Abstract
In order to evaluate the corrosive action of microorganisms on 316L metal exposed directly to a marine environment, a system was designed to immerse coupons in seawater. After periods of 30, 60 and 90 days, the coupons were recovered, the corrosion rates evaluated and the biofilm samples on their surface were analyzed by 16S rRNA gene sequencing. The results of the corrosion rate showed an acceleration over the entire experimental period. Alpha diversity measurements showed higher rates after 60 days of the experiment, while abundance measurements showed higher rates after 90 days of exposure to the marine environment. The beta-diversity results showed a clear separation between the three conditions and proximity in the indices between replicates of the same experimental condition. The results of 16S rRNA gene sequencing showed that after 30 days of exposure to seawater, there was massive representativeness of the pioneer bacteria, Gamma and Alphaproteobacteria, with emphasis on the genera Alcanivorax, Oceanospirillum and Shewanella. At the 60-day analysis, the Gammaproteobacteria class remained dominant, followed by Alphaproteobacteria and Flavobacteria, and the main representatives were Flexibacter and Pseudoalteromonas. In the last analysis, after 90 days, a change in the described bacterial community profile was observed. The Gammaproteobacteria class was still the largest in diversity and OTUs. The most predominant genera in number of OTUs were Alteromonas, Bacteriovorax and, Nautella. Our results describe a change in the microbial community over coupons directly exposed to the marine environment, suggesting a redirection to the formation of a mature biofilm. The conditions created by the biofilm structure suggest said condition favor biocorrosion on the analyzed coupons.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The deterioration of metallic as well non-metallic materials by corrosion processes is a problem widely present in many industry sectors, as well as having enormous impacts on public safety and the environment (Hamilton 2003; Muyzer and Stams 2008). The economic losses due to corrosion approximate US$2.5 trillion per annum, equivalent to roughly 3% of global Gross Domestic Product (GDP) (Koch et al. 2002). Although numerous efforts are made to prevent corrosion, for example by coating with paint or other metal covers, or even cathodic protection, the problem persists (Chandler 1985). Corrosion is an electrochemical process characterized by the dissolution of the zero-valence metal at an anode site, but, in turn, a secondary reaction is necessary, which consists of a cathodic site that acts as an external acceptor of electrons, to complement the reaction as a whole. The tendency of a material to act as an anodic or cathodic site, or a donor or electron acceptor, respectively, is dependent on the electrochemical potential of such material. The propensity of the anodic site undergoing corrosion will be dependent on the potential difference between the anode and the cathode sites. However, this difference in potential is necessary, but not sufficient for metals to corrode. In fact, for corrosion to occur, there must be a kinetic path available to facilitate the flow of electrons between the anode and the cathode, and the rate of that flux determines the rate of corrosion. This process is particularly accelerated in microbially stimulated corrosion when biofilms accumulate at the cathodic site (Hamilton 2003).
Microbiologically Influenced Corrosion (MIC) is a commonly related problem in damage to metal infrastructure, including in marine environments, with losses that can reach billions of USD worldwide (Koch et al. 2002). MIC over metallic surfaces is an interfacial process involving interactions between microbes and structures, by means of strong interactions of microbial communities, named biofilms (Bonifay et al. 2017). The adhered microbes can induce, facilitate or accelerate metallic damage as a result of the requirement for an energy source, electron donor or electron acceptor (Videla and Herrera 2005; Procópio 2019). The scientific literature describes corrosion as frequently linked to the activity of Sulfate-Reducing Bacteria (SRB) and Thiosulfate-Reducing Bacteria (TRB) in anaerobic systems (Boudaud et al. 2010). More recently, attention has been directed to corrosion processes, especially metal corrosion, by the presence of lithotrophic Fe-Oxidizing Bacteria (FeOB) (Lee and Newman 2003; Marty et al. 2014; McBeth and Emerson 2016). FeOB-marine members are commonly aerobic, lithotrophic and capable of oxidizing iron under neutral pH conditions (Emerson et al. 2010). This group has been more studied in recent years in different marine environments, such as near-shore and deep-marine environments (Edwards et al. 2003; Emerson and Moyer 2002; McBeth and Emerson 2016). FeOB participates in the biogeochemical cycle of environments with high Fe levels, through redox reactions, and have been widely described as participants in the early stages of acceleration of carbon steel corrosion. As a consequence of its colonization, surface rust is formed, which enables the subsequent growth of anaerobic bacteria in the formed microenvironment (McBeth et al. 2011). Microorganisms related to iron oxidation accelerate the process through the mechanisms of pH changing, acid productions, secretion of corrosive metabolites or extracellular enzymes (Kato 2016; Little et al. 2007). The basis of an aerobic corrosion reaction is characterized by electrochemical coupling between iron oxidation (anodic reaction) and oxygen reduction (cathodic reaction) (Lee and Newman 2003). The product of this reaction is ferrous iron Fe(II), which is further oxidized to ferric iron, Fe(III). Fe(III) serves as a final electron acceptor in the Electron Transport Chain (ETC), allowing the flow of electrons through the ATPase enzyme, thus generating electrochemical strength for the synthesis of ATP, coupling the corrosion reactions with Fe(III)-respiring bacteria (Kato 2016; Tremblay et al. 2017).
Metallic structures are materials commonly used in marine engineering and industrial facilities, as well as in the construction of piers, ships, and bridges near marine environments. As marine environments are very conducive to corrosive processes, such as chemical and microbiologically influenced corrosion, the safety and conservation of these structures are of great global concern (Dang et al. 2011). Submerged metallic surfaces in aquatic environments are rapidly colonized by microorganisms, which adhere strongly, forming biofilm structures (Procópio 2019). The bacterial growth and subsequent formation of biofilm structures over metallic surfaces modify the oxygen concentration gradient, essentially due to microbial respiration, which generates conditions that can induce and accelerate cathodic corrosion reactions (Lee and Newman 2003; Marty et al. 2014). Despite the fact that Fe(II) is their primary energy source and they are frequently identified in corrosion biofilms, the role of the FeOB group in MIC is poorly described and the understanding of the role of FeOB in marine MIC is largely unvalued (Mumford et al. 2016). Studies have shown FeOB are able to colonize and grow either on metallic coupons introduced into the environment (Dang et al. 2011; McBeth et al. 2011), or are present on corroded steel associated with ALWC (Accelerated Low Water Corrosion) structures (Marty et al. 2014). Even structures made with metal alloys described as more resistant to corrosion are deteriorated by the action of microorganisms (Li et al. 2016; Sun et al. 2017). The 316L metal alloy is widely used in structures due to being more resistant to corrosion. Its application in structures close to marine environments is in the construction of deck components for boats and ships e.g. deck eyes, brackets for anchor ropes, housings for equipment, shackles, and handrails, among others.
Establishing a correlation between the action of bacteria and the corrosive processes of metal alloys is a challenge in biocorrosion studies. Culture techniques usually do not reflect the actual composition present in the sample in studies, owing to the innumerable non-cultivated species or because they are limited to a single-species study. Today, large-scale sequencing techniques, such as Illumina MiSeq, enable the investigation of microbial communities in different environments, as well as tracking succession over a given period. Moreover, the analysis of environmental samples in situ, allow more realistic approaches, once the real composition of the bacterial community is identified (Procópio 2020a, b). One reflection of this is the growing number of metagenomic studies which became available in the past few years (An et al. 2016; Li et al. 2017a; Liang et al. 2016; McBeth and Emerson 2016; Ramírez et al. 2016).
