Abstract
Objectives
2,3-Butanediol (2,3-BD) is widely used in several chemical syntheses as well as the manufacture of plastics, solvents, and antifreeze formulations, and can be manufactured by microbial glucose fermentation. Conventional (2,3-BD) fermentation typically has low productivity, yield, and purity, and is expensive for commercial applications. We aimed to delete the lactate dehydrogenase and acetate kinase (ldhA and ack) genes in Klebsiella pneumoniae HD79 by using λRed homologous recombination technology, to eliminate by-products and thereby improve (2,3-BD) production. We also analyzed the resulting gene changes by using transcriptomics.
Results
The yield of (2,3-BD) from the mutant Klebsiella strain was 46.21 g/L, the conversion rate was 0.47 g/g, and the productivity was 0.64 g/L·h, which represented increases of 54.9%, 20.5%, and 106.5% respectively, compared to (WT) strains. Lactate and acetate decreased by 48.2% and 62.8%, respectively. Transcriptomics analysis showed that 4628 genes were differentially expressed (404 significantly up-regulated and 162 significantly down-regulated). Moreover, the (2,3-BD) operon genes were differentially expressed.
Conclusion
Our data showed that the biosynthesis of (2,3-BD) was regulated by inducers (lactate and acetate), a regulator (BudR), and carbon flux. Elimination of acidic by-products by ldhA and ack knockdown significantly improved (2,3-BD) production. Our results provide a deeper understanding of the mechanisms underlying (2,3-BD) production, and form a molecular basis for the improvement this process by genetic modification in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
2,3-Butanediol (2,3-BD) is widely used in many chemical syntheses (Białkowska 2016; Harvianto et al. 2018), as well as in the production of plastics, solvents, and antifreeze formulations (Taeyeon et al. 2016). Increasing attention to environmental problems and fossil fuel resources have resulted in a growing interest in the production of 2,3-BD by microbial fermentation. However, conventional 2,3-BD fermentation processes show low productivity, yield, and purity, and are expensive for commercial applications (Yang and Zhang 2018).
Several microorganisms such as Bacillus, Enterobacter, Serratia, Paenibacillus, Klebsiella, Saccharomyces cerevisiae, and Escherichia coli produce relatively large amounts of 2,3-BD (Gao et al. 2018; Ishii et al. 2018). Bacillus subtilis, B. licheniformis, and B. amyloliquefaciens exhibit good 2,3-BD production (Ge et al. 2016; Sikora et al. 2015). However, Klebsiella strains appear to be the most suited to fermentative 2,3-BD production. In fed-batch glycerol fermentation with K. pneumoniae, the yield of 2,3-BD was 49.2 g/L (Petrov and Petrova 2009). Park et al. (2015) reported that K. oxytoca (ΔldhA ΔpflB ΔBudC::PBDH) with intermittent glucose feeding showed a (R, R)-2,3-BD production of 106.7 g/L. Rathnasingh et al. (2016) deleted genes associated with lactate, ethanol, and acetate formation in K. pneumoniae, which resulted in higher titers and yields of 2,3-BD. K. pneumoniae HD79 produces 2,3-BD by a typical production machinery.
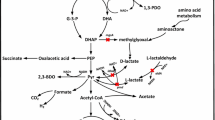
Glucose is used as a substrate in microbial fermentation for the synthesis of several proteins, nucleic acids, and other substances required for cell growth. Microbial fermentation of glucose is usually accompanied by the formation of various metabolites and by-products (Yang et al. 2017). The typical synthetic route for microbial 2,3-BD production is conversion of polysaccharide to pyruvate, α-acetolactate, acetoin, and ultimately 2,3-BD. This process produces ethanol, acetate, lactate, succinate, 1,3-propanediol (1,3-PD), and other by-products, which consume energy, and are not conducive to 2,3-BD (Park et al. 2016) production. Thus, 2,3-BD production could be improved by eliminating or controlling the production of metabolic by-products.
In this work, λRed homologous recombination technology was used to knock out the lactate dehydrogenase and acetate kinase genes ldhA and ack from K. pneumoniae HD79, in order to eliminate lactate and acetate formation, and thereby increase 2,3-BD production. We then studied the effects of ldh and ack on the production of 2,3-BD from glucose fermentation by comparing the transcriptomic profiles of the knockout and wild-type (WT) strains, and identified significantly up- and down-regulated genes.
Materials and methods
Bacterial strains and growth conditions
All strains and plasmids used or created in this work, along with their sources and characteristics are shown in Table 1. E. coli DH5α was grown in Luria–Bertani (LB) medium (1% tryptone, 0.5% yeast extract, and 1% NaCl; w/v). K. pneumoniae HD79, designated the WT strain, was grown at 30 °C in medium containing 150 g/L glucose, 5 g yeast extract, 0.05 g FeSO4·7H2O, 0.001 g ZnSO4·7H2O, 0.001 g MnSO4·H2O, 0.001 g CaCl2·2H2O, 0.25 g MgSO4·7H2O, 6.6 g (NH4)2SO4, 8.7 g K2HPO4, and 6.8 g KH2PO4 in 1 L H2O. The initial pH was adjusted to 7.0 by adding 5 M KOH. The loading volume was 125 mL/250 mL. Flask cultures were maintained at 150 rpm for 12 h. K. pneumoniae HD79-01 was grown with 150 µg/mL chloramphenicol in the aforementioned medium. K. pneumoniae HD79-02 was grown with 150 µg/mL chloramphenicol and 125 µg/mL kanamycin in the same medium.
