Abstract
There is a strong relationship between the dysregulation of epidermal growth factor receptor (EGFR) and the development of epithelial-derived cancers. Therefore, EGFR has usually been considered the desired target for gene therapy. Here, we propose an approach for targeting EGFR-expressing cells by phage particles capable of displaying EGF and GFP as tumor-targeting and reporting elements, respectively. For this purpose, the superfolder GFP-EGF (sfGFP-EGF) coding sequence was inserted at the N-terminus of the pIII gene in the pIT2 phagemid. The capability of the constructed phage to recognize EGFR-overexpressing cells was monitored by fluorescence microscopy, fluorescence-activated cell sorting (FACS), and cell-based ELISA experiments. FACS analysis showed a significant shift in the mean fluorescence intensity (MFI) of the cells treated with phage displaying sfGFP-EGF compared to phage displaying only sfGFP. The binding of phage displaying sfGFP-EGF to A-431 cells, monitored by fluorescence microscopy, indicated the formation of the sfGFP-EGF-EGFR complex on the surface of the treated cells. Cell-based ELISA experiments showed that phages displaying either EGF or sfGFP-EGF can specifically bind EGFR-expressing cells. The vector constructed in the current study has the potential to be engineered for gene delivery purposes as well as cell-based imaging for tumor detection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tyrosine kinase receptors (TKRs) are a group of regulatory proteins that play important roles in the normal functions of epithelial tissues in the skin, lung, pancreas, gastrointestinal tract, and central nervous system (Ferguson 2008; Normanno et al. 2006; Sebastian et al. 2006). The EGFR family of TKRs includes four members: EGFR (epidermal growth factor receptor) or ErbB1 (the symbol is derived from the name of erythroblastic leukemia viral oncogene), ErbB2 (HER2), ErbB3 (HER3) and ErbB4 (HER4). EGFR dysregulation can lead to the transformation of normal cells into malignant cells (Sebastian et al. 2006; Wieduwilt and Moasser 2008). Therefore, inhibition of EGFR activation using various strategies, such as the application of anti-EGFR monoclonal antibodies (mAbs) and tyrosine kinase inhibitors (TKIs), is considered a promising cancer treatment regimen (Azzazy and Highsmith 2002; Hynes and Lane 2005; Morgillo and Lee 2005; Wheeler et al. 2010; Wieduwilt and Moasser 2008). Human EGFR (hEGFR) is a transmembrane glycoprotein with a molecular weight of 170 kDa containing 1186 amino acids resulting from its precursor polypeptide (Ogiso et al. 2002; Jorissen et al. 2003). From a conformational point of view, the extracellular region of inactive EGFR has a “tethered” configuration. Receptor dimerization is induced by EGF binding to the extracellular region of the receptor and the formation of a 2:2 complex from EGF and EGFR. Dimerization of the receptor is followed by the activation of a tyrosine kinase domain, phosphorylation of tyrosine residues, and triggering many downstream signaling pathways (Dawson et al. 2007; Klein et al. 2004; Lemmon 2009; Ogiso et al. 2002). Given the substantial evidence, the overexpression and mutation of EGFR are implicated in several human epithelial malignancies. Therefore, this receptor has been considered a potential therapeutic target for drug discovery and gene delivery in cancer therapy (Xu et al. 2017). The most crucial step in gene therapy is choosing an appropriate vector for efficient delivery of therapeutic agents that can specifically target cancer cells with minimal toxicity in normal tissues. Eukaryotic viruses and nonviral vehicles are two main types of vectors for the delivery and expression of genes in mammalian cells, each type with some benefits and pitfalls (Mali 2013; Sung and Kim 2019). Recently, bacteriophages have received great attention as novel vehicles for gene delivery due to their lack of tropism for mammalian cells and high safety profile. Bacteriophages also provide the possibility of being manipulated genetically and chemically to improve the efficiency of gene transfer (Clark and March 2006; Dabrowska et al. 2005). It has circular single-stranded DNA (ss-DNA) consisting of 11 genes responsible for producing several proteins, which have been classified based on their functions (Felici et al. 1995; Marvin et al. 2014; Straus and Bo 2018; Wang et al. 2006). Important coat proteins for displaying the proteins and peptides on the phage surface are pIII and pVIII, known as minor and major coat proteins, respectively. They are responsible for virion infection, assembly, termination, and stabilization (Marvin et al. 2014; Straus and Bo 2018; Wang et al. 2006). Phagemids are Ff-phage-derived vectors containing genes for only one of the coat fusion proteins. In this case, the entire phage-derived coat proteins necessary for phagemid genome encapsulation, as well as the enzymes needed for phage replication, are provided by a helper phage (Dani 2001; Qi et al. 2012).
Due to the merits of phage for gene delivery, phage-based vectors have been successfully used for transferring TNF-α, IL-12, IL-15, CRISPR/Cas9, and HSVtk coding genes to cancer cells (Petrov et al. 2022). Another example is the successful application of bacteriophage T7 for the targeted delivery of the mammalian granulocyte–macrophage colony-stimulating factor (GM-CSF)-encoding gene to mouse melanoma cells and its feasibility for in vitro and in vivo imaging as well as therapeutic purposes (Hwang and Myung 2020).
The current study aims to target EGFR-overexpressing cells based on the phage pIII display system. EGF was used as the targeting agent, and superfolder green fluorescent protein (sfGFP) was used to monitor protein localization, as well as protein‒protein and phage-cell interactions. Here, phagemid pIT2 was used for the construction of the vector of interest in which phage pIII coat protein was used for displaying EGF and sfGFPs linked to each other by a (GGGGS)3 linker. In the current study, phages displaying either EGF or sfGFP were also produced as controls by inserting their corresponding coding genes at the N-terminus of the phage pIII coding sequence. The constructed vectors were amplified to produce a complete phage, in which the desired proteins were expressed attached to the phage pIII coat-protein. To investigate EGF-mediated binding, fluorescence microscopy, ELISA, and FACS analyses were employed.
