Abstract
In recent years, several strains capable of degrading 1,4-dioxane have been isolated from the genera Pseudonocardia and Rhodococcus. This study was conducted to evaluate the 1,4-dioxane degradation potential of phylogenetically diverse strains in these genera. The abilities to degrade 1,4-dioxane as a sole carbon and energy source and co-metabolically with tetrahydrofuran (THF) were evaluated for 13 Pseudonocardia and 12 Rhodococcus species. Pseudonocardia dioxanivorans JCM 13855T, which is a 1,4-dioxane degrading bacterium also known as P. dioxanivorans CB1190, and Rhodococcus aetherivorans JCM 14343T could degrade 1,4-dioxane as the sole carbon and energy source. In addition to these two strains, ten Pseudonocardia strains could degrade THF, but no Rhodococcus strains could degrade THF. Of the ten Pseudonocardia strains, Pseudonocardia acacia JCM 16707T and Pseudonocardia asaccharolytica JCM 10410T degraded 1,4-dioxane co-metabolically with THF. These results indicated that 1,4-dioxane degradation potential, including degradation for growth and by co-metabolism with THF, is possessed by selected strains of Pseudonocardia and Rhodococcus, although THF degradation potential appeared to be widely distributed in Pseudonocardia. Analysis of soluble di-iron monooxygenase (SDIMO) α-subunit genes in THF and/or 1,4-dioxane degrading strains revealed that not only THF and 1,4-dioxane monooxygenases but also propane monooxygenase-like SDIMOs can be involved in 1,4-dioxane degradation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
1,4-Dioxane is a cyclic ether which is an industrially important solvent used in paints, lacquers, cosmetics, deodorants, fumigants and detergents. It is also formed as a by-product during the manufacture of polyesters. 1,4-Dioxane has high water solubility, low volatility, low adsorbability to solids, and low susceptibility to chemical and biological reactions (Wolfe and Jeffers 2000; Agency for Toxic Substances and Disease Registry (ATSDR) 2012; Sei et al. 2013a; Stepien et al. 2014). Because of these properties, once 1,4-dioxane is released into water environments, it can persist for a long period.
1,4-Dioxane contamination has been extensively detected in surface water, groundwater, and landfill leachate throughout the world (Lesage et al. 1990; Abe 1999; Isaacson et al. 2006; Fujiwara et al. 2008; Agency for Toxic Substances and Disease Registry (ATSDR) 2012; Chiang et al. 2012; Sei et al. 2013a; Stepien et al. 2014). The major causes of the water contamination are illegal dumping of industrial wastes, and incomplete treatment and leakage of landfill leachate and industrial wastewater. Because 1,4-dioxane is a group 2B (possible) human carcinogen (International Agency for Research on Cancer (IARC) 1999), appropriate cleanup of 1,4-dioxane-contaminated water is an important public issue. However, most of the conventional physical and chemical remediation methods are not effective to decontaminate 1,4-dioxane (Adams et al. 1994). Although advanced oxidation processes (AOPs) such as the combination of ozone and hydrogen peroxide treatments can efficiently decompose 1,4-dioxane (Adams et al. 1994; Kim et al. 2006; Kishimoto et al. 2008), their application for the cleanup of a 1,4-dioxane-contaminated environment would be unrealistic in light of cost- and energy-effectiveness. Consequently, the development of low cost, energy efficient and environmentally friendly alternatives is strongly desired.
Bioremediation is a promising remediation technique because of its cost effectiveness, inherent eco-friendly properties, and the potential for complete decomposition of harmful compounds. Although 1,4-dioxane had been recognized as a recalcitrant to biodegradation, bacterial strains capable of degrading 1,4-dioxane as the sole carbon and energy source or co-metabolically with tetrahydrofuran (THF), methane, propane, and toluene have been isolated and characterized during the last two decades (Table S1 in Online Resource 1). Furthermore, several recent studies have confirmed the intrinsic 1,4-dioxane biodegradation potential of contaminated environments by microcosm studies (Li et al. 2010; Sei et al. 2010; Li et al. 2014; 2015) or field surveys (Chiang et al. 2012). These findings suggested the feasibility of in situ bioremediation for 1,4-dioxane contaminated environments. However, little is known concerning 1,4-dioxane degrading microorganisms compared with what is known about strains that can degrade petroleum hydrocarbons like benzene, toluene, ethylbenzene, and xylene (Alvarez and Vogel 1991; Cao et al. 2009; Weelink et al. 2010; Sun and Cupples 2012), and chlorinated solvents like trichloroethylene (Damborský 1999; Shukla et al. 2014) for which in situ bioremediation technologies have been well established. To establish effective bioremediation strategies for 1,4-dioxane contamination, further knowledge of 1,4-dioxane degrading microorganisms is necessary.
