Abstract
The Cas9 nuclease initiates double-stranded breaks at the target position in DNA, which are repaired by the intracellular restoration pathways to eliminate or insert pieces of DNA. CRISPR-Cas9 is proficient and cost-effective since cutting is guided by a piece of RNA instead of protein. Emphasis on this technology, in contrast with two recognized genome editing platforms (i.e., ZFNs and TALENs), is provided. This review evaluates the benefits of chemically synthesized gRNAs as well as the integration of chemical amendments to improve gene editing efficiencies. CRISPR is an indispensable means in biological investigations and is now as well transforming varied fields of biotechnology and agriculture. Recent advancement in targetable epigenomic-editing tools allows researchers to dispense direct functional and transcriptional significance to locus-explicit chromatin adjustments encompassing gene regulation and editing. An account of diverse sgRNA design tools is provided, principally on their target competence prediction model, off-target recognition algorithm, and generation of instructive annotations. The modern systems that have been utilized to deliver CRISPR-Cas9 in vivo and in vitro for crop improvement viz. nutritional enhancement, production of drought-tolerant and disease-resistant plants, are also highlighted. The conclusion is focused on upcoming directions, biosafety concerns, and expansive prospects of CRISPR technologies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genome editing is a kind of genetic engineering mechanism wherein DNA is introduced, obliterated, modified or substituted in the genome of a living individual. Homologous recombination is the foundation of genome engineering, but its occurrence at low frequencies limits the editing efficiency (Chen et al. 2019). To improve editing frequency, researchers took over the utility of enzyme endonucleases that intricate to restore DNA double-stranded breaks (DSB). There are various genome altering technologies like zinc finger nuclease (ZFN) and transcription activator-like effector nuclease (TALEN) already been discussed for targeted modifications of the genome (Zhang et al. 2010; Adli 2018).
Clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated9 (Cas9) technology is being widely used to incorporate high specificity and activity, at the preferred target locus. As endonucleases, Cas proteins are known to use a single guide RNA (sgRNA) to make complementary base pairs with target DNA followed by cutting the DNA at explicit sites (Agrotis and Ketteler 2015). Supposedly, using CRISPR a method can be developed to engineer just about any DNA sequence in the genome as it offers flexibility, easy multiplexing, and scaling. Nowadays, its applications have reached a variety of fields, counting biotechnology, biological investigation, human medicinal application, and agricultural research (Veres et al. 2014; Chen et al. 2019).
The guide RNA (gRNA) of the CRISPR-Cas9 system is the RNA element that possibly comprises either the chimeric sgRNA or the dual RNAs (crRNA:tracrRNA) (Arroyo et al. 2016). The gRNAs can be swiftly created by the use of chemical synthesis methods and offer correspondent characteristics and advantages, such as integration of chemical modifications to improve on-target precision, gene editing proficiencies, and genome-scale high throughput range analysis for practical genomic studies (Ryan et al. 2018). The CRISPR-Cas9 genome complex and epigenome editing can be introduced into living cells for precise and dynamic manipulation of an epigenetic state that would facilitate its employment in plants.
Abiotic stresses such as drought, salinity, etc., can decrease crop yield up to 50% (Afzal et al. 2019). CRISPR technology has been used to study some significant drought stress-related genes such as AREB1 and OsSAPK2 in Arabidopsis and rice, respectively (Shinwari et al. 2020). It has been documented that CRISPR is used to manipulate the genome of different plant species, including Arabidopsis, Medicago truncatula, tomato, potato, wheat, corn, rice, and mushroom (Khatodia et al. 2016; Gong et al. 2002; Papikian et al. 2019). Some mainstream problems allied with nucleic acid-based application analysis are off-target effects, ethical concerns, and a need for safe and proficient delivery systems. Although several methods have been developed to detect the off-target mutations such as SITE-seq, Digenome-seq, GUIDE-seq, and DISCOVER-seq, etc. (Wada et al. 2020) yet these major bottlenecks exist in plant system. Hence the emphasis is given on the current modern systems developed to transport and consequently deliver CRISPR in vivo and in vitro for a variety of advantageous applications.
In the present communication, the salient features of the CRISPR-Cas9 system, a comprehensive comparison, as well as chemical synthesis and modifications of the sgRNA elements are discussed. A brief description of the bioinformatics tools used to design sgRNA is also mentioned. Epigenetic changes, regulation mechanisms, and their possible implications in the plants are highlighted. Finally, the focus is laid on possible delivery strategies and genome editing applications in plants.
A comparative mechanism of genome editing by CRISPR-Cas9, ZFNs, and TALENs
Technologies for the introduction of site-specific alterations and amendments in the genome of cells and individuals remain exclusive. Supplementary examples of programmable genome editing machinery consist of TALENs and ZFNs (Fig. 1). TALENs and ZFNs function as dimers and only the protein components are required.
A diagrammatic evaluation of various pliable sequence explicit genome editing nucleases that cleaves adjoining DNA sequences to generate nicks on corresponding strands: A Zinc-finger nucleases (ZFNs) are dimer, with every monomer comprising of DNA binding domain (3–6 zinc finger recurs identifying 9–18 nucleotides) and type II restriction endonuclease Fok1 domain. B Transcription activator-like nucleases (TALENs) are dimers with every subunit consisting of DNA binding domain (conserved, 23–28 amino acid sequence explicit for each nucleotide) and Fok1 nuclease domain. C CRISPR/Cas9: Cas9 naturally evolved, RNA-guided endonuclease directed by sgRNA, (crRNA and tracrRNA) for precise objective cleavage. It recognizes about 20 nucleotide recognition spot upstream of protospacer adjacent motif (PAM) of its DNA target
A ZFN is a heterodimer in which every subunit comprises a zinc finger domain and a FokI endonuclease domain (Urna et al. 2010). Genome editing by ZFNs has been demonstrated in plants, including rice and Arabidopsis (Ainley et al. 2013; Gallego-Bartolome et al. 2019). ZFNs are effectual genome editing elements; however, they were not extensively adopted because of the complexity in nature of the contact between zinc fingers and DNA. Other limitations include the inherent difficulty in designing, interest-dependent specificity, and difficulty in authenticating such proteins for a particular DNA locus of context (Sander et al. 2011).
TALENs are dimeric transcription nucleases or factor built from arrays of 33 to 35 amino acid modules, each one of which is targeted to a single nucleotide. Researchers can easily design TALENs because there is a one-to-one recognition convention among protein repeats and nucleotide sequences; hence it can target nearly any sequence of interest presently by assembling the arrays (Luo et al. 2019). TALENs has been used to edit the genomes of a wide variety of plants, including barley (Budhagatapalli et al. 2015), rice (Shan et al. 2015), soybean (Du et al. 2016), sugarcane (Jung and Altpeter 2016), maize (Char et al. 2015), and potato (Clasen et al. 2016). TALENs were simpler to construct and authenticate, facilitating an inexpensive, faster method of genome editing; however, the difficulties in synthesis, protein designing, and corroboration remained an obstruction to its extensive adoption in genome editing applications.
