Abstract
Over-expression of the proto-oncogene pleomorphic adenoma gene 1 (PLAG1) plays a crucial role in the formation of pleomorphic adenoma, which is the most common type of salivary gland tumor. To understand the molecular mechanisms governing PLAG1-mediated tumorigenesis, we used a microarray-based approach to identify PLAG1 target genes. We validated the expression of several genes, including Bax, Fas, p53, p21, p16, Cyclin D1, Egfr, Trail-R/DR5, c-Fos, c-myc and Igf2, by real-time RT-PCR or western blotting. Using luciferase reporter gene assays, we determined that the promoters of Bax, Fas, p53, TRAIL-R/DR5, and c-Fos were transactivated by PLAG1. PLAG1 not only activates genes that promote cell proliferation and tumor formation but also genes that inhibit these cellular processes. Therefore, we conclude that PLAG1 may play a dual role in tumor formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pleomorphic adenoma (PA), which accounts for more than 50 % of all salivary gland neoplasms, or 87–93 % of benign epithelial salivary gland tumors, is the most common type of salivary gland tumor. While most tumors are surgically removed by radical resection, recurrences can occur even after several decades. Some patients may even undergo malignant transformation (carcinoma ex pleomorphic adenoma) (Enlund et al. 2002; Landrette et al. 2005; Voz et al. 2004) with cervical lymphatic and/or distant metastasis. Therefore, pleomorphic adenoma, which is traditionally thought of as a benign lesion, displays several characteristics of a malignant tumor.
Based on current studies of salivary gland tumors, pleomorphic adenoma gene 1 (PLAG1) on chromosome 8q12 plays a critical role in the pathogenesis and development of pleomorphic adenomas. Ectopic PLAG1 activation has also been found in lipoblastomas, hepatoblastomas, leiomyomas, and leiomyosarcomas. PLAG1 is upregulated in approx. 20 % of human acute myeloid leukemia (AML) (Landrette et al. 2005). Thus, PLAG1 plays an important role in tumorigenesis, not only in the formation of pleomorphic adenomas.
PLAG1 is a transcription factor that contains seven canonical C2H2 zinc-finger domains and a serine-rich C-terminus. It binds DNA at a bipartite consensus binding site which contains a core sequence (GRGGC) and a G-cluster (RGGK) that are separated by six to eight random nucleotides (Voz et al. 2004). Igf2 had been identified be activated by PLAG1, and we had also identified four PLAG1 consensus binding sites in the promoter region of mouse β-catenin, a component of the Wingless/Wnt signaling pathway. Additionally, early growth response-1 (Egr-1) (Shen et al. 2011) was significantly upregulated in tumors raised from our PLAG1 transgenic mice. These findings suggest that Igf2, β-catenin, and Egr-1 may be PLAG1 target genes and these genes may be critical downstream mediators of PLAG1-induced pleomorphic adenomas.
We previously established transgenic mice expressing human PLAG1 to evaluate the oncogenic activity of PLAG1 in vivo: the salivary gland tumors that developed in our transgenic mice shared the main pathologic features with human pleomorphic adenomas (Zhao et al. 2006). To identify target genes of PLAG1 and the molecular events occurring throughout the development of pleomorphic adenomas, we performed a signal transduction-focused microarray, which contained 96 genes involved in 15 different signaling pathways. Finally, we found that PLAG1 induces expression of multiple genes, including those that promote apoptosis and those that promote cell proliferation and tumor formation.
Materials and methods
MMTV–PLAG1 transgenic mice
MMTV–PLAG1 transgenic mice were generated at the Experimental Animal Center, Department of Medical Genetics and were maintained on the C57BL/6 background. All transgenic mice used in this study were heterozygous for the PLAG1 transgene and were all genotyped via PCR. The whole experimental protocol was approved by the Animal Care and Experiment Committee of Ninth People’s Hospital affiliated to Shanghai Jiao Tong University, School of Medicine (China).
Tissue samples and quantitative real-time PCR analysis
Normal gland tissue from wild-type mice (WT), tumor tissue (T), and normal gland tissue from PLAG1 transgenic mice (G) were obtained from four mice aged about 9 weeks. Total RNA was isolated by homogenizing the bulk tissue samples with TRIzol (Invitrogen) according to the manufacturer’s instructions. RNA samples were reverse transcribed into cDNA with AMV reverse transcriptase(TaKaRa, Shiga, Japan). Real-time PCR was performed on the ABI 7300 with SYBR® Premix Ex Taq™ (TaKaRa, Shiga, Japan). Primers for PLAG1 have been previously described. A list of all primer sequences used for real-time PCR is displayed in Supplementary Table 1.
Western blotting
Homogenates from tumor or gland tissue samples were extracted by lysis buffer [10 mM Tris/HCl (pH 8.0), 150 mM NaCl, 1 % NP-40] containing 1 % (v/v) proteinase inhibitor cocktail (Sigma-Aldrich). The homogenates were filtered through a mesh and centrifuged. The resulting supernatants were subjected to SDS–PAGE gels and electroblotted onto nitrocellulose membranes. Western blot analysis was performed as previously described. Actin was used as a control for equal loading. Antibodies for Igf2, c-myc, cyclinD1, p53, p21, Fas were purchased from Santa Cruz Biotechnology, antibodies for Bax and Trail-R/DR5 were from Abcam, and actin from Sigma.
