Abstract
Objectives
Although abnormal expression of early growth response-1 (Egr1) has been revealed in various human solid tumors, the functions and potential mechanisms of Egr1 in the progression of salivary gland pleomorphic adenoma (SGPA) are not entirely understood.
Results
An elevated expression of Egr1 was observed both in the human salivary gland pleomorphic adenoma tissues and tumor-initiating cell (TIC) cells, when compared with control group. By loss-of-function assay, the proliferation and invasion capacities of TICs were inhibited, while the cell apoptosis was promoted, which were further evidenced by the protein expression analysis of several key apoptosis-related regulators. Furthermore, TICs with Mithramycin A (an Egr1 inhibitor) treatment achieved the same effects of endogenous Egr1 knockdown.
Conclusions
All these data collectively suggest that Egr1 act as an oncogenic factor in salivary gland pleomorphic adenoma, which may be a potential target for the treatment of SGPA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pleomorphic adenoma (PA), also known as “mixed tumor”, is the most common salivary gland tumor, which accounts for 40–70% of all salivary gland tumors. Despite the recent progress in diagnosis and management of PA, the pathogenesis of PA, particularly the recurrent pleomorphic adenoma or carcinoma ex pleomorphic adenoma, remains poorly understood.
Early growth response 1 (Egr1), one of the zinc finger transcription factors, has a crucial role in various diseases processes, such as tumor development and progression (Zhang et al. 2014), chronic obstructive pulmonary diseases (Chen et al. 2008), hypertension (Pagel and Deindl 2012), and myocardial fibrosis (Shen et al. 2019). It has been evidenced that Egr1 may play a dual role (an oncogenic gene or tumor-suppressor gene) in tumor formation, which is tumor-type specific (Chen et al. 2017; Lei et al. 2019; Wu et al. 2002). Our previous study has indicated that Egr1 is a downstream target gene of PLAG1 and that Egr1 may participate in the developing and progression of SGPA (Shen et al. 2011). However, the detailed mechanisms of Egr1’s involvement in SGPA tumorigenesis warrant further explorations.
Investigations of SGPA had been hindered by the lack of appropriate cell lines that resemble the in vivo biology of SGPA. PALG1-transgenic mice develop spontaneous submandibular tumors which closely resemble the human SGPA. More interestingly, CD44hi cells isolated from the PLAG1 transgenic mice submandibular tumors have been characterized as “tumor-initiating cells” TICs. The isolation of TICs from PLAG1 transgenic mice offers a reliable tool for SGPA research (Shen et al. 2011).
To the best of our knowledge, there have been no studies addressing the association between Egr1 and cellular biological behaviors in SGPA. The objective of the present study is to analyze the expression and intervention endogenously or exogenously of Egr1 in SGPA, aiming to shed new insights into the pathogenesis of SGPA or provide novel therapeutic target of SGPA.
Methods and materials
Patient population and generation of stable cells
A total of 18 pairs of tissue samples were retrieved from patients with primary PA of the parotid gland who had received surgical treatment at the Shanghai Ninth People’s Hospital. The present study was approved by the Ethics Committee of Ninth People’s Hospital, Shanghai Jiao Tong University School of Medicine. Informed consent was obtained from each patient. The PLAG1 transgenic mice were generated and maintained as previously described (Wang et al. 2013; Xu et al. 2019). All the procedures described were approved by the Animal Use and Care Committee of Shanghai Jiao Tong University School of Medicine. Then, single cell suspensions from the tumors of the transgenic mice were obtained via a process of mincing, digestion, blocking, filtering, centrifugation and resuspension. Subsequently, the cells were cultured in RPMI 1640 medium (Invitrogen, CA, USA) supplemented with 10% FBS, 20 mM Hepes and 2 mM L glutamine. Cells were routinely incubated at 37 °C and 5% CO2 under a humidified atmosphere. Following digestion with 0.25% trypsin EDTA (Invitrogen), the single cell suspensions were subsequently sorted into CD44hi cells (TICs) and CD44low cells (Control) by flow cytometric analysis.
