM, Kay AR, Tank DW (1990) Brain magnetic resonance imaging with contrast dependent on blood oxygenation. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.87.24.9868
Ohara S, Mima T, Baba K, Ikeda A, Kunieda T, Matsumoto R, Yamamoto J, Matsuhashi M, Nagamine T, Hirasawa K, Hori T, Mihara T, Hashimoto N, Salenius S, Shibasaki H (2001) Increased synchronization of cortical oscillatory activities between human supplementary motor and primary sensorimotor areas during voluntary movements. J Neurosci 21:9377–9386. https://doi.org/10.1523/JNEUROSCI.21-23-09377.2001
Oldfield RC (1971) The assessment and analysis of handedness. Neuropsychologia 9:97–113
Portnova GV, Tetereva A, Balaev V, Atanov M, Skiteva L, Ushakov V, Ivanitsky A, Martynova O (2018) Correlation of BOLD signal with linear and nonlinear patterns of EEG in resting state EEG-informed fMRI. Front Hum Neurosci 11:1–12. https://doi.org/10.3389/fnhum.2017.00654
Rangaprakash D, Wu G-R, Marinazzo D, Hu X, Deshpande G (2018) Hemodynamic response function (HRF) variability confounds resting-state fMRI functional connectivity. Magn Reson Med 80:1697–1713. https://doi.org/10.1002/mrm.27146
Ritter P, Moosmann M, Villringer A (2009) Rolandic alpha and beta EEG rhythms’ strengths are inversely related to fMRI-BOLD signal in primary somatosensory and motor cortex. Hum Brain Mapp 30:1168–1187. https://doi.org/10.1002/hbm.20585
Rosa MJ, Daunizeau J, Friston KJ (2010a) EEG-fMRI integration: a critical review of biophysical modeling and data analysis approaches. J Integr Neurosci 9:453–476. https://doi.org/10.1142/S0219635210002512
Rosa MJ, Kilner J, Blankenburg F, Josephs O, Penny W (2010b) Estimating the transfer function from neuronal activity to BOLD using simultaneous EEG-fMRI. Neuroimage 49:1496–1509. https://doi.org/10.1016/j.neuroimage.2009.09.011
Rosa MJ, Kilner JM, Penny WD (2011) Bayesian comparison of neurovascular coupling models using EEG-fMRI. PLoS Comput Biol 7:e1002070. https://doi.org/10.1371/journal.pcbi.1002070
Rospiral R, Kramer N (2006) Overview and recent advances in partial least squares 43–51. doi: https://doi.org/10.1075/aals.6.03ch3
Ryali S, Glover GH, Chang C, Menon V (2009) Development, validation, and comparison of ICA-based gradient artifact reduction algorithms for simultaneous EEG-spiral in/out and echo-planar fMRI recordings. Neuroimage 48:348–361. https://doi.org/10.1016/j.neuroimage.2009.06.072
Sato JR, Rondinoni C, Sturzbecher M, de Araujo DB, Amaro E Jr (2010) From EEG to BOLD: brain mapping and estimating transfer functions in simultaneous EEG-fMRI acquisitions. Neuroimage 5:1416–1426
Scheeringa R, Bastiaansen MCM, Petersson KM, Oostenveld R, Norris DG, Hagoort P (2008) Frontal theta EEG activity correlates negatively with the default mode network in resting state. Int J Psychophysiol 67:242–251. https://doi.org/10.1016/j.ijpsycho.2007.05.017
Scheeringa R, Fries P, Petersson KM, Oostenveld R, Grothe I, Norris DG, Hagoort P, Bastiaansen MCM (2011) Neuronal dynamics underlying high- and low-frequency EEG oscillations contribute independently to the human BOLD signal. Neuron 69:572–583. https://doi.org/10.1016/j.neuron.2010.11.044
Scheeringa R, Koopmans PJ, van Mourik T, Jensen O, Norris DG (2016) The relationship between oscillatory EEG activity and the laminar-specific BOLD signal. Proc Natl Acad Sci USA 113:6761–6766. https://doi.org/10.1073/pnas.1522577113
Sclocco R, Tana MG, Visani E, Gilioli I, Panzica F, Franceschetti S, Cerutti S, Bianchi AM (2014) EEG-informed fMRI analysis during a hand grip task: estimating the relationship between EEG rhythms and the BOLD signal. Front Hum Neurosci 8:186. https://doi.org/10.3389/fnhum.2014.00186
Shmuel A, Leopold DA (2008) Neuronal correlates of spontaneous fluctuations in fMRI signals in monkey visual cortex: implications for functional connectivity at rest. Hum Brain Mapp 29:751–761. https://doi.org/10.1002/hbm.20580