CRISPR-Cas consists of a single distinct monomeric protein and a chimeric RNA unit. Unlike ZFNs or TALENs, CRISPR-Cas is like a DNA-targeted form of RNA interference. CRISPR-Cas has revolutionized the genome-editing field as it is simple, inexpensive, easily programmed. It is also well efficient, as only 20 nucleotides in the gRNA need to be customized to identify a diverse target. The targeting of endonucleases to a specific locus results in DNA cleavage and induces the cell to undertake homology-directed repair (HDR), microhomology-mediated end joining (MMEJ) or non-homologous end joining (NHEJ). HDR occurs as a repair template-specific desired genomic modification that enables precise editing (Bassett et al. 2013). MMEJ is an error-prone repair system that involves the arrangement of microhomologous sequences internal to broken ends prior to joining and is coupled with insertions and deletions (Yanik et al. 2018). In the case of NHEJ, no DNA repair template is provided, and its error-prone nature often leads to inactivating mutations (Chen et al. 2019) (Fig. 2). Some other repair mechanisms also exist like single-stranded annealing (SSA) pathway of HDR, which requires only a single DNA duplex and uses the repeat sequences as the identical sequences as in HDR (Yanik et al. 2018). A specialized form of MMEJ is known as polymerase theta-mediated end joining (TMEJ) and can repair breaks using ≥ 1 bp of homology (Schimmel et al. 2019).
A diagrammatic representation of Cas9 in genomic editing with endogenous cellular site-specific nucleases: The double stranded breaks (DSBs) generated by CRISPR/Cas9 system be repaired by non-homologous end joining (NHEJ), microhomology-mediated end joining (MMEJ) or homologous recombination (HR) pathways. NHEJ produces random insertions or deletions (indels) of random base pairs as a result of homozygous, heterozygous or biallelic mutations. Diminutive microhomologies (~ 5–25 bp) bordering DSB recombine through MMEJ, resulting in deletion amid homology arms. HDR can produce desired precise nucleotide substitution mutations or indels by homologous recombination guided through donor DNA digested with the identical endonuclease following related overhangs
Chemical synthesis of the guide RNA
The RNA unit of the CRISPR-Cas9 complex can be created enzymatically or via the chemical synthesis process (Helm et al. 1999). Enzymatic synthesis is a cost-effective method, and the process of in vitro transcription requires a DNA template, T7, T3, or SP6RNA polymerases and ribonucleoside triphosphates. A 5′-triphosphate remains on the gRNA after transcription that necessitates elimination by phosphatase enzyme following purification (Cho et al. 2013). Solid-phase synthesis chemistry is used to create synthetic gRNAs. There is greater flexibility in time consumption, yield, length, and higher precision in the synthesized RNAs with no obligation for several cloning and sequencing steps. Chemical production of gRNAs employs amalgamation (solid phase) through nucleoside phosphoramidite structure blocks for constructing gRNA (Kelly et al. 2016). 2′-Silyl (2′-TBDMS, 2′-TOM), 2′-O-thionocarbamate (TC) (Cullot et al. 2019) and 2′-bis(acetoxyethoxy)-methyl ether (2′-ACE) (Scaringe et al. 1998) are some of the RNA synthesis chemistries offered.
Researchers successfully generated gRNA sequences intended for Streptococcus pyogenes Cas9 structures using a chemical synthesis approach (Anderson et al. 2015). A two-RNA approach with a crRNA and tracrRNA to program Cas9 or a sgRNA approach can be used. Conventional chemistries such as 2′-TOM or TBDMS (Pitsch et al. 2001) are capable of synthesizing RNA > 70 bases whereas long RNA (~ 150 nucleotides) are characteristically synthesized employing TC or 2′-ACE chemistries (Cullot et al. 2019). Jinek et al. (2012) reported that in S. pyogenes, the crRNA, tracrRNA, and sgRNA are in the order of 40, 70, and 100 nucleotides in length, respectively. Hence use of 2′-ACE or TC chemistries is ideal for synthesis providing high throughput, greater purity, rapid coupling rates, and higher production than any other RNA chemistries.
The tools used for CRISPR/Cas9 designing
The precision of the CRISPR-Cas system depends on well-designed sgRNA as it is a critical aspect of the successful editing of target genes. The design tools differ in parameters and design specifications, predominantly highlighting the on-target efficacy calculation models and off-target calculation algorithms to ultimately improve sgRNA specificity (Zhu 2015).
Various computational tools have been created to design sgRNA with improved specificity and efficiency. Certain representatives are described (Table 1) that has been developed to assemble information and offer useful purpose to study CRISPR-Cas organization. Wong et al. (2015) developed WU-CRISPR; the program is suggested for its ease of use and proficient sgRNA design using a machine learning technique. The tool recognizes several sequences and structural arrangements from Doench’s dataset (Doench et al. 2014) and constructs a sgRNA effective estimate model with SVM. Chari dataset was used to assess and compare the tool for superlative performance (Chari et al. 2015). The preceding off-target scoring process cannot be quickly contained through organisms, so the researchers wished for a novel procedure to evaluate the off-target action and termed it CASPER (Mendoza and Trinh 2017). The model was derived from Hsu–Zhang matrix and appraised for its off-target activity even in the absence of adequate experimental statistics (Hsu et al. 2013).
Additionally, several other tools are also reported, such as Cas-OFFinder (Baltes et al. 2014), SSFinder (Upadhyay and Sharma 2014), CRISPR-P (Lei et al. 2015), and Cas OT (Xiao et al. 2014) that eases the sgRNA designing process. The assembly of expression vectors and delivery of those vectors in plant systems involves the use of diverse methodologies essential for amplifying editing efficiencies. There is still a great deal to be experimented and optimized for the exploitation of CRISPR-Cas9 in plant systems.
CRISPR mediated epigenetic regulations in plants
Examining the usage of the Cas9 system to inspect regulatory sequences that can transform gene expression through epigenetic mechanisms, and chromatin modifications are the outcomes of the discovery of a versatile RNA-guided DNA-targeting platform (Naito et al. 2015). Canver et al. (2017) showed that the majority of the gRNAs did not affect gene expression regulation when aimed to create indel mutations in recognized enhancer regions. This led to the understanding that only a few critical domains are significant for enhancer function (Oldridge et al. 2015; Maurano et al. 2012). Sumoylation has majorly been linked to transcriptional repression mechanism (Decque et al. 2016). However, its functional roles are focused on proteasome-dependent proteolysis, activation of DNA damage signaling cascades (Wu et al. 2014), cellular localization, and assembly (Deshaies and Joazeiro 2009) fitting in the general principle of the dCas9-KRAS system.
Numerous reports simultaneously established the concept for up- or down-regulation of target genes drawn out by dCas9 fusions to catalytic domains, directing methylation and demethylation of CpG islands that span promoter regions (McDonald et al. 2016; Vojta et al. 2016). Similarly, evidence of the production of synergistic effects leading to increased methylation of the promoter by targeting multiple promoter sites and by co-expression of diverse sgRNAs is provided. The study was demonstrated using a dCas9-DNMT3A fusion (with a Gly4Ser flexible linker). Which showed competent and precise CpG island methylation of the BACH2 and IL6ST promoters (Vojta et al. 2016). In a Transgenic Arabidopsis plant overexpression of a chromatin remodeling gene AtCHR12 shows evidence of growth termination in stem and bud. On the contrary, the response under unfavorable conditions was fewer in the AtCHR12-knockout mutant than in the wild type plants (Mlynarova et al. 2007). The current advancement in understanding various epigenetic control mechanisms and in developing effective and flexible tools to study these procedures makes it easy to exploit it for crop management and improvement (Fig. 3).