RNA extraction and real time-PCR
Total RNA was extracted from cells using TRIzol reagent (Invitrogen), reverse transcribed and amplified as previously reported (Xu et al. 2017). The primer sequences were: Egr1, 5′-CCACAACAACAGGAGACCTG-3′/5′-AGGCTTTAATAGTGGATAGTGGAG-3′, and β-actin, 5′-AACCCTAAGGCCAACCGTGAAAAG-3′/5′-TGGCGTGAGGGAGAGCATAGC-3′. The relative quantification of gene expression was performed using 2−ΔΔCT method (Xu et al. 2017).
Western blotting analysis
Whole protein lysates were obtained from harvested cells using protein RIPA lysis buffer (Beyotime, Shanghai, China) containing 10 mM phenylmethylsulphonyl fluoride as a protease inhibitor (Beyotime Institute of Biotechnology) on ice for 30 min. Equal amounts of protein samples were loaded and separated by SDS PAGE and transferred to PVDF membranes (EMD Millipore), which were subsequently blocked with 5% non-fat dried milk powder in 0.05% Tris buffered saline and Tween 20 (TBST) for 1 h at room temperature. These blots were incubated at 4 °C overnight with primary antibodies against Egr1 (all from Abcam; 1:1000 dilution), p21, p53, Bax, Bim, Caspase 3 and Caspase 9 (all from Bioss; 1:1000 dilution), β-actin (Bioss, 1:3000 dilution). The secondary antibodies conjugated with horseradish peroxidase label (ThermoFisher), were diluted at 1:1000, and the membranes were incubated for 1 h at room temperature. Finally, the immunoreactive bands were scanned and analyzed. The relative levels of each protein were quantified with Quantity One software (Bio Rad Laboratories, Inc.), and the levels were normalized against β-actin, which served as a control.
Immunohistochemistry
Immunohistochemistry analysis was conducted with indicated antibodies as previously described (Zhao et al. 2016). Briefly, Paraffin-embedded tissue sections were deparaffinized and rehydrated. The tissue sections were incubated with a polyclonal anti-human Egr1 antibody, and then stained using the Dako Real™ EnVision™ horseradish peroxidase-3, 3′-daiminobenzidine detection system.
Flow cytometric analysis
Flow cytometry was utilized for assessment of cell sorting and apoptotic analysis. Cell sorting was performed using a BD FACS Aria™ cell sorter (BD Medical Technology). Briefly, the single cell suspensions were sorted into CD44 hi and CD44low cells with APC anti CD44 allophycocyanin (eBioscience). Once sorted, the collected cells were cultured in complete media supplemented with 1% penicillin–streptomycin solution. With respect to the apoptosis assay, the prepared cells were collected and stained using a PE Annexin V Apoptosis Detection kit (BD Pharmingen) according to the manufacturer’s protocol. The apoptotic rate of the cells for each experimental group were counted and analyzed.
Cell immunofluorescence
Immunofluorescence analyses of Egr1 were performed using a standard protocol (Xu et al. 2017). The TICs or CD44 low cells were seeded and grown on glass coverlips 24 h prior to the staining experiment, fixed with 4% paraformaldehyde, permeabilized with 0.2% Triton-X 100 (Sigma), and incubated with primary antibody against Egr1, then secondary antibody. Immunofluorescence was visualized and captured under fluorescence microscope.
Egr1 vectors construction and cell transfection
The CDS (Coding sequence) of mouse Egr1 (NM_007913.5,1602 bp) was amplified using the primers 5′-GGAATTCATGGCAGCGGCCAAGGCCG-3′ and 5′-CGGGATCCTTAGCAAATTTCAATTGTCCTGG-3′, which contained an EcoRI cutting site and kozak sequence (5′-GCCACC-3′) and a BamHI cutting site, respectively. The recombinant vector was named pEgr1. A siRNA sequence that complementarily bound to Egr1 was chosen. The target sequences of siRNA (5′-GCCCTTCCAGTGTCGAATC-3′) were homologous to Egr1 (pshEgr1). The products of the vectors were confirmed by DNA sequencing. One day before the transfection, TICs in logarithmic phase were seeded in 6-well plates at 5 × 105 cells/well. Then 2 μg of expression vectors or shRNA were transfected using Lipofectamine 2000 (Invitrogen, CA, USA) in accordance with the manufacturer’s protocol.