RNA labeling and hybridization
RNA, 5 μg, from matched tumor/normal sample was reverse transcribed into cDNA using MMLV Reverse Transcriptase (Promega) with RT Primer (SuperArray Bioscience) and biotin-16-dUTP (Roche). The cDNAs were hybridized to the cDNA array using GEAhyb hybridization solution (SuperArray Bioscience). After an overnight hybridization at 60 °C, slides were washed and detected using a chemiluminescent detection kit (SuperArray Bioscience). After exposing the membrane to X-ray film, we used a desktop scanner to convert the image into a gray scale TIFF file. Spots within the image were converted to numerical data using the free ScanAlyze software. We used the GEArray Analyzer software for background subtraction and data normalization. Each GEArrayTM Q Series membrane was spotted with negative controls (pUC18 DNA and blanks) and housekeeping genes, including those for β-actin, GAPDH, cyclophilin A and ribosomal protein L13a. All raw signal intensities were corrected for background by subtracting the minimum value to avoid the problem of negative numbers. All signal intensities were also normalized to the intensities of a housekeeping gene. These corrected, normalized signals were then used to estimate the relative abundance of particular transcripts.
Transfection and reporter gene assays
NIH3T3 cells were transfected with plasmid DNA using FuGENE HD (Roche). Promoter fragments of mouse p53 (−1919 to 150), Bax (−1526 to 100), Trail-R/DR5 (−1636 to 100), Fos (−1601 to 55), and Fas (−1062 to 58) were cloned into the pGL3-basic vector (Promega,). Primer sequences for promoter analyses are displayed in Supplementary Table 2. Luciferase activity was measured 48 h after transfection with a luciferase assay system. Relative luciferase activity was normalized with β-galactosidase activity derived from pSV-β-galactosidase control vector.
Results
Sample characteristics
Transgenic mouse lines 42 and 9 displayed the highest levels of PLAG1 transgene expression in the salivary gland. Additionally, these lines displayed the highest rates of tumor incidence (100 % in line 42 and 68 % in line 9). We analyzed six samples in total, including two tumors (9-174T and 42-502T), two normal salivary glands from PLAG1 transgenic mice (9-174G and 42-502G), and two normal salivary glands from wide-type mice (9-178G and 42-500G)—all from transgenic lines 42 and 9. Tumor samples and matched normal salivary glands were obtained from the same mouse. They were categorized into three groups and gene expression profiles were compared against each other.
Aberrant gene expression in pleomorphic adenomas (PA) compared to normal salivary glands from both PLAG1 transgenic mice (PSG) and wild-type mice (WTSG)
The expression of 96 genes involved in 15 different signal transduction pathways was analyzed, and the signal variance was calculated.
Comparison of PA to WTSG revealed that 31 genes were upregulated at least twofold in both tumor samples; and 7 genes were upregulated in just one tumor sample. Additionally, 16 genes were downregulated in both tumors, and 8 genes were downregulated in a single tumor sample (Supplementary Table 3). Gene ontology revealed that many aspects of cellular biology are altered in PLAG1 tumors compared to normal tissue. We also classified these differentially expressed genes based on their functional annotation within the SWISSPROT database (data not shown). We found simultaneous induction of genes that promote apoptosis and cell cycle arrest with genes that drive cellular proliferation. Similar expression patterns were also identified in the other comparisons. (Supplementary Tables 4, 5).
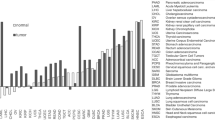
Expression profile analysis revealed that 17 genes (Akt-1, Cyclin D1, p21Waf1/p21cip, p16ink4a, p15INK4b, Egfr, c-fos, HMOX1, Hsp25, Hspb2, ICAM-1, Igfbp3, IL-4R, N4wbp4T/MEPAI, Glucose transporter 1, TRAIL-R/DR5, and Wig1) were upregulated in PA compared to PSG and WTSG. When WTSG was compared to the other two groups, 7 genes (Bax, p18, Beta-casein, Mdm2, Patched, Transferrin, and Tfr1) were found to be upregulated. Expression of 15 genes (Bcl-2, NAIP2, NAIP5, BMP2, BMP 4, p57Kip2, MIG/Scyb9, Edn1, autotaxin (ATX), Folh1, Hip, Kallikrein 3, INOS, Progesterone receptor, and Trail) was decreased in tumors compared to the other two classes. Finally the expression of 8 genes (ABCB4, HNF3B/forkhead box A2, A1 m, Tcf1, TRAF1, ELM1, Wnt1, and Wnt2) was reduced in PA and PSG relative to WTSG. Differentially expressed genes were grouped according to their signaling pathway. Clustering results for differentially expressed genes are shown in Fig. 1 and Supplementary Fig. 1.