Although recent research revealed that a relatively wide range of bacterial species possess the ability to degrade 1,4-dioxane, the majority are nocardioform actinomycetes, belonging to genera such as Pseudonocardia (Parales et al. 1994; Kohlweyer et al. 2000; Kämpfer and Kroppenstedt 2004; Prabahar et al. 2004; Vainberg et al. 2006; Sei et al. 2013a; Matsui et al. 2016) and Rhodococcus (Bernhardt and Diekmann 1991; Deeb and Alvarez-Cohen 1999; Sei et al. 2013b) (Table S1 in Online Resource 1). Thus, an understanding of the 1,4-dioxane degrading potential and properties of bacterial species of these genera is important for developing bioremediation technologies for 1,4-dioxane contamination. Therefore, this study was conducted to evaluate the 1,4-dioxane degradation potential of phylogenetically diverse members of Pseudonocardia and Rhodococcus. The ability of the test strains to degrade 1,4-dioxane as the sole carbon and energy source and co-metabolically degrade 1,4-dioxane in the presence of THF was evaluated. The genes encoding soluble di-iron monooxygenase (SDIMO), which is known to be associated with the initial oxidation of 1,4-dioxane (Li et al. 2013), in the test strains were also analyzed.
Materials and methods
Test strains
A total of 13 and 12 phylogenetically diverse species of Pseudonocardia and Rhodococcus, respectively, were selected to evaluate their 1,4-dioxane degradation potential (Figs. S1 and S2 in Online Resource 1). Type strains for the 25 species (Table 1) were provided by the RIKEN BRC through the National Bio-Resource Project of the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan, and were used as test strains in this study. Pseudonocardia dioxanivorans JCM 13855T, which is known as P. dioxanivorans CB1190 (Parales et al. 1994), was used as the positive control as it was capable of degrading 1,4-dioxane as a sole carbon and energy source.
Culture media
MGY medium (malt extract 10 g/L, D(+)-glucose 4 g/L, yeast extract 4 g/L, pH 7.3) and ISP medium 2 (Becton, Dickinson and Company, Sparks, MD, USA) were used as liquid and solid media, respectively, for routine cultivation of test strains. Basal salt medium (BSM) (Parales et al. 1994) adjusted to pH 7.0 was used for 1,4-dioxane degradation experiments.
1,4-Dioxane degradation experiments
Prior to the 1,4-dioxane degradation experiments, test strains were precultivated in MGY medium at 28 °C, except for Pseudonocardia acaciae JCM 16707T and Pseudonocardia thermophila JCM 3095T, which were precultivated at 37 °C. The cells of the precultivated strains were harvested by centrifugation (8500×g, 4 °C, 5 min) and washed twice with sterilized 0.85 % (w/v) NaCl. Then, the washed cells were inoculated in 50-ml glass vials containing 20 ml of BSM to give a final cell density (determined by optical density at 600 nm (OD600)) of 1.0. 1,4-Dioxane was added to BSM at 20 mg/L as the sole carbon and energy source for 1,4-dioxane utilization experiments. 1,4-Dioxane and THF were added at 20 and 50 mg/L, respectively, for the 1,4-dioxane co-metabolic degradation experiments. Control systems without bacterial inoculation were also prepared for each experiment. The cultures were incubated at 28 °C with rotary shaking at 150 rpm, while P. acaciae JCM 16707T and P. thermophila JCM 3095T were incubated at 37 °C, excepting for the screening experiments to select THF degrading strains in the 1,4-dioxane co-metabolic degradation experiments. 1,4-Dioxane utilization experiments were conducted for 14 days. For the 1,4-dioxane co-metabolic degradation experiments, first, screening experiments to select the THF degrading strains were conducted for 7 days, and then experiments to examine the co-metabolic degradation of 1,4-dioxane with THF were carried out for 14 days on the selected strains. Aliquots (0.5 ml) were periodically collected, centrifuged (20,000×g, 4 °C, 5 min), filtered through a cellulose acetate filter (pore size 0.45 µm, Advantec, Tokyo, Japan), and subjected to 1,4-dioxane and THF quantification. All of the degradation experiments were conducted in duplicate. Statistical significance for THF and 1,4-dioxane degradation by test strains was determined by Student’s t test with p < 0.05 using the Prism 6J for Windows program (GraphPad Software, La Jolla, CA, USA).