MMA treatment
Cells of MMA treatment group was added to medium at a final concentration (0.03, 0.3 or 3 μM) of MMA (Sigma, USA). Cells cultured for 24 h under normal conditions after transfection or MMA-treatment were used for proliferative activity assay, apoptosis and invasion assay, and related protein expression detection.
Cell proliferation assay
Cell proliferation was determined using the cell counting kit 8 (CCK 8) assay. Briefly, cells were plated at a density of 5 × 104 cells/well in 96 well plates. At the indicated time points (24, 48 and 72 h), cells were treated with 10 µl CCK8 solution (Dojindo Molecular Technologies, Inc., Shanghai, China) and subsequently incubated for 4 h at 37 °C. The absorption at 450 nm was measured using an automatic enzyme linked immunosorbent assay reader (BioTek Instruments, Inc.). Each experiment was performed at least three times.
Cell invasion assay
Cell invasion experiments were performed as previously described (Zhao et al. 2016). The transwell (6.5 mm) with 8 μm pore polycarbonate membrane inserts (Corning, Albany, NY) were imbedded with 120 μg and 100 μg gelatin in DMEM. The differently treated TIC cells were added to the inserts in serum-free medium, then the inserts were placed into the bottom chambers containing 10% FBS. The cells in the upper chamber were carefully removed with cotton swab, and the cells that invaded through matrigel were stained with 0.1% crystal violet (Beyotime, China). Images of at least 10 random fields per chamber were photographed, counted and analyzed.
Statistically analysis
All data showed here are expressed as the mean ± SD. Statistical analyses were conducted using Student’s t test or ANOVA unless otherwise stated. A value of p < 0.05 was considered statistically significant.
Results
Overexpression of Egr1 in human SGPA specimens
To investigate the role of Egr1 in human SGPA, we first examined the mRNA expression of Egr1 in fresh human SGPA tissues (Tumor group) and paired normal gland tissues (Gland group) with real time-PCR. As shown in Fig. 1a, the mRNA of Egr1 was significantly up-regulated in the SGPA tissues. Consistent with this, Egr1 protein levels, as determined by western blots, were markedly elevated, when compared with the normal gland tissues (Fig. 1b). Moreover, we conducted immuno-histochemical staining with 18 pairs of formalin-fixed, paraffin-embedded tumor or gland tissues. An negative or weakly positive staining of Egr1 was detected in the normal gland tissues; while in the 18 cases of PA tumor specimens, 14 cases (14/18, 77.8%) showed an overexpression of Egr1. The representative images of Egr1 staining in Gland or Tumor group were presented in Fig. 1c. These data showed that the expression of Egr1 was significantly increased in human SGPA tissues.
Elevated expression of Egr1 in the human SGPA specimens. a Relative Egr1 mRNA expression was detected by RT-PCR in salivary gland or tumor tissues. b Western blot analysis of the protein expression of Egr1 in salivary gland or tumor tissues. c Representative immunohistochemical staining of Egr1 in salivary gland (left) or PA tumor (right) specimens. × 100, *p < 0.05
Expression of Egr1 in TICs of SGPA
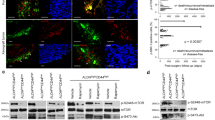
Next, the expressions of Egr1 in TICs (CD44hi cells) were analyzed by real-time PCR, western blot and immunofluorescence assays, using the CD44low cells as control. According to the real-time PCR (Fig. 2a), western bolt (Fig. 2b) and immunofluorescence (Fig. 2c) results, when compared with CD44low cells, the expression of Egr1 was markedly upregulated in TICs, which was consistent with the overexpression of Egr1 in human SGPA tissues (Fig. 1).
