Abstract
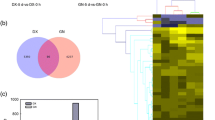
Elymus nutans is an important alpine perennial forage of the Pooideae subfamily, which can survive subzero temperatures. To understand the molecular mechanisms underlying cold tolerance in E. nutans, we performed the transcriptional analysis by RNA-Seq in two genotypes, the tolerant Damxung (DX) and the sensitive Gannan (GN), under cold stress. The new E. nutans transcriptomes comprised 200 520/200 836 and 181 331/211 973 transcripts in leaves/crowns of DX and GN, respectively. More cold-stress-related genes were identified in leaves than in crowns of both genotypes throughout the whole cold stress. The most prominent functional category in leaves of both genotypes at 3 h of stress was transcriptional regulation. Brassinosteroid and jasmonic acid mediated signalling pathways played central roles in regulating downstream protective responses in DX after 24 h of cold stress. Prolonged cold stress caused more severe transcriptome responses in crowns and leaves of DX compared to GN. The most significant transcriptomic changes in both genotypes were associated with the response to abiotic stresses and the oxidation-reduction processes, implying reprogramming of the cellular metabolism as an adaptation to cold stress. This study reveals mechanisms of genotype- and organ-specific cold stress response in E. nutans and thus provides a basis for future breeding strategies aimed at improving the tolerance of cold-sensitive plants.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- BR:
-
brassinosteroid
- CBF:
-
C-repeat binding factor
- COG:
-
orthologous groups
- COR:
-
cold-responsive genes
- DEGs:
-
differentially expressed genes
- DX:
-
Damxung
- FEH:
-
fructan exohydrolase
- GN:
-
Gannan
- GO:
-
gene ontology
- HSPs:
-
heat shock proteins
- JA:
-
jasmonic acid
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- nr:
-
non-redundant
- nt:
-
nucleotide sequence
- TFs:
-
transcription factors
- 1-SST:
-
sucrose: sucrose-1-fructosyltransferase
References
Abeynayake, S.W., Byrne, S., Nagy, I., Jonaviciene, K., Etzerodt, T.P., Boelt, B., Asp, T.: Changes in Lolium perenne transcriptome during cold acclimation in two genotypes adapted to different climatic conditions. - BMC Plant Biol. 15: 250, 2015.
Barrs, H.D., Weatherley, P.E.: A re-examination of the relative turgidity techniques for estimating water deficits in leaves. - Aust. J. biol. Sci. 15: 413–428, 1962.
Bates, L.S., Waldren, R.P., Teare, I.D.: Rapid determination of free proline for water-stress studies. - Plant Soil 39: 205–207, 1973.
Calzadilla, P.I., Maiale, S.J., Ruiz, O.A., Escaray, F.J.: Transcriptome response mediated by cold stress in Lotus japonicus. - Front. Plant Sci. 7: 374, 2016.
Chan, Z.L., Wang, Y.P., Cao, M.J., Gong, Y.H., Mu, Z.X., Wang, H.Q., Hu, Y.L., Deng, X., He, X.J., Zhu, J.K.: RDM4 modulates cold stress resistance in Arabidopsis partially through the CBF-mediated pathway. - New Phytol. 209: 1527–1539, 2015.
Chen, S.Y., Huang, X., Yan, X.Q., Liang, Y., Wang, Y.Z., Li, X.F., Peng, X.J., Ma, X.Y., Zhang, L.X., Cai, Y.Y., Ma, T., Cheng, L.Q., Qi, D.M., Zheng, H.J., Yang, X.H., Li, X.X., Liu, G.S.: Transcriptome analysis in sheepgrass (Leymus chinensis): a dominant perennial grass of the Eurasian steppe. - PLoS ONE 8: e67974, 2013.
Chinnusamy, V., Zhu, J., Zhu, J.K.: Cold stress regulation of gene expression in plants. - Trends Plant Sci. 12: 444–451, 2007.
Clauw, P., Coppens, F., De Beuf, K., Dhondt, S., Van Daele, T., Maleux, K., Storme, V., Clement, L., Gonzalez, N., Inzé, D.: Leaf responses to mild drought stress in natural variants of Arabidopsis. - Plant Physiol. 167: 800–816, 2015.