Generation of ldh and ack knockout mutants
The ldhA (GenBank accession number JX104342.1) of K. pneumoniae HD79 was amplified by polymerase chain reaction (PCR) by using the following primers: ldhAL-forward (5′-GGAAGATCTCAGTACGACAAGAAGTATCTG-3′, BglII), ldhAL-reverse (5′-CCGCTCGAGACCGAAGCCTTTAAGAATGCGCAGC-3′, XhoI); ldhAR-forward (5′-CCGCTCGAGCTTCGGTATGCGCCTGCT-3′, XhoI), and ldhAR-reverse (5′-GGAGGATCCTCAGACCAGCGCGTTAGGG-3′, BamHI). To clone Cmr, pGP704-Cm was used as a template with the following primers: Cmr-forward (5′-CCGCTCGAGGCTTGCGCAGACCAAAACG-3′, XhoI) and Cmr-reverse (5′-CCGCTCGAGGCAGGCATGCAAGCTTGGT-3′, XhoI). The ldhA-L and ldhA-R genes were cloned into pMD18-T (TaKaRa, Dalian, China) to construct pldhA-L and pldhA-R, respectively. The Cmr was ligated into pMD18-T to construct pT-Cmr. Disrupted ldhA was constructed by inserting a Cmr cassette into the middle of ldhA obtained from K. pneumoniae HD79. The pldhA-LR was constructed by inserting the XhoI–EcoRI fragment containing ldhA-R into the XhoI–EcoRI sites of pldhA-L. After using restriction enzyme XhoI to digest pT-Cmr and pldhA-LR, the Cmr fragment was inserted into the XhoI site of pldhA-LR. Finally, pT-LCR containing interrupted ldhA (i.e. ldhAL-Cmr-ldhAR) was linearized with restriction enzymes BglII and BamHI. The fragment ldhAL-Cmr-ldhAR was used in the λRed replacement approach to transform K. pneumoniae HD79 and obtain K. pneumoniae HD79-01.
The ack (GenBank accession number CP003218.1) of K. pneumoniae HD79-01 was amplified by PCR with the following primers: ackL-forward (5′-GGAAGATCTTGAACTGCGGTAGCTCCTCTCTGAA-3′, BglII), ackL-reverse (5′-CCGCTCGAGCTGAATTACGGACTCGTCGATCACC-3′, XhoI); ackR-forward (5′-CCGCTCGAGCAGACCATGCCGGAAGAATCCTATC-3′, XhoI), and ackR-reverse (5′-CGCGGATCCGGTGTCGTGCAGATGGAAGATGATG-3′, BamHI). To clone Kanr, pET-28a(+) was used as a template with the following primers: Kanr-forward (5′-CCGCTCGAGTACATAAACAGTAATACAAGGGGTG-3′, XhoI) and Kanr-reverse (5′-CCGCTCGAGATTAATTCTTAGAAAAACTCATCGA-3′, XhoI). The ack-L and ack-R were cloned into pMD18-T to construct pack-L and pack-R, respectively. The Kanr was ligated into pMD18-T to construct pT-Kanr. Disrupted ack was constructed by inserting the Kanr cassette into the middle of ack obtained from K. pneumoniae HD79-01. The pack-LR was constructed by inserting a XhoI-HindI fragment containing ack-R into the XhoI-HindI sites of pack-L. After using the restriction enzyme XhoI to digest pT-Kanr and pack-LR, the Kanr was inserted into the XhoI site of pack-LR. Finally, pT-LKR containing the interrupted ack (i.e. ackL–Kanr–ackR) was linearized with restriction enzymes BglII and BamHI. The fragment ackL–Kanr–ackR was used in the same approach to transform K. pneumoniae HD79-01 to obtain the K. pneumoniae HD79-02.
Transformation by electroporation
We then introduced linear target fragment ldhAL–Cmr–ldhAR into the genome of K. pneumoniae HD79 by homologous recombination. First, the plasmid pKD46 was transformed into K. pneumoniae HD79; cells were cultured in medium containing 5 mM L-arabinose, inducing expression of Exo, Bet, and Gam. The linear target fragment ldhAL–Cmr–ldhAR was then transformed into K. pneumoniae HD79, which was plated on agar containing 150 μg/mL chloramphenicol. Finally, the target fragment ldhAL–Cmr–ldhAR was amplified by PCR.
The same approach was used to introduce the target fragment ackL–Kanr–ackR into the genome of K. pneumoniae HD79-01, which was plated on agar containing 150 μg/mL chloramphenicol and 125 μg/mL kanamycin. The target fragment ackL–Kanr–ackR was amplified by PCR.
Real-time quantitative polymerase chain reaction to measure ldh and ack
RNA was extracted from K. pneumoniae HD79 and K. pneumoniae HD79-02 by using the RNA Prep Pure Bacteria Kit (Tiangen Biotech Co. Ltd., China). Reverse transcription was performed using the BioRT cDNA First Strand Synthesis Kit (Bioer Technology Co. Ltd., China; Mayer et al. 1995). Real-time Quantitative Polymerase Chain Reaction (qRT-PCR) was performed with at least three biological replicates using SYBR Green-based detection in triplicate in a 7500 Real-Time PCR System (Applied Biosystems, Inc., USA). The 16S rRNA gene was chosen as an internal control to normalize RNA amounts. The following primers were used: ldh (forward 5′-CCGCTCGAGCTTCGGTATGCGCCTGCT-3′, reverse 5′-GGAGGATCCTCAGACCAGCGCGTTAGGG-3′); ack (forward 5′-CCGCTCGAGCAGACCATGCCGGAAGAATCCTATC-3′, reverse 5′-CGCGGATCCGGTGTCGTGCAGATGGAAGATGATG-3′); 16S rRNA (forward 5′-AGAAGAAGCACCGGCTAACTC-3′, reverse 5′-CTCTACGCATTTCACCGCTAC-3′). Diluted cDNA equivalent to 1 ng RNA starting material was used as the template for qRT-PCR amplification. The transcriptional levels of ldh and ack were normalized to the transcriptional level of the 16S rRNA gene. The relative changes (x-fold) at the transcriptional level in different samples were calculated.