Up-regulated Egr1 in SGPA tumor-initiating cells. a Relative Egr1 mRNA expression was detected by RT-PCR in CD44low or CD44hi (TICs) cells. b Western blot analysis of the protein expression of Egr1 in CD44low or CD44hi (TICs) cells. c Representative immune-fluorescence staining of Egr1 in CD44low or CD44hi (TICs) cells, *p < 0.05, **p < 0.01
Effects of upregulation or downregulation of endogenous Egr1 on the proliferation and invasion of TICs
To further investigate the specific role of Egr1 in SGPA, we selectively overexpressed or depleted endogenous Egr1 through transfection with pEgr1 or pshEgr1 (shRNA) in TICs, respectively. As shown in Fig. 3a, the western bolt results indicated that the Egr1 protein expression was significantly increased in the pEgr1 group, when compared with or without empty vector group. The protein levels of Egr1 were significantly decreased in the pshEgr1 group, when compared to the Control group. Subsequently, the biological behavior of TICs with up- or down-regulation of Egr1 was interrogated. The proliferative capacity of tumor cells was examined with CCK-8 assay, while the invasion capacity was determined with transwell assay. As displayed in Fig. 3b and c, the result indicated that overexpression of Egr1 in the pEgr1 group promoted cell proliferation and invasion, while the shRNA-mediated Egr1 knockdown resulted in impaired cell proliferation and invasion. These results indicate that Egr1 could serve as a potential oncogenic regulator for the tumorigenesis of pleomorphic adenoma, and that genetic manipulation of Egr1 may perturbate the proliferation and invasion abilities of TICs from SGPA.
Effects of up- or down- regulation of endogenous Egr1 on the proliferation and invasion capacities of the TICs. a Western blot analysis of the protein expression of Egr1 in the Control, Vector, pEgr1 and pshEgr1 group. b TICs were transfected with siRNAs for 24 h before the proliferation assays. The CCK8 assay detection of the differently treated TICs groups. c TICs were transfected with siRNAs for 24 h, starved for another 24 h, and then invasion was analyzed using transwell assays. d Quantitative results for the invasion abilities for different group cells. Data showed here are mean ± SD from three independent experiments, *p < 0.05, **p < 0.01, #p > 0.05
Susceptibility of TICs to reduced proliferation or invasion induced by MMA treatment
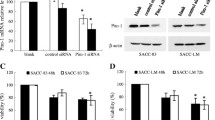
It has been demonstrated that MMA was an effective inhibitor of Egr1 (Yu et al. 2018; Zhao et al. 2015). To determine whether this inhibitory effect of MMA on the Egr1 expression applies to SGPA, we determined the Egr1 level changes in TICs after MMA treatment. The TICs were incubated with different concentrations (0.03 μM, 0.3 μM or 3 μM) of MMA and then subjected to western blot analysis. As shown in Fig. 4a, the MMA induced a dose-dependent decrease of Egr1 in TICs. We then analyzed the proliferation or invasion capacities alteration in TICs following MMA treatment. The CCK-8 assay results indicated that the proliferative ability was significantly decreased following 3 μM MMA incubation (Fig. 4b). Similarly, the invasive properties of TICs were reduced pronouncedly when treated with MMA as determined by the Matrigel invasion assay (Fig. 4c, d). Taken together, these data suggest that MMA inhibits cell proliferation and invasion probably through downregulation of Egr1.