Cock, P.J., Fields, C.J., Goto, N., Heuer, M., Rice, P.M.: The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. - Nucl. Acids Res. 38: 1767–1771, 2010.
Dhindsa, R.S., Plumb-dhindsa, P., Thorpe, T.A.: Leaf senescence: correlated with increased levels of membrane permeability and lipid peroxidation and decreased levels of superoxide dismutase and catalase. - J. exp. Bot. 32: 93–101, 1981.
Diallo, A.O., Agharbaoui, Z., Badawi, M.A., Ali-Benali, M.A., Moheb, A., Houde, M., Sarhan, F.: Transcriptome analysis of an mvp mutant reveals important changes in global gene expression and a role for methyl jasmonate in vernalization and flowering in wheat. - J. exp. Bot. 65: 2271–2286, 2014.
Dubois, M., Skirycz, A., Claeys, H., Maleux, K., Dhondt, S., De Bodt, S., Van den Bossche, R., De Milde, L., Yoshizumi, T., Matsui, M., Inzé, D: ETHYLENE RESPONSE FACTOR 6 acts as a central regulator of leaf growth under water limiting conditions in Arabidopsis. - Plant Physiol. 162: 319–332, 2013.
Fernie, A.R., Carrari, F., Sweetlove, L.J.: Respiratory metabolism: glycolysis, the TCA cycle and mitochondrial electron transport. - Curr. Opin. Plant Biol. 7: 254–261, 2004.
Foyer, C.H., Noctor, G.: Redox regulation in photosynthetic organisms: signaling, acclimation and practical implications. - Antioxid. Redox Sign. 11: 862–905, 2009.
Fu, J.J., Miao, Y.J., Shao, L.H., Hu, T.M., Yang, P.Z.: De novo transcriptome sequencing and gene expression profiling of Elymus nutans under cold stress. - BMC Genomics 17: 870, 2016b.
Fu, J.J., Xu, Y.F., Miao, Y.J., Hu, T.M.: Altitude variation in chilling tolerance among natural populations of Elymus nutans in the Tibetan Plateau. - Acta Physiol. Plant. 38: 218, 2016a.
Gao, Q., Li, X., Jia, J., Zhao, P.C., Liu, P.P., Liu, Z.J., Ge, L., Chen, S.Y., Qi, D., Deng, B., Lee, B.H., Liu, G.S., Cheng, L.Q.: Overexpression of a novel cold-responsive transcript factor LcFIN1 from sheepgrass enhances tolerance to lowtemperature stress in transgenic plants. - Plant Biotechnol. J. 14: 861–874, 2016.
Gibson, S.I.: Sugar and phytohormone response pathways: navigating a signalling network. - J. exp. Bot. 55: 253–264, 2004.
Grabherr, M.G., Haas, B.J., Yassour, M., Levin, J.Z., Thompson, D.A., Amit, I., Adiconis, X., Fan, L., Raychowdhury, R., Zeng, Q.D., Chen, Z.H., Mauceli, E., Hacohen, N., Gnirke, A., Rhind, N., Di Palma, F., Birren, B.W., Nusbaum, C., Lindblad-Toh, K., Friedman, N.: Fulllength transcriptome assembly from RNA-Seq data without a reference genome.–Natur. Biotechnol. 29: 644–652, 2011.
Hu, Y., Jiang, L., Wang, F., Yu, D.: Jasmonate regulates the INDUCER OF CBF EXPRESSION-C-REPEAT BINDING FACTOR/DRE BINDING FACTOR1 cascade and freezing tolerance in Arabidopsis. - Plant Cell 25: 2907–2924, 2013.
Joshi, V., Joung, J.G., Fei, Z., Jander, G.: Interdependence of threonine, methionine and isoleucine metabolism in plants: accumulation and transcriptional regulation under abiotic stress. - Amino Acids 39: 933–947, 2010.