Tadel F, Baillet S, Mosher JC, Pantazis D, Leahy RM (2011) Brainstorm: a user-friendly application for MEG/EEG analysis. Comput Intell Neurosci. https://doi.org/10.1155/2011/879716
Thornton RC, Rodionov R, Laufs H, Vulliemoz S, Vaudano A, Carmichael D, Cannadathu S, Guye M, McEvoy A, Lhatoo S, Bartolomei F, Chauvel P, Diehl B, De Martino F, Elwes RDC, Walker MC, Duncan JS, Lemieux L (2010) Imaging haemodynamic changes related to seizures: comparison of EEG-based general linear model, independent component analysis of fMRI and intracranial EEG. Neuroimage 53:196–205. https://doi.org/10.1016/j.neuroimage.2010.05.064
Tyvaert L, LeVan P, Grova C, Dubeau F, Gotman J (2008) Effects of fluctuating physiological rhythms during prolonged EEG-fMRI studies. Clin Neurophysiol 119:2762–2774. https://doi.org/10.1016/j.clinph.2008.07.284
Uji M, Wilson R, Francis ST, Mullinger KJ, Mayhew SD (2018) Exploring the advantages of multiband fMRI with simultaneous EEG to investigate coupling between gamma frequency neural activity and the BOLD response in humans. Hum Brain Mapp 39:1673–1687
Van Veen BD, Van Drongelen W, Yuchtman M, Suzuki A (1997) Localization of brain electrical activity via linearly constrained minimum variance spatial filtering. IEEE Trans Biomed Eng 44:867–880. https://doi.org/10.1109/10.623056
van Wijk BCM, Beek PJ, Daffertshofer A (2012) Differential modulations of ipsilateral and contralateral beta (de)synchronization during unimanual force production. Eur J Neurosci 36:2088–2097. https://doi.org/10.1111/j.1460-9568.2012.08122.x
Wager TD, Nichols TE (2003) Optimization of experimental design in fMRI: a general framework using a genetic algorithm. Neuroimage 182:293–309. https://doi.org/10.1016/S1053-8119(02)00046-0
Wan X, Riera J, Iwata K, Takahashi M, Wakabayashi T, Kawashima R (2006) The neural basis of the hemodynamic response nonlinearity in human primary visual cortex: implications for neurovascular coupling mechanism. Neuroimage 32:616–625. https://doi.org/10.1016/j.neuroimage.2006.03.040
Westwick DT, Kearney RE (2003) Identification of nonlinear physiological systems. Wiley, New York
Wirsich J, Bénar C, Ranjeva JP, Descoins M, Soulier E, Le Troter A, Confort-Gouny S, Liégeois-Chauvel C, Guye M (2014) Single-trial EEG-informed fMRI reveals spatial dependency of BOLD signal on early and late IC-ERP amplitudes during face recognition. Neuroimage 100:325–336. https://doi.org/10.1016/j.neuroimage.2014.05.075
Wu G-R, Liao W, Stramaglia S, Ding J-R, Chen H, Marinazzo D (2013) A blind deconvolution approach to recover effective connectivity brain networks from resting state fMRI data. Med Image Anal 17:365–374. https://doi.org/10.1016/j.media.2013.01.003
Xifra-Porxas A, Niso G, Larivière S, Kassinopoulos M, Baillet S, Mitsis GD, Boudrias M-H (2019) Older adults exhibit a more pronounced modulation of beta oscillations when performing sustained and dynamic handgrips. Neuroimage 201:116037. https://doi.org/10.1016/J.NEUROIMAGE.2019.116037
Funding
This work was supported by funds from the Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grants RGPIN-2019-06638 [GDM], and RGPIN-2017-05270 [MHB], the Fonds de la Recherche du Quebec—Nature et Technologies (FRQNT) Team Grant 254680-2018 [GDM], the Canadian Foundation for Innovation grant numbers 34362 [GDM] and 34277 [MHB]. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
PCP: Conceptualization, Investigation, Data curation, Methodology, Software, Formal analysis, Visualization, Writing—original draft. AX-P: Investigation, Data curation, Methodology, Formal analysis, Writing—reviewing and editing. MK: Investigation, Data curation, Formal analysis, Writing—reviewing and editing. M-HB: Conceptualization, Methodology, Resources, Writing—reviewing and editing, Funding acquisition. GDM: Conceptualization, Methodology, Resources, Writing—reviewing and editing, Project administration, Funding acquisition, Supervision.