Chemical analysis
Bacterial cell density (OD600) was determined with a PD-3000UV spectrophotometer (Apel, Saitama, Japan) or a UVmini-1240 spectrophotometer (Shimadzu, Kyoto, Japan). The concentrations of 1,4-dioxane and THF were determined using a gas chromatograph GC2014 (Shimadzu) equipped with a flame-ionization detector (FID) and a 2.1-m (3.2-mm i.d.) glass column packed with Gaskuropack 54 (GL Science, Tokyo, Japan). Nitrogen gas was applied as the carrier gas at a flow rate of 55 ml/min. The injector and column oven temperature was set at 200 °C, while the detector temperature was at 230 °C. The injection volume of the filtered samples was 5 µl, and the detection limit of 1,4-dioxane and THF was 1 mg/L.
Analysis of soluble di-iron monooxygenase genes
1,4-Dioxane degradation enzymes are included in the large SDIMO family, which includes multicomponent enzymes that catalyze the initial oxidation of a variety of hydrocarbons (Coleman et al. 2006; Li et al. 2013). The presence of SDIMO genes in test strains was examined by PCR using the NVC57 and NVC66 primer sets, which was specifically designed to detect the conserved region in the SDIMO α-subunit genes (Coleman et al. 2006). Pseudonocardia sp. D17 [a 1,4-dioxane utilizing bacterium (Sei et al. 2013a)] and Rhodococcus ruber T1 and T5 [THF degrading bacteria capable of co-metabolically degrading 1,4-dioxane (Sei et al. 2013b)] were also analyzed for their SDIMO genes. PCR was conducted using the following thermal profile: initial denaturation at 94 °C for 5 min, 35 cycles of denaturation at 94 °C for 30 s, annealing at 57 °C for 30 s and extension at 72 °C for 1 min, and final extension at 72 °C for 5 min. This protocol gives PCR products of approximately 420 bp. The PCR products were analyzed by electrophoresis on a 1.5 % (w/v) agarose gel stained with SYBR Green I (Takara Bio, Shiga, Japan), after which they were purified with NucleoSpin Gel and PCR Clean-up (Macherey–Nagel, Düren, Germany) and sequenced on an Applied Biosystems 3730XL DNA analyzer (Applied Biosystems, Foster City, CA, USA) with primers NVC57 and NVC66 at Macrogen Janan (Tokyo, Japan). The nucleotide sequences obtained from the sequencing with both primers were combined to yield single sequences. The nucleotide sequences determined for the test strains were compared with those in the NCBI database using the BLAST search program (http://www.ncbi.nlm.nih.gov/blast/). Then, the nucleotide sequences of the test strains in this study and reference strains in the NCBI database were aligned using CLUSTAL W (Eddy 1995), and a phylogenetic tree was produced using TreeView X (Page 1996).
Nucleotide sequence accession numbers
The partial sequences of the SDIMO α-subunit genes determined in this study were deposited in the DDBJ/EMBL/GenBank databases under accession numbers LC114132 to LC114146.
Results
1,4-Dioxane utilization ability
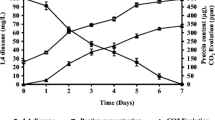
In the 1,4-dioxane utilization experiments, where 1,4-dioxane was added as the sole carbon and energy source, 1,4-dioxane was significantly degraded by P. dioxanivorans JCM 13855T and Rhodococcus aetherivorans JCM 14343T (p < 0.05), while 1,4-dioxane was not significantly degraded by the other 23 test strains within 14 days (Table 1). P. dioxanivorans JCM 13855T completely degraded 20 mg/L of 1,4-dioxane within 4 days after a 2 days lag period (Fig. 1a), which agreed with the previous finding that this strain has inducible 1,4-dioxane degradation enzymes (Kelly et al. 2001). R. aetherivorans JCM 14343T degraded 20 mg/L of 1,4-dioxane completely within 1 day after a 9 h lag period (Fig. 1b).