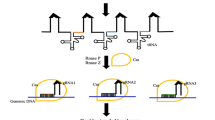
Method of CRISPR/Cas9 action and epigenetic manipulation based on the probability to allocate chromatin modifiers: CRISPR loci after incorporation of foreign DNA is transcribed into prime transcript and progressed into crRNA by aid of tracrRNA, later Cas9 intricate with a crRNA, cleaves foreign DNA. a Targeted relocation of transcriptional regulator-enzymes accountable for modification in the DNA methylation; DNMT, DNA methyltransferase; TET, ten-eleven translocation enzymes. b Targeted relocation of transcriptional regulator-histone modifiers; HDM, histone demethylase; HAT, histone acetyltransferase; HMT, histone methyltransferase; HDAC, histone deacyetylase; HUbq, histone ubiquitin ligase. c Gene knockout modification
Gene/ genome editing applications using guide RNA for crop plant improvement
Genome editing by CRISPR is adaptable to edit any gene in any monocot or dicot plant species. CRISPR-Cas9 has already been used to improve tolerance to biotic pathogens (fungal, viral or bacterial), or abiotic stresses (cold, heat, drought, salt), enhance metabolic pathways, improve nutritional value, grain quality, increase shelf life, obtain haploid seeds, and upsurge agricultural yield (Wang et al. 2014, 2016). Decline of phytic acid content in maize (Liang et al. 2008) and the formation of acrylamide free potatoes (Halterman et al. 2016) has also been reported. Representative applications of CRISPR-Cas in plant improvement have been discussed in the subsequent section (Table 2).
In dicotyledons
Characteristic and evident research work has been done for the production of non-browning apples, and potatoes employing Polyphenol oxidase (PPO) gene mutant (Halterman et al. 2016). In a recent study, Ortigosa et al. (2018) reported the creation of a tomato variety resistant to the bacterial speck disease caused by Pseudomonas syringae pv. tomato (PtoDC3000) without reducing resistance to necrotrophs. The functional ortholog of AtJAZ2 in tomato favorably aggregates in stomata showing that SlJAZ2 is a key co-receptor of coronatine (COR) in stomatal guard cells. Using CRISPR-Cas9 SlJAZ2 was modified to create dominant JAZ2 repressors that lacked the C-terminal Jas domain (SlJAZ2Δjas) and disallowed stomatal reopening by COR providing resistance to PtoDC3000. Furthermore, it also established a novel CRISPR-Cas-built tactic for crop protection that could be employed in the field.
Li et al. (2017a) worked on a Chinese herb Salvia miltiorrhiza with documented vasorelaxation and antiarrhythmic properties. The researchers targeted the diterpene synthase gene (SmCPS1) concerned in the biosynthesis of tanshinone that utilizes geranylgeranyl diphosphate (GGPP) as substrate. The tanshinone biosynthesis metabolic flux was switched to the taxol synthesis pathway by using SmCPS1 knockout (post-GGPP synthesis step) mutants as GGPP is also a substrate for taxol biosynthesis. Three homozygous mutants with zero tanshinone accumulation and a decreased proportion of eight chimeric mutants were produced. Using CRISPR/Cas9-Agrobacterium rhizogenes mediated alteration from twenty-six independent transgenic hairy root lines of Salvia. Malzahn et al. (2019) demonstrated CRISPR-Cas12a mediated genome editing in two target genes (TT4 and GL2) in transgenic Arabidopsis. Cas12a was also used for targeted genome editing in Nicotiana benthamiana, Solanum lycopersicum, and Arabidopsis thaliana (Bernabé‐Orts et al. 2019).
In an experiment, researchers demonstrated Cpf1-mediated gene targeting in protoplasts isolated from wild tobacco and soybean. The result led to effective mutational induction in AOC in wild tobacco and FAD2 paralogues in soybean (Kim et al. 2017). In Solanum tuberosum CRISPR-Cas9 was used to knock out the gene encoding granule-bound starch synthase (GBSS) by a single transfection. It resulted in the generation of amylopectin producing potato, which is an extremely required marketable trait (Andersson et al. 2017). An experimental study on CRISPR-Cas9 targeted modification in Citrus sinensis for disease resistance against Xanthomonas citri causing citrus canker was conducted. Deletion of the intact EBEPthA4 sequence series from susceptibility gene Lateral organ boundaries 1 (CsLOB1) alleles examined the intensity of resistance to wanjincheng orange as CsLOB1 promoter augments disease resistance (Peng et al. 2017). Chandrasekaran et al. (2016) created non-transgenic homozygotic mutant cucumber plants that were resistant to several viruses such as cucumber vein yellowing virus, papaya ringspot mosaic virus, etc. The researchers inactivated elF4E (eukaryotic translation initiation factor gene) using the CRISPR-Cas9 system.
In monocotyledons
Li et al. (2016) evidenced that multiple regulators of significant traits can be edited in a single rice cultivar Zhonghua 11 by CRISPR-Cas9. They used the CRISPR system to mutate the genes controlling grain number, grain size, panicle, and plant architecture, i.e., Gn1a, GS3, DEP1, and IPA1, respectively. The second generation of the gn1a, dep1, and gs3 mutants showed a higher grain number, dense, panicles, and large grain size, respectively. Besides, semi-dwarf and grain with a lengthy-awn phenotype were also detected in dep1 and gs3 mutants, correspondingly. The ipa1 mutants presented two distinct phenotypes, having either fewer or more tillers. Such studies facilitate the separation of complex gene regulatory systems in the same genomic background and the assembling of vital traits in cultivated varieties. Another study was conducted in rice plants using three engineered gRNAs with a 20–22 nucleotide seed region customized to pair with distinctive rice genomic locations. The experimental analysis led to the conclusion that the mismatch site involving target DNA and gRNA seed is a substantial determinant of the Cas9 targeting exactitude. The resulting mutational proficiency of the target site was expected to be 3–8% (Khlestkina and Shumny 2016).
Lawrenson et al. (2015) targeted two copies of HvPM19 using Cas9 genome editing in barley (Hordeum vulgare). The researchers observed Cas9-induced mutations in the first generation of the lines. Wang et al. (2014) have utilized CRISPR-Cas9 technology to generate transgenic Triticum aestivum plants conferring resistance to powdery mildew. This report has provided a methodological framework to improve polyploid crops. The researchers have showed that TaMLO-A1 allele (TALEN-induced mutation in MILDEW-RESISTANCE LOCUS (MLO) proteins) in barley plant confers heritable broad-spectrum resistance to powdery mildew. Zhou et al. (2014) reported large chromosomal segment deletions (115–245 kb) induced by Cas9 as well as the inheritance of genome edits in multiple generations, by targeting four sugar efflux transporter (OsSWEET) genes in rice. Up to 87–100% editing efficiency was observed in T0 transgenic plants, all with di-allelic edits.
Wang et al. (2016) indicated that gene modification via CRISPR-Cas9 is a useful approach for enhancing blast resistance in rice. The researchers reported the improvement of rice blast resistance by targeting the OsERF922 gene in rice. Among 50 T0 transgenic plants twenty-one mutant plants were identified, and several Indel mutations at the target site were revealed by Sanger sequencing. Moreover, six second generation homozygous mutant lines were additionally studied for a blast resistance phenotype and various agronomic traits viz. plant height, panicle length, number of grains per panicle, flag leaf length, and width etc. It was also observed that the number of blast lesions formed after pathogen infection was decreased in mutant lines as compared to wild-type plants. Some other noteworthy reports include the enhanced resistance to herbicides (Endo et al. 2016) and thermosensitive genic male sterility in maize and wheat (Li et al. 2017b; Okada et al. 2019). Producing genetic resistance to viruses has huge potential to manage diseases for which no natural resistance has been reported, such as maize lethal necrosis disease (Luria et al. 2017). These results infer advance aspects in molecular breeding to enhance plant function utilizing optimized CRISPR/Cas9-plant systems.