Materials and methods
Reagents
All of the chemicals used in this work were of biological grade. Tryptone, yeast extract, isopropyl-b-D-thio galactopyranoside (IPTG), Triton X-100, trypsin, phenylmethylsulfonyl fluoride (PMSF), N,N,N',N'-tetra methyl ethylene diamine (TEMED), ortho phosphoric acid, Coomassie brilliant blue G-250, anhydrous D-glucose, potassium dihydrogen phosphate, disodium hydrogen phosphate, sodium azide (NaN3) and urea were purchased from AppliChem (Darmstadt, Germany). Glycine and TMB (3,3′,5,5′-Tetramethylbenzidine) were purchased from Sigma, USA. NaCl and polyethylene glycol (PEG) 8000 were obtained from Scharlau (Barcelona, Spain). β-Mercaptoethanol, absolute ethanol, methanol, tris, sodium dodecyl sulfate, sodium hydroxide, agar, hydrochloric acid, sulfuric acid, and acrylamide were obtained from Merck (Darmstadt, Germany). DNA ladders, 6 × DNA loading dye, lysozyme, restriction enzymes (SalI, XhoI, NotI, and NcoI) and T4 DNA ligase were received from Fermentas (USA). The primers used in this work were supplied by Bioron (Germany) and ordered via FAZA Biotech (Tehran, Iran). The Qiagen plasmid maxi and mini extraction kits were purchased from Qiagen (Hilden, Germany). A gel purification kit was obtained from Yekta Tajhiz (Iran). Ni-Sepharose 6 fast flow was prepared from GE Healthcare Life Sciences (Sweden). Phagemid vector (pIT2) was obtained from the Tomlinson Phage-display antibody library by MRC HGMP Resource Centre. Anti-M13 HRP-conjugated monoclonal antibody was prepared from Sino Biological Inc. (Beijing, P. R. China). Mouse monoclonal His-probe antibody and goat anti-mouse IgG-HRP secondary antibody were prepared from Santa Cruz Biotechnology (USA). Agarose was obtained from Invitrogen Ltd. (Paisley, UK). N,N'-Methylene-bis-acrylamide and PCR master kits were purchased from CinnaGen (Tehran, Iran). Ultrapure Milli-Q water (Millipore Corporation, Bradford, MA, USA) was used for the preparation of all solutions.
The epidermal carcinoma cell line (A-431; originated from an 85-year-old female with epidermal carcinoma, ATCC: CRL-1555) and the Chinese Hamster Ovary Cell Line (CHO; ATCC CRL-9606), used in the cell culture experiments, were purchased from Pasteur Institute of Iran. Dulbecco’s modified Eagle’s medium (DMEM) with L-glutamine and RPMI 1640 with l-glutamine were obtained from Biosera (East Sussex, UK). Fetal bovine serum (FBS) was obtained from GIBCO. Penicillin and streptomycin were purchased from Dana Pharmaceutical Co.
All tissue culture flasks and plates were from SPL, South Korea. Sterile pipettes were purchased from ALP (Chorges, France). Filters (0.22 µm) for sterilization of a small amount of solution were from Sartorius (Gottingen, Germany). For sterilization of media for cell culture, 0.22 µm filters were used (CORNING, Japan).
sfGFP-EGF gene design
The DNA coding sequences of sfGFP and EGF were extracted from the Universal Protein Resource (UniProt) database. The EGF gene was linked to the N-terminus of the sfGFP coding sequence by a (GGGGS)3 linker. The restriction sites of SalI and NotI were introduced at the 5´ and 3´ ends of the sfGFP gene, respectively. Furthermore, the restriction site of NcoI was incorporated at the 5´ end of EGF. The designed gene was synthesized by Generay Biotech Co., Ltd., USA.
Construction of vectors
For the construction of a genetic construct harboring pIT2-sfGFP-EGF, the coding sequence of sfGFP-EGF was amplified from the synthesized plasmid using the F1 and R1 primers listed in Table 1. The PCR product was digested using NcoI and NotI restriction enzymes and inserted into the pIT2 genome cut with the same enzymes. The pIT2 vector containing sfGFP was constructed by amplifying the coding sequence of sfGFP using F1 and R1 primers (Table 1), followed by PCR product digestion using SalI and NotI restriction enzymes and insertion into the pIT2 vector. For the construction of the pIT2-EGF vector, PCR was performed to introduce the NotI restriction site at the 3′ end of the EGF sequence while amplifying the coding sequence using the plasmid synthesized by the F2 and R2 primers (Table 1). The PCR product was digested using NcoI and NotI restriction enzymes and inserted into the pIT2 vector cut with the same enzymes. To prove the constructed vectors, colony PCRs were performed using pIT-F and pIT-R primers according to the following conditions: initial denaturation at 95 °C for 3 min, 35 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 1 min, extension at 72 °C for 1 min and final extension at 72 °C for 8 min. The positive clones were sent out for sequencing.
Expression and purification of recombinant proteins
The pIT2 constructs containing EGF, sfGFP-EGF, and sfGFP with a 6 × His tag were transformed into E. coli Origami B (DE3) competent cells (F–ompT hsdSB(rB–mB–) gal dcm lacY1 aphC (DE3) gor522::Tn10 trxB (KanR, TetR) under a standard procedure. LB medium supplemented with 1% glucose and 100 µg mL−1 ampicillin was inoculated with the transformed bacteria and incubated overnight at 37 °C. The overnight culture was diluted at a ratio of 1:50 in 2 × TY medium supplemented with 100 µg mL−1 ampicillin and incubated at 37 °C with vigorous shaking until an OD600 of 0.9. Isopropyl β-D-1-thiogalactopyranoside (IPTG) was added to the culture at a final concentration of 1 mM and incubated overnight at 25 °C with shaking at 150 rpm. The next day, the culture was centrifuged at 5000 rpm for 10 min, and the harvested bacterial pellet was lysed using lysis buffer containing 50 mM Tris, 100 mM NaCl, 0.1% β-mercaptoethanol, 0.1 mg/mL lysozyme, and 1.4 mM PMSF. Subsequently, the lysate was freeze‒thawed three times and sonicated on ice five times for 30 s with 30-s pauses. The lysate was centrifuged at 10,000 rpm at 4 °C for 20 min. Then, the supernatant was subjected to a Ni-Sepharose affinity column and incubated for 1 h at 4 °C with gentle rocking. The flowthrough was removed, and the column was washed thoroughly with TBS buffer containing 20 mM imidazole. After several washing steps, elution buffer (pH 7.4) containing 20 mM sodium phosphate, 500 mM NaCl, and 500 mM imidazole was added to the column and incubated for 1 h. The procedure was repeated one more time, and the obtained samples were dialyzed in TBS buffer. Then, the samples were analyzed by SDS‒PAGE. The Bradford assay was used for the determination of protein concentrations during the production and purification processes.