MMA inhibits cell growth and invasion capacities of TICs. a MMA inhibits Egr1 in a dose-dependent manner in TICs. b The CCK8 assay of the proliferation abilities of TICs with different concentrations of MMA. c Invasion potentials of TICs treated with MMA or vehicle were determined by transwell assays. d Quantitative results for the invasion abilities for four differently treated groups of TICs. Data showed here are mean ± SD from three independent experiments, *p < 0.05, **p < 0.01, #p > 0.05
Effects of selectively overexpression or knockdown of Egr1 and MMA treatment on the apoptosis of TICs
Egr1 is a pivotal gene in cell signaling, which is a potent transcription activator of key genes involved in cell death pathway, such as p53 (Ahmed et al. 1997) and TNF-α (Ahmed et al. 1996). Therefore, we next analyzed the role of Egr1 on apoptosis of TICs. Again, we used genetic tools to overexpress (pEgr1) or decrease (pshEgr1) Egr1 expression, as well as pharmacological inhibition of Egr1 using MMA. The results from apoptotic assay showed that TICs with decreased Egr1 levels through pshEgr1 exhibited significant increase of cell apoptosis, while TICs with increased Egr1 levels through pEgr1 exhibited a remarkable decrease of cell apoptosis (Fig. 5a). The proportions of apoptotic cells decreased from 20.3 ± 3.0% (Control group) to 12.3 ± 1.7% in the pEgr1 group, but increase to 39.2 ± 3.9% in the pshEgr1 group (Fig. 5b). The relative abundance of several key apoptosis-related proteins (including p21, p53, Bax, Bim, caspase-3 and caspase-9) were measured by western blot. The results demonstrated that the expression levels of these proteins were significantly dampened in the pEgr1 group; while increased markedly in the pshEgr1 group (Fig. 5c). Moreover, as shown in Fig. 5d, e, MMA treatment of TICs significantly increased apoptosis (Vehicle: 18.8 ± 3.7%; 0.03 μM: 20.3 ± 4.9%; 0.3 μM: 30.1% ± 3.3; 3 μM: 52.2 ± 6.4%). The abundance of apoptosis-related proteins was also increased in a dose-dependent manner in TICs incubated with MMA, which indicated that Egr1 inhibition by MMA phenocopies the effects of Egr1 knockdown using shRNA pshEgr1 (Fig. 5f). Taken together, these data suggested that both Egr1 knockdown using shRNA and inhibition using MMA promoted the apoptosis of TICs, presumably through Egr1’s influence on apoptosis-related proteins.
a Flow cytometry of apoptosis in TICs 72 h after treatment with siRNAs. b The percentages of TIC cells undergoing apoptosis were quantitatively analyzed. c Western blot analysis of apoptosis-related proteins in differently treated TICs. d Apoptosis of TICs 72 h after incubation with MMA or vehicle were measured by flow cytometry. e The percentages of TIC cells undergoing apoptosis were analyzed accordingly. f Western blot analysis of apoptosis-related proteins in TICs after vehicle or MMA treatment. Data showed here are mean ± SD from three independent experiments, *p < 0.05, **p < 0.01, #p > 0.05
Discussion
Early growth response 1 (Egr1) is a multifunctional transcription factor, which influences cell proliferation, apoptosis, immune activation, and matrix degradation (Li et al. 2019). In the present study, we provided evidence that Egr1 was overexpressed both in human SGPA specimens and TIC cells from PLAG1 transgenic mice. Moreover, the oncogenic role of Egr1 in PA was interrogated through genetic and pharmacological approaches.
There is accumulating evidence indicates that the expression of Egr1 is tumor-specific and characterized by spatio-temporal specificity. On one hand, in gastric cancer, Egr1 mRNA expression was significantly higher in gastric cancer tissues and cultured gastric cancer cells, which revealed the function of Egr1 as a key regulatory factor (Kobayashi et al. 2002). A high expression of Egr1 protein was also observed in laryngeal and hypopharyngeal squamous cell carcinoma (Yoon et al. 2015). On the other hand, Chen and colleagues reported that the expression levels of Egr1 in human gliomas are significantly reduced in contrast to normal brain tissues (Chen et al. 2017). It’s also been established that Egr1 expression was down-regulated in nasopharyngeal carcinoma tissues when compared with normal nasopharynx tissue specimens (Zhong et al. 2017). Herein, we demonstrated that the expression of Egr1 was elevated both in human SGPA tissues and the TIC cells from PLAG1 transgenic mice. This is consistent with our previous result, which suggested that Egr1 could be targeted by PLAG1, a key mediator in PA tumorigenesis (Shen et al. 2011).