Delivery strategies with special emphasis on plants
One of the critical challenges in targeting cells in plant systems is the secure and competent transfer of CRISPR-Cas9 genome-editing complex (Joung et al. 2017). Hence, an emphasis is given on the modern systems developed to transport CRISPR-Cas9 in-vivo and in-vitro (Han et al. 2017). Genome editing using CRISPR-Cas9 is performed by three strategies. The primary and foremost approach utilizes a simple and suitable plasmid-based CRISPR-Cas system, programming the Cas protein with sgRNA from the identical vector (Ran et al. 2013). Cas9 protein can be delivered using electroporation, microinjection, and lipid nanoparticle strategies (Qin et al. 2015). The next approach is based on carrying the fusion of the sgRNA and Cas9 mRNA. It offers improvements in off-target effects and limits the time of gene-editing (Niu et al. 2014). Delivery strategies such as microinjection, electroporation, and lipid nanoparticles (Zuris et al. 2015) can be classified under this strategy. The third approach is based on delivering the combination of the Cas protein and the sgRNA. It is used widely due to several advantages such as elevated editing efficiency, quick action, and no requirement of promoter choice or codon optimization (Kim et al. 2014). This combination of Cas protein and sgRNA can be delivered using electroporation, cell-penetrating peptide (CPP), and gold nanoparticles (Zuris et al. 2015).
Physical delivery approaches
Physical delivery strategies employ temporary disruption of physical barriers and allow cargo to reach its targeted location. Electroporation is an extensively used strategy, and it offers high transfection efficiency and usage in the in-vitro and in-vivo analysis (Tebas et al. 2014). The inadequacy of electroporation is that the plasmid DNA is barely assimilated into approximately 0.01% of target cells. Moreover, it induces substantial cell death and also leads to nonspecific transfection. Microinjection is another physical delivery approach where cargoes are injected to the target site using a 0.5–5.0 µm diameter needle (Raveux et al. 2017). Protoplast transformation has been confirmed advantageous for the evaluation of the efficiency of CRISPR/Cas9 designs where plasmids can be delivered into protoplasts using electroporation and microinjection (Malnoy et al. 2016). Using particle bombardment technique that offers high transformation efficiency researchers have succeeded in delivering exotic DNA into scutellar tissues of maize, epidermal tissues of Allium cepa, and leaf and cell culture of several other crops (Maggio et al. 2014).
Non-viral delivery approaches
The non-viral vectors offer advantages of availability, safety, lack of size limitation, and cost-effectiveness (Glass et al. 2018). Agrobacterium-mediated plant transformation is an extremely multifaceted, evolved, and widely used method that utilizes genetic determinants of bacterium and host plant cells mutually (Gelvin 2003). Vector ZH11 was transformed via A. tumefaciens-mediated callus transformation. Additionally, A. rhizogenes mediated-hairy roots are an excellent transformation model system for species of fabaceae. The transient assay can be implemented to test the CRISPR genome editing ability (Hiei et al. 1994). PEG mediated transformation is a simple reproducible and highly competent strategy for the transformation of plant protoplasts (Liu and Vidali 2011). Nanoparticles composed of mesoporous silica (Cunningham et al. 2018), gold, layered double hydroxides (Mitter et al. 2017), and polyethylenemine (Cunningham et al. 2018) are widely used as carriers. Carbon nanotubes have been used as a delivery vehicle to transfer DNA for successful protein expression in mature plant leaves (Demirer et al. 2018). The commonly used explants in plant transformations include calli, i.e., unorganized cell mass (monocots and eudicot), leaf cuttings (eudicot), and zygotic embryos (monocots) (Ikeuchi et al. 2016).
Viral delivery approaches
Despite the safety distress and the chances of introduction of undesirable mutations, viral delivery systems are the most proficient method to carry plasmid-based nucleic acids to cells in the in-vitro and in-vivo analysis (Koike-Yusa et al. 2014). Virus mediated genome editing has been reported in both inoculated and non-inoculated leaves. In a recent report, the authors developed a tobacco rattle RNA virus-mediated genome transduction method for N. benthamiana (Mahas et al. 2019). Bean yellow dwarf virus, begomovirus, cabbage leaf curl virus, and wheat dwarf virus are some of the most widely used DNA viruses for gene transduction. Bean yellow dwarf virus has been used to target stALS1, stALS2 (Solanum tuberosum acetolactate synthase1) and P-GUS:NPTII (Promoter of GUS and neomycin phosphotransferase) gene in S. tuberosum and Nicotiana tabacum respectively (Butler et al. 2015; Baltes et al. 2014). The wheat dwarf virus has also been used as a viral vector to target Ubi, MLO, GFP (β-glucuronidase [GUS] reporter controlling gene, MildewLocusO, green fluorescent protein) in T. aestivum (Gil-Humanes et al. 2017). These methods provide several advantages such as immense infection efficiency, broad cell tropism, and long-term gene expression (Zaidi and Mansoor 2017). However, there are disadvantages of difficulty in production, limited packaging size, and potential for insertional mutagenesis.
Conclusion and future prospective
It is improbable for traditional plant breeding to meet the growing food demands as well as other ecological challenges. On the contrary, CRISPR-Cas technology is removing genome editing barriers and has the potential to revolutionize plant breeding. What has been achieved so far with this technology is just the tip of the iceberg. The CRISPR system can be used for several futuristic applications in plant systems, such as for studying abiotic stress responses or adaptation pathways. Likewise, activation or suppression of genes can be regulated by utilizing CRISPR as a binding tool to stimulate repressors or activators to induce traits. CRISPR also has the capability for gene shuffling, i.e., assembling desirable traits in the genome that would group even in traditional crossbreeding. This technology will allow the emerging genomic and systems biological data to be exploited more comprehensively in gene discovery as well as novel trait development in countless plant species. CRISPR has been used for improved screening for genes and traits in human health via guide molecule libraries. This could be potentially used in plants to screen for characters contributing to crop yield, pest, and disease resistance. Application in orthogonal gene targeting is another aspect which is so far not been tested in plant systems. Hence, it is crucial to present parallel studies in plants to guarantee the adaptability to different species.
Bioinformatic gRNA design tools can be used to increase efficiency and decrease off-target effects. The tools depend on the activity prediction models and off-target detection algorithms; therefore, there is a need for additional CRISPR-Cas datasets for the development of new design tools. A substantial bottleneck to the implementation of CRISPR tools in agriculture is the effective packaging and delivery of CRISPR-Cas complex to the targeted plant cells. Novel delivery methods need to be established to achieve high-efficiency genome editing in plants. Thus, the outlook for improvement in reducing the size of presented Cas proteins or the innovation of smaller Cas9 proteins is needed.