1,4-Dioxane co-metabolic degradation ability with THF
Prior to the evaluation of the ability of the test strains to degrade 1,4-dioxane co-metabolically with THF, we first screened their THF degradation potential. Among the 13 Pseudonocardia strains, 11 strains except for Pseudonocardia ammonioxydans JCM 12462T and Pseudonocardia chloroethenivorans JCM 12679T were capable of significantly degrading THF within 7 days (Table 1). Among the 12 Rhodococcus strains, THF was significantly degraded within 7 days by R. aetherivorans JCM 14343T, while THF degradation did not occur within 7 days by the other 11 test strains.
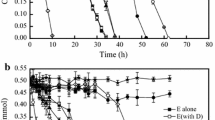
The 12 THF degrading strains were then examined for their 1,4-dioxane co-metabolic degradation ability with THF for 14 days. In addition to P. dioxanivorans JCM 13855T and R. aetherivorans 14343T, both of which were capable of utilizing 1,4-dioxane for their growth, P. acaciae JCM 16707T and Pseudonocardia asaccharolytica JCM 10410T could significantly degrade 1,4-dioxane subsequent to THF degradation (Table 1). P. acaciae JCM 16707T completely degraded THF within 12 days without a lag period, and also completely degraded 1,4-dioxane within 14 days after a lag period of 10 days (Fig. 2a). P. asaccharolytica JCM 10410T degraded nearly 80 % of the initial THF within 14 days without a lag period (Fig. 2b). 1,4-Dioxane degradation by this strain was initiated after a lag period of 12 days, and nearly 45 % of the initial 1,4-dioxane was degraded after 14 days. Co-metabolic degradation of 1,4-dioxane with THF did not occur within 14 days by the other eight strains (Table 1; Fig. 2c, d, and Fig. S3 in Online Resource 1).
Co-metabolic degradation of 1,4-dioxane with tetrahydrofuran (THF) by Pseudonocardia acaciae JCM 16707T (a) and Pseudonocardia asaccharolytica JCM 10410T (b). Examples of THF degradation without co-metabolic degradation of 1,4-dioxane by Pseudonocardia hydrocarbonoxydans JCM 3392T (c) and Pseudonocardia thermophila JCM 3095T (d) are also shown. Error bars indicate the data range in duplicate experiments
Possession of SDIMO genes
The presence of SDIMO genes in the 12 strains capable of degrading THF and/or 1,4-dioxane, and three recently isolated 1,4-dioxane degrading strains (Pseudonocardia sp. D17, R. ruber T1 and R. ruber T5) were examined by a PCR assay specific to the SDIMO α-subunit genes. PCR products with anticipated size were obtained from all of the strains. The phylogenetic tree constructed based on the nucleotide sequences of the PCR products from the 15 strains determined in this study and those of known SDIMO α-subunit genes is shown in Fig. 3. From the nucleotide sequence of the SDIMO α-subunit gene of the control strain P. dioxanivorans JCM 13855T (P. dioxanivorans CB1190), the presence of a putative 1,4-dioxane monooxygenase α-subunit gene (dxmA gene) was confirmed. The nucleotide sequences of the SDIMO α-subunit genes of Pseudonocardia sp. D17, R. ruber T1, and R. ruber T5 were 100 % identical to that of the THF monooxygenase α-subunit gene (thmA) of Rhodococcus sp. YYL.
Phylogenetic tree constructed based on the nucleotide sequences of soluble di-iron monooxygenase α-subunit genes of the test strains and the ones previously reported for cultured strains. Sequences for the strains indicated in boldface were determined in this study. Numbers adjacent to the branches indicate the bootstrap values based on 1000 replicates. Bar indicates 0.1 substitutions per sequence position
The nucleotide sequences of the SDIMO α-subunit genes of the other 11 strains (i.e., R. aetherivorans JCM 14343T and 10 Pseudonocardia strains with THF degradation ability) were closely related to the sequences of putative propane monooxygenase hydroxylase large subunit genes (prmA-like genes) of previously characterized strains (Fig. 3). The SDIMO α-subunit gene of R. aetherivorans JCM 14343T was identical to the prmA gene of Rhodococcus sp. RR1, and allocated into a prmA gene subcluster for Rhodococcus strains. It was highly different from the SDIMO α-subunit genes classified into the thm/dxm gene cluster (similarity: 58.4–58.6 %). The SDIMO α-subunit genes of the 10 Pseudonocardia strains had 89.2–95.6 % similarity to each other. Of these, that of P. acaciae JCM 16707T had the highest similarity (91.0 %) to the prm1A gene of Pseudonocardia sp. TY-7, while those of the other nine strains had the highest similarity of 92.0–95.9 % to the prmA-like gene located on the chromosome of P. dioxanivorans CB1190.