Stainless steel is widely used in various structures close to marine environments. The reason for its use is that 316L steel has corrosion-resistant properties. However, despite its stainless property, corrosive deterioration processes, such as pitting, are commonly described in this metal exposed to corrosive environments (Marconnet et al. 2008). In addition, although microbial corrosion of metal structures has been widely studied in recent decades, most of these studies have been conducted under controlled laboratory conditions. This metagenomic study is the first attempt to assess corrosion influenced by microorganisms in an actual marine environment. Our results allowed us to evaluate the true conditions, with the numerous factors in the conduction of biocorrosion. The goal of our study was to evaluate the action of bacteria grown over a period of up to 90 days on 316L stainless steel coupons dipped in seawater and the resultant corrosion rates due to biofilm growth over them. For this, we analyzed, by means of 16S rRNA gene sequencing, the dynamics of the bacterial community during the time of exposure to the marine environment and verified the attendant corrosion of the metal coupon.
Materials and methods
Experiment design
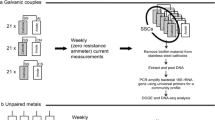
The experiment was conducted in situ, in Guanabara Bay, Rio de Janeiro, RJ, Brazil, near the naval shipyard of the Brazilian Navy, at a depth of roughly 5 m. A 3-cubic meter (m3) polypropylene box was designed, opened at the top. Fifteen stainless coupons of 3 cm2 surface area and 0.1 cm thickness were placed in the box, suspended by nylon strings and separated from each other. The box was fixed to the bottom of the sea using ropes and weights to prevent it being turned or dragged by the tide. The experiment was conducted for 90 days, and at intervals of 30, 60 and 90 days the box was retrieved, and 5 coupons were removed, immediately transferred in an icebox to the Microbial Corrosion Laboratory, Estacio University (UNESA), Rio de Janeiro-Brazil, to evaluate the mass loss and analyze the presence of microbial communities over the coupon using 16S DNA gene sequencing.
Corrosion rate analysis
Stainless steel 316L coupons were used to evaluate microbial corrosion in a marine environment, with maximum chemical composition of 0.030% C, 2.00% Mn, 0.75% Si, 0.045% P, 0.030% S, from 16.00 to 18.00% Cr, from 10.00 to 14.00% Ni, from 2.00 to 3.00% Mo. The stainless steel was cut into coupons of approximately 3cm2, with a small hole cut close to one of the edges, permitting them to be secured by nylon strings in the polypropylene box described above. Initially, the stainless steel coupons were sanded sequentially using sandpaper with decreasing grains. Then, they were then incubated for thirty minutes in absolute ethanol to degrease the surface, washed twice with acetone to remove any organic matter and finally dried at 70 °C for thirty minutes and stored in a desiccator. Subsequently, all coupons were autoclaved, cooled and identified. The coupons were weighed twice employing precision scales with 0.00001 readability in order to evaluate any weight loss after exposure to seawater.
At each timepoint, five stainless coupons were carefully removed of the box, and each coupon was stored in a previously sterilized 200 ml glass filled with local seawater, then transported to the Microbial Corrosion Laboratory of Estácio University. Two of the coupons were separated to determine microbial community by 16S rRNA gene sequencing, and the other three coupons were used for mass loss analysis.
The biofilm formed over the coupons and corrosion debris were scraped and immediately immersed in acid pickling (15% HCl) to remove all surface corrosion products according to ASTM G1 (Standard Practice for Preparing, Cleaning, and Evaluating Corrosion Test Specimens) (ASTM G1-03 2017). The acid reaction was ended by the application of thiourea solution for 5 s, then the coupons were washed with distilled water, neutralized with 10% NaOH for 5 s, and finally immersed in acetone for the same period. Following this process, the coupons were dried and used to determine corrosion rates. Evaluation of corrosion progress was undertaken using the weight-loss method. The results of weight-loss measurement were employed to calculate the value of the corrosion rate (CR) following the equation:
where the W is the decrease in metal weight the during time period analyzed, K is the constant (8.76 × 104), D represents the metal density in g/cm3, the coupon area (mm2) and T the exposure time in days (ASTM G1-03 2017).
16S rRNA gene-based community analysis
The coupons immersed in the marine environment of each experimental condition were carefully removed and, rinsed with sterile water to remove residues loosely adhered to the coupons. Following this, the biofilm growth over the coupons surfaces were scraped with a swab, previously moistened to facilitate collection and, kept refrigerated in a thermal box at 0 °C. Sequentially, the replicates were immediately sent to Neoprospecta Microbiome Technologies (Florianópolis-SC, Brazil). The DNA total was extracted using the commercial DNeasy PowerSoil Kit (Qiagen), respecting the conditions suggested by the manufacturer. The sample preparation and sequencing were performed by Neoprospecta Microbiome Technologies. It consisted of the 16S rRNA V3/V4 region, which was amplified using 341F (5′-CCTACGGGRSGCAGCAG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) primers, employing Illumina adapters. Amplification consisted of 35 cycles at 50 °C annealing temperature of the primers. In each sample, the process was carried out in triplicate. Then, sequencing was performed by Illumina MiSeq using V2 kits, with a 300 nts run.
The bioinformatics analyses were performed by Neoprospecta Microbiome (Florianopolis-SC, Brazil). The primer and adapter sequences were trimmed from the reads and only sequences with 275nt or more were used in downstream analysis. Then, all reads with one or more indeterminate “N” bases and truncated sequences with two or more consecutive bases with quality scores below Q20 were eliminated. OTUs (Operational Taxonomic Units) were performed using BLASTN 2.2.28 against the GreenGenes 13.8 database (DeSantis et al. 2006). For the purpose of attributing taxonomy, only sequences with 99% identity hits and over 99% alignment coverage were considered. The phylogenetic tree was constructed initially using an alignment with high abundance OTUs by MUSCLE method (Edgar 2004a, b). Next, a tree was generated using the UPGMA method and MEGA6 software (Tamura et al. 2013), with 1000 bootstraps for branching, and the tree topology by 1000 re-samplings. The OTUs sequences are available in the NCBI Sequence Read Archive (SRA) database under accession no. PRJNA494887, in SAMN10184473-8 Biosample accession numbers.
Statistical analysis and processing data
The abundances of phylum, class, genus and species levels, as well as their statistical differences in terms of the microbial community throughout the analyzed period were delineated in R language using Studio R software. Bar graphs were produced by the Studio R software, while a Heatmaps graph based on the relative abundance of OTUs at genetic levels was constructed using the ggplot2 (Wickham 2016) packet R program. Statistical analyzes of corrosion rates were performed using the parametric t test, as well as the standard error of the means were calculated and using the program R.