SDS-PAGE
SDS-PAGE was performed using the protocol of Bio-Rad Laboratories, Inc. (Hercules, CA, USA). Protein samples were subjected to electrophoresis (JY-SCZ2+ , Beijing JUNYI Electrophoresis Co. Ltd., Beijing, China) at a constant 140 V until the tracking dye reached the bottom of the gel. The gels were stained with Coomassie Brilliant Blue for 15 min and destained with a solution containing 20% methanol and 10% acetic acid. Finally, the target protein bands were viewed and the gray value (the quantified color of target protein band) was analyzed by AlphaImager HP gel imaging system (AlphaImager HP, Protein Simple, USA).
Enzyme assays
The precipitate from 6 mL fermentation broth was centrifuged at 5000 rpm for 10 min at 4 °C, washed twice with potassium phosphate buffer (100 mM, pH 7.4), and resuspended in the same buffer. The supernatant from broken cells was obtained by ultrasonic treatment in an ice-water bath and subsequent centrifugation (10,000 rpm, 10 min, and 4 °C), and used as crude liquor for the measurement of enzyme activities. An enzyme unit is defined as the amount of enzyme required to consume 1 μmol of substrate per minute.
Fermentation kinetics
Batch fermentation was performed in 500 mL flasks containing 150 mL fermentation medium with an initial pH 6.8, with glucose as the substrate. Flasks were cultured at 30 °C with shaking at 150 rpm for 156 h. Samples were harvested every 12 h and transcriptome analysis was performed at 6 h. The sample was diluted tenfold and the optical density (OD) measured at 600 nm was used to describe cell growth. The concentrations of glucose, 1,3-PD, 2,3-BD, ethanol, acetate, succinate, and lactate obtained from the batch fermentation process were determined using a High Performance Liquid Chromatography (HPLC) system (Shimadzu Corp. Kyoto, Japan). The sample was diluted 100-fold, and an Aminex HPX-87H column (Bio-Rad Laboratories) was used with 0.005 M H2SO4 as the elution solvent at a flow rate of 0.8 mL/min, with the column oven temperature maintained at 65 °C with a refractive index detector.
Library preparation
Total RNA was treated with DNase I prior to library construction, and poly-(A) mRNA and double-stranded cDNA were purified using AMPure XP beads. Double-stranded cDNA was further subjected to end-repair with A tailing and ligated to the U-adaptor by using T4 DNA polymerase, Klenow fragment, and T4 polynucleotide kinase, followed by single dNTP base addition using the 3′ to 5′ exo-polymerase activity of Klenow fragment, and ligation to an adapter or index adapter using T4 DNA ligase. Adaptor-ligated fragments were separated on 1% agarose gel, and cDNA fragments of the desired size were excised. The first chain cDNA containing U was degraded using the USER enzyme, and PCR was performed to selectively enrich and amplify cDNA fragments. After validation with an Agilent 2100 Bioanalyzer and ABI StepOnePlus Real-Time PCR System, the cDNA library was subjected to Solexa sequencing using an Illumina HiSeq2000 sequencing platform.
De novo assembly and annotation
Illumina HiSeq 2000/MiSeq was performed to generate sequencing reads, and the NGS QC Toolkit (https://59.163.192.90:8080/ngsqctoolkit/) was used to analyze raw count data, which was compared with the reference genome and reference genes using Bowtie2 (https://bowtie-bio.sf.net/bowtie2; Theissinger et al. 2016). Sequences were annotated using sequential BLAST searches designed to identify the most descriptive annotation for each sequence. Assembled unique transcripts were compared with sequences using the BLAST algorithm, and GO accessions of the highest hits were retrieved and mapped to GO terms according to molecular function, biological process, and cellular component ontologies (https://www.geneontology.org/; Harney et al. 2016). Remaining sequences encoding putative proteins were searched against the SwissProt (https://www.expasy.ch/sprot), KEGG, and COG (https://www.ncbi.nlm.nih.gov/COG) databases, with a typical E-value threshold of less than 10–5 (Kanehisa et al. 2017). Samples from different compartments were analyzed with EdgeR (https://www.bioconductor.org/packages/release/bioc/html/edgeR.html) to determinate the statistical significance of differential expression (Dimont et al. 2015).
Validation of differentially expressed genes by qRT-PCR
qRT-PCR was performed to exclude false positives from high-throughput mRNA sequencing (RNA-seq). Four differentially expressed genes involved in 2,3-BD biosynthesis were chosen; the primers up (5′-GGATCCATGGAACTTCGTTATCTTCGCT-3′) and down (5′-GTCGACTCAGAACATCGCCAGAAACG-3′) were used for amplification.
Statistical analysis
Each experiment was repeated independently three times. JMP 10.0 software was used for statistical analyses. One-way analysis of variance (ANOVA) was used to analyze the differences between the experimental data. P < 0.05 (represented by *) was considered a significant difference; P < 0.01 (represented by **) was considered an extremely significant difference. Sigmaplot 12.5 software was used for statistical data analysis and chart analysis.