Western blotting
Western blotting was carried out to detect the expressed EGF, sfGFP, and sfGFP-EGF proteins. Purified proteins were denatured and loaded on a 12% SDS‒PAGE gel. Afterward, the proteins were transferred to PVDF (polyvinylidene fluoride) membranes. The membrane was blocked with buffer supplemented with 5% skim milk overnight at 4 °C and then incubated sequentially with anti-His (with a ratio of 1:3000) and goat anti-mouse HRP-conjugated antibodies (with a ratio of 1:9000) as primary and secondary antibodies in TBS buffer supplemented with 3% skim milk, respectively. TBS with tween (0.1%) was used for washing between different steps. A BM chemiluminescence western blotting kit was used to visualize the protein bands.
Phage amplification
The pIT2 vectors containing EGF, sfGFP-EGF, and sfGFP coding genes were transformed into the E. coli DH5α strain, and the bacteria were cultured in 10 mL of 2 × TY overnight at 37 °C. The overnight culture was diluted in 2 × TY medium at a ratio of 1:100 and further incubated at 37 °C with vigorous shaking until an OD600 of 0.4 was reached. From the culture, 10 mL was taken, and 5.00E + 10 KM13 helper phage was added. After incubation at 37 °C in a water bath for 30 min, the mixture was pelleted by centrifugation at 5500 rpm for 10 min and resuspended in 50 mL 2 × TY containing 50 µg mL−1 kanamycin, 100 µg mL−1 ampicillin and 0.1% glucose. After overnight incubation at 30 °C with vigorous shaking, the overnight bacterial culture was centrifuged at 6000 rpm min, and the supernatant was PEG precipitated twice by adding 20% volume of 20% PEG 8000 in 2.5 M NaCl on ice. Phage particles were centrifuged and resuspended in a final volume of 0.5 mL PBS.
Titration of the amplified phage was carried out by preparing tenfold serial dilutions and adding 10 µL from each to 90 µL of ER2738 culture at OD600 0.4. After incubation for 30 min at 37 °C, the infected bacteria were spread on LB plates supplemented with 100 µg mL−1 ampicillin and 1% glucose and incubated overnight. The plates with countable well-isolated colonies were used to determine the titer of the amplified phage.
Fluorescence spectroscopy
Fluorescence experiments were performed using a Jasco FP-750 spectrofluorometer at 25 °C equipped with an external thermal controller. The fluorescence spectra were measured in a 3-mm quartz cuvette with slit widths of 5 and 10 nm for emission and excitation beams, respectively. From the phage displaying EGF, sfGFP, and sfGFP-EGF, 1.00E + 12 particles/mL were excited at 480 nm, and the emission spectra were recorded between 500 and 560 nm.
Flow cytometry
To analyze whether the sfGFP-EGF proteins and phage particles bind to EGFR cell surface receptors, 1.00E + 05 cells were grown and seeded in a 6-well plate in DMEM with 10% FBS and incubated at 37 °C in a humidified atmosphere containing 5% CO2 to achieve 70% confluency. Then, the cells were serum-starved for 4 h and treated with 1.00E + 12 phage displaying sfGFP-EGF, EGF, and sfGFP as well as sfGFP-EGF and sfGFPs. The cells were incubated at 4 °C for 30 min with samples and then washed 6 × 5 min with cold PBS. After the final wash, the cells were placed at 37 °C for 15 min to allow protein and phage internalization. Subsequently, the cells were detached by using trypsin/EDTA and resuspended in 0.5 mL PBS. Fluorescence measurements were performed using fluorescence-activated cell sorting (FACS). Living cells were confirmed and gated according to the forward scatter/side scatter (FSC/SSC) plot.
Cellular imaging and fluorescence microscopy
To visualize the binding of sfGFP-EGF-displaying phages and proteins to EGFR, fluorescence microscopy was used. A-431 cells were grown in DMEM supplemented with 10% FBS, seeded in a sterile 6-well plate (1.00E + 05 cells/well) and incubated until 70% confluency. After serum starvation for 4 h, the medium was aspirated, and the cells were washed two times with PBS. Then, the cells were exposed to 1.00E + 12 sfGFP-EGF and sfGFP-displaying phages and 20 ng sfGFP-EGF and sfGFPs prepared in cold PBS buffer. After a 1 h incubation at 4 °C, the cells were washed six times with cold PBS and fixed using 4% formaldehyde prepared in PBS for 20 min at room temperature without shaking. Next, the formaldehyde solution was removed gently, and the cells were washed three times with 1 mL of PBS. After the final wash, fluorescence microscopy was used for cellular imaging.
Cell-based ELISA
To investigate the binding affinity of the produced phage to EGFR, a cell-based ELISA was carried out. To do so, the phage particles displaying sfGFP-EGF and sfGFP were amplified and titered. Afterward, A-431 and CHO cells were cultivated in DMEM with 10% FBS at 37 °C in a humidified atmosphere containing 5% CO2. Cells were harvested and seeded in a sterile 96-well plate (7.00E + 04 cells/well) and incubated until 70% confluency. Then, the medium was removed gently, and the cells were washed two times with PBS. Subsequently, the cells were incubated with phages displaying 1.00E + 12 sfGFP and sfGFP-EGF for 30 min.