The alpha-diversity analysis was calculated using Shannon and Simpson, whereas the non-parametric abundance Chao1. All these indices employed the EstimateS software. The rarefaction curves based on the six samples of the three conditions were constructed using the Past3 software. The stacked bars graphs, Venn diagram and Heatmap were made using the ggplot2 and Vegan packages available for the R Studio software. The beta-diversity index was used to verify the differences among the communities employing the Bray–Curtis index. The plots used to show these results were the principal component analysis (PCA) and the non-metric multidimensional scaling (NMDS) analysis, both using the Past3 software.
Results
Corrosion rate measurement
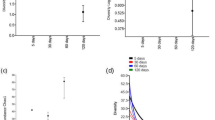
The coupons were submerged in seawater at a depth of about 5 m and recovered after 30, 60 and 90 days to evaluate corrosion rates. Deposits of rust on the surface were removed and then the coupon underwent the acid pickling process as described above. After the process, the coupons were weighed to compare the mass before and after the period of exposure to seawater. Even after the first collection and analysis at 30 days, biofouling was visible on the whole surface, and this characteristic deposit persisted, and gradually increased until the last analytical period. The results of the mass loss measures indicated a clear progression in the corrosion rate throughout the experiment. Figure 1 shows the corrosion rate over the assessed period of up to 90 days, showing a small loss of mass only after 90 days of exposure.
Richness of OTUs among the samples
The Shannon and Simpson indices were used to measure the alpha heterogeneity in the samples evaluated. Both index values indicated a great diversity in all the samples. The values presented in Table 1 show an increase in diversity between 30 and 60 days of exposure to the coupons in the marine environment. In fact, the condition where the greatest diversity occurred, according to the alpha diversity index was at 60 days. Although after 90 days there was a decrease in relation to the previous period of 60 days, a high diversity index was still maintained. The indices of abundance calculated by Chao1 also presented a profile where there is higher richness in the condition of 60 days of incubation in seawater, followed by 30 and 90 days (Table 1). The rarefaction curve analysis showing all replicates of the three analyzed conditions confirm the greatest richness after 60 days, while conditions at 30 and 90 days show similar indices (Fig. 2). The analysis of the structures of the three communities using Non-Metrric Multidimensional Scaling (NMDS) showed a distinct separation between them (Fig. 3).
The Illumina MiSeq sequencing generated a total of 307,183 high-quality OTUs sequences within the six sequenced samples of the three time points analyzed. In the first analysis, after 30 days of exposure in the marine environment, 163,356 OTUs were generated between the two sequenced coupons. At the time of the second analysis, there was a substantial decrease in the number of OTUs generated, 38,833 sequences in total for both coupons. And in the last two sequenced samples which refer to 90 days’ exposure to seawater, there was an increase in the number of sequences, in which 104,994 OTUs were detected. The Venn diagram shows the OTU distribution and overlap for the different microbial communities throughout the study (Fig. 4a).
The results of the OTUs similarity analysis enabled the identification of 229 different bacterial species (Fig. 4b). When comparing microbial diversity during the period in which the coupons were exposed in the marine environment, it was noted that between the first 30 and 60 days, there was a small increase in the number of species detected, from 127 to 135 bacteria, although the number of OTUs was lower at the second time point. The analysis of the species detected throughout the experiment shows that the largest number was identified at 60 days, followed by 30 days. Despite the fact that number of OTUs present exclusively after 90 days was the highest when compared to those at 30 and 60 days, the number of bacteria species present only at day 90 was the lowest (Fig. 4b).
Annotated diversity analysis
The results showed that, in general, there were consistent patterns of taxonomic distributions between replications of sequences in coupons, as can be seen in taxonomic analyses of the phylum and class groups. At the phylum level, Proteobacteria, Bacteroidetes, Firmicutes, and Verrucomicrobia, were represent (Table 2). Members of Proteobacteria phyla were dominant in all samples sequenced. In this group alone, 288,856 OTUs were detected, constituting a total of 94.03% of the total detected sequences. The next most abundant group was Bacteroidetes, which comprised about 5.62% of the sequences, with 17,252 OTUs. Firmicutes (0.34%/1035 OTUs) and Verrucomicrobia (0.01%/40 OTUs) were detected only at 30 and 60 days of incubation, respectively (Fig. 4). In general, when analyzing the core of class-level sequencing, community composition was overwhelmingly dominated by Gammaproteobacteria (86.8%), Alphaproteobacteria came second with 6.3%, and Flavobacteriia with 3,6% was the third most predominant class. Saprospiria (1.2%), Oligoflexia (1%), Bacilli (0.4%), Epsilonproteobacteria (0.05%) and Cytophagia (0.9%) were detected in lesser proportions.
After 30 days of seawater incubation, 132 different species were indentified growing over coupons. The main group was Gammaproteobacteria, with 153,353 OTUs (93.9%) distributed among 88 different representatives. The second most prevalent group was Alphaproteobacteria, with 4620 OTUs (2.8%), and 24 different members, with a value well below the first group. Next, Flavobacteria class presented 4210 OTUs (2.6%) and 18 different species (Fig. 5). The Bacilli (1035 OTUs) and Epsilonproteobacteria (138 OTUs) classes were detected only at this point of analysis, Bacillus subtilis and Arcobacter bivalviorum (138 OTUs) being the only representatives of each class, respectively (Fig. 6). The dominant genus in this analysis was Vibrio, with 38 different species, followed by Pseudoalteromonas, with 13 species, both representing 42.4% and 38.8%, respectively. However, when observing the number of sequences between the two groups, Pseudoalteromonas had 63,308 sequences, in close proximity to Vibrio, which despite having almost double the number of representatives, accounted for 69,284 sequences detected. Moreover, the bacterium Pseudoalteromonas mariniglutinosa was present in 37,736 sequences, which alone accounted for 23% of all sequences. The Alcanivorax borkumensis was represented by 5644 OTUs, with 3.5% of the detected sequences, while other more representative genera were Shewanella, with 3.7% of OTUs, Oceanospirillum was present in 1.3% and Alteromonas in 1.2%, and all other genera had values below 1% (Fig. 6).
With 60 days of exposure to seawater 135 different species was identified. The most representative group, in contrast to the first time point, was the class Alphaproteobacteria with 54 different species. The second group was members from class Gammaproteobacteria, with 45 members, followed by the Flavobacteria class represented in the 31 different genera. Members of the Cytophagia, Sapropiria and Verrucomicrobiae classes were identified for the first time at this timepoint. Although class Alphaproteobacteria was the most diverse, Gammaproteobacteria still accounted for the largest number of OTUs, 21,761 versus 9951 OTUs detected for Alphaproteobacteria (Fig. 5). The Flexibacter echinicida of the Flavobacteria class was one of the bacteria that showed the greater number of OTUs, 3504. The Vibrio genus, despite still being the most abundant genus, had a clear reduction in the number of OTUs and species, from 36 representatives detected at 30 days to 15 at this timepoint. Other Gammaproteobacteria members detected were represented by Aestuariibacter halophilus (5.1%), A. borkumensis (4.9%) and Alteromonas macleodii (4.1%). Pseudoalteromonas and Shewanella were present in five representatives each, being the most notable Pseudoalteromonas spongiae (3.5%) and Shewanella waksmanii (3.1%). Also, the oxyhydroxide-reducing Ferrimonas (0.7%), and the hydrocarbon degrader Marinobacter hydrocarbonoclasticus (0.2%) were detected. Along with M. hydrocarbonoclasticus, two other species of this genus were also identified, Marinobacter coastalis and Marinobacter xestospongiae, both wit 0.2% of representativeness. Among the Alphaproteobacteria, the Roseovarius genus was represented with seven species, followed by Erythrobacter and Ruegeria genera, with five and four representatives, respectively. The Sulfitobacter genus was represented by three different members, Sulfitobacter donghicola (0.15%), Sulfitobacter mediterraneus (0.1%), and Sulfitobacter pontiacus (0.9%). 16S DNAs from the Donghicola, Hyphomonas, Marivita, Pseudoruegeria, Shimia, Thalassobius, and Tropicibacter genera were also represented. Finally, the extremophilic bacterium Antarctobacter heliothermus was detected in this analysis with 0.2% representativity (Fig. 7).