Corresponding author
Ethics declarations
Conflict of interest
Not applicable.
Ethical Approval
This study was performed in accordance with the McGill University Ethical Advisory Committee.
Consent to Participate
Informed consent was obtained from all individual participants included in the study.
Additional information
Handling Editor: Louis Lemieux.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendices
Appendix
Identification of Hammerstein and Wiener-Hammerstein Models
The Hammerstein model (Fig. 1a; Eq. (1)) can be estimated efficiently from the input–output data using orthonormal basis functions for the representation of the LTI block (Gómez and Baeyens 2004), which is given by
where \(\left\{{\text{B}}_{\text{j}}\left(\text{n}\right);\text{j}=0,\dots ,\text{ L}-1;\text{n}=0,\dots ,\text{M}\right\}\) is a set of \(\text{L}\) orthonormal basis functions, and \({\text{b}}_{\text{j}}\) is the unknown expansion coefficient of the \(\text{j}\)-th order basis function. The use of orthonormal bases reduces the number of required free parameters in the model and allows parameter estimation using least-squares regression. This leads to increased estimation accuracy in the presence of noise even from short experimental data-records. Combining Eqs. (1, 2) and (6), the input–output relationship can be written as
where \({\text{v}}_{\text{i},\text{j}}^{\left(\text{p}\right)}\left(\text{n}\right)\) denotes the convolution between the \(\text{p}\)-th polynomial power of the \(\text{i}\)-th input with the \(\text{j}\)-th basis function. Equation (7) can be re-expressed as a linear regression problem
where \(\mathbf{V}=\left[{\text{v}}_{\text{1,0}}^{\left(1\right)},\dots , {\text{v}}_{\text{Q},0}^{\left(1\right)}, \dots ,{\text{v}}_{1,\text{L}-1}^{\left(1\right)},\dots ,{\text{v}}_{\text{Q},\text{L}-1}^{\left(1\right)} ,\dots \dots , {\text{v}}_{\text{1,0}}^{\left(\text{P}\right)},\dots , {\text{v}}_{\text{Q},0}^{\left(\text{P}\right)}, \dots ,{\text{v}}_{1,\text{L}-1}^{\left(\text{P}\right)},\dots ,{\text{v}}_{\text{Q},\text{L}-1}^{\left(\text{P}\right)}\right]\), and \({\mathbf{c}} = \left[ {{\text{a}}_{{1,1}} {\text{b}}_{0} , \ldots ,{\text{a}}_{{1,1}} {\text{b}}_{{{\text{L}} - 1}} , \ldots ,{\text{a}}_{{{\text{Q}},1}} {\text{b}}_{0} , \ldots ,{\text{a}}_{{{\text{Q}},1}} {\text{b}}_{{{\text{L}} - 1}} , \ldots \ldots ,{\text{a}}_{{1,{\text{P}}}} {\text{b}}_{0} , \ldots ,{\text{a}}_{{1,{\text{P}}}} {\text{b}}_{{{\text{L}} - 1}} , \ldots ,{\text{a}}_{{{\text{Q,P}}}} {\text{b}}_{0} , \ldots ,{\text{a}}_{{{\text{Q,P}}}} {\text{b}}_{{{\text{L}} - 1}} } \right]^{{\text{T}}}\) is a vector of the unknown model parameters.