Afterward, the cells were fixed with 4% formaldehyde in PBS for 20 min at room temperature without shaking. In the next step, the formaldehyde solution was removed, and the cells were washed three times with PBS and blocked with blocking buffer (PBS supplemented with 2% BSA) for 1 h with rocking. Subsequently, the cells were washed three times with PBS, and then 100 µL of 1:5000 diluted HRP-conjugated anti-M13 monoclonal antibody in PBS was added to each well, and the plate was incubated for 1.5 h at room temperature with gentle shaking. After washing four times with PBS, the wells were treated with a solution containing 100 µg mL−1 TMB prepared in potassium acetate (100 mM, pH 6.0) and hydrogen peroxide (0.006% v/v). The enzymatic reaction was terminated after 15 min using 50 µL of 1 M H2SO4. The absorbance was measured at 450 nm using an ELISA reader.
Results
sfGFP-EGF gene design
In the current study, we aimed to construct an EGF-displaying phage system as a tumor-targeting molecule harboring GFP as a biomarker to provide a suitable detection and targeting vehicle. To do so, the DNA coding for sfGFP-EGF was designed, and after optimization for expression in the E. coli system, it was synthesized by Generay Biotech Co., Ltd.
Construction of vectors
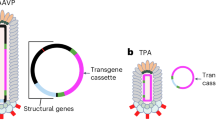
PCR on the ordered DNA sequence using F1 and R2 primers resulted in the amplified EGF DNA sequence with a size of 183 bp. For the amplification of the GFP sequence, F2 and R1 primers were used to obtain a 737 bp DNA sequence. The EGF-GFP sequence (954 bp) was obtained by performing PCR using R1 and F1 primers. Using these DNA fragments, three pIT2-based vectors including EGF, GFP, and EGF-GFP coding genes were constructed according to the method presented in the Materials and Methods section (Fig. 1). These vectors were verified by DNA sequencing and used for conducting the experiments.
Gene maps for the synthesized sequence and the pIT2-based vectors constructed in the current study. Panel a corresponds to the EGF-GFP DNA sequence, linked to each other by a linker (L), and ordered to be synthesized. The primer positions and expected sizes are indicated in the figure. The sequences for the primers are available in Table 1. Panels b, c, and d indicate the constructed pIT2-based vector harboring EGF-GFP, GFP, and EGF DNA sequences
Expression and purification of EGF, sfGFP, and sfGFP-EGF proteins
Three pIT2-based vectors including EGF, GFP, and EGF-GFP coding genes were constructed. EGF, sfGFP, and sfGFP-EGF proteins were expressed using the constructed pIT2-based vectors and purified using a Ni-Sepharose affinity column. Figure 2a demonstrates the purification results for the sfGFP-EGF protein shown as a band at approximately 38 kDa. In Fig. 2b, purified sfGFP is observed at approximately 32 kDa. SDS‒PAGE analysis of purified EGF with a molecular weight of approximately 12 kDa is shown in Fig. 2c. To confirm the expression of EGF, sfGFP, and sfGFP-EGF proteins, western blotting was carried out. Since the purified proteins contain a 6 × His-tag, an anti-His antibody was used for immunoblotting. The results are shown in Fig. 2d, where the unique protein bands at approximately 12 kDa are attributed to EGF, and the protein bands at approximately 32 kDa and 38 kDa represent sfGFP and sfGFP-EGF proteins, respectively.
SDS‒PAGE and western blot analyses of the produced sfGFP-EGF, EGF and sfGFPs proteins. In panel a, the band at approximately 38 kDa represents purified sfGFP-EGF. Panel b is related to the purified sfGFP. The band at approximately 32 kDa shows purified sfGFP from the affinity column. The purified EGF with a molecular weight of ~ 12 kDa is illustrated in Panel (c). Panel d is the western blot analysis of proteins using anti-His primary antibody and goat anti-mouse HRP-conjugated secondary antibody. Unique bands of sfGFP, sfGFP-EGF and EGF are seen at ~ 32 kDa, ~ 38 kDa and ~ 12 kDa in lanes 1, 2, and 3, respectively
Detection of displayed sfGFP-EGF protein on phage surface using fluorescence spectroscopy
To investigate whether the proteins of interest have been correctly folded and displayed on the phage surface, the fluorescence emission of phages displaying EGF, sfGFP, and sfGFP-EGF was measured (Velappan et al. 2010). For this, phage solutions containing 1.00E + 12 phage particles/mL were excited at 480 nm, and the emission spectra were recorded between 500 and 560 nm (Supplementary Data). Phages displaying sfGFP and sfGFP-EGF fluoresce with maximum emission at 510 nm, which can be attributed to the displayed sfGFP. However, the phage displaying only EGF showed no fluorescence emission under the same conditions.
Binding of EGF-displaying phages to cell-surface EGFR
Flow cytometry analysis
To evaluate the binding of phage displaying sfGFP-EGF to EGFR, flow cytometry analysis was performed (Fig. 3). A-431 cells expressing EGFR were allowed to bind EGF fused to sfGFP displayed on the phage surface. Phages displaying sfGFP as well as sfGFP-EGF and sfGFPs proteins were used as controls. The analysis of flow cytometry data showed a significant increase in the fluorescence intensity (right shift) in the cells treated with sfGFP-EGF protein (MFI of 22.58, P < 0.0001) and phage displaying sfGFP-EGF (MFI of 5.17, P < 0.025) compared with the control cells treated with sfGFP protein and phage displaying sfGFP, respectively.