The last time point analyzed was after 90 days of exposure to the marine environment. At this point, despite the fact the number of OTUs was the largest in this study (104,994), there was a decrease in the diversity of bacteria, 83 different species. The dominant group was class Gammaproteobacteria, with 55 members identified, and 87.4% representativeness, followed by Alphaproteobacteria (4.4%) and Flavobacteria (2.4%) (Fig. 5). The Vibrio genus was again the most prevalent on the analyzed coupons. Even though, the number of species was 16, similar to 60 days, there was an increase in the number of OTUs detected in the group, with 58.3% representativeness. Seven members of the Pseudoalteromonas genus were detected with 14,551 sequences (13.9%), and three species of Pseudomonas, including the petroleum-degrading Pseudomonas putida. Additionally, the genus Ferrimonas, previously described, was detected again, with the Ferrimonas futtsuensis (0.02%), and Ferrimonas kyonanensis (0.2%) species. Other Gammaproteobacteria members, previously described in other analyses, such as Aestuariibacter (2.0%), Alcanivorax (0.3%), Alteromonas (2.0%), Shewanella (1.3%), Thalassolituus (0.3%), persisted over metallic coupons until the end of the experiment. The Ruegeria genus belonging to the class Alphaproteobacteria continued to be detected, with four members and 1065 OTUs (1%). Futhermore, other genera that persisted as shown by 16S rRNAs gene sequencing were Celeribacter and Thalassobius genera. Cytophagia class contained two identified members, Algoriphagus vanfongensis, Algoriphagus zhangzhouensis, and the Saprospiria class with a single representative, Saprospira grandis (3.3%/3485 OTUs). Among the members of the Flavobacteria class, Bizionia palithoae, Corallibacter vietnamensis and Meridianimaribacter flavus predominated in our analyses. Finally, the two representatives of the Tenacibaculum genus, detected in the 60-day analysis, persisted to this point with a representativeness of 1.1% of the sequences (Fig. 8).
Discussion
After each period analyzed, the corrosion rate measurements indicated an acceleration of the deterioration of the metal exposed to the marine environment. Initially, the low corrosion rate in the first analysis was supplanted by the later results. Due to the resistance characteristics inherent in 316L stainless steel, because of the protective layer of Cr-oxides, the corrosive processes are normally slower (Landolt 2007). However, studies have shown that the presence of bacterial biofilms accelerates its corrosion by reducing oxygen in the cathodic zone (Videla 1994). As a consequence, pitting damage is identified in these situations. In a recent study on the effects of tidal cycles on the corrosion of 316L stainless steel, the acceleration of the corrosion rate influenced by the biofilm present on the surface was demonstrated (Daille et al. 2020). Furthermore, in this study three coupons were used to measure corrosion rates after 1 day, 5 and 15 weeks. Similarly, in our analysis, three coupons were also evaluated for each measure. However, the choice to start corrosion assay after 30 days was due to the prior knowledge that mass loss is insignificant in the first weeks of exposure.
One of the major challenges in understanding how the corrosion process occurs in a natural environment is to establish a relationship between the microorganisms associated with the process, the intrinsic environmental factors and the characteristics of the deteriorated material. The pivot for this understanding lies in deciphering the role of the microbial community throughout this process. Experiments carried out in controlled laboratory environments greatly aid in this task, but in situ experiments shed light on the evolution of the microbial taxa involved in the corrosive process, despite bringing more factors to be considered. Taxonomic information provides valuable insights into the microbial structure formed on the coupon by establishing functional relationships of the biofilm structure grown on the structure. Thus, our experiment was conducted in situ deep-water over a 90-day period in a marine environment. During this period, coupons immersed in seawater were analyzed at predetermined intervals by sequencing the 16S rRNA gene in order to determine the microbial taxonomy grown, evaluating for bacterial succession over time. This approach was made possible by Next-generation sequencing techniques, which allows for an easy and inexpensive analysis of microbial communities present in different environments (Parada et al. 2016). A key factor for the success of this analysis is the choice of primers for the amplification of the 16S genes. Criteria for choosing the most satisfactory primers include depth of sequencing, high coverage of taxa, including Bacteria and Archaea, ability to compare the results obtained and, clear data that permit inferring phylogenetic relationships (Parada et al. 2016). The 16S rRNA gene contains nine hypervariable regions (V1–V9) commonly used in taxonomic studies (Yang et al. 2016). The choice of the V3–V4 region of the 16S gene was due to its high sensitivity and accuracy. The amplification products of this region were 283 bp, and the following steps of the pipeline allowed confidence in sequenced reads, not considering sequencing errors, chimeras or PCR artifacts in the bacterial taxonomic evaluation.
In analyzing the sequencing results, the predominance of certain groups over the 90 days of exposure to seawater was clear. Proteobacteria phyla was widely distributed in all metagenomic analyses of replicate samples. Several other similar studies describe Proteobacteria as dominant at the core, based on 16S rRNA gene analyses (Bermont-Bouis et al. 2007; Dang et al. 2011; Garcia and Procópio 2020; Jones et al. 2007; Li et al. 2017b; McBeth and Emerson 2016; Moura et al. 2018; Vigneron et al. 2017). Although most of these studies have been directed to metagenomic analysis on offshore oil production facilities, a relationship between the microbial community structure present in both conditions is possible, especially when we analyze the initial periods in the microbial colonization on the rusting metallic surfaces. One of the possible reasons for this similar ecological behavior is suggested by Li et al. (2017b), who describe the members of the phylum Proteobacteria as pioneers in the colonization of surfaces, including metal surfaces. This step is central to the emergence of new bacterial species and the development and stability of the biofilm structure formation (Slightom and Buchan 2009).