Genome editing is a promising technology with the ability to contribute to food generation for the use of the rising population. However, the biosafety, social and ethical concerns remain about the usage of genome editing in plants. The major concern is the risk of creating undesirable genetic changes in plants due to off-target mutations. Fragments of the CRISPR-Cas9 might be inserted into expected or unexpected sites during the DNA repair mechanism or degraded into filler DNA. Substantial work is being required including improving gRNA design strategies, protein engineering, ribonucleoprotein delivery, using spatiotemporally controlled Cas9, or gRNAs through chemical or environmental inducers, that can modify CRISPR function. The human population has been subjected to Cas9 protein homologs long before the utilization of CRISPR-Cas9 in genome editing. The amino acid sequence of the Cas9 protein from S. pyogenes has ~ 58%, 35%, and ≥ 80% similarity to Cas9 protein from S. thermophilus (probiotic), Lactobacillus plantarum (probiotic and in food production) and human commensal and pathogenic bacteria such as S. dysgalactiae subsp. equisimilis, Staphylococcus aureus, Klebsiella pneumonia, respectively (El-Mounadi et al. 2020). Nevertheless, there is a need to revise the regulations of genome-edited plants and to enlighten the general community about their characteristics. A sustainable future for agriculture can now be imagined along with the responsibility of continuously resolving both scientific and public concerns about its usage.
References
Adli M (2018) The CRISPR tool kit for genome editing and beyond. Nature Commun. https://doi.org/10.1038/s41467-018-04252-2
Afzal S, Sirohi P, Yadav AK, Singh MP, Kumar A, Singh NK (2019) A comparative screening of abiotic stress tolerance in early flowering rice mutants. J Biotechnol 20(302):112–122
Agrotis A, Ketteler R (2015) A new age in functional genomics using CRISPR/Cas9 in arrayed library screening. Front Genet 6:300. https://doi.org/10.3389/fgene.2015.00300
Ainley WM, Sastry-Dent L, Welter ME, Murray MG, Zeitler B, Amora R et al (2013) Trait stacking via targeted genome editing. Plant Biotechnol J 11:1126–1134. https://doi.org/10.1111/pbi.12107
Ali Z, Abulfaraj A, Idris A, Ali S, Tashkandi M, Mahfouz MM (2015) CRISPR/Cas9-mediated viral interference in plants. Genom Biol 16(1):238. https://doi.org/10.1186/s13059-015-0799-6
Anderson EM, Haupt A, Schiel JA, Chou E, Machado HB, Strezoska Z et al (2015) Systematic analysis of CRISPR–Cas9 mismatch tolerance reveals low levels of off-target activity. J Biotechnol 211:56–65. https://doi.org/10.1016/j.jbiotec.2015.06.427
Andersson M, Turesson H, Nicolia A, Fält AS, Samuelsson M, Hofvander P (2017) Efficient targeted multiallelic mutagenesis in tetraploid potato (Solanum tuberosum) by transient CRISPR-Cas9 expression in protoplasts. Plant Cell Rep 36(1):117–128. https://doi.org/10.1007/s00299-016-2062-3
Arroyo JD, Jourdain AA, Calvo SE, Ballarano CA, Doench JG, Root DE, Mootha VK (2016) A genome-wide CRISPR death screen identifies genes essential for oxidative phosphorylation. Cell Metab 24(6):875–885. https://doi.org/10.1016/j.cmet.2016.08.017
Baltes NJ, Gil-Humanes J, Cermak T, Atkins PA, Voytas DF (2014) DNA replicons for plant genome engineering. Plant Cell 26(1):151–163. https://doi.org/10.1105/tpc.113.119792
Bassett AR, Tibbit C, Ponting CP, Liu JL (2013) Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep 4(1):220–228. https://doi.org/10.1016/j.celrep.2013.06.020
Bernabé-Orts JM, Casas-Rodrigo I, Minguet EG, Landolfi V, Garcia-Carpintero V, Gianoglio S, Vázquez-Vilar M, Granell A, Orzaez D (2019) Assessment of Cas12a-mediated gene editing efficiency in plants. Plant Biotechnol J. https://doi.org/10.1111/pbi.13113
Brooks C, Nekrasov V, Lippman ZB, Van Eck J (2014) Efficient gene editing in tomato in the first generation using the clustered regularlyinterspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol 166(3):1292–1297. https://doi.org/10.1104/pp.114.247577
Budhagatapalli N, Rutten T, Gurushidze M, Kumlehn J, Hensel G (2015) Targeted modification of gene function exploiting homology-directed repair of TALEN-mediated double-strand breaks in Barley. G3 (Bethesda) 5:1857–1863
Butler NM, Atkins PA, Voytas DF, Douches DS (2015) Generation and inheritance of targeted mutations in potato (Solanum tuberosum L.) using the CRISPR/Cas system. PLoS ONE 10:e0144591. https://doi.org/10.1371/journal.pone.0144591
Canver MC, Lessard S, Pinello L, Wu Y, Ilboudo Y, Stern EN et al (2017) Variant-aware saturating mutagenesis using multiple Cas9 nucleases identifies regulatory elements at trait-associated loci. Nat Genet 49(4):625. https://doi.org/10.1038/ng.3793
Cermak T, Baltes NJ, Cegan R, Zhang Y, Voytas DF (2015) High-frequency, precise modification of the tomato genome. Genom Biol 16(1):232. https://doi.org/10.1186/s13059-015-0796-9
Chandrasekaran J, Brumin M, Wolf D, Leibman D, Klap C, Pearlsman M et al (2016) Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol Plant Pathol 17(7):1140–1153. https://doi.org/10.1111/mpp.12375
Char SN, Unger-Wallace E, Frame B, Brigss SA, Main M, Spalding MH et al (2015) Heritable site-specific mutagenesis using TALENs in maize. Plant Biotechnol J 13:1002–1010
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697. https://doi.org/10.1146/annurev-arplant-050718-100049
Cho SW, Kim S, Kim JM, Kim JS (2013) Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31(3):230. https://doi.org/10.1038/nbt.2507
Clasen BM, Stoddard TJ, Luo S, Demorest ZL, Li J, Cedrone F et al (2016) Improving cold storage and processing traits in potato through targeted gene knockout. Plant Biotechnol J 14:169–176
Cullot G, Boutin J, Toutain J, Prat F, Pennamen P, Rooryck C et al (2019) CRISPR-Cas9 genome editing induces megabase-scale chromosomal truncations. Nat Commun 10(1):1136. https://doi.org/10.1038/s41467-019-09006-2
Cunningham FJ, Goh NS, Demirer GS, Matos JL, Landry MP (2018) Nanoparticle-mediated delivery towards advancing plant genetic engineering. Trends Biotechnol 36:P882–P897. https://doi.org/10.1016/j.tibtech.2018.03.009
Decque A, Joffre O, Magalhaes JG, Cossec JC, Blecher-Gonen R, Lapaquette P et al (2016) Sumoylation coordinates the repression of inflammatory and anti-viral gene-expression programs during innate sensing. Nat Immunol 17(2):140
Demirer GS, Zhang H, Matos JL, Goh N, Cunningham F et al (2018) High aspect ratio nanomaterials enable delivery of functional genetic material without DNA integration in mature plants. bioRxiv. https://doi.org/10.1101/179549