Results
Assay of ldh and ack knockout mutants
Characterization of the ldhA and ack knockout mutants is shown in Fig. 1. We observed a clear band of 1765 bp (ldhAL–Cmr–ldhAR), which likely represents homologous recombination of the fragment into the chromosome of K. pneumoniae HD79 (Fig. 1a), and a clear band about 876 bp (Cmr), which was absent in the control strain (Fig. 1b). The mutant strain K. pneumoniae HD79-01 was therefore obtained by integration of the Cmr cassette into the genome of K. pneumoniae HD79.
Amplification of ldhA and Cmr (a, b) and ack and Kanr (c, d). M is the marker of 2000; a Lanes 1–4 represent the genome of K. pneumoniae HD79-01 as template. b Lanes 1–4 represent the genome of K. pneumoniae HD79-01 as the template, Lane 5 represents the genome of K. pneumoniae HD79 as the template. c Lanes 1–4 represent the genome of K. pneumoniae HD79-02 as the template. d Lanes 1–4 show the genome of K. pneumoniae HD79-02 as the template, and lane 5 represents the genome of K. pneumoniae HD79-01 as the template
As shown in Fig. 1c, we observed a clear band of about 1433 bp (ackL-Kanr-ackR), which indicates homologous recombination of this fragment into the chromosome of K. pneumoniae HD79-01. We also observed a clear band of about 813 bp (Kanr), which was not seen in the control strain (Fig. 1d). The mutant strain K. pneumoniae HD79-02 was obtained therefore by integration of the Kanr cassette into the genome of K. pneumoniae HD79-01.
Expression of ldh and ack mRNA
There was a significant reduction in ldh mRNA expression in K. pneumoniae HD79-02 compared to that in K. pneumoniae HD79. The transcription levels at 24 h, 48 h, and 72 h decreased by 76%, 74%, and 66% respectively, indicating successful disruption of ldh expression. There was also a significant reduction in ack mRNA levels in K. pneumoniae HD79-02 compared to those in K. pneumoniae HD79. The transcription levels at 24 h, 48 h, and 72 h decreased by 95.3%, 79.2%, and 76% respectively, indicating successful disruption of ack expression.
Protein expression
The protein expression pattern in K. pneumoniae HD79-02 was studied. Expression of LDH and ACK were analyzed by grayscale values using a gel imaging system. LDH and ACK levels decreased by 72.2% and 59% respectively compared with those in K. pneumoniae HD79 (P < 0.05). Thus, the disruption of ldhA and ack in the recombinant strain K. pneumoniae HD79-02 successfully reduced the expression of LDH and ACK.
Enzyme assays
The LDH and ACK activity in K. pneumoniae HD79-02 were reduced by 55.9% and 73.4% respectively compared with that in K. pneumoniae HD79. Thus, the disruption of ldh and ack in the recombinant strain K. pneumoniae HD79-02 successfully reduced enzyme activity.
Fermentation kinetics analysis
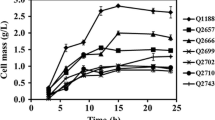
The batch fermentation kinetics of K. pneumoniae HD79 and K. pneumoniae HD79-02 are shown in Fig. 2a. The growth of the WT strain was superior to that of the recombinant strain. The rate of glucose consumption was also significantly higher in the parental strain (Fig. 2b). The concentration of 2,3-BD produced by the mutant strains reached a maximum of 46.21 g/L at 72 h, whereas 2,3-BD concentration for the WT strain was 29.83 g/L (96 h); the 2,3-BD concentration for the recombinant strain was 1.55 times greater than that for the WT strain (P < 0.05). At 72 h, 2,3-BD productivity was 0.64 g/L·h in the recombinant strain and the conversion rate of glucose was 0.47 g/g. For the WT strain, 2,3-BD productivity was 0.31 g/L·h and the conversion rate was 0.39 g/g; these values increased by 106.5% (2,3-BD productivity) and 20.5% (conversion rate of glucose), respectively in the mutant (P < 0.05). The lactate and acetate production in the recombinant strain was significantly lower than that in the WT strain (Fig. 2c). The maximum observed acetate production by the WT strain was 4.58 g/L at 144 h, and that in the mutant strain was 1.59 g/L. Thus, the acetate production in the recombinant strain decreased by 62.8% compared with that in the WT strain (P < 0.05). The output of ethanol by recombinant strain K. pneumoniae HD79-02 was higher than that in the WT strain (maximum 8.11 g/L at 72 h for the mutant strain, 6.36 g/L at 72 h for the parental strain; an increase of 27.5% [P < 0.05]).
Batch fermentation of K. pneumoniae HD79 and K. pneumoniae HD79-02. a black circle, cell growth of K. pneumoniae HD79; open circle, cell growth of K. pneumoniae HD79-02; black triangle, pH of K. pneumoniae HD79 culture; open triangle, pH of K. pneumoniae HD79-02 culture. b black circle, glucose residue for K. pneumoniae HD79; open circle, glucose residue for K. pneumoniae HD79-02; black triangle, concentration of 2,3-BD for K. pneumoniae HD79; open triangle, concentration of 2,3-BD for K. pneumoniae HD79-02. c black square, concentration of lactate produced by K. pneumoniae HD79; open square, concentration of lactate produced by K. pneumoniae HD79-02; black triangle, concentration of ethanol produced by K. pneumoniae HD79; open triangle, concentration of ethanol produced by K. pneumoniae HD79-02; black circle, concentration of acetate produced by K. pneumoniae HD79; open circle, concentration of acetate produced by K. pneumoniae HD79-02. The values are the mean of three independent samples
The typical end products in microaerobic batch fermentations are shown in Table 2 (average values with 95% confidence intervals). Maximum 2,3-BD production by K. pneumoniae HD79-02 was increased by 54.9% compared to that by K. pneumoniae HD79, and occurred 24 h earlier in K. pneumoniae HD79-02. The highest yield of lactate and acetate were reduced by 48.2% and 62.8% respectively in K. pneumoniae HD79-02. Simultaneously, the highest ethanol output increased by 27.5% in K. pneumoniae HD79-02. Production of other by-products such as succinate and 1,3-PD was similar in the mutant and WT strains.