Among the representatives of Proteobacteria, the predominant class consisted of Gammaproteobacteria members. As in other similar studies, this group was present in much higher numbers than others described in all the replicates analyzed, especially members of the Alteromonadales, Oceanospirillales, and Vibrionales families (Dang et al. 2011; McBeth and Emerson 2016; Moura et al. 2018). The scientific literature describes these bacteria as having important roles in the initial formation of the structure of biofilms on surfaces, being detected soon after the metal surface is exposed to the experimental conditions, such as coastal and marine environments (Dang and Lovell 2015; Dang et al. 2008, 2011; Moura et al. 2018; Slightom and Buchan 2009). The genus Pseudoalteromonas, described in the literature as potential marine FeOB (Ramírez et al. 2016), was extensively detected in all 16S rRNA sequencing of our samples. In a study evaluating the participation of P. lipolytica and B. subtillis demonstrated the corrosive action of P. lipolytica, especially in the formation of pitting, as a result of the inhibitory action on steel corrosion (Guo et al. 2017). Members of the Shewanella genus were detected in all samples collected. These bacteria are widely associated with microbial corrosion is different conditions and metal (Miller et al. 2018; Philips et al. 2018). In our study, the species Shewanella loihica was identified after 60 days of exposure to seawater. The halophilic S. loihica has been described as facultative anerobic and iron-reducing, having an active action on the corrosion of the metallic surface (Kooli et al. 2018). Shewanella algae were only detected in the last (60 days) analysis. Although S. algae are not directly related to the corrosion of metal surfaces, they have been described as an iron-reducing bacteria, suggesting their corrosive role in this study (Gandhi et al. 2002). The high number of specimens was also accompanied by a high number of OTUs in all replicates. Three different species of the Marinobacter genus were detected in this study, with greater prevalence after 90 days of exposure to seawater. Marinobacter species are commonly identified in similar studies and are associated with metal corrosion processes, mainly in offshore oil structures (Cluff et al. 2014; Brauer et al. 2015; Vigneron et al. 2016). The Ruegeria genus was present consistently in all three conditions analyzed in this experiment. This genus was also identified in a study on the colonization by bacteria of different metals and rust (Zhang et al. 2019). Some of the representatives detected belonging to the class Gammaproteobacteria are described as hydrocarbon-degrading, such as A. borkumensis (Konieczna et al. 2018), Pseudomonas (Brown et al. 2017; Gomila et al. 2017; Li et al. 2016; Maia et al. 2019; Rajasekar et al. 2010; Xia et al. 2015; Xu et al. 2017), Acinetobacter venetianus and Thalassolituus (Choi and Cho 2013, Golyshin et al. 2013). The presence of these bacteria can be explained by proximity to the location where the Brazilian naval ships are maintained, and the attendant oil leakage into the water.
The second group with the highest number of Bacteria identified and OTUs in the analysis was Alphaproteobacteria. Members of this group are commonly reported in marine biocorrosion studies (Dang and Lovell 2015; Lenhart et al. 2014; McBeth and Emerson 2016; Vigneron et al. 2016). In a microcosm experiment simulating the marine environment, Alphaproteobacteria were also extensively identified, especially Rhodobacteriales (Moura et al. 2018). In homology analysis, 90 different specimens were detected in all the replicates analyzed, and were present in the first analysis at 30 days, suggesting a preponderant role in the colonization and formation of the biofilm structure over the coupons exposed to seawater. Similar to Alphaproteobacteria described in marine corrosion studies, Alphaproteobacteria are suggested as polysaccharide producers used in the formation of biofilms, thus creating a microenvironment between the metallic surfaces and the bacterial community present (Elifantz et al. 2013; Kip and van Veen 2015; Videla and Herrera 2005). Additionally, among the Alphaproteobacteria, the bacteria Shimia marina was identified, which are reported in the literature as a hydrocarbon-degrading strain, including aromatic compounds (Rodrigo-Torres et al. 2016). Another sequence identified was Roseobacter denitrificans, whose genome sequencing indicates the possibility of being related to sulphur oxidation under oxic and anoxic conditions (Lenk et al. 2012). Two different species of Hyphomonas were detected in the 60-day experimental period. The Hyphomonas genus is not related to the corrosion of metals, although it is found in corrosive biofilms, probably due to its ability to manufacture a polysaccharide capsule that allows it to become adhesive in relation to mineral oxides (Bhosle et al. 1998). A single Epsilonproteobacteria were detected in the 16S rRNA gene sequencing of the 316L submerged coupons. The A. bivalviorum was identified in the samples at 30 days, but was no longer present at other timepoints. Although the genus Arcobacter is commonly detected in environments of greater marine depth, being related to hydrothermal sediments (Alain et al. 2004; An et al. 2016; López-García et al. 2003), the A. bivalviorum is related to colonization of seafood (Levican et al. 2012).
Flavobacteria represented a considerable community in analyzed sequences, both in the number of representatives and in OTU sequences. Flavobacteria were detected in all samples with special representativity after 60 days. This group has already been described in a previous experiment under controlled laboratory conditions, although in a lower number of representativeness (Moura et al. 2018). However, in several other studies on metallic corrosion in marine environments Bacteroidetes phylum are always highly represented (Li et al. 2017a, b). Microbiological succession analysis of mild steel in marine and estuary environments showed high percentages of representative indices, reaching up to 75% of the sequences when analyzed within a period of up to 24 days (McBeth and Emerson 2016). Bacteroidetes have an important influence on the other bacteria and the maintenance of the formed structure on surfaces. Predator specimens of Bacteroidetes act on the degradation of polymers present in biofilms, thus contributing to the maintenance of aerobic microenvironments within biofilms (Kirchman 2002).
In the analysis of 16S rRNA gene sequences in this study, only one representative of the B. subtilis was detected, though the number of OTUs was high. Firmicutes are constantly described in corrosion studies of metals in marine environments (Guo et al. 2017; Lenhart et al. 2014; Li et al. 2017a, b; Rajasekar et al. 2010). Normally, these studies demonstrate the role of Bacillus genus acting as pioneers in bacterial adhesion on metallic surfaces and thus acting in the initial processes of formation of bacterial biofilms (Karn et al. 2017; Wan et al. 2018). Moreover, the role of species of the Bacillus genus in the corrosion of metals is widely described in scientific literature, whether acting on corrosion actively or as a process inhibitor (Guo et al. 2019; Kang et al. 2019; Suma et al. 2019; Wang et al. 2018). Recently, the corrosive action by marine B. subtilis was demonstrated in a study with 10MnNiCrCu steel (Wang et al. 2018). Metagenomic survey on the bacterial community involved in the corrosion of petroleum pipelines also detected the presence of species of the genus Bacillus, including B. subtillis in the corrosive biofilm (Rajasekar et al. 2010). Finally, the genus Bacillus was also identified in a study concerning bacterial community participating in the corrosion of carbon steel under microcosm conditions (Procópio 2020a, b).
Conclusions
Based on the analyses of the sample sequencing, it was possible to describe in detail all the dynamics of the microbiological succession that occurred on the coupons throughout the entire period of the experiment. Initially, pioneer specimens that formed the bacterial biofilm structure were identified. However, over the course of the experiment, there was a change in the profile of the community present, characterized by microorganisms related to the maintenance of the biofilm and the appearance of specimens related to corrosive processes. Although it is not possible to relate the loss of mass of the coupons with the presence of the growing bacterial community, changing the bacterial profile over time seems to exert an influence on the microenvironment formed, which favors the conditions necessary to initiate or accelerate a corrosive process of the analyzed metal.