The specific function of Egr1 in oncogenesis is controversial, including both oncogenic and tumor-suppressive potentials. For example, in patients with nasopharyngeal carcinoma, the survival rate with high Egr1 expression was higher than those with low/negative Egr1 expression and low Egr-1 expression was a risk factor for poor prognosis (Zhong et al. 2017). In rhabdomyosarcoma, it’s reported that Egr1 may inhibit the expression of cell cycle regulators (including p21, PTEN, NDRG1 and CST6) through interaction with TBX2 and activate two pro-apoptotic factors (BAX and dephosphorylated BAD) (Mohamad et al. 2018). These findings suggest that Egr1 could function as a tumor-suppressive gene in certain tumors. However, in glioma cell lines, silencing of Egr1 gene inhibited proliferation and induced G1 phase arrest through directly targeting CCND1 (Chen et al. 2017). It has also been demonstrated that Egr1 could promote tumor metastasis via regulating angiogenic and osteoclastogenic factors in prostate cancer (Li et al. 2019), which indicated an oncogenic role of Egr1 in the development and progression of prostatic cancer. In the current study, Egr1 was shown to act as an oncogenic gene in PA tumorigenesis, including promoting cell proliferation and invasion, using gain- and loss-of-function assays as well as pharmacological inhibition approaches. More recently, studies indicate that Egr1 could exert its function through interaction with different long non-coding RNAs or micro-RNAs, such as taurine upregulated 1 (TUG1) (Shi et al. 2019), hepatocyte nuclear factor 1 homeobox A antisense RNA 1 (HNF1A-AS1) (Liu et al. 2018), miR-150-5p (Shen et al. 2019) and miR-301b (Yan et al. 2017), which might underlie the different function of Egr1 in different types of tumors. Our unpublished data also indicated that in salivary gland pleomorphic adenoma, there may be close interaction between Egr1 and several long non-coding RNAs and micro-RNAs. The more specific regulatory mechanism of Egr1 in the tumorigenesis of SGPA warrants further investigation.
Previous studies have indicated that Egr1 may function as a pivotal regulator in cell apoptosis process. Xie et al. established that Bim may act as an Egr1 target gene in neurons, uncovering a novel Egr1/Bim pathway in neuronal apoptosis (Xie et al. 2011). In pancreatic cancer cells, Egr1 may act as a proapoptotic factor via induction of Bax (Wang et al. 2015). Indeed, our data indicated that genetic knockdown or pharmacological inhibition lead to enhanced apoptosis of tumor cells while overexpression of Egr1 inhibited the apoptosis of TICs (Fig. 5). This result was further supported by the protein expression of six key regulators in cell apoptosis, including p21, p53, Bax, Bim, Caspase-3 and Caspase-9. Among these, Bax is a chief executioner protein of the apoptotic pathway, which induces apoptosis due to a caspase-mediated mitochondrial pathway (Walensky 2019). Caspase-3 and Caspase-9 are belonging to the caspases family, which are considered as the primary hallmarks of apoptosis (Van Opdenbosch and Lamkanfi 2019). Taken together, our results suggest that Egr1 might exert an oncogenic role in the development of SGPA partially through inhibition of cell apoptosis.
Mounting evidence has demonstrated that Egr1 may play a crucial role in radiation-signaling since the promoter of Egr1 contains radiation-inducible DNA sequences (Oben et al. 2017; Yoon et al. 2015). Ahmed et al. have reported that Egr1 may function as a key gene that initiates early signal transduction events in response to ionizing radiation leading to either cell growth arrest or cell death in tumor cells (Ahmed 2004). Other investigators reported that silencing Egr1 could induce tumor cells death while attenuate radiation-induced apoptosis in normal tissues, suggesting that knockdown of Egr1 could enhance tumor local control rates while minimize the normal tissue complications associated with radiotherapy (Zhao et al. 2015). Together with these findings, it’s plausible that Egr1 may be a potential therapeutic target in recurrent PA or carcinoma ex pleomorphic adenoma, which needs radiotherapy as a supplementary treatment option in addition to surgery.
Conclusion
In summary, the present study indicates that Egr1 is overexpressed and plays an oncogenic role in SGPA, which may be accomplished through promoting proliferation, invasion and inhibiting apoptosis of TICs. Our study thus sheds new light on the pathogenesis of SGPA.