Flow cytometry data of sfGFP florescence in A-431 cells. Panel a shows the flow cytometry histograms of A-431 cells treated with 20 ng sfGFP and sfGFP-EGF proteins for 1 h. The calculated MFI for A-431 cells, which were treated with sfGFP and sfGFP-EGF proteins, is illustrated in Panel b. Flow cytometry histograms of A-431 cells treated with 1.00E + 12 phage particles displaying sfGFP and sfGFP-EGF are shown in Panel (c). Panel d shows the calculated MFI for the A-431 cells treated with phage displaying sfGFP and sfGFP-EGF. All data are the means of triplicate ± SD
Cell imaging and fluorescence microscopy
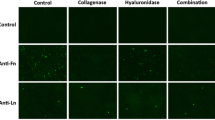
The epidermal carcinoma cell line A-431 was used to visualize the binding of the constructed phage displaying sfGFP-EGF to these EGFR-expressing cells. The cells were grown and, after counting, seeded in a 6-well plate. Then, the cells were serum-starved and treated with phage displaying sfGFP-EGF. The results shown in Fig. 4 indicated that sfGFP-EGF protein and phage displaying sfGFP-EGF can bind to EGFR-expressing A-431 cells and visualize them. In contrast, sfGFP and phage displaying sfGFP were not capable of binding to A-431 cells.
ELISA-based functionality assay
To estimate the affinity of phage displaying sfGFP-EGF protein toward EGFR-expressing cells, a cell-based ELISA was performed using A-431 cells. CHO cells, as non-EGFR-expressing cells, were used as the control (Fig. 5). The ELISA results showed that the binding of phage displaying EGF and sfGFP-EGF to EGFR-expressing A-431 cells was significantly higher than that to non-EGFR-expressing CHO cells (unpaired t test, P < 0.0012 and P < 0.0035, respectively).
Cell-based ELISA experiment using phage displaying EGF and sfGFP-EGF. A-431 and CHO cells were incubated with 1.00E + 12 phage displaying EGF and sfGFP-EGF. Subsequently, anti-M13 HRP-conjugated antibodies were used for phage detection. Each data point is the average of three independent experiments. The error bars represent the standard deviation (SD)
Discussion
EGFR, also known as the ERBB1 or HER1 receptor, belongs to the receptor tyrosine kinase (RTK) family of proteins. Constitutive tyrosine kinase activity has been confirmed in several solid tumors, including lung, liver, breast, and bladder cancers and glioblastoma. This makes EGFR an important target for cancer drug development and gene delivery (Dutta et al. 2007; Liang et al. 2003; Sebastian et al. 2006; Wujcik 2006). Pharmaceuticals that are currently available in the market for inhibiting EGFR-mediated signaling pathways are divided into two classes: (i) monoclonal antibodies that bind to the extracellular component of EGFR, preventing EGF binding to the receptor, and (ii) tyrosine kinase inhibitors that bind to the tyrosine kinase domain of EGFR, suppressing its activity. Cetuximab (C225, Erbitux®) and panitumumab (ABX-Vectibix®) are anti-EGFR antibodies, while gefitinib (ZD1839, Iressa®), erlotinib (OSI-774, Tarceva “) and lapatinib (GW572016, Tykerb/Tyverb®) are examples of approved tyrosine kinase inhibitors (Jimeno and Hidalgo 2005; Wheeler et al. 2010; Bratkovic 2010).
One of the approaches recently used for cancer diagnosis and treatment is the application of hybrid vectors (Hajitou et al. 2006), in which cancer-targeting proteins and biomarkers are displayed on a vector such as a bacteriophage (Khalili et al. 2018; Zhu et al. 2019). In recent years, several researchers have established vectors based on bacteriophage structure (Kia et al. 2012; Willats 2002; Xu et al. 2017; Yata et al. 2014). A bacteriophage is a virus that selectively infects the bacterial host and can be used for designing novel gene carriers. Contrary to other viruses, bacteriophage has received great attention as a novel vehicle for gene delivery due to its lack of tropism for mammalian cells, high safety profile, and the possibility of being manipulated genetically and chemically to improve the efficiency of gene transfer (Clark and March 2006; Dabrowska et al. 2005). Furthermore, modification of phage DNA that carries the mammalian gene expression cassette can be useful for targeting eukaryotic cells. In a study, Larocca et al. developed a phage displaying FGF2 that targeted mammalian cells (Larocca et al. 2001, 2002). In the current study, a phage displaying GFP-EGF was developed based on the pIII display method to evaluate the possibility of EGFR recognition on A-431 cancer cells by EGF as a targeting agent and GFP as the reporting molecule. Successful application of EGF as the targeting agent for the delivery of M13 phages coding siRNA against focal adhesion kinase (FAK) to H1299 lung carcinoma cells has been reported elsewhere (Cai et al. 2008), indicating the feasibility of the designed system in the current study for detection and gene delivery to EGFR-expressing cancer cells. Bacterial secretion systems involve cell membrane proteins that contribute to the secretion of substances such as virulence factors in the environment. In biological research, general secretion (Sec) is a frequently used bacterial secretory pathway that is important for proteins that need an oxidizing environment for disulfide bridge formation and proper folding (Green and Mecsas 2016; Lycklama and Driessen 2012). In this work, the pIT2 phagemid includes the PelB leader, which is a kind of Sec secretory pathway that enables translocation of translated proteins to the periplasm. Here, GFP-EGF protein was expressed as a protein fused to phage pIII protein. After protein synthesis in the cytoplasm, it translocates into the periplasmic space for phage assembly and release. We designed a phage-displaying construct based on a superfolder variant of GFP (sfGFP). Pédelacq et al. developed sfGFP, which contains mutations of both cycle-3 GFP, enhanced GFP (EGFP), and six new mutations that improved its folding kinetics and stability (Pedelacq et al. 2006). This FP shows an improved maturation rate after production in the cytoplasm and retains its fluorescent features when it is translocated to the periplasm via bacterial posttranslational Sec-mediated transport (Velappan et al. 2010; Aronson et al. 2011; Dinh and Bernhardt 2011). In the current study, the pIT2 phagemid was included with the sfGFP-EGF gene and SRP signal peptide with a flexible (GGGGS)3 linker inserted between EGF and sfGFP to allow individual folding of each protein and prevent steric hindrance in ligand‒receptor interactions. The sfGFP-EGF DNA coding sequence was amplified and cloned into pIT2 using HindIII and NotI restriction enzymes to produce the pIT2 phagemid containing the sfGFP-EGF gene at the N-terminus of the pIII coat protein. The constructed vector was transformed into E. coli origami to explore the expression pattern of EGF-sfGFP. However, following the routine protein expression and purification mentioned in the Materials and Methods section did not result in correctly folded and fluorescent EGF-sfGFP. This problem was solved by replacing the SRP signal peptide with the PelB leader, which resulted in the production of a fluorescent sfGFP-EGF fusion protein. The produced protein was purified by a Ni-Sepharose affinity column and analyzed by SDS‒PAGE, western blotting, and fluorescence spectroscopy to confirm the successful expression of sfGFP-EGF in the PelB secretory system (Figs. 1 and 2). The same method was used for the production of sfGFP and EGF proteins. These verified vectors were amplified and used in the experiments.