Library construction
A cDNA library was obtained from an equal mixture of RNA isolated from the K. pneumoniae HD79 and HD79-02 samples, which was used for Illumina 90 bp pair-end sequencing, yielding 14,266,260 reads, 1,283,963,400 bp, and 13,419,787 (94.07%) mapped reads for HD79-02, and 14,202,252 reads, 1,278,202,680 bp, and 13,386,385 (94.26%) mapped reads for HD79. Genome sequences were assembled using Trinity software.
Analysis of differentially expressed genes
Klebsiella pneumoniae HD79 and HD79-02 samples for sequencing were collected at 6 h of culture. Following comparative analysis with EdgeR, differentially expressed genes in HD79 and HD79-02 revealed obvious differences between the strains. A total of 4628 differentially expressed genes were identified, of which 404 (of 2312) up-regulated genes and 162 (of 2316) down-regulated genes showed statistically significant differences in expression. The differentially expressed genes were analyzed using MA plot (Fig. 3a), and volcano plot (Fig. 3b) was used to show the overall distribution of differential genes and the abundance in the two transcripts. Genes with unknown function or predicted general functions were excluded, and 27 and 24 kinds of functionally annotated genes were identified.
MA plot (a), Volcano plot (b), the top 20 enriched pathways (c), and GO classification (d) analysis of K. pneumoniae HD79 and HD79-02. a The Y-axis represents the log2 times of each differential gene, the X-axis represents the average of the log2 of the gene. b The Y-axis represents the reported error rate, the X-axis represents the log2 times of each differential gene. Genes that were identified as significantly differentially expressed at a maximum of 0.1% FDR are colored red
Significant enrichment by pathway allowed the identification of the most important biochemical metabolic pathways and signal transduction pathways in the differentially expressed transcripts (Fig. 3c). The first 20 metabolic pathways of differentially expressed genes were enriched. According to the number of differentially expressed genes in the annotation of metabolic pathway genes, the biosynthesis of amino acids metabolic pathway ranked first, followed by the carbon metabolism pathway, the phosphotransferase system, and pyruvate metabolism.
Annotation of predicted proteins
A sequence similarity search was performed against the non-redundant (Nr), UniProtKB/Swiss-Prot (SwissProt), Gene Ontology (GO), Clusters of Orthologous Groups (COG), and Kyoto Encyclopaedia of Genes and Genomes (KEGG) databases, and GO assignments were used to classify the predicted functions of Sativa mycorrhizal genes. Based on sequence homology, 7839 genes were categorized into 28 functional groups (Fig. 3d). The biological function of these genes included the following molecular processes, intracellular/extracellular enzymes, and structural proteins including hypothetical proteins: phosphofructokinase family protein, pullulanase secretion envelope protein, phosphotransferase system associated protein, propanediol utilization protein, LysR family transcriptional regulator, ethanolamine utilization proteins, ABC transporter permease, translocator protein, LysE family, AraC family transcriptional regulator, major facilitator family transporter, DNA-binding proteins, and other proteins associated with 2,3-BD production. All clusters were then searched against the Cluster of Orthologous Groups (COG) database for functional prediction and classification, which enabled assignment of the eight Nr hits as COG classifications. Among the 28 COG categories, the ‘General function prediction’ cluster represented the largest group, followed by ‘replication, recombination, and repair’ and ‘transcription’, while ‘nuclear structure’ and ‘organelle’ were the smallest groups. In each of the three main categories (biological process, cellular component, and molecular function) of the GO classification, metabolic processes, diverse cellular activities, and catalytic activity terms were dominant. Biological processes were related to biological regulation, metabolic regulation, and nuclear transport processes; cell composition, cell membrane composition, and nuclear composition were all functions related to cellular components; and molecular functions were mostly related to nucleic acid transcriptional activators and nuclear transport activities.
Validation of differentially expressed genes by qRT-PCR
The 2,3-BD-related genes KPK_1166, KPK_2399, KPK_4670, and KPK_2279 were selected for verification by qRT-PCR, and their expression in K. pneumoniae HD79-02 was higher than that in HD79, as expected (Fig. 4a). The four genes associated with BudR (KPK_1166, KPK_2399, KPK_4670, KPK_2279) were all clearly up-regulated in HD79-02 compared with the parent strain after 6 h of cell growth, and the amount of acidic by-products were relatively low at this stage. Expression of 2,3-BD biosynthesis-related genes clearly differed between HD79 and HD79-02. Additionally, these genes encoded propanediol and ethanolamine-related proteins that indirectly influenced 2,3-BD production, and were highly expressed in HD79-02.