Keunen, E., Peshev, D., Vangronsveld, J., Van den Ende, W., Cuypers, A.: Plant sugars are crucial players in the oxidative challenge during abiotic stress: extending the traditional concept. - Plant Cell Environ. 36: 1242–1255, 2013.
Kwasniewski, M., Daszkowska-Golec, A., Janiak, A., Chwialkowska, K., Nowakowska, U., Sablok, G., Szarejkol, I.: Transcriptome analysis reveals the role of the root hairs as environmental sensors to maintain plant functions under water-deficiency conditions. - J. exp. Bot. 67: 1079–1094, 2016.
Leyva-Pérez, M.D., Valverde-Corredor, A., Valderrama, R., Jiménez-Ruiz, J., Muñoz-Merida, A., Trelles, O., Barroso, J.B., Mercado-Blanco, J., Luque, F.: Early and delayed long-term transcriptional changes and short-term transient responses during cold acclimation in olive leaves. - DNA Res. 22: 1–11, 2015.
Li, C., Rudi, H.D., Stockinger, E.J., Cheng, H.M., Cao, M.J., Fox, S.E., Mockler, T.C., Westereng, B., Fjellheim, S., Rognli, O., Sandve, S.R.: Comparative analyses reveal potential uses of Brachypodium distachyon as a model for cold stress responses in temperate grasses. - BMC Plant Biol. 12: 65, 2012.
Livak, K.J., Schmittgen, T.D.: Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. - Methods 25: 402–408, 2001.
Santner, A., Estelle, M.: Recent advances and emerging trends in plant hormone signalling. - Nature 459: 1071–1078, 2009.
Shi, H., Ye, T., Chen, F., Cheng, Z., Wang, Y., Yang, P., Zhang, Y., Chan, Z.L.: Manipulation of arginase expression modulate abiotic stress tolerance in Arabidopsis: effect on arginine metabolism and ROS accumulation. - J. exp. Bot. 64: 1367–1379, 2013.
Shi, Y.T., Ding, Y.L., Yang, S.H.: Cold signal transduction and its interplay with phytohormones during cold acclimation. - Plant Cell Physiol. 56: 7–15, 2015.
Sun, P.P., Mao, Y.X., Li, G.Y., Cao, M., Kong, F.N., Wang, L., Bi, G.Q.: Comparative transcriptome profiling of Pyropia yezoensis (Ueda) M.S. Hwang & H.G. Choi in response to temperature stresses. - BMC Genomics 16: 463, 2015.
Tarazona, S., García-Alcalde, F., Dopazo, J., Ferrer, A., Conesa, A.: Differential expression in RNA-seq: a matter of depth. - Genome Res. 21: 4436, 2011.
Thomashow, M.F.: Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. - Ann. Rev. Plant Biol. 50: 571–599, 1999.
Triantaphylidès, C., Havaux, M.: Singlet oxygen in plants: production, detoxification and signaling. - Trends Plant Sci. 14: 219–228, 2009.
Tunnacliffe, A., Wise, M.J.: The continuing conundrum of the LEA proteins. - Sci. Natur. 94: 791–812, 2007.
Verelst, W., Bertolinid, E., De Bodt, S., Vandepoele, K., Demeulenaere, M., Pèd, M.E., Inzéa, D.: Molecular and physiological analysis of growth-limiting drought stress in Brachypodium distachyon leaves. - Mol. Plant 6: 311–322, 2013.
Vogt, T.: Phenylpropanoid biosynthesis. - Mol. Plant 3: 2–20, 2010.
Wang, W., Vinocur, B., Shoseyov, O., Altman, A.: Role of plant heat shock proteins and molecular chaperones in the abiotic stress response. - Trends Plant Sci. 9: 244–252, 2004.
Wang, W.S., Zhao, X.Q., Li, M., Huang, L.Y., Xu, J.L., Zhang, F., Cui, Y.R., Fu, B.Y., Li, Z.K.: Complex molecular mechanisms underlying seedling salt tolerance in rice revealed by comparative transcriptome and metabolomic profiling. - J. exp. Bot. 67: 405–419, 2016.