References
Ahmed MM (2004) Regulation of radiation-induced apoptosis by early growth response-1 gene in solid tumors. Curr Cancer Drug Targets 4:43–52
Ahmed MM et al (1996) EGR-1 induction is required for maximal radiosensitivity in A375-C6 melanoma cells. J Biol Chem 271:29231–29237. https://doi.org/10.1074/jbc.271.46.29231
Ahmed MM et al (1997) Ionizing radiation-inducible apoptosis in the absence of p53 linked to transcription factor EGR-1. J Biol Chem 272:33056–33061. https://doi.org/10.1074/jbc.272.52.33056
Chen ZH et al (2008) Egr-1 regulates autophagy in cigarette smoke-induced chronic obstructive pulmonary disease. PLoS ONE 3:e3316. https://doi.org/10.1371/journal.pone.0003316
Chen DG et al (2017) Inhibition of EGR1 inhibits glioma proliferation by targeting CCND1 promoter. J Exp Clin Cancer Res 36:186. https://doi.org/10.1186/s13046-017-0656-4
Kobayashi D et al (2002) Overexpression of early growth response-1 as a metastasis-regulatory factor in gastric cancer. Anticancer Res 22:3963–3970
Lei T, Zhu X, Zhu K, Jia F, Li S (2019) EGR1-induced upregulation of lncRNA FOXD2-AS1 promotes the progression of hepatocellular carcinoma via epigenetically silencing DKK1 and activating Wnt/beta-catenin signaling pathway. Cancer Biol Ther 20:1007–1016. https://doi.org/10.1080/15384047.2019.1595276
Li L et al (2019) EGR1 regulates angiogenic and osteoclastogenic factors in prostate cancer and promotes metastasis. Oncogene. https://doi.org/10.1038/s41388-019-0873-8
Liu HT, Liu S, Liu L, Ma RR, Gao P (2018) EGR1-mediated transcription of lncRNA-HNF1A-AS1 promotes cell-cycle progression in gastric cancer. Cancer Res 78:5877–5890. https://doi.org/10.1158/0008-5472.can-18-1011
Mohamad T, Kazim N, Adhikari A, Davie JK (2018) EGR1 interacts with TBX2 and functions as a tumor suppressor in rhabdomyosarcoma. Oncotarget 9:18084–18098. https://doi.org/10.18632/oncotarget.24726
Oben KZ et al (2017) Radiation induced apoptosis of murine bone marrow cells is independent of early growth response 1 (EGR1). PLoS ONE 12:e0169767. https://doi.org/10.1371/journal.pone.0169767
Pagel JI, Deindl E (2012) Disease progression mediated by egr-1 associated signaling in response to oxidative stress. Int J Mol Sci 13:13104–13117. https://doi.org/10.3390/ijms131013104
Shen S et al (2011) Tumor-initiating cells are enriched in CD44(hi) population in murine salivary gland tumor. PLoS ONE 6:e23282. https://doi.org/10.1371/journal.pone.0023282
Shen J et al (2019) MiR-150-5p retards the progression of myocardial fibrosis by targeting EGR1. Cell Cycle 18:1335–1348. https://doi.org/10.1080/15384101.2019.1617614
Shi B et al (2019) Upregulation of long noncoding RNA TUG1 by EGR1 promotes adenomyotic epithelial cell migration and invasion through recruiting EZH2 and suppressing TIMP2. Mol Reprod Dev 86:239–247. https://doi.org/10.1002/mrd.23099
Van Opdenbosch N, Lamkanfi M (2019) Caspases in cell death, inflammation, and disease. Immunity 50:1352–1364. https://doi.org/10.1016/j.immuni.2019.05.020
Walensky LD (2019) Targeting BAX to drug death directly. Nat Chem Biol 15:657–665. https://doi.org/10.1038/s41589-019-0306-6
Wang Y et al (2013) Opposing functions of PLAG1 in pleomorphic adenoma: a microarray analysis of PLAG1 transgenic mice. Biotechnol Lett 35:1377–1385. https://doi.org/10.1007/s10529-013-1213-7