References
Alain K, Zbinden M, Le Bris N, Lesongeur F, Quérellou J, Gaill F, Cambon-Bonavita MA (2004) Early steps in microbial colonization processes at deep-sea hydrothermal vents. Environ Microbiol 6:227–241. https://doi.org/10.1111/j.1462-2920.2003.00557.x
An D, Dong X, An A, Park HS, Strous M, Voordouw G (2016) Metagenomic analysis indicates Epsilonproteobacteria as a potential cause of microbial corrosion in pipelines injected with bisulfite. Front Microbiol 7:28. https://doi.org/10.3389/fmicb.2016.00028
ASTM G1–03 (2017) Standard practice for preparing, cleaning, and evaluating corrosion test specimens. ASTM International, West Conshohocken
Bermont-Bouis D, Janvier M, Grimont PA, Dupont I, Vallaeys T (2007) Both sulfate-reducing bacteria and Enterobacteriaceae take part in marine biocorrosion of carbon steel. J Appl Microbiol 102(1):161–168. https://doi.org/10.1111/j.1365-2672.2006.03053.x
Bhosle N, Suci PA, Baty AM, Weiner RM, Geesey GG (1998) Influence of divalent cations and pH on adsorption of a bacterial polysaccharide adhesin. J Colloid Interface Sci 205:89–96
Bonifay V, Wawrik B, Sunner J, Snodgrass EC, Aydin E, Duncan KE, Callaghan AV, Oldham A, Liengen T, Beech I (2017) Metabolomic and metagenomic analysis of two crude oil production pipelines experiencing differential rates of corrosion. Front Microbiol 8:99. https://doi.org/10.3389/fmicb.2017.00099
Boudaud N, Coton M, Coton E, Pineau S, Travert J, Amiel C (2010) Biodiversity analysis by polyphasic study of marine bacteria associated with biocorrosion phenomena. J Appl Microbiol 109(1):166–179. https://doi.org/10.1111/j.1365-2672.2009.04643.x
Brauer JI, Makama Z, Bonifay V, Aydin E, Kaufman ED, Beech IB, Sunner J (2015) Mass spectrometric metabolomic imaging of biofilms on corroding steel surfaces using laser ablation and solvent capture by aspiration. Biointerphases 10:019003
Brown LM, Gunasekera TS, Ruiz ON (2017) Draft genome sequence of Pseudomonas stutzeri strain 19, an isolate capable of efficient degradation of aromatic hydrocarbons. Gen Announc 5(49):e01373–e1417. https://doi.org/10.1128/genomeA.01373-17
Chandler KA (1985) Marine and offshore corrosion. Butterworth-Heinemann Ltd, Chicago
Choi A, Cho JC (2013) Thalassolituus marinus sp. nov., a hydrocarbon-utilizing marine bacterium. Int J Syst Evol Microbiol 63(6):2234–2238. https://doi.org/10.1099/ijs.0.046383-0
Cluff MA, Hartsock A, MacRae JD, Carter K, Mouser PJ (2014) Temporal changes in microbial ecology and geochemistry in produced water from hydraulically fractured Marcellus shale gas wells. Environ Sci Technol 48:6508–6517
Daille LK, Aguirre J, Fischer D, Galarce C, Armijo F, Pizarro GE, Walczak M, De la Iglesia R, Vargas IT (2020) Effect of tidal cycles on bacterial biofilm formation and biocorrosion of stainless steel AISI 316L. J Mar Sci Eng 8:124–139. https://doi.org/10.3390/jmse8020124
Dang H, Lovell CR (2015) Microbial surface colonization and biofilm development in marine environments. Microbiol Mol Biol Rev 80(1):91–138. https://doi.org/10.1128/MMBR.00037-15
Dang H, Li T, Chen M, Huang G (2008) Cross-ocean distribution of Rhodobacterales bacteria as primary surface colonizers in temperate coastal marine waters. Appl Environ Microbiol 74(1):52–60. https://doi.org/10.1128/AEM.01400-07
Dang H, Chen R, Wang L, Shao S, Dai L, Ye Y, Guo L, Huang G, Klotz MG (2011) Molecular characterization of putative biocorroding microbiota with a novel niche detection of Epsilon- and Zetaproteobacteria in Pacific Ocean coastal seawaters. Environ Microbiol 13(11):3059–3074. https://doi.org/10.1111/j.1462-2920.2011.02583.x
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72(7):5069–5072. https://doi.org/10.1128/AEM.03006-05
Edgar RC (2004a) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform 5:113. https://doi.org/10.1186/1471-2105-5-113
Edgar RC (2004b) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
Edwards KJ, Rogers DR, Wirsen CO, McCollom TM (2003) Isolation and characterization of novel psychrophilic, neutrophilic, Fe-oxidizing, chemolithoautotrophic Alpha- and Gammaproteobacteria from the deep sea. Appl Environ Microbiol 69(5):2906–2913. https://doi.org/10.1128/AEM.69.5.2906-2913.2003
Elifantz H, Horn G, Ayon M, Cohen Y, Minz D (2013) Rhodobacteraceae are the key members of the microbial community of the initial biofilm formed in Eastern Mediterranean coastal seawater. FEMS Microbiol Ecol 85(2):348–357. https://doi.org/10.1111/1574-6941.12122
Emerson D, Fleming EJ, McBeth JM (2010) Iron-oxidizing bacteria: an environmental and genomic perspective. Annu Rev Microbiol 64:561–583. https://doi.org/10.1146/annurev.micro.112408.134208
Emerson D, Moyer CL (2002) Neutrophilic Fe-oxidizing bacteria are abundant at the Loihi Seamount hydrothermal vents and play a major role in Fe oxide deposition. Appl Environ Microbiol 68(6):3085–3093. https://doi.org/10.1128/AEM.68.6.3085-3093.2002
Gandhi S, Oh BT, Schnoor JL, Alvarez PJ (2002) Degradation of TCE, Cr(VI), sulfate, and nitrate mixtures by granular iron in flow-through columns under different microbial conditions. Water Res 36(8):1973–1982. https://doi.org/10.1016/s0043-1354(01)00409-2
Garcia M, Procópio L (2020) Distinct profiles in microbial diversity on carbon steel and different welds in simulated marine microcosm. Curr Microbiol. https://doi.org/10.1007/s00284-020-01898-4
Golyshin PN, Werner J, Chernikova TN, Tran H, Ferrer M, Yakimov MM, Teeling H, Golyshina OV (2013) Genome sequence of Thalassolituus oleivorans MIL-1 (DSM 14913T). Genome Announc 1(2):e0014113. https://doi.org/10.1128/genomeA.00141-13
Gomila M, Mulet M, Lalucat J, García-Valdés E (2017) Draft Genome sequence of Pseudomonas pachastrellae strain CCUG 46540(T), a deep-Sea bacterium. Genome Announc 5(14):e00136–e217. https://doi.org/10.1128/genomeA.00136-17
Guo Z, Liu T, Cheng YF, Guo N, Yin Y (2017) Adhesion of Bacillus subtilis and Pseudoalteromonas lipolytica to steel in a seawater environment and their effects on corrosion. Colloids Surf B 157:157–165. https://doi.org/10.1016/j.colsurfb.2017.05.045
Guo Z, Pan S, Liu T, Zhao Q, Wang Y, Guo N, Chang X, Liu T, Dong Y, Yin Y (2019) Bacillus subtilis inhibits Vibrio natriegens-induced corrosion via biomineralization in seawater. Front Microbiol 10:1111. https://doi.org/10.3389/fmicb.2019.01111