Deshaies RJ, Joazeiro CA (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78:399–434. https://doi.org/10.1146/annurev.biochem.78.101807.093809
Doench JG, Hartenian E, Graham DB, Tothova Z, Hegde M, Smith I et al (2014) Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat Biotechnol 32(12):1262. https://doi.org/10.1038/nbt.3026
Du H, Zeng X, Zhao M, Cui X, Wang Q, Yang H et al (2016) Efficient targeted mutagenesis in soybean by TALENs and CRISPR/Cas9. J Biotechnol 217:90–97. https://doi.org/10.1016/j.jbiotec.2015.11.005
El-Mounadi K, Morales-Floriano ML, Garcia-Ruiz H (2020) Principles, applications, and biosafety of plant genome editing using CRISPR-Cas9. Front Plant Sci 11:56. https://doi.org/10.3389/fpls.2020.00056
Endo M, Mikami M, Toki S (2016) Biallelic gene targeting in rice. Plant Physiol 170:667–677. https://doi.org/10.1104/pp.15.01663
Fauser F, Schiml S, Puchta H (2014) Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsisthaliana. Plant J 79(2):348–359. https://doi.org/10.1111/tpj.12554
Gallego-Bartolome J, Liu W, Kuo PH, Feng S, Ghoshal B, Gardiner J et al (2019) Co-targeting RNA polymerases IV and V promotes efficient de novo DNA methylation in Arabidopsis. Cell 176:1068–1082. https://doi.org/10.1016/j.cell.2019.01.029
Gao J, Wang G, Ma S, Xie X, Wu X, Zhang X et al (2015) CRISPR/Cas9-mediated targeted mutagenesis in Nicotiana tabacum. Plant Mol Biol 87(1–2):99–110. https://doi.org/10.1007/s11103-014-0263-0
Gelvin SB (2003) Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol Mol Biol Rev 67(1):16–37. https://doi.org/10.1128/MMBR.67.1.16-37.2003
Gil-Humanes J, Wang Y, Liang Z, Shan Q, Ozuna CV, Sanchez-Leon S et al (2017) High-efficiency gene targeting in hexaploid wheat using DNA replicons and CRISPR/Cas9. Plant J 89:1251–1262. https://doi.org/10.1111/tpj.13446
Glass Z, Lee M, Li Y, Xu Q (2018) Engineering the delivery system for CRISPR-based genome editing. Trends Biotechnol 36(2):173–185. https://doi.org/10.1016/j.tibtech.2017.11.006
Gong Z, Morales-Ruiz T, Ariza RR, Roldán-Arjona T, David L, Zhu JK (2002) ROS1, a repressor of transcriptional gene silencing in Arabidopsis, encodes a DNA glycosylase/lyase. Cell 111(6):803–814. https://doi.org/10.1016/S0092-8674(02)01133-9
Haeussler M, Schonig K, Eckert H, Eschstruth A, Mianne J, Renaud JB et al (2016) Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genom Biol 17(1):148. https://doi.org/10.1186/s13059-016-1012-2
Halterman D, Guenthner J, Collinge S, Butler N, Douches D (2016) Biotech potatoes in the 21st century: 20 years since the first biotech potato. Am J Potato Res 93(1):1–20. https://doi.org/10.1007/s12230-015-9485-1
Han K, Jeng EE, Hess GT, Morgens DW, Li A, Bassik MC (2017) Synergistic drug combinations for cancer identified in a CRISPR screen for pairwise genetic interactions. Nat Biotechnol 35(5):463. https://doi.org/10.1038/nbt.3834
Heigwer F, Kerr G, Boutros M (2014) E-CRISP: fast CRISPR target site identification. Nat Methods 11(2):122. https://doi.org/10.1038/nmeth.2812
Helm M, Brule H, Giege R, Florentz C (1999) More mistakes by T7 RNA polymerase at the 5′ ends of in vitro-transcribed RNAs. RNA 5(5):618–621. https://doi.org/10.1017/S1355838299982328
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V et al (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31(9):827. https://doi.org/10.1038/nbt.2647
Ikeuchi M, Ogawa Y, Iwase A, Sugimoto K (2016) Plant regeneration: cellular origins and molecular mechanisms. Development 143:1442–1451
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res 41(20):e188. https://doi.org/10.1093/nar/gkt780
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA–guided DNA endonuclease in adaptivebacterial immunity. Science 337(6096):816–821. https://doi.org/10.1126/science.1225829
Joung J, Konermann S, Gootenberg JS, Abudayyeh OO, Platt RJ, Brigham MD et al (2017) Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat Protoc 12(4):828. https://doi.org/10.1038/nprot.2017.016
Jung JH, Altpeter F (2016) TALEN mediated targeted mutagenesis of the caffeic acid O-methyltransferase in highly polyploid sugarcane improves cell wall composition for production of bioethanol. Plant Mol Biol 92:131–142
Kelley ML, Strezoska Z, He K, Vermeulen A, van Brabant SA (2016) Versatility of chemically synthesized guide RNAs for CRISPR-Cas9 genome editing. J Biotechnol 233:74–83. https://doi.org/10.1016/j.jbiotec.2016.06.011
Khatodia S, Bhatotia K, Passricha N, Khurana SMP, Tuteja N (2016) The CRISPR/Cas genome-editing tool: application in improvement of crops. Front Plant Sci 7:506. https://doi.org/10.3389/fpls.2016.00506
Khlestkina EK, Shumny VK (2016) Prospects for application of breakthrough technologies in breeding: the CRISPR/Cas9 system for plant genome editing. Russ J Genet 52(7):676–687. https://doi.org/10.1134/S102279541607005X
Kim H, Ishidate T, Ghanta KS, Seth M, Conte D, Shirayama M et al (2014) A co-CRISPR strategy for efficient genome editing in Caenorhabditiselegans. Genetics 197(4):1069–1080. https://doi.org/10.1534/genetics.114.166389
Kim H, Kim ST, Ryu J, Kang BC, Kim JS, Kim SG (2017) CRISPR/Cpf1-mediated DNA-free plant genome editing. Nat Commun 8:14406. https://doi.org/10.1038/ncomms14406
Koike-Yusa H, Li Y, Tan EP, Velasco-Herrera MDC, Yusa K (2014) Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat Biotechnol 32(3):267. https://doi.org/10.1038/nbt.2800
Labun K, Montague TG, Gagnon JA, Thyme SB, Valen E (2016) CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res 44(W1):W272–W276. https://doi.org/10.1093/nar/gkw398
Lawrenson T, Shorinola O, Stacey N, Li C, Ostergaard L, Patron N et al (2015) Induction of targeted, heritable mutations in barley and Brassica oleracea using RNA-guided Cas9 nuclease. Genome Biol 16(1):258. https://doi.org/10.1186/s13059-015-0826-7
Li B, Cui G, Shen G, Zhan Z, Huang L, Chen J, Qi X (2017a) Targeted mutagenesis in the medicinal plant Salvia miltiorrhiza. Sci Rep 7:43320. https://doi.org/10.1038/srep43320
Li J, Zhang H, Si X, Tian Y, Chen K, Liu J et al (2017b) Generation of thermosensitive male-sterile maize by targeted knockout of the ZmTMS5 gene. J Genet Genom 44:465–468. https://doi.org/10.1016/j.jgg.2017.02.002
Li JF, Norville JE, Aach J, McCormack M, Zhang D, Bush J et al (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31(8):688. https://doi.org/10.1038/nbt.2654
Li M, Li X, Zhou Z, Wu P, Fang M, Pan X et al (2016) Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front Plant Sci 7:377. https://doi.org/10.3389/fpls.2016.00377