Discussion
In this study, we observed different growth conditions in the recombinant and WT strains. The reason for this could be that acetate kinase catalyzes the formation of acetate from acetyl phosphate, accompanied by the formation of adenosine triphosphate (ATP). With the deletion of ack, the available energy of the recombinant strain was reduced to some extent, thus affecting the growth rate of the recombinant strain (Förster and Gescher 2014). Another reason could be that the carbon flux of the recombinant strain changed, which affected the distribution and transportation of nutrients in the recombinant strain, thereby affecting the supply of raw materials necessary for growth and acetate metabolism (You et al. 2008). The culture pH of the recombinant strain was slightly higher than that of the WT strain. This could be because of reduced lactate and acetate production due to disruption of ldhA and ack, resulting in a slight increase in the pH of the fermentation broth. Further, glucose consumption was increased; this may be due to the consumption of more substrate to maintain cell growth. On the other hand, it could be due to the elimination of by-products, which increased the availability of the carbon source for the 2,3-BD metabolic pathway and thereby increased the yield of 2,3-BD Kim et al. (2013). successfully knocked out the ldh of K. pneumoniae GSC12206 by λRed homologous recombination and carried out fermentation with glucose as the substrate. The result showed that compared with the WT strain, the lactate production was 92.86% lower than that of the WT strain, while the productivity and yield of 2,3-BD increased by 13% and 60%. Suwannakham et al. (2010) used the λRed homologous recombination technique to knock out ack in P. acidipropionici, which was used in fermentation with glucose as a substrate. They showed that the yield of propionic acetate increased by 13%, while the yield of acetate decreased by 17%. When ldhA and aldH were deleted in K. pneumoniae 2–1, 1,3-PD production was enhanced and the concentration of by-products was reduced (Chen et al. 2016). In the context of these results, we achieved a good increase in 2,3-BD production. Therefore, the knockout effect was very obvious, and it can better explain the relationship between lactic acid and acetic acid and 2,3-BD production.
Lactate and acetate are typical acidic by-products of K. pneumoniae HD79 fermentation to produce 2,3-BD (Guo et al. 2014; Ji et al. 2011) with glucose as a substrate. The production of lactate and acetate consumes energy and inhibits the growth of the host strain. Deletions in recombinant strains are often designed with the intent of maintaining normal physiological metabolic functions; gene knockout therefore often sdoes not cause complete loss of traits or related products, but may considerably reduce their expression. The recombinant strains in this study did not exhibit complete loss of lactate and acetate function; a small amount of lactate and acetate were still produced under conditions of low LDH and ACK activity. This could be because ldh had multiple sequences in different genomic regions of K. pneumoniae HD79, which belong to the multi-copy functional gene (Markert et al. 1975). Therefore, not all ldh was lost, and a small number of genes were still expressed. In addition, lactate may have been produced to balance NADH/NAD+ in the environment. Considering the recombination efficiency factors, it was difficult to screen out multiple cloned knockout clones. After ack knock out, the recombinant strain K. pneumoniae HD79-02 produces a small amount of acetate by other pathways to maintain normal physiological metabolic function. For example, pyruvate may produce a small amount of acetate via pyruvate oxidase (poxB pathway). A small amount of acetate could promote the formation of 2,3-BD; maintaining the concentration of acetate at a lower range was therefore a beneficial result (Sang et al. 2017). Succinate is produced by the action of fumarate reductase on fumaric acid. 1,3-PD is produced by the action of 1,3-PD oxidoreductase on 3-hydroxypropionaldehyde. When the genes of certain products are knocked out, there are repercussions in the production of other products. This could be because of changes in the flow of carbon sources (Nguyen et al. 2018).
In this study, all data were above 88%, indicating a high base reading accuracy during sequencing, and sequencing data were adequate to provide functional information. The GC content from the two strains was similar, which confirmed that the sequencing time was sufficient (Kiran Gopinath et al. 2015). Li et al. used transcriptomics to analyze the vulnerable and stable genes in carotid atherosclerotic plaques; 318 genes were up-regulated by and 363 were down-regulated. Volcanic maps were used to identify differential gene expression between the two groups. These were more illustrative of the authenticity and reliability of the results (Li et al. 2018). Guo et al. changed the expression level and genotype of the rpoD gene to increase the tolerance of K. pneumoniae to xylose and improve the production of 2,3-BD. Differentially expressed genes were mainly involved in signal transduction, membrane transport, carbohydrate metabolism, and energy metabolism (Guo et al. 2018). In summary, by analyzing the differentially expressed genes, it is possible to determine what changes occur in the metabolic pathway in which the gene is involved. A transcriptome analysis of five K. pneumoniae strains showed that differences in gene expression mainly occurred in seven aspects including genes coding 2,3-BD biosynthesis (BudA, BudB, BudC), and resulted in metabolism changes (Lee et al. 2015). This is similar to the results of this study wherein 2,3-BD-related genes were differentially expressed and affected the production of 2,3-BD.