Results of cluster analysis. Clustering results for differentially expressed genes between pleomorphic adenomas and normal salivary glands from wide-type mice. Each column represents an individual sample and each row represents a different gene. The expression level of each gene is indicated by color. The intensity of the red is proportional to the degree of expression above the mean, and the intensity of the green is proportional to the degree of expression below the mean. Gray indicates no detectable expression of the gene. Genes that were up-regulated or down-regulated over two-fold in both experiments were hierarchically clustered with The Institute for Genomic Research (TIGR) multi-experiment viewer
Validation of candidate genes using quantitative real-time RT-PCR and Western blot
Total RNA from trangenic samples was used for real time RT-PCR analysis to validate the microarray data. Eleven genes (Bax, Fas, p53, p21, p16, Cyclin D1, Egfr, Trail-R/DR-5, c-Fos, c-myc and Igf2) were selected for validation. As expected, PLAG1 and Igf2 transcripts were highly elevated in tumors compared to normal tissue. The expression of the other genes was similar to the microarray data. The results showed that the data obtained from western blot analysis matched those of real time RT-PCR analysis and the microarray data (Fig. 3), demonstrating that the data obtained from the microarray assay were reliable. The expression of PLAG1 in normal salivary glands is not as high as it is in tumors. Additionally, in accordance with the microarray data, high expression of all these genes was detected in every tumor but not in all normal salivary glands from PLAG1 transgenic mice at both RNA and protein levels (Figs. 2, 3). Experiments were repeated and the results matched in all cases.
Validation of microarray data by RT-PCR. a PLAG1 transcript levels were determined by real-time RT-PCR in wild-type and transgenic salivary glands and tumor tissues. Igf2 is a previously identified PLAG1 target gene and was used as a control in our experiment. b Quantification of differential expression of randomly selected genes by real-time RT-PCR in transgenic mouse lines9 and 42. Asterisks indicate significant difference (P < 0.05) compared to WT using the Student’s t test. Abbreviations: WT, wild-type salivary gland; G, PLAG1 transgenic salivary gland; T, tumor. Bars represent the mean ± SD (n = 3 individuals)
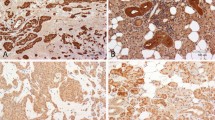
Protein levels detected by western blot analysis. a Expression of Igf2, CyclinD1 and c-myc were analyzed in tumors, matched normal salivary glands and wild-type salivary glands. As expected, the expression of these genes was induced not so greatly in PLAG1 transgenic normal salivary glands as in tumors. b Expression of p53, p21, Bax, Fas and Trail-R/DR5 was analyzed by the same method, and the western blot results were consistent with those yielded by microarray analysis and real-time RT-PCR. Abbreviations as described before
Identification of PLAG1 target genes
We next analyzed the promoter regions of differentially expressed genes for PLAG1 consensus binding sites. Putative PLAG1 binding sites were analyzed based on the JASTAR database (http://jaspar.genereg.net/cgi-bin/jaspar_db.pl). We limited our analysis to genes that were upregulated at least ten-fold based on microarray data to limit inclusion of false-positives. We cloned promoter sequences of multiple genes, including Bax, Fas, p53, TRAIL-R/DR5, and c-Fos upstream of the luciferase coding sequence. We found that co-transfection of PLAG1 expression vector (pCI-neo-PLAG1) with each of these promoter-luciferase reporter constructs resulted in a dose-dependent increase in luciferase activity, indicating that PLAG1 can activate transcription of these genes (Fig. 4; Supplementary Fig. 2).
Transactivation of promoters by PLAG1. Part labels a, b, c and d show putative PLAG1 consensus binding sites in the promoters of mouse p53, Bax, Fas and Trail-R/DR5 and transactivation of these promoters by PLAG1. The expression vector pCl-neo-PLAG1 was co-transfected with luciferase promoter reporter vectors into NIH3T3 cells. Relative luciferase activity was normalized to β-galactosidase activity derived from the pSV-β-galactosidase control vector. The results shown are the average of three independent experiments. Bars represent the mean ± SD (n = 3 individuals)
Discussion
PLAG1 is a transcription factor that can either activate or repress target genes. Here, we use a signal transduction microarray-based approach to show that PLAG1 overexpression is associated with gene expression changes in transgenic mice.
One of the largest classes of upregulated genes was the PI-3 Kinase/AKT pathway. Gene within this group included c-myc, c-Jun, c-Fos, and CyclinD1. Expression of c-myc and CyclinD1 was found to be increased in tumors (Figs. 2, 3), which is in accordance with our previous findings (Zhao et al. 2006). Together, c-Jun and c-Fos form the AP-1 early response transcription factor and expression of the components of AP-1 has been detected in numerous cancers lending to the tumorigenic potential of the two genes.
Not only did PLAG1 induce expression of genes involved in proliferation and tumor growth but it also induced expression of several genes that are growth inhibitory. The induction of tumor suppressor gene Tp53 is the most surprising example. p53 also acts as a transcription factor that regulates p21 and p27 to induce cell cycle arrest by inhibiting the kinase activity of cyclin/cyclin-dependent kinase complexes. Our microarray data showed that the expression of p53 and p21 was upregulated in the tumors. We