To evaluate the EGFR binding and detection ability of the constructed phage displaying sfGFP-EGF, flow cytometry analysis was performed (Fig. 3). For this, A-431 cells were treated with phage displaying sfGFP-EGF, phage displaying sfGFP, as well as sfGFP and sfGFP-EGF fusion proteins. The results showed significant intracellular fluorescence accumulation in A-431 cells treated with sfGFP-EGF protein and phage displaying sfGFP-EGF (Fig. 3) compared to the cells treated with sfGFP protein and phage displaying sfGFP. Accordingly, it can be concluded that the constructed phage was able to specifically bind to cell surface EGFR. Further selectivity and binding ability evaluation of the produced phage displaying sfGFP-EGF were carried out by fluorescence microscopy, where A-431 cells were treated with the phage as well as a control phage displaying only sfGFP. The fluorescence emitted from cells was captured using a fluorescence microscope, indicating that the constructed phage displaying sfGFP-EGF can recognize the EGF receptors on the A-431 cell surface (Fig. 4). Although the cells treated with phage displaying sfGFP-EGF were less fluorescent than cells treated with sfGFP-EGF protein, the observed signal (fluorescent intensity) was more pronounced compared to the negative controls. The lower fluorescence intensity of phage displaying sfGFP-EGF compared to sfGFP-EGF protein can be attributed to the low number of sfGFP-EGF-pIII coat proteins on the phage surface. Phagemid pIT2 only possesses a DNA sequence corresponding to the pIII coat protein, and other proteins needed for its assembly and packaging are provided by a helper phage. In the process of phage assembly, three to five pIII proteins are displayed on the phagemid surface. These pIII proteins can be wild-type pIII proteins provided by helper phage or recombinant pIII fused to sfGFP-EGF originating from the pIT2 construct. There might even be phage without recombinant pIII protein. This may lead to the production of phage particles with a low number of pIII coat proteins bearing sfGFP-EGF and result in low fluorescence intensity compared to the soluble form of sfGFP-EGF protein in the fluorescence imaging experiment. Moreover, to test the binding ability and specificity of the derived phage displaying sfGFP-EGF, a cell-based ELISA experiment was carried out using EGFR-overexpressing cells, while CHO cells were used as the negative control. The results showed that the constructed phage was able to specifically bind EGF receptors on the A-431 cell surface. As shown in Fig. 5, phage displaying sfGFP-EGF binds less tightly to EGFR-expressing cells than phage displaying only EGF. This lower binding may be attributed to the steric hindrance imposed by GFP on EGF binding to EGFR. As the molecular weight of a protein increases, its possibility of being displayed on a phage decreases (Imai et al. 2008). This may also contribute to the observed lower binding of phage displaying sfGFP-EGF to A-431 cells. In most similar studies, the reporting agent has been chemically fused to phages; therefore, their fluorescence yields are high. For example, HK97 phage was chemically equipped with a labeling dye (fluorescein) and a targeting moiety (Tf) for the diagnosis and therapy of Tf receptor-overexpressing tumor cells (Huang et al. 2011). Additionally, an anti-prostate-specific membrane antigen (PSMA) antibody fused with the gp3 protein of M13 phages and near infrared (NIR)-fluorescent single-walled carbon nanotubes (SWNTs) linked to the pVIII protein of M13 phages were successfully employed as imaging agents of prostate cancer cells in mice (Yi et al. 2012). Although the fluorescence yield of expressed GFP on phage in the current study was lower than that of chemically fused reporting agents in other studies, the process of production of phage needed no extra chemical reactions.
Conclusion
The relationship between EGFR dysregulation and the development of epithelial-derived cancers makes EGFR an appropriate target for gene therapy. In the current study, a novel phage-mediated system for targeting EGFR in mammalian cells was developed. The recognition and specificity of the developed vector toward EGFR were elucidated using different methods, such as flow cytometry and ELISA experiments. Additionally, EGFR-overexpressing cell detection was performed using a fluorescence-based imaging system. We believe that the vector constructed in the current study has the potential to be engineered for gene delivery purposes as well as tumor detection.