Hamilton WA (2003) Microbially influenced corrosion as a model system for the study of metal microbe interactions: a unifying electron transfer hypothesis. Biofouling 19(1):65–76. https://doi.org/10.1080/0892701021000041078
Jones PR, Cottrell MT, Kirchman DL, Dexter SC (2007) Bacterial community structure of biofilms on artificial surfaces in an estuary. Microb Ecol 53(1):153–162. https://doi.org/10.1007/s00248-006-9154-5
Kang Y, Li L, Li S, Zhou X, Xia K, Liu C, Qu Q (2019) Temporary inhibition of the corrosion of AZ31B magnesium alloy by formation of Bacillus subtilis biofilm in artificial seawater. Materials 12(3):523. https://doi.org/10.3390/ma12030523
Karn SK, Fang G, Duan J (2017) Bacillus sp. acting as dual role for corrosion induction and corrosion inhibition with carbon steel (CS). Front Microbiol 8:2038. https://doi.org/10.3389/fmicb.2017.02038
Kato S (2016) Microbial extracellular electron transfer and its relevance to iron corrosion. Microb Biotechnol 9(2):141–148. https://doi.org/10.1111/1751-7915.12340
Kip N, van Veen JA (2015) The dual role of microbes in corrosion. ISME J 9(3):542–551. https://doi.org/10.1038/ismej.2014.169
Kirchman DL (2002) The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microbiol Ecol 39(2):91–100. https://doi.org/10.1111/j.1574-6941.2002.tb00910.x
Koch GH, Brongers MPH, Thompson NG, Virmani YP, Payer JH (2002) Corrosion cost and preventive strategies in the United States. National Technical Information Service, Alexandria
Konieczna M, Olzog M, Naether DJ, Chrzanowski Ł, Heipieper HJ (2018) Membrane fatty acid composition and cell surface hydrophobicity of marine hydrocarbonoclastic Alcanivorax borkumensis SK2 grown on diesel, biodiesel and rapeseed oil as carbon sources. Molecules 23(6):E1432. https://doi.org/10.3390/molecules23061432
Kooli WM, Comensoli L, Maillard J, Albini M, Gelb A, Junier P, Joseph E (2018) Bacterial iron reduction and biogenic mineral formation for the stabilisation of corroded iron objects. Sci Rep 8(1):764. https://doi.org/10.1038/s41598-017-19020-3
Landolt D (2007) Corrosion and surface chemistry of metals in engineering sciences: materials. EPFL Press, Lausanne
Lee AK, Newman DK (2003) Microbial iron respiration: impacts on corrosion processes. Appl Microbiol Biotechnol 62(2):134–139. https://doi.org/10.1007/s00253-003-1314-7
Lenhart TR, Duncan KE, Beech IB, Sunner JA, Smith W, Bonifay V, Biri B, Suflita JM (2014) Identification and characterization of microbial biofilm communities associated with corroded oil pipeline surfaces. Biofouling 30(7):823–835. https://doi.org/10.1080/08927014.2014.931379
Lenk S, Moraru C, Hahnke S, Arnds J, Richter M, Kube M, Reinhardt R, Brinkhoff T, Harder J, Amann R, Mußmann M (2012) Roseobacter clade bacteria are abundant in coastal sediments and encode a novel combination of sulfur oxidation genes. ISME J 6(12):2178–2187. https://doi.org/10.1038/ismej.2012.66
Levican A, Collado L, Aguilar C, Yustes C, Diéguez AL, Romalde JL, Figueras MJ (2012) Arcobacter bivalviorum sp. nov. and Arcobacter venerupis sp. nov., new species isolated from shellfish. Syst Appl Microbiol 35(3):133–138. https://doi.org/10.1016/j.syapm.2012.01.002
Li H, Zhou E, Zhang D, Xu D, Xia J, Yang C, Feng H, Jiang Z, Li X, Gu T, Yang K (2016) Microbiologically influenced corrosion of 2707 hyper-duplex stainless steel by marine Pseudomonas aeruginosa biofilm. Sci Rep 6:20190. https://doi.org/10.1038/srep20190
Li X, Duan J, Xiao H, Li Y, Liu H, Guan F, Zhai X (2017a) Analysis of bacterial community composition of corroded steel immersed in Sanya and Xiamen seawaters in China via method of Illumina MiSeq sequencing. Front Microbiol 8:1737. https://doi.org/10.3389/fmicb.2017.01737
Li XX, Yang T, Mbadinga SM, Liu JF, Yang SZ, Gu JD, Mu BZ (2017b) Responses of microbial community composition to temperature gradient and carbon steel corrosion in production water of petroleum reservoir. Front Microbiol 8:2379. https://doi.org/10.3389/fmicb.2017.02379
Liang R, Davidova IA, Marks CR, Stamps BW, Harriman BH, Stevenson BS, Duncan KE, Suflita JM (2016) Metabolic capability of a predominant Halanaerobium sp. in hydraulically fractured gas wells and its implication in pipeline corrosion. Front Microbiol 7:988. https://doi.org/10.3389/fmicb.2016.00988
Little B, Lee J, Ray R (2007) A review of 'green' strategies to prevent or mitigate microbiologically influenced corrosion. Biofouling 23(1–2):87–97. https://doi.org/10.1080/08927010601151782
López-García P, Duperron S, Philippot P, Foriel J, Susini J, Moreira D (2003) Bacterial diversity in hydrothermal sediment and epsilonproteobacterial dominance in experimental microcolonizers at the Mid-Atlantic Ridge. Environ Microbiol 5(10):961–976. https://doi.org/10.1046/j.1462-2920.2003.00495.x
Maia M, Capão A, Procópio L (2019) Biosurfactant produced by oil-degrading Pseudomonas putida AM-b1 strain with potential for microbial enhanced oil recovery. Bioremediat J 23:302–310. https://doi.org/10.1080/10889868.2019.1669527
Marconnet C, Dagbert C, Roy M, Féron D (2008) Stainless steel ennoblement in freshwater: from exposure tests to mechanism. Corros Sci 50:2342–2352. https://doi.org/10.1016/j.corsci.2008.05.007
Marty F, Gueuné H, Malard E, Sánchez-Amaya JM, Sjögren L, Abbas B, Quillet L, van Loosdrecht MC, Muyzer G (2014) Identification of key factors in accelerated low water corrosion through experimental simulation of tidal conditions: influence of stimulated indigenous microbiota. Biofouling 30(3):281–297. https://doi.org/10.1080/08927014.2013.864758
McBeth JM, Emerson D (2016) In situ microbial community succession on mild steel in estuarine and marine environments: exploring the role of iron-oxidizing bacteria. Front Microbiol 7:767. https://doi.org/10.3389/fmicb.2016.00767
McBeth JM, Little BJ, Ray RI, Farrar KM, Emerson D (2011) Neutrophilic iron-oxidizing "Zetaproteobacteria" and mild steel corrosion in nearshore marine environments. Appl Environ Microbiol 77(4):1405–1412. https://doi.org/10.1128/AEM.02095-10
Miller RB 2nd, Lawson K, Sadek A, Monty CN, Senko JM (2018) Uniform and pitting corrosion of carbon steel by Shewanella oneidensis MR-1 under nitrate-reducing conditions. Appl Environ Microbiol 84(12):e00790–e818. https://doi.org/10.1128/AEM.00790-18