Discussion
Several strains of the genus Pseudonocardia are capable of degrading 1,4-dioxane for growth (Parales et al. 1994; Kämpfer and Kroppenstedt 2004; Sei et al. 2013a; Matsui et al. 2016), or co-metabolically (Kohlweyer et al. 2000; Vainberg et al. 2006). In this study, although 1,4-dioxane degradation as the sole carbon and energy source by P. dioxanivorans JCM 13855T (also known as P. dioxanivorans CB1190 (Parales et al. 1994)) was confirmed, none of the other 12 Pseudonocardia strains could degrade 1,4-dioxane as the sole carbon and energy source (Table 1). It was surprising that Pseudonocardia carboxydivorans JCM 14827T was not capable of degrading 1,4-dioxane for its growth or by co-metabolism with THF, although its 16S rRNA gene was 100 % homologous to Pseudonocardia sp. RM-31, a strain that was very recently isolated as a novel 1,4-dioxane assimilating bacterium (Matsui et al. 2016). Different 1,4-dioxane degradation abilities in the two P. carboxydivorans strains suggest that their 1,4-dioxane assimilation ability is strain-specific rather than species-specific. By contrast, 10 of the 12 Pseudonocardia strains other than P. dioxanivorans JCM 13855T were capable of degrading THF, and two of them (P. acaciae JCM 16707T and P. asaccharolytica JCM 10410T) enabled co-metabolic 1,4-dioxane degradation with THF. To our knowledge, this is the first study to report the THF degradation and co-metabolic 1,4-dioxane degradation potential of these Pseudonocardia strains. Taken together, the results of this study indicated that limited Pseudonocardia strains possess the ability to degrade 1,4-dioxane degradation as the sole carbon and energy source. Additionally, our results indicated that phylogenetically diverse Pseudonocardia species/strains commonly possess the potential to degrade THF for growth and some can co-metabolically degrade 1,4-dioxane with THF.
Within the genus Rhodococcus, 1,4-dioxane degradation as the sole carbon and energy source was first reported for R. ruber 219 (Bernhardt and Diekmann 1991). It was recently reported that R. ruber T1 and T5 can co-metabolically degrade 1,4-dioxane with THF (Sei et al. 2013b). However, R. ruber JCM 3205T examined in this study could not degrade either 1,4-dioxane or THF. This suggested that 1,4-dioxane and THF degradation ability is a strain-specific property in R. ruber. By contrast, this study found that R. aetherivorans JCM 14343T could degrade 1,4-dioxane as the sole carbon and energy source. Also, we have confirmed in another study that the strain can utilize 1,4-dioxane for its growth, with the cell yield of 0.031 mg-protein/mg-1,4-dioxane (unpublished data). R. aetherivorans JCM 14343T (originally named strain 10bc312T) was isolated from an enrichment obtained from the petrochemical biotreater sludge of a chemical effluent treatment plant as a methyl tert-butyl ether degrading strain (Goodfellow et al. 2004), but its degradation ability for cyclic ethers has not been reported. Although another strain of R. aetherivorans was reported to degrade THF (Tajima et al. 2012), toluene (Hori et al. 2009), and a spectrum of petroleum compounds (Auffret et al. 2009), this is the first study to clarify that a R. aetherivorans strain can degrade 1,4-dioxane as the sole carbon and energy source. Additionally, the evidence from previous studies and this study indicates that R. aetherivorans may be a specific species in Rhodococcus that is capable of degrading both cyclic and non-cyclic recalcitrant ether compounds. It is likely that 1,4-dioxane degradation ability, including both utilization for growth and co-metabolic degradation with THF, is not distributed widely among the genus Rhodococcus, but is possessed by specific strains in limited species of the genus. Based on the phylogenetic composition of Rhodococcus (Fig. S2 in Online Resource 1), its 1,4-dioxane degradation ability is likely a specific function of some strains in a subcluster consisting of R. aetherivorans and R. ruber.