Wang, W.Y., Wang, Q.J., Wang, H.C.: The effect of land management on plant community composition, species diversity, and productivity of alpine Kobersia steppe meadow. - Ecol. Res. 21: 181–187, 2006.
Xiu, Y., Iqbal, A., Zhu, C., Wu, G., Chang, Y., Li, N., Cao, Y., Zhang, W., Zeng, H., Chen, S., Wang, H.: Improvement and transcriptome analysis of root architecture by overexpression of Fraxinus pennsylvanica DREB2A transcription factor in Robinia pseudoacacia L.‘Idaho. - Plant Biotechnol. J. 14: 1456–1469, 2016.
Xu, W.R., Li, R.M., Zhang, N.B., Ma, F.L., Jiao, Y.T., Wang, Z.P.: Transcriptome profiling of Vitis amurensis, an extremely cold-tolerant Chinese wild Vitis species, reveals candidate genes and events that potentially connected to cold stress. - Plant mol. Biol. 86: 527–541, 2014.
Yonekurasakakibara, K., Urano, K., Suzuki, M., Suzuki, M., Yamada, Y., Nishizawa, T., Matsuda, F., Kojima, M., Sakakibara, H., Shinozaki, K., Michael, A.J., Tohge, T., Yamazaki, M., Saito, K.: Enhancement of oxidative and drought tolerance in Arabidopsis by overaccumulation of antioxidant flavonoids. - Plant J. 77: 367–379, 2014.
Yu, L.J., Luo, Y.F., Liao, B., Xie, L.J., Chen, L., Xiao, S., Li, J.T., Hu, S.N., Shu, W.S.: Comparative transcriptome analysis of transporters, phytohormone and lipid metabolism pathways in response to arsenic stress in rice (Oryza sativa). - New Phytol. 195: 97–112, 2012.
Yue, C., Cao, H.L., Wang, L., Zhou, Y.H., Huang, Y.T., Hao, X.Y., Wang, Y.C., Wang, B., Yang, Y.J., Wang, X.C.: Effects of cold acclimation on sugar metabolism and sugar-related gene expression in tea plant during the winter season. - Plant mol. Biol. 88: 591–608, 2015.
Zeng, Y., Yu, J., Cang, J., Liu, L.J., Mu, Y.C., Wang, J.H., Zhang, D.: Detection of sugar accumulation and expression levels of correlation key enzymes in winter wheat (Triticum aestivum) at low temperatures. - Biosci. Biotechnol. Biochem. 75: 681–687, 2011.
Zhao, M.L., Wang, J.N., Shan, W., Fan, J.G., Kuang, J.F., Wu, K.Q., Li, X.P., Chen, W.X., He, F.Y., Chen, J.Y., Lu, W.J.: Induction of jasmonate signalling regulators MaMYC2s and their physical interactions with MaICE1 in methyl jasmonate-induced chilling tolerance in banana fruit. - Plant Cell Environ. 36: 30–51, 2013.
Zhao, W.C., Yang, X.Y., Yu, H.J., Jiang, W.J., Sun, N., Liu, X.R., Liu, X.L., Zhang, X.M., Wang, Y., Gu, X.F.: RNASeq-based transcriptome profiling of early nitrogen deficiency response in cucumber seedlings provides new insight into the putative nitrogen regulatory network. - Plant Cell Physiol. 56: 455–467, 2015.
Zhao, Z.G., Tan, L.L., Dang, C.Y., Zhang, H., Wu, Q.B., An, L.Z.: Deep-sequencing transcriptome analysis of chilling tolerance mechanisms of a subnival alpine plant, Chorispora bungeana. - BMC Plant Biol. 12: 222, 2012.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Acknowledgment: This study was funded by the China Agriculture Research System (CARS-34), the major Project for Tibetan forage industry (Z2015C02N02), and the PhD research startup foundation of the Northwest A&F University (2452017195).
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Fu, JJ., Geng, J.C., Miao, YJ. et al. Transcriptomic analyses reveal genotype- and organ-specific molecular responses to cold stress in Elymus nutans. Biol Plant 62, 671–683 (2018). https://doi.org/10.1007/s10535-018-0812-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-018-0812-5