References
Aronson DE, Costantini LM, Snapp EL (2011) Superfolder GFP is fluorescent in oxidizing environments when targeted via the Sec translocon. Traffic (Copenhagen, Denmark) 12:543–548. https://doi.org/10.1111/j.1600-0854.2011.01168.x
Azzazy HM, Highsmith WE Jr (2002) Phage display technology: clinical applications and recent innovations. Clin Biochem 35:425–445
Bratkovic T (2010) Progress in phage display: evolution of the technique and its application. Cell Mol Life Sci 67:749–767
Cai X-M, Xie H-L, Liu M-Z, Zha X-L (2008) Inhibition of cell growth and invasion by epidermal growth factor-targeted phagemid particles carrying siRNA against focal adhesion kinase in the presence of hydroxycamptothecin. BMC Biotechnol 8:74. https://doi.org/10.1186/1472-6750-8-74
Clark JR, March JB (2006) Bacteriophages and biotechnology: vaccines, gene therapy and antibacterials. Trends Biotechnol 24:212–218. https://doi.org/10.1016/j.tibtech.2006.03.003
Dabrowska K, Switala-Jelen K, Opolski A, Weber-Dabrowska B, Gorski A (2005) Bacteriophage penetration in vertebrates. J Appl Microbiol 98:7–13. https://doi.org/10.1111/j.1365-2672.2004.02422.x
Dani M (2001) Peptide display libraries: design and construction. J Recept Signal Transduct Res 21:469–488. https://doi.org/10.1081/rrs-100107927
Dawson JP, Bu Z, Lemmon MA (2007) Ligand-induced structural transitions in ErbB receptor extracellular domains. Structure 15:942–954. https://doi.org/10.1016/j.str.2007.06.013
Dinh T, Bernhardt TG (2011) Using superfolder green fluorescent protein for periplasmic protein localization studies. J Bacteriol 193:4984–4987. https://doi.org/10.1128/JB.00315-11
Dutta PR, Maity A (2007) Cellular responses to EGFR inhibitors and their relevance to cancer therapy. Cancer Lett 254:165–177. https://doi.org/10.1016/j.canlet.2007.02.006
Felici F, Luzzago A, Monaci P, Nicosia A, Sollazzo M, Traboni C (1995) Peptide and protein display on the surface of filamentous bacteriophage. Biotechnol Annu Rev 1:149–183
Ferguson KM (2008) Structure-based view of epidermal growth factor receptor regulation. Annu Rev Biophys 37:353–373. https://doi.org/10.1146/annurev.biophys.37.032807.125829
Green ER, Mecsas J (2016) Bacterial secretion systems: an overview. Microbiol Spectr. https://doi.org/10.1128/microbiolspec.VMBF-0012-2015
Hajitou A, Trepel M, Lilley CE, Soghomonyan S, Alauddin MM, Marini FC 3rd, Restel BH, Ozawa MG, Moya CA, Rangel R et al (2006) A hybrid vector for ligand-directed tumor targeting and molecular imaging. Cell 125:385–398. https://doi.org/10.1016/j.cell.2006.02.042
Huang RK, Steinmetz NF, Fu CY, Manchester M, Johnson JE (2011) Transferrin-mediated targeting of bacteriophage HK97 nanoparticles into tumor cells. Nanomedicine (London) 6:55–68. https://doi.org/10.2217/nnm.10.99
Hwang YJ, Myung H (2020) Engineered bacteriophage T7 as a potent anticancer agent in vivo. Front Microbiol 11:491001. https://doi.org/10.3389/fmicb.2020.491001
Hynes NE, Lane HA (2005) ERBB receptors and cancer: the complexity of targeted inhibitors. Nat Rev Cancer 5:341–354. https://doi.org/10.1038/nrc1609
Imai S, Mukai Y, Takeda T, Abe Y, Nagano K, Kamada H, Nakagawa S, Tsunoda S, Tsutsumi Y (2008) Effect of protein properties on display efficiency using the M13 phage display system. Pharmazie 63:760–764
Jain R, Joyce P, Molinete M, Pa H, Su G (2001) Oligomerization of green fluorescent protein in the secretory pathway of endocrine cells. Biochem J. https://doi.org/10.1042/bj3600645
Jimeno A, Hidalgo M (2005) Blockade of epidermal growth factor receptor (EGFR) activity. Crit Rev Oncol Hematol 53:179–192. https://doi.org/10.1016/j.critrevonc.2004.10.005
Jorissen RN, Walker F, Pouliot N, Garrett TP, Ward CW, Burgess AW (2003) Epidermal growth factor receptor: mechanisms of activation and signaling. Exp Cell Res 284:31–53
Khalili S, Rasaee MJ, Bamdad T, Mard-Soltani M, Asadi Ghalehni M, Jahangiri A, Pouriayevali MH, Aghasadeghi MR, Malaei F (2018) A novel molecular design for a hybrid phage-DNA construct against DKK1. Mol Biotechnol 60:833–842. https://doi.org/10.1007/s12033-018-0115-2
Kia A, Przystal JM, Nianiaris N, Mazarakis ND, Mintz PJ, Hajitou A (2012) Dual systemic tumor targeting with ligand-directed phage and Grp78 promoter induces tumor regression. Mol Cancer Ther 11:2566–2577. https://doi.org/10.1158/1535-7163.mct-12-0587
Klein P, Mattoon D, Lemmon MA, Schlessinger J (2004) A structure-based model for ligand binding and dimerization of EGF receptors. Proc Natl Acad Sci USA 101:929–934. https://doi.org/10.1073/pnas.0307285101
Larocca D, Jensen-Pergakes K, Burg MA, Baird A (2001) Receptor-targeted gene delivery using multivalent phagemid particles. Mol Ther 3:476–484. https://doi.org/10.1006/mthe.2001.0284
Larocca D, Burg MA, Jensen-Pergakes K, Ravey EP, Gonzalez AM, Baird A (2002) Evolving phage vectors for cell targeted gene delivery. Curr Pharm Biotechnol 3:45–57
Lemmon MA (2009) Ligand-induced ErbB receptor dimerization. Exp Cell Res 315:638–648. https://doi.org/10.1016/j.yexcr.2008.10.024
Liang K, Ang KK, Milas L, Hunter N, Fan Z (2003) The epidermal growth factor receptor mediates radioresistance. Int J Radiat Oncol Biol Phys 57:246–254
Lycklama ANJA, Driessen AJ (2012) The bacterial Sec-translocase: structure and mechanism. Philos Trans R Soc Lond B 367:1016–1028. https://doi.org/10.1098/rstb.2011.0201