Li Z, Liu ZB, Xing A, Moon BP, Koellhoffer JP, Huang L et al (2015) Cas9-guide RNA directed genome editing in soybean. Plant Physiol 169(2):960–970. https://doi.org/10.1104/pp.15.00783
Liang J, Han BZ, Nout MR, Hamer RJ (2008) Effects of soaking, germination and fermentation on phytic acid, total and in vitro soluble zinc in brown rice. Food Chem 110(4):821–828. https://doi.org/10.1104/pp.15.00783
Liang Z, Zhang K, Chen K, Gao C (2014) Targeted mutagenesis in Zea mays using TALENs and the CRISPR/Cas system. J Genet Genom 41(2):63–68. https://doi.org/10.1016/j.jgg.2013.12.001
Liu YC, Vidali L (2011) Efficient polyethylene glycol (PEG) mediated transformation of the moss Physcomitrella patens. JoVE 50:e2560. https://doi.org/10.3791/2560
Luo M, Li H, Chakraborty S, Morbitzer R, Rinaldo A, Upadhyaya N, Bhatt D, Louis S, Richardson T, Lahaye T, Ayliffe M (2019) Efficient TALEN mediated gene editing in wheat. Plant Biotechnol J. https://doi.org/10.1111/pbi.13169
Luria N, Smith E, Reingold V, Bekelman I, Lapidot M, Levin I et al (2017) A new Israeli Tobamovirus isolate infects tomato plants harboring Tm-22 resistance genes. PloS one 12(1):e0170429. https://doi.org/10.1371/journal.pone.0170429
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R et al (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8(8):1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Maggio I, Holkers M, Liu J, Janssen JM, Chen X, Gonçalves MA (2014) Adenoviral vector delivery of RNA-guided CRISPR/Cas9 nuclease complexes induces targeted mutagenesis in a diverse array of human cells. Sci Rep 4:5105. https://doi.org/10.1038/srep05105
Mahas A, Ali Z, Tashkandi M, Mahfouz MM (2019) Virus-Mediated cxgenome editing in plants using the CRISPR/Cas9 system. In: Qi Y (ed) Plant genome editing with crispr systems methods in molecular biology. Press, New York
Malnoy M, Viola R, Jung MH, Koo OJ, Kim S, Kim JS et al (2016) DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front Plant Sci 7:1904. https://doi.org/10.3389/fpls.2016.01904
Malzahn AA, Tang X, Lee K, Ren Q, Sretenovic S, Zhang Y, Chen H, Kang M, Bao Y, Zheng X, Deng K (2019) Application of CRISPR-Cas12a temperature sensitivity for improved genome editing in rice, maize, and Arabidopsis. BMC Biol 17(1):9. https://doi.org/10.1186/s12915-019-0629-5
Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H et al (2012) Systematic localization of common disease-associated variation in regulatory DNA. Science 337(6099):1190–1195. https://doi.org/10.1126/science.1222794
McDonald JI, Celik H, Rois LE, Fishberger G, Fowler T, Rees R et al (2016) Reprogrammable CRISPR/Cas9-based system for inducing site-specific DNA methylation. Biol Open 5(6):866–874. https://doi.org/10.1242/bio.019067
Mendoza BJ, Trinh CT (2017) Enhanced guide-RNA design and targeting analysis for precise CRISPR genome editing of single and consortia of industrially relevant and non-model organisms. Bioinformatics 34(1):16–23. https://doi.org/10.1093/bioinformatics/btx564
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X et al (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 23(10):1233. https://doi.org/10.1038/cr.2013.123
Mitter N, Worrall EA, Robinson KE, Li P, Jain RG et al (2017) Clay nanosheets for topical delivery of RNAi for sustained protection against plant viruses. Nat Plants 3:16207. https://doi.org/10.1038/nplants.2016.207
Mlynarova L, Nap JP, Bisseling T (2007) The SWI/SNF chromatin-remodeling gene AtCHR12 mediates temporary growth arrest in Arabidopsis thaliana upon perceiving environmental stress. Plant J 51(5):874–885. https://doi.org/10.1111/j.1365-313X.2007.03185.x
Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E (2014) CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res 42(W1):W401–W407. https://doi.org/10.1093/nar/gku410
Naito Y, Hino K, Bono H, Ui-Tei K (2015) CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31(7):1120–1123. https://doi.org/10.1093/bioinformatics/btu743
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31(8):691. https://doi.org/10.1038/nbt.2655
Niu Y, Shen B, Cui Y, Chen Y, Wang J, Wang L et al (2014) Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156(4):836–843. https://doi.org/10.1016/j.cell.2014.01.027
Okada A, Arndell T, Borisjuk N, Sharma N, Watson-Haigh NS, Tucker EJ et al (2019) CRISPR/Cas9-mediated knockout of Ms1 enables the rapid generation of male-sterile hexaploid wheat lines for use in hybrid seed production. Plant Biotechnol J 17:1905–1913. https://doi.org/10.1111/pbi.13106
Oldridge DA, Wood AC, Weichert-Leahey N, Crimmins I, Sussman R, Winter C et al (2015) Genetic predisposition to neuroblastoma mediated by a LMO1 super-enhancer polymorphism. Nature 528(7582):418. https://doi.org/10.1038/nature15540
Ortigosa A, Gimenez-Ibanez S, Leonhardt N, Solano R (2018) Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol J 17:665–673. https://doi.org/10.1111/pbi.13006
Papikian A, Liu W, Gallego-Bartolome J, Jacobsen SE (2019) Site-specific manipulation of Arabidopsis loci using CRISPR-Cas9 SunTag systems. Nat Commun 10(1):729. https://doi.org/10.1038/s41467-019-08736-7
Peng A, Chen S, Lei T, Xu L, He Y, Wu L et al (2017) Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene Cs LOB 1 promoter in citrus. Plant Biotechnol J 15(12):1509–1519. https://doi.org/10.1111/pbi.12733
Pitsch S, Weiss PA, Jenny L, Stutz A, Wu X (2001) Reliable chemical synthesis of oligoribonucleotides (RNA) with 2′-O-[(triisopropylsilyl) oxy] methyl (2′-O-tom)-protected phosphoramidites. Helv Chim Acta 84(12):3773–3795. https://doi.org/10.1002/1522-2675(20011219)84:12<3773:AID-HLCA3773>3.0.CO;2-E
Pliatsika V, Rigoutsos I (2015) “Off-Spotter”: very fast and exhaustive enumeration of genomic lookalikes for designing CRISPR/Cas guide RNAs. Biol Direct 10(1):4. https://doi.org/10.1186/s13062-015-0035-z
Pyott DE, Sheehan E, Molnar A (2016) Engineering of CRISPR/Cas9-mediated potyvirus resistance in transgene-free Arabidopsis plants. Mol Plant Pathol 17(8):1276–1288. https://doi.org/10.1111/mpp.12417
Qin W, Dion SL, Kutny PM, Zhang Y, Cheng AW, Jillette NL et al (2015) Efficient CRISPR/Cas9-mediated genome editing in mice by zygote electroporation of nuclease. Genetics 200(2):423–430. https://doi.org/10.1534/genetics.115.176594
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8(11):2281. https://doi.org/10.1038/nprot.2013.143
Raveux A, Vandormael-Pournin S, Cohen-Tannoudji M (2017) Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote. Sci Rep 7:42661. https://doi.org/10.1038/srep42661