Moura V, Ribeiro I, Moriggi P, Capão A, Salles C, Bitati S, Procópio L (2018) The influence of surface microbial diversity and succession on microbiologically influenced corrosion of steel in a simulated marine environment. Arch Microbiol 200(10):1447–1456. https://doi.org/10.1007/s00203-018-1559-2
Mumford AC, Adaktylou IJ, Emerson D (2016) Peeking under the iron curtain: development of a microcosm for imaging the colonization of steel surfaces by Mariprofundus sp. strain DIS-1, an oxygen-tolerant Fe-oxidizing bacterium. Appl Environ Microbiol 82(22):6799–6807. https://doi.org/10.1128/AEM.01990-16
Muyzer G, Stams AJ (2008) The ecology and biotechnology of sulphate-reducing bacteria. Nat Rev Microbiol 6:441–454. https://doi.org/10.1038/nrmicro1892
Parada AE, Needham DM, Fuhrman JA (2016) Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18(5):1403–1414. https://doi.org/10.1111/1462-2920.13023
Philips J, Van den Driessche N, De Paepe K, Prévoteau A, Gralnick JA, Arends JBA, Rabaey K (2018) A novel Shewanella isolate enhances corrosion by using metallic iron as the electron donor with fumarate as the electron acceptor. Appl Environ Microbiol 284(20):e01154–e1218. https://doi.org/10.1128/AEM.01154-18
Procópio L (2019) The role of biofilms in the corrosion of steel in marine environments. World J Microbiol Biotechnol 35(5):73. https://doi.org/10.1007/s11274-019-2647-4
Procópio L (2020a) Microbial community profiles grown on 1020 carbon steel surfaces in seawater-isolated microcosm. Ann Microbiol 70:13. https://doi.org/10.1186/s13213-020-01547-y
Procópio L (2020b) The era of 'omics' technologies in the study of microbiologically influenced corrosion. Biotechnol Lett 42(3):341–356. https://doi.org/10.1007/s10529-019-02789-w
Rajasekar A, Anandkumar B, Maruthamuthu S, Ting YP, Rahman PK (2010) Characterization of corrosive bacterial consortia isolated from petroleum-product-transporting pipelines. Appl Microbiol Biotechnol 85(4):1175–1188. https://doi.org/10.1007/s00253-009-2289-9
Ramírez GA, Hoffman CL, Lee MD, Lesniewski RA, Barco RA, Garber A, Toner BM, Wheat CG, Edwards KJ, Orcutt BN (2016) Assessing marine microbial induced corrosion at Santa Catalina Island, California. Front Microbiol 7:1679. https://doi.org/10.3389/fmicb.2016.01679
Rodrigo-Torres L, Pujalte MJ, Arahal DR (2016) Draft genome sequence of Shimia marina CECT 7688(T). Mar Genom 28:83–86. https://doi.org/10.1016/j.margen.2016.01.006
Slightom RN, Buchan A (2009) Surface colonization by marine roseobacters: integrating genotype and phenotype. Appl Environ Microbiol 75(19):6027–6037. https://doi.org/10.1128/AEM.01508-09
Suma MS, Basheer R, Sreelekshmy BR, Riyas AH, Bhagya TC, Sha MA, Shibli SMA (2019) Synergistic action of Bacillus subtilis, Escherichia coli and Shewanella putrefaciens along with Pseudomonas putida on inhibiting mild steel against oxygen corrosion. Appl Microbiol Biotechnol 103(14):5891–5905. https://doi.org/10.1007/s00253-019-09866-0
Sun H, Shi B, Yang F, Wang D (2017) Effects of sulfate on heavy metal release from iron corrosion scales in drinking water distribution system. Water Res 114:69–77. https://doi.org/10.1016/j.watres.2017.02.021
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 60. Mol Biol Evol 30(12):2725–2729. https://doi.org/10.1093/molbev/mst197
Tremblay PL, Angenent LT, Zhang T (2017) Extracellular electron uptake: among autotrophs and mediated by surfaces. Trends Biotechnol 35(4):360–371. https://doi.org/10.1016/j.tibtech.2016.10.004
Videla H (1994) Biofilms and corrosion interactions on stainless steel in seawater. Int Biodeterior Biodegrad 34:245–257
Videla HA, Herrera LK (2005) Microbiologically influenced corrosion: looking to the future. Int Microbiol 8(3):169–180
Vigneron A, Alsop EB, Chambers B, Lomans BP, Head IM, Tsesmetzis N (2016) Complementary microorganisms in highly corrosive biofilms from an offshore oil production facility. Appl Environ Microbiol 82(8):2545–2554. https://doi.org/10.1128/AEM.03842-15
Vigneron A, Alsop EB, Lomans BP, Kyrpides NC, Head IM, Tsesmetzis N (2017) Succession in the petroleum reservoir microbiome through an oil field production lifecycle. Isme J 11:2141
Wan H, Song D, Zhang D, Du C, Xu D, Liu Z, Ding LX (2018) Corrosion effect of Bacillus cereus on X80 pipeline steel in a Beijing soil environment. Bioelectrochemistry 121:18–26. https://doi.org/10.1016/j.bioelechem.2017.12.011
Wang YS, Liu L, Fu Q, Sun J, An ZY, Ding R, Li Y, Zhao XD (2018) Effect of Bacillus subtilis on corrosion behavior of 10MnNiCrCu steel in marine environment. Sci Rep 10(1):5744. https://doi.org/10.1038/s41598-020-62809-y
Wickham H (2016) ggplot2: Elegant graphics for data analysis. Springer, New York
Xia J, Yang C, Xu D, Sun D, Nan L, Sun Z, Li Q, Gu T, Yang K (2015) Laboratory investigation of the microbiologically influenced corrosion (MIC) resistance of a novel Cu-bearing 2205 duplex stainless steel in the presence of an aerobic marine Pseudomonas aeruginosa biofilm. Biofouling 31(6):481–492. https://doi.org/10.1080/08927014.2015.1062089
Xu D, Xia J, Zhou E, Zhang D, Li H, Yang C, Li Q, Lin H, Li X, Yang K (2017) Accelerated corrosion of 2205 duplex stainless steel caused by marine aerobic Pseudomonas aeruginosa biofilm. Bioelectrochemistry 113:1–8. https://doi.org/10.1016/j.bioelechem.2016.08.001
Yang B, Wang Y, Qian PY (2016) Sensitivity and correlation of hypervariable regions in 16S rRNA genes in phylogenetic analysis. BMC Bioinform 17:135. https://doi.org/10.1186/s12859-016-0992-y
Zhang Y, Ma Y, Duan J, Li X, Wang J, Hou B (2019) Analysis of marine microbial communities colonizing various metallic materials and rust layers. Biofouling 35(4):429–442. https://doi.org/10.1080/08927014.2019.1610881
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The current research work did not involve either of human or animal studies.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Capão, A., Moreira-Filho, P., Garcia, M. et al. Marine bacterial community analysis on 316L stainless steel coupons by Illumina MiSeq sequencing. Biotechnol Lett 42, 1431–1448 (2020). https://doi.org/10.1007/s10529-020-02927-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02927-9