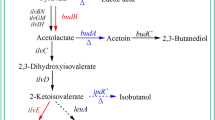
The Klebsiella species were screened particularly closely by whole-genome sequencing by Macrogen Inc. The amino acid sequence of the putative LysR family transcriptional regulatory protein BudR was obtained. Annealing of this sequence with K. pneumoniae MGH 78,578 sequence generated hits based on Cluster of Orthologous Groups (COG) (Heum et al. 2012; Soojin et al. 2013). The 873-nucleotide BudR was located slightly upstream of the BudA (Patel and Jain 2012), and fell into the post translational modification, protein turnover, and chaperones group. In most microorganisms producing 2,3-BD, the key genes (BudB, BudA and BudC) for 2,3-BD production are organized into an operon controlled by the neighboring and divergently transcribed BudR (Ji et al. 2008). In our study, LysR transcriptional regulators were up-regulated in K. pneumoniae HD79-02, suggesting that regulation of BudR expression is important in 2,3-BD biosynthesis. However, the ldh and ack genes encoding proteins involved in the biosynthesis of acidic by-products were down-regulated during the log and stationary phases in HD79-02. Thus, biosynthesis of 2,3-BD likely played a role in diverting pyruvate flux away from the production of acidic substrates such as lactate, formate, and acetate to prevent the culture medium form becoming too acidic for cell growth (Sun et al. 2009; Wang et al. 2019). While these proteins were not directly involved in production of 2,3-BD, they likely shifted the carbon flux towards the biosynthesis of alcohols. The transcriptional regulator BudR regulated the expression of BudA, BudB, BudC, which directly affected the viability of key enzymes producing 2,3-BD, and therefore increased production of 2,3-BD (Fig. 4b). ldh and ack can directly regulate the production of lactic acid and acetic acid. The production of 2,3-BD can be greatly increased by reducing acetic acid and lactic acid. BudR is related to three key genes, BudA, BudB, and BudC, which produce 2,3-BD. As a transcriptional activator to increase 2,3-BD production. In addition, these genes encode propylene glycol and ethanolamine-related proteins, which indirectly affect 2,3-BD production, and they are highly expressed in K. pneumoniae HD79-02 for increasing 2,3-BD production. The lower concentration of acetic acid can promote 2,3-BD production, and the transcriptional activator BudR can also induce the formation of 2,3-BD under acidic conditions. In other studies, we have confirmed acetic acid can indeed promote the expression of BudA and BudB genes at a certain concentration. Therefore, acetic acid may act on the BudR gene, which may increase 2,3-BD production. The biosynthetic pathway of 2,3-BD therefore seems to be regulated by three main factors: the inducers (lactate and acetate), regulator (BudR), and carbon flux. Although the changes in 2,3-BD, lactate, and acetate levels reported in this work were relatively moderate, our results could significantly reduce the production cost of 2,3-BD. Our results greatly improve our understanding of the mechanism of 2,3-BD production and the interaction between acids and alcohols, and provide a sound theoretical basis for further improvement of 2,3-BD production.
References
Białkowska AM (2016) Strategies for efficient and economical 2,3-butanediol production: new trends in this field. World J Microbiol Biotechnol 32:1–14
Chen L, Ma C, Wang R, Yang J, Zheng H (2016) Deletion of ldhA and aldH genes in Klebsiella pneumoniae to enhance 1,3-propanediol production. Biotechnol Lett 38:1769–1774
Dimont E, Shi J, Kirchner R, Hide W (2015) edgeRun: an R package for sensitive, functionally relevant differential expression discovery using an unconditional exact test. Bioinform 31:2589–2590
Förster AH, Gescher J (2014) Metabolic engineering of Escherichia coli for production of mixed-acid fermentation end products. Biotechnol Bioeng 2:16
Gao Y, Huang H, Chen S, Qi G (2018) Production of optically pure 2,3-butanediol from Miscanthus floridulus hydrolysate using engineered Bacillus licheniformis strains. World J Microbiol Biotechnol 34:66
Ge Y, Li K, Li L, Chao G, Zhang L, Ma C, Ping X (2016) Contracted but effective: production of enantiopure 2,3-butanediol by thermophilic and GRAS Bacillus licheniformis. Green Chem 18:4693–4703
Guo X, Cao C, Wang Y, Li C, Wu M, Chen Y, Zhang C, Pei H, Xiao D (2014) Effect of the inactivation of lactate dehydrogenase, ethanol dehydrogenase, and phosphotransacetylase on 2,3-butanediol production in Klebsiella pneumoniae strain. Biotechnol Biofuels 7:44
Guo XW, Zhang Y, Li LL, Guan XY, Guo J, Wu DG, Chen YF, Xiao GD (2018) Improved xylose tolerance and 2,3-butanediol production of Klebsiella pneumoniae by directed evolution of rpoD and the mechanisms revealed by transcriptomics. Biotechnol Biofuels 11:307
Harney E, Dubief B, Boudry P, Basuyaux O, Schilhabel MB, Huchette S, Paillard C, Nunes FLD (2016) De novo assembly and annotation of the European abalone Haliotis tuberculata transcriptome. Mar Genom 28:11–16
Harvianto GR, Haider J, Hong J, Long NVD, Shim JJ, Cho MH, Kim WK, Lee M (2018) Purification of 2,3-butanediol from fermentation broth: process development and techno-economic analysis. Biotechnol Biofuels 11:18
Heum SS, Sewhan K, Jae Young K, Soojin L, Youngsoon U, Min-Kyu O, Young-Rok K, Jinwon L (2012) Complete genome sequence of the 2,3-butanediol-producing Klebsiella pneumoniae strain KCTC 2242. J Bacteriol 194:2736–2737
Ishii J, Morita K, Ida K, Kato H, Kinoshita S, Hataya S, Shimizu H, Kondo A, Matsuda F (2018) A pyruvate carbon flux tugging strategy for increasing 2,3-butanediol production and reducing ethanol subgeneration in the yeast Saccharomyces cerevisiae. Biotechnol Biofuels 11:180