SDIMOs are multicomponent enzymes that catalyze the initial oxidation of a variety of hydrocarbons such as chlorinated solvents, aromatic hydrocarbons, alkanes and alkenes in phylogenetically and physiologically diverse bacteria (Coleman et al. 2006; Li et al. 2013). Previous studies have reported that THF and 1,4-dioxane monooxygenases (THM and DXM, respectively) are involved in 1,4-dioxane degradation for growth or by co-metabolism, and some propane monooxygenases (PMOs) can also co-metabolize 1,4-dioxane (Mahendra and Alvarez-Cohen 2006; Li et al. 2013). Nevertheless, our SDIMO gene analysis revealed that the SDIMO genes possessed by R. aetherivorans JCM 14343T, P. acaciae JCM 16707T and P. asaccharolytica JCM 10410T, whose 1,4-dioxane degrading abilities were revealed in this study for the first time, were closely related to prm genes, and clearly separated from thm/dxm genes. This is the first study to identify the presence of PMO-like SDIMOs that are possibly involved in 1,4-dioxane degradation as the sole carbon and energy source (i.e., SDIMO of R. aetherivorans JCM 14343T). PMO-like SDIMOs are diverse in light of their 1,4-dioxane degradation abilities; some enable 1,4-dioxane degradation for the growth of host strains or by co-metabolism with primary substrates such as THF and propane, while the others cannot catalyze 1,4-dioxane degradation even by co-metabolism. This indicates the importance of not only THM/DXM-possessing microorganisms but also PMO-possessing microorganisms in the implementation of 1,4-dioxane bioremediation. Further genetic and enzymatic studies are needed to obtain a deeper understanding of the 1,4-dioxane degradation abilities of SDIMOs including THM/DXM and PMOs.
In conclusion, the 1,4-dioxane degradation potential of strains of Pseudonocardia and Rhodococcus was evaluated in this study. Our results revealed that 1,4-dioxane can be degraded by selected strains of Pseudonocardia and Rhodococcus, indicating that the 1,4-dioxane degradation potential by natural attenuation or biostimulation cannot by evaluated by the occurrence of those genera in 1,4-dioxane-contaminated sites. Another novel finding that PMO-like SDIMOs are possibly capable of degrading 1,4-dioxane not only by co-metabolism but also for growth suggests that the molecular tool previously developed based on THM/DXM genes (Li et al. 2014) would underestimate the 1,4-dioxane degradation potential of microbial communities in contaminated sites. Thus, a comprehensive molecular tool that can detect all of the SDIMO genes that enable 1,4-dioxane degradation is needed for adequate evaluation of the 1,4-dioxane degradation potential of microbial communities. Nevertheless, the evidence for the presence of phylogenetically diverse 1,4-dioxane degrading strains reported in previous studies (Table S1 in Online Resource 1) and this study would suggest that in situ bioremediation of 1,4-dioxane-contaminated water environments such as groundwater may be possible by dominance and/or selective activation of indigenous degraders through injecting appropriate growth and/or primary substrates even though the degraders may be minor constituents in the environment.
References
Abe A (1999) Distribution of 1,4-dioxane in relation to possible sources in the water environment. Sci Total Environ 227:41–47
Adams CD, Scanlan PA, Secrist ND (1994) Oxidation and biodegradability enhancement of 1,4-dioxane using hydrogen peroxide and ozone. Environ Sci Technol 28:1812–1818
Agency for Toxic Substances and Disease Registry (ATSDR) (2012) Toxicological profile for 1,4-dioxane. United States Department of Health and Human Services, Public Health Service, Atlanta, GA. http://www.atsdr.cdc.gov/toxprofiles/tp.asp?id=955&tid=199. Accessed 28 February 2016
Alvarez PJJ, Vogel TM (1991) Substrate interactions of benzene, toluene, and para-xylene during microbial degradation by pure cultures and mixed culture aquifer slurries. Appl Environ Microbiol 57:2981–2985