Ron M, Kajala K, Pauluzzi G, Wang D, Reynoso MA, Zumstein K et al (2014) Hairy root transformation using Agrobacterium rhizogenes as a tool for exploring cell type-specific gene expression and function using tomato as a model. Plant Physiol 166(2):455–469. https://doi.org/10.1104/pp.114.239392
Ryan DE, Taussig D, Steinfeld I, Phadnis SM, Lunstad BD, Singh M et al (2018) Improving CRISPR–Cas specificity with chemical modifications in single-guide RNAs. Nucleic Acids Res 46(2):792–803. https://doi.org/10.1093/nar/gkx1199
Sander JD, Dahlborg EJ, Goodwin MJ, Cade L, Zhang F, Cifuentes D et al (2011) Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat Methods 8(1):67. https://doi.org/10.1038/nmeth.1542
Scaringe SA, Wincott FE, Caruthers MH (1998) Novel RNA synthesis method using 5′-O-Silyl-2′-O-orthoester protecting groups. J Am Chem Soc 120(45):11820–11821. https://doi.org/10.1021/ja980730v
Schiml S, Fauser F, Puchta H (2014) The CRISPR/C as system can be used as nuclease for in planta gene targeting and as paired nickases for directed mutagenesis in Arabidopsis resulting in heritable progeny. Plant J 80(6):1139–1150. https://doi.org/10.1111/tpj.12704
Schimmel J, van Schendel R, den Dunnen JT, Tijsterman M (2019) Templated insertions: a smoking gun for polymerase theta-mediated end joining. Trends Genet 35(9):632–644. https://doi.org/10.1016/j.tig.2019.06.001
Shan Q, Zhang Y, Chen K, Zhang K, Gao C (2015) Creation of fragrant rice by targeted knockout of the OsBADH2 gene using TALEN technology. Plant Biotechnol J 13:791–800
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z et al (2013) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31(8):686. https://doi.org/10.1038/nbt.2650
Shinwari ZK, Jan SA, Nakashima K, Yamaguchi-Shinozaki K (2020) Genetic engineering approaches to understanding drought tolerance in plants. Plant Biotechnol Rep 1:1–2
Stemmer M, Thumberger T, del Sol KM, Wittbrodt J, Mateo JL (2015) CCTop: an intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS ONE 10(4):e0124633. https://doi.org/10.1371/journal.pone.0124633
Svitashev S, Young JK, Schwartz C, Gao H, Falco SC, Cigan AM (2015) Targeted mutagenesis, precise gene editing, and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol 169(2):931–945. https://doi.org/10.1104/pp.15.00793
Tebas P, Stein D, Tang WW, Frank I, Wang SQ, Lee G et al (2014) Gene editing of CCR5 in autologous CD4 T cells of persons infected with HIV. N Engl J Med 370(10):901–910. https://doi.org/10.1056/NEJMoa1300662
Upadhyay SK, Kumar J, Alok A, Tuli R (2013) RNA-guided genome editing for target gene mutations in wheat. G3 3(12):2233–2238. https://doi.org/10.1534/g3.113.008847
Upadhyay SK, Sharma S (2014) SSFinder: high throughput CRISPR-Cas target sites prediction tool. BioMed Res Int. https://doi.org/10.1155/2014/742482
Urna FD, Reber EJ, Holmes MC et al (2010) Genome editing with engineering zinc finger nucleases. Nat Rev Genet 11:636–646. https://doi.org/10.1038/nrg2842
Veres A, Gosis BS, Ding Q, Collins R, Ragavendran A, Brand H et al (2014) Low incidence of off-target mutations in individual CRISPR-Cas9 and TALEN targeted human stem cell clones detected by whole-genome sequencing. Cell Stem Cell 15(1):27–30. https://doi.org/10.1016/j.stem.2014.04.020
Vojta A, Dobrinic P, Tadic V, Bockor L, Korac P, Julg B et al (2016) Repurposing the CRISPR-Cas9 system for targeted DNA methylation. Nucleic Acids Res 44(12):5615–5628. https://doi.org/10.1093/nar/gkw159
Wada N, Ueta R, Osakabe Y, Osakabe K (2020) Precision genome editing in plants: state-of-the-art in CRISPR/Cas9-based genome engineering. BMC Plant Biol 20(1):1–2
Wang F, Wang C, Liu P, Lei C, Hao W, Gao Y et al (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS ONE 11(4):e0154027. https://doi.org/10.1371/journal.pone.0154027
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32(9):947. https://doi.org/10.1038/nbt.2969
Wong N, Liu W, Wang X (2015) WU-CRISPR: characteristics of functional guide RNAs for the CRISPR/Cas9 system. Genom Biol 16(1):218. https://doi.org/10.1186/s13059-015-0784-0
Wu CS, Ouyang J, Mori E, Nguyen HD, Maréchal A, Hallet A et al (2014) SUMOylation of ATRIP potentiates DNA damage signaling by boosting multiple protein interactions in the ATR pathway. Genes Dev 28(13):1472–1484. https://doi.org/10.1101/gad.238535.114
Xiao A, Cheng Z, Kong L, Zhu Z, Lin S, Gao G, Zhang B (2014) CasOT: a genome-wide Cas9/gRNA off-target searching tool. Bioinformatics 30(8):1180–1182. https://doi.org/10.1093/bioinformatics/btt764
Xie K, Minkenberg B, Yang Y (2015) Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc Natl Acad Sci USA 112(11):3570–3575. https://doi.org/10.1073/pnas.1420294112
Yanik M, Ponnam SPG, Wimmer T, Trimborn L, Muller C, Gambert I et al (2018) Development of a reporter system to explore MMEJ in the context of replacing large genomic fragments. Mol Ther Nucleic Acids 11:407–415. https://doi.org/10.1016/j.omtn.2018.03.010
Zaidi SS, Mansoor S (2017) Viral vectors for plant genome engineering. Front Plant Sci 11(8):539. https://doi.org/10.3389/fpls.2017.00539
Zhang J, Abadia E, Refregier G, Tafaj S, Boschiroli ML, Guillard B et al (2010) Mycobacterium tuberculosis complex CRISPR genotyping: improving efficiency, throughput and discriminative power of ‘spoligotyping’ with new spacers and a microbead-based hybridization assay. J Med Microbiol 59(3):285–294. https://doi.org/10.1099/jmm.0.016949-0
Zhou H, Liu B, Weeks DP, Spalding MH, Yang B (2014) Large chromosomal deletions and heritable small genetic changes induced by CRISPR/Cas9 in rice. Nucleic Acids Res 42(17):10903–10914. https://doi.org/10.1093/nar/gku806
Zhu LJ (2015) Overview of guide RNA design tools for CRISPR-Cas9 genome editing technology. Front Biol 10(4):289–296. https://doi.org/10.1007/s11515-015-1366-y
Zuris JA, Thompson DB, Shu Y, Guilinger JP, Bessen JL, Hu JH et al (2015) Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat Biotechnol 33(1):73. https://doi.org/10.1038/nbt.3081
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
NKS wrote the initial draft of the manuscript; SA and PS helped with the literature review; SA prepared the manuscript; SA and PS contributed to the several revisions of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Afzal, S., Sirohi, P. & Singh, N.K. A review of CRISPR associated genome engineering: application, advances and future prospects of genome targeting tool for crop improvement. Biotechnol Lett 42, 1611–1632 (2020). https://doi.org/10.1007/s10529-020-02950-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02950-w