Ji XJ, Huang H, Li S, Du J, Lian M (2008) Enhanced 2,3-butanediol production by altering the mixed acid fermentation pathway in Klebsiella oxytoca. Biotechnol Lett 30:731–734
Ji XJ, Huang H, Ouyang PK (2011) Microbial 2,3-butanediol production: a state-of-the-art review. Biotechnol Adv 29:351–364
Kanehisa M, Furumichi M, Mao T, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45:D353–D361
Kim DK, Rathnasingh C, Song H, Lee HJ, Seung D, Yong KC (2013) Metabolic engineering of a novel Klebsiella oxytoca strain for enhanced 2,3-butanediol production. J Biosci Bioeng 116:186–192
Kiran Gopinath B, Vivek Nagaraj T, Rohit Nandan S, Madavan V (2015) Ameliorated de novo transcriptome assembly using illumina paired end sequence data with trinity assembler. Genomics 5:352–359
Lee S, Kim B, Jeong D, Oh M, Um Y, Kim YR, Kim J, Lee J (2013) Observation of 2,3-butanediol biosynthesis in Lys regulator mutated Klebsiella pneumoniae at gene transcription level. J Biotechnol 168:520–526
Lee S, Kim B, Yang J, Jeong D, Park S, Sang HS, Kook JH, Yang KS, Lee J (2015) Comparative whole genome transcriptome and metabolome analyses of five Klebsiella pneumonia strains. Bioprocess Biosyst Eng 38:2201–2219
Li T, Zhang X, Zhang J, Liu R, Gu C (2018) The transcriptome difference between vulnerable and stable carotid atherosclerotic plaque. Int J Clin Exp Med 11:8988–9004
Markert CL, Shaklee JB, Whitt GS (1975) Evolution of a gene. multiple genes for LDH isozymes provide a model of the evolution of gene structure, function and regulation. Science 189:102–114
Mayer D, Schlensog V, Ck AB (1995) Identification of the transcriptional activator controlling the butanediol fermentation pathway in Klebsiella terrigena. J Bacteriol 177:5261–5269
Nguyen AD, Hwang IY, Lee OK, Kim D, Kalyuzhnaya MG, Mariyana R, Hadiyati S, Kim MS, Lee EY (2018) Systematic metabolic engineering of Methylomicrobium alcaliphilum 20Z for 2,3-butanediol production from methane. Metab Eng 47:323–333
Park JM, Rathnasingh C, Song H (2015) Enhanced production of (R, R)-2,3-butanediol by metabolically engineered Klebsiella oxytoca. J Ind Microbiol Biotechnol 42:1419–1425
Park SW, Lee YJ, Lee WJ, Jee Y, Choi WY (2016) One-step reverse transcription-polymerase chain reaction for ebola and marburg viruses. Int J Env Res Pub Health 7:205–209
Patel RK, Jain M (2012) NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS ONE 7:e30619
Petrov K, Petrova P (2009) High production of 2,3-butanediol from glycerol by Klebsiella pneumoniae G31. Appl Microbiol Biotechnol 84:659–665
Rathnasingh C, Park JM, Kim DK, Song H, Yong KC (2016) Metabolic engineering of Klebsiella pneumoniae and in silico investigation for enhanced 2,3-butanediol production. Biotechnol Lett 38:975–982
Sang JL, Han SC, Chan KK, Thapa LP, Park C, Kim SW (2017) Process strategy for 2,3-butanediol production in fed-batch culture by acetate addition. J Ind Eng Chem 56:157–162
Sikora B, Kubik C, Kalinowska H, Gromek E, Białkowska A, Jędrzejczak-Krzepkowska M, Schüett F, Turkiewicz M (2015) Application of byproducts from food processing for production of 2,3-butanediol using Bacillus amyloliquefaciens TUL 308. Prep Biochem Biotech 46:610–619
Sun LH, Wang XD, Dai JY, Xiu ZL (2009) Microbial production of 2,3-butanediol from Jerusalem artichoke tubers by Klebsiella pneumoniae. Appl Microbiol Biot 82:847–852
Suwannakham S, Huang Y, Yang ST (2010) Construction and characterization of ack knock-out mutants of Propionibacterium acidipropionici for enhanced propionic acid fermentation. Biotechnol Bioeng 94:383–395
Taeyeon K, Sukhyeong C, Sun-Mi L, Min WH, Jinwon L, Youngsoon U, Jin-Ho S (2016) High production of 2,3-Butanediol (2,3-BD) by Raoultella ornithinolytica B6 via optimizing fermentation conditions and overexpressing 2,3-BD synthesis genes. PLoS ONE 11:0165076
Theissinger K, Falckenhayn C, Blande D, Toljamo A, Gutekunst J, Makkonen J, Jussila J, Lyko F, Schrimpf A, Schulz R, Kokko H (2016) De novo assembly and annotation of the freshwater crayfish Astacus astacus transcriptome. Mar Genomics 28:7–10
Wang P, Zhang J, Feng J, Wang S, Guo L, Wang Y, Lee YY, Taylor S, McDonald T, Wang Y (2019) Enhancement of acid re-assimilation and biosolvent production in Clostridium saccharoperbutylacetonicum through metabolic engineering for efficient biofuel production from lignocellulosic biomass. Bioresource Technol 281:217–225
Yang Z, Zhang Z (2018) Production of (2R, 3R)-2,3-butanediol using engineered Pichia pastoris: strain construction, characterization and fermentation. Biotechnol Biofuels 11:35
Yang T, Rao Z, Zhang X, Xu M, Xu Z, Yang ST (2017) Metabolic engineering strategies for acetoin and 2,3-butanediol production: advances and prospects. Crit Rev Biotechnol 37:1–16
You KO, Park SH, Seol EH, Kim SH, Mi SK, Hwang JW, Dewey DY (2008) Carbon and energy balances of glucose fermentation with hydrogen-producing bacterium Citrobacter amalonaticus Y19. J Microbiol Biotechnol 18:532–538
Acknowledgement
This work was supported by the National Natural Science Foundation of China (Grant Nos 31570492, 31770544) and Heilongjiang Provincial Key Laboratory of Plant Genetic Engineering and Biological Fermentation Engineering for Cold Region.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ge, J., Wang, J., Ye, G. et al. Disruption of the lactate dehydrogenase and acetate kinase genes in Klebsiella pneumoniae HD79 to enhance 2,3-butanediol production, and related transcriptomics analysis. Biotechnol Lett 42, 537–549 (2020). https://doi.org/10.1007/s10529-020-02802-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02802-7