Mali S (2013) Delivery systems for gene therapy. Indian J Hum Genet 19:3–8. https://doi.org/10.4103/0971-6866.112870
Marvin DA, Symmons MF, Straus SK (2014) Structure and assembly of filamentous bacteriophages. Prog Biophys Mol Biol 114:80–122. https://doi.org/10.1016/j.pbiomolbio.2014.02.003
Morgillo F, Lee HY (2005) Resistance to epidermal growth factor receptor-targeted therapy. Drug Resist Updat 8:298–310. https://doi.org/10.1016/j.drup.2005.08.004
Normanno N, De Luca A, Bianco C, Strizzi L, Mancino M, Maiello MR, Carotenuto A, De Feo G, Caponigro F, Salomon DS (2006) Epidermal growth factor receptor (EGFR) signaling in cancer. Gene 366:2–16. https://doi.org/10.1016/j.gene.2005.10.018
Ogiso H, Ishitani R, Nureki O, Fukai S, Yamanaka M, Kim JH, Saito K, Sakamoto A, Inoue M, Shirouzu M et al (2002) Crystal structure of the complex of human epidermal growth factor and receptor extracellular domains. Cell 110:775–787
Pedelacq JD, Cabantous S, Tran T, Terwilliger TC, Waldo GS (2006) Engineering and characterization of a superfolder green fluorescent protein. Nat Biotechnol 24:79–88. https://doi.org/10.1038/nbt1172
Petrov G, Dymova M, Richter V (2022) Bacteriophage-mediated cancer gene therapy. Int J Mol Sci. https://doi.org/10.3390/ijms232214245
Qi H, Lu H, Qiu HJ, Petrenko V, Liu A (2012) Phagemid vectors for phage display: properties, characteristics and construction. J Mol Biol 417:129–143. https://doi.org/10.1016/j.jmb.2012.01.038
Sebastian S, Settleman J, Reshkin SJ, Azzariti A, Bellizzi A, Paradiso A (2006) The complexity of targeting EGFR signaling in cancer: from expression to turnover. Biochim Biophys Acta 1766:120–139. https://doi.org/10.1016/j.bbcan.2006.06.001
Straus SK, Bo HE (2018) Filamentous bacteriophage proteins and assembly. Subcell Biochem 88:261–279. https://doi.org/10.1007/978-981-10-8456-0_12
Sung YK, Kim SW (2019) Recent advances in the development of gene delivery systems. Biomater Res 23:8. https://doi.org/10.1186/s40824-019-0156-z
Velappan N, Fisher HE, Pesavento E, Chasteen L, D’Angelo S, Kiss C, Longmire M, Pavlik P, Bradbury ARM (2010) A comprehensive analysis of filamentous phage display vectors for cytoplasmic proteins: an analysis with different fluorescent proteins. Nucleic Acids Res 38:e22–e22. https://doi.org/10.1093/nar/gkp809
Wang YA, Yu X, Overman S, Tsuboi M, Thomas GJ Jr, Egelman EH (2006) The structure of a filamentous bacteriophage. J Mol Biol 361:209–215. https://doi.org/10.1016/j.jmb.2006.06.027
Wheeler DL, Dunn EF, Harari PM (2010) Understanding resistance to EGFR inhibitors-impact on future treatment strategies. Nat Rev Clin Oncol 7:493–507. https://doi.org/10.1038/nrclinonc.2010.97
Wieduwilt MJ, Moasser MM (2008) The epidermal growth factor receptor family: biology driving targeted therapeutics. Cell Mol Life Sci 65:1566–1584. https://doi.org/10.1007/s00018-008-7440-8
Willats WG (2002) Phage display: practicalities and prospects. Plant Mol Biol 50:837–854
Wujcik D (2006) EGFR as a target: rationale for therapy. Semin Oncol Nurs 22:5–9. https://doi.org/10.1016/j.soncn.2006.01.010
Xu MJ, Johnson DE, Grandis JR (2017) EGFR-targeted therapies in the postgenomic era. Cancer Metastasis Rev 36:463–473. https://doi.org/10.1007/s10555-017-9687-8
Yata T, Lee KY, Dharakul T, Songsivilai S, Bismarck A, Mintz PJ, Hajitou A (2014) Hybrid nanomaterial complexes for advanced phage-guided gene delivery. Mol Ther Nucleic Acids 3:e185. https://doi.org/10.1038/mtna.2014.37
Yi H, Ghosh D, Ham MH, Qi J, Barone PW, Strano MS, Belcher AM (2012) M13 phage-functionalized single-walled carbon nanotubes as nanoprobes for second near-infrared window fluorescence imaging of targeted tumors. Nano Lett 12:1176–1183. https://doi.org/10.1021/nl2031663
Zhu J, Tao P, Mahalingam M, Sha J, Kilgore P, Chopra Ashok K, Rao V (2019) A prokaryotic-eukaryotic hybrid viral vector for delivery of large cargos of genes and proteins into human cells. Science Advances 5:e0064. https://doi.org/10.1126/sciadv.aax0064
Acknowledgements
The work was supported by the Research Office of Tabriz University of Medical Sciences under Grant Number 62787. The authors would like to thank the Biotechnology Research Center of Tabriz University of Medical Sciences for providing financial and facility support.
Funding
The work was supported by Grant Number 62787. Author Siavoush Dastmalchi has received a Grant from the Research Office of Tabriz University of Medical Sciences.
Author information
Authors and Affiliations
Contributions
SD and AAA contributed to the study conception and design. Material preparation, data collection and analysis were performed by MS, AAA and MHM. The first draft of the manuscript was written by AAA, and all authors commented on previous versions of the manuscript. All the authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors have no relevant financial or nonfinancial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sharifi, M., Alizadeh, A.A., Mivehroud, M.H. et al. Construction of a bacteriophage-derived vector with potential applications in targeted drug delivery and cell imaging. Biotechnol Lett 46, 147–159 (2024). https://doi.org/10.1007/s10529-023-03455-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-023-03455-y