Power fluctuations within distinct EEG frequency bands are highly correlated as previously reported in the literature (de Munck et al. 2009). As a result, the columns in \(\mathbf{V}\) are strongly collinear, which makes estimation of \(\mathbf{c}\) using ordinary least-squares numerically unstable due to ill-conditioning of the Gram matrix \(\left[{\mathbf{V}}^{\mathbf{T}}\mathbf{V}\right]\). Therefore, to obtain a numerically more stable estimate of the unknown parameter vector \(\mathbf{c}\), we employed partial least-squares regression (PLSR) (Rospiral and Kramer 2006).
PLSR is performed in three phases. In phase 1, the algorithm finds projections of \(\mathbf{V}\) and \(\mathbf{y}\) to a new co-ordinate system such that the covariance of these projections is maximized. This is achieved using a linear decomposition of both \(\mathbf{V}\) and \(\mathbf{y}\) into a set of orthonormal latent variables (scores) and loadings given by
where \(\mathbf{T}\) and \(\mathbf{U}\) are matrices of latent variables associated with \(\mathbf{V}\) and \(\mathbf{y}\), respectively. \(\mathbf{P}\) and \(\mathbf{W}\) are the corresponding loadings for each latent variable matrix, and \({\mathbf{e}}_{1,2}\) are error terms. The decomposition of \(\mathbf{V}\) and \(\mathbf{y}\) is performed such that the covariance between \(\mathbf{T}\) and \(\mathbf{U}\) is maximized. In phase 2, the algorithm performs ordinary least-squares regression analysis between the latent variables \(\mathbf{T}\) and system output \(\mathbf{y}\)
where \({\varvec{\uptheta}}={\left[{\uptheta }_{1},\dots , {\uptheta }_{\text{Q}\times \text{L}\times \text{P}}\right]}^{\text{T}}\) is a vector of the regression coefficients. Note that in this case the Gram matrix \(\left[{\mathbf{T}}^{\mathbf{T}}\mathbf{T}\right]\) is well-conditioned since the columns in \(\mathbf{T}\) are orthonormal. In phase 3, the estimated \({\widehat{{\varvec{\uptheta}}}}_{\mathbf{L}\mathbf{S}\mathbf{E}}\) coefficients are projected back to the original parameter space yielding unbiased estimates of the original model parameters \({\widehat{\mathbf{c}}}_{\mathbf{P}\mathbf{L}\mathbf{S}}\) (die Jong 1993).
To uniquely identify the unknown parameters of the Hammerstein model described by (1, 2) and (6, 7), the bilinear parameter vector \(\mathbf{c}\) needs to be dissociated into its constituent \({\text{a}}_{\text{i},\text{p}}\) and \({\text{b}}_{\text{j}}\) parameters. The parameter vector \(\mathbf{c}\) can be reshaped into a block-column matrix \({\mathbf{c}}_{\mathbf{a}\mathbf{b}}\), such that
where \(\mathbf{a}={\left[{\text{a}}_{\text{1,1}},\dots ,{\text{a}}_{\text{Q},1},{\text{a}}_{\text{1,2}},\dots \dots , {\text{a}}_{\text{Q},\text{P}-1}, {\text{a}}_{1,\text{P}},\dots ,{\text{a}}_{\text{Q},\text{P}}\right]}^{\text{T}}\) and \(\mathbf{b}={\left[{\text{b}}_{0},\dots ,{\text{b}}_{\text{L}-1}\right]}^{\text{T}}\). An optimal, in the least-squares sense, estimate of the model parameters \({\widehat{\mathbf{a}}}_{\mathbf{L}\mathbf{S}\mathbf{E}}\) and \({\widehat{\mathbf{b}}}_{\mathbf{L}\mathbf{S}\mathbf{E}}\) can be obtained solving the following constrained minimization problem
where \(\text{h}\left(\text{m}\right)\) is given by (6). Note that as a result of normalizing the polynomial coefficients \(\mathbf{a}\), the estimate \({\widehat{\mathbf{a}}}_{\mathbf{L}\mathbf{S}\mathbf{E}}\) reflects a relative rather than absolute contribution of individual EEG bands to the BOLD signal variance. A solution to (A9) is provided by the singular value decomposition (SVD) of matrix \({\mathbf{c}}_{\mathbf{a}\mathbf{b}}\) (Gómez and Baeyens 2004). Specifically,
where \({\mathbf{U}}_{1}\) is the first left singular vector, \({\mathbf{V}}_{1}\) the first right singular vector, and \({{\varvec{\Sigma}}}_{1}\in {\mathbb{R}}\) the first singular value of the SVD of \({\widehat{\mathbf{c}}}_{\text{ab}}\).