Wang C et al (2015) EGR-1/Bax pathway plays a role in vitamin E delta-tocotrienol-induced apoptosis in pancreatic cancer cells. J Nutr Biochem 26:797–807. https://doi.org/10.1016/j.jnutbio.2015.02.008
Wu MY, Liang YR, Wu XY, Zhuang CX (2002) Relationship between Egr-1 gene expression and apoptosis in esophageal carcinoma and precancerous lesions. World J Gastroenterol 8:971–975. https://doi.org/10.3748/wjg.v8.i6.971
Xie B, Wang C, Zheng Z, Song B, Ma C, Thiel G, Li M (2011) Egr-1 transactivates Bim gene expression to promote neuronal apoptosis. J Neurosci 31:5032–5044. https://doi.org/10.1523/JNEUROSCI.5504-10.2011
Xu WL et al (2017) In situ release of VEGF enhances osteogenesis in 3D porous scaffolds engineered with osterix-modified adipose-derived stem cells. Tissue Eng Part A 23:445–457. https://doi.org/10.1089/ten.TEA.2016.0315
Xu W, Liu L, Lu H, Fu J, Zhang C, Yang W, Shen S (2019) Dysregulated long noncoding RNAs in pleomorphic adenoma tissues of pleomorphic adenoma gene 1 transgenic mice. Mol Med Rep 19:4735–4742. https://doi.org/10.3892/mmr.2019.10149
Yan L, Wang Y, Liang J, Liu Z, Sun X, Cai K (2017) MiR-301b promotes the proliferation, mobility, and epithelial-to-mesenchymal transition of bladder cancer cells by targeting EGR1. Biochem Cell Biol 95:571–577. https://doi.org/10.1139/bcb-2016-0232
Yoon TM et al (2015) EGR1 regulates radiation-induced apoptosis in head and neck squamous cell carcinoma. Oncol Rep 33:1717–1722. https://doi.org/10.3892/or.2015.3747
Yu Q, Huang Q, Du X, Xu S, Li M, Ma S (2018) Early activation of Egr-1 promotes neuroinflammation and dopaminergic neurodegeneration in an experimental model of Parkinson’s disease. Exp Neurol 302:145–154. https://doi.org/10.1016/j.expneurol.2018.01.009
Zhang H et al (2014) EGR1 decreases the malignancy of human non-small cell lung carcinoma by regulating KRT18 expression. Sci Rep 4:5416. https://doi.org/10.1038/srep05416
Zhao DY, Jacobs KM, Hallahan DE, Thotala D (2015) Silencing Egr1 attenuates radiation-induced apoptosis in normal tissues while killing cancer cells and delaying tumor growth. Mol Cancer Ther 14:2343–2352. https://doi.org/10.1158/1535-7163.MCT-14-1051
Zhao LY et al (2016) The overexpressed functional transient receptor potential channel TRPM2 in oral squamous cell carcinoma. Sci Rep 6:38471. https://doi.org/10.1038/srep38471
Zhong JT, Wang HJ, Yu J, Zhang JH, Wang SF, Yang X, Su W (2017) Correlations of the expressions of c-Jun and Egr-1 proteins with clinicopathological features and prognosis of patients with nasopharyngeal carcinoma. Cancer Biomark 19:213–220. https://doi.org/10.3233/CBM-161710
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81302359), Fundamental research program funding of Ninth People’s Hospital affiliated to Shanghai Jiao Tong University School of Medicine (JYZZ086B) and the 6th session of hospital outstanding Youth Funding of Ninth People’s Hospital affiliated to Shanghai Jiao Tong University School of Medicine.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Supporting information
Supplementary Table 1—The sequences used in this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shen, S., Zhu, Y., Lu, H. et al. Transcription factor early growth response-1 plays an oncogenic role in salivary gland pleomorphic adenoma. Biotechnol Lett 42, 197–207 (2020). https://doi.org/10.1007/s10529-019-02776-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-019-02776-1