Auffret M, Labbé D, Thouand G, Greer CW, Fayolle-Guichard F (2009) Degradation of a mixture of hydrocarbons, gasoline, and diesel oil additives by Rhodococcus aetherivorans and Rhodococcus wratislaviensis. Appl Environ Microbiol 75:7774–7782
Bernhardt D, Diekmann H (1991) Degradation of dioxane, tetrahydrofuran and other cyclic ethers by an environmental Rhodococcus strain. Appl Microbiol Biotechnol 36:120–123
Cao B, Nagarajan K, Loh KC (2009) Biodegradation of aromatic compounds: current status and opportunities for biomolecular approaches. Appl Microbiol Biotechnol 85:207–228
Chiang SYD, Mora R, Diguiseppi WH, Davis G, Sublette K, Gedalanga P, Mahendra S (2012) Characterizing the intrinsic bioremediation potential of 1,4-dioxane and trichloroethene using innovative environmental diagnostic tools. J Environ Monit 14:2317–2326
Coleman NV, Bui NB, Holmes AJ (2006) Soluble di-iron monooxygenase gene diversity in soils, sediments and ethene enrichments. Environ Microbiol 8:1228–1239
Damborský J (1999) Tetrachloroethene-dehalogenating bacteria. Folia Microbiol 44:247–262
Deeb RA, Alvarez-Cohen L (1999) Temperature effects and substrate interactions during the aerobic biotransformation of BTEX mixtures by toluene-enriched consortia and Rhodococcus rhodochrous. Biotechnol Bioeng 62:526–536
Eddy SR (1995) Multiple alignment using hidden Markov models. In: Rawlings C, Clark D, Altman R, Hunter L, Lengauer T, Wodak S (eds.) Proceedings of the third international conference on intelligent systems for molecular biology. AAAI Press, Menlo Park, pp 114–120
Fujiwara T, Tamada T, Kurata Y, Ono Y, Kose T, Ono Y, Nishimura F, Ohtoshi K (2008) Investigation of 1,4-dioxane originating from incineration residues produced by incineration of municipal solid waste. Chemosphere 71:894–901
Goodfellow M, Jones AL, Maldonado LA, Salanitro J (2004) Rhodococcus aetherivorans sp. nov., a new species that contains methyl t-butyl ether-degrading actinomycetes. Syst Appl Microbiol 27:61–65
Hori K, Kobayashi A, Ikeda H, Unno H (2009) Rhodococcus aetherivorans IAR1, a new bacterial strain synthesizing poly(3-hydroxybutyrate-co-3-hydroxyvalerate) from toluene. J Biosci Bioeng 107:145–150
International Agency for Research on Cancer (IARC) (1999) 1,4-Dioxane. In: IARC monographs on the evaluation of carcinogenic risks to humans, vol. 71, Re-evaluation of some organic chemicals, hydrazine and hydrogen peroxide. IARC, Lyon, pp 589–602
Isaacson C, Mohr TKG, Field JA (2006) Quantitative determination of 1,4-dioxane and tetrahydrofuran in groundwater by solid phase extraction GC/MS/MS. Environ Sci Technol 40:7305–7311
Kämpfer P, Kroppenstedt RM (2004) Pseudonocardia benzenivorans sp. nov. Int J Syst Evol Microbiol 54:749–751
Kelly SL, Aitchison EW, Deshpande M, Schnoor JL, Alvarez PJJ (2001) Biodegradation of 1,4-dioxane in planted and unplanted soil: effect of bioaugmentation with Amycolata sp. CB1190. Water Res 35:3791–3800
Kim CG, Seo HJ, Lee BR (2006) Decomposition of 1,4-dioxane by advanced oxidation and biochemical process. J Environ Sci Health A 41:599–611
Kishimoto N, Nakagawa T, Asano M, Abe M, Yamada M, Ono Y (2008) Ozonation combined with electrolysis of 1,4-dioxane using a two-compartment electrolytic flow cell with solid electrolyte. Water Res 42:379–385
Kohlweyer U, Tiemer B, Schräder T, Andreesen JR (2000) Tetrahydrofuran degradation by a newly isolated culture of Pseudonocardia sp. strain K1. FEMS Microbiol Lett 186:301–306
Lesage S, Jackson RE, Priddle MW, Riemann PG (1990) Occurrence and fate of organic solvent residues in anoxic groundwater at the Gloucester Landfill, Canada. Environ Sci Technol 24:559–566
Li M, Fiorenza S, Chatham JR, Mahendra S, Alvarez PJJ (2010) 1,4-Dioxane biodegradation at low temperatures in Arctic groundwater samples. Water Res 44:2894–2900