Hammerstein-Wiener Model Identification
The MISO Hammerstein-Wiener (HW) described by Eq. (3) can be re-expressed in a compact matrix form as
where \(\mathbf{F}\) denotes a matrix the columns of which are polynomial powers of \({\text{y}}_{\text{H}}\), and \(\mathbf{z}\) is a vector of the unknown polynomial coefficients, which can be estimated using ordinary least-squares
Orthonormal Basis Functions
There are several sets of orthonormal basis functions that can be used for modeling the impulse response function of the LTI block in the Hammerstein and Wiener-Hammerstein model configuration (Heuberger et al. 2005). The selection of the appropriate basis set depends on the dynamic behavior of the system to be modelled. One basis set that has been extensively used in the literature for modeling of physiological systems is the Laguerre basis. Laguerre basis functions exhibit exponentially decaying structure and constitute an orthonormal set in \(\left[0,\infty \right)\), which makes them suitable for modeling causal systems with finite memory (Marmarelis 1993).
In this work we employ a smoother variant of the Laguerre basis functions that is based on the spherical Laguerre basis functions (Leistedt and McEwen 2012), which allow obtaining robust HRF estimates in single voxels even during resting conditions where the signal-to-noise ratio (SNR) is particularly low. The \(\text{j}\)-th spherical Laguerre basis function \({\text{b}}_{\text{j}}\left(\text{n}\right); \text{j}=0,\dots ,\text{L}-1;\text{ n}=1,\dots ,\text{M}\) is given by
where \(\alpha\in {\mathbb{R}}_{+}\) is a parameter that determines the rate of exponential asymptotic decline of \({\text{b}}_{\text{j}}\left(\text{n}\right)\), and \({\text{K}}_{\text{j}}\left(\text{n}\right)\) is the \(\text{j}\)-th generalized Laguerre polynomial of order two, defined as
The \(\text{j}\)-th spherical Laguerre basis function \({\text{b}}_{\text{j}}\left(\text{n}\right)\) were convolved with a Gaussian kernel G(μ,τ), with μ > 0 and τ = 1. The μ > 0 parameter controls how late \({\text{b}}_{\text{j}}\left(\text{n}\right)\) will start to fluctuate. The range of this parameter was set to 2.5 < μ < 5 corresponding to HRF estimates with time-to-peak ranging between 3 and 10 s (Fig. S12). The orthonormal properties of the derived basis set was ensured by entering the result of this convolution into an orthogonalization process based on the Gram-Schmidt orthogonalization algorithm.
Rights and permissions
About this article
Cite this article
Prokopiou, P.C., Xifra-Porxas, A., Kassinopoulos, M. et al. Modeling the Hemodynamic Response Function Using EEG-fMRI Data During Eyes-Open Resting-State Conditions and Motor Task Execution. Brain Topogr 35, 302–321 (2022). https://doi.org/10.1007/s10548-022-00898-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10548-022-00898-w