Li M, Mathieu J, Yang Y, Fiorenza S, Deng Y, He Z, Zhou J, Alvarez PJJ (2013) Widespread distribution of soluble di-iron monooxygenase (SDIMO) genes in Arctic groundwater impacted by 1,4-dioxane. Environ Sci Technol 47:9950–9958
Li M, Mathieu J, Liu Y, Van Orden ET, Yang Y, Fiorenza S, Alvarez PJJ (2014) The abundance of tetrahydrofuran/dioxane monooxygenase genes (thmA/dxmA) and 1,4-dioxane degradation activity are significantly correlated at various impacted aquifers. Environ Sci Technol Lett 1:122–127
Li M, Van Orden ET, DeVries DJ, Xiong Z, Hinchee R, Alvarez PJ (2015) Bench-scale biodegradation tests to assess natural attenuation potential of 1,4-dioxane at three sites in California. Biodegradation 26:39–50
Mahendra S, Alvarez-Cohen L (2006) Kinetics of 1,4-dioxane biodegradation by monooxygenase-expressing bacteria. Environ Sci Technol 40:5435–5442
Matsui R, Takagi K, Sakakibara F, Abe T, Shiiba K (2016) Identification and characterization of 1,4-dioxane-degrading microbe separated from surface seawater by the seawater-charcoal perfusion apparatus. Biodegradation. doi:10.1007/s10532-016-9763-8
Page RDM (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Parales RE, Adamus JE, White N, May HD (1994) Degradation of 1,4-dioxane by an actinomycete in pure culture. Appl Environ Microbiol 60:4527–4530
Prabahar V, Dube S, Reddy GSN, Shivaji S (2004) Pseudonocardia antarctica sp. nov. an Actinomycetes from McMurdo Dry Valleys, Antarctica. Syst Appl Microbiol 27:66–71
Sei K, Kakinoki T, Inoue D, Soda S, Fujita M, Ike M (2010) Evaluation of the biodegradation potential of 1,4-dioxane in river, soil and activated sludge samples. Biodegradation 21:585–591
Sei K, Miyagaki K, Kakinoki T, Fukugasako K, Inoue D, Ike M (2013a) Isolation and characterization of bacterial strains that have high ability to degrade 1,4-dioxane as a sole carbon and energy source. Biodegradation 24:665–674
Sei K, Oyama M, Kakinoki T, Inoue D, Ike M (2013b) Isolation and characterization of tetrahydrofuran-degrading bacteria for 1,4-dioxane-containing wastewater treatment by co-metabolic degradation. J Water Environ Technol 11:11–19
Shukla AK, Upadhyay SN, Dubey SK (2014) Current trends in trichloroethylene biodegradation: a review. Crit Rev Biotechnol 34:101–114
Stepien DK, Diehl P, Helm J, Thoms A, Püttmann W (2014) Fate of 1,4-dioxane in the aquatic environment: from sewage to drinking water. Water Res 48:406–419
Sun W, Cupples AM (2012) Diversity of five anaerobic toluene-degrading microbial communities investigated using stable isotope probing. Appl Environ Microbiol 78:972–980
Tajima T, Hayashida N, Matsumura R, Omura A, Nakashimada Y, Kato J (2012) Isolation and characterization of tetrahydrofuran-degrading Rhodococcus aetherivorans strain M8. Process Biochem 47:1665–1669
Vainberg S, McClay K, Masuda H, Root D, Condee C, Zylstra GJ, Steffan RJ (2006) Biodegradation of ether pollutants by Pseudonocardia sp. strain ENV478. Appl Environ Microbiol 72:5218–5224
Weelink SAB, van Eekert MHA, Stams AJM (2010) Degradation of BTEX by anaerobic bacteria: physiology and application. Rev Environ Sci Biotechnol 9:359–385
Wolfe NL, Jeffers PM (2000) Hydrolysis. In: Boethling RS, Mackay D (eds) Handbook of property estimation methods for chemicals: environmental and health sciences. Lewis Publishers, Chelsea, pp 311–334
Acknowledgments
This study was supported by Environment Research and Technology Development Fund (5B-1201) of the Ministry of the Environment, Japan, by the Adaptable and Seamless Technology Transfer Program through Target-driven R&D (A-STEP) of the Japan Science and Technology (JST), Japan, and by the Kurita Water and Environment Foundation (15A066).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Inoue, D., Tsunoda, T., Sawada, K. et al. 1,4-Dioxane degradation potential of members of the genera Pseudonocardia and Rhodococcus . Biodegradation 27, 277–286 (2016). https://doi.org/10.1007/s10532-016-9772-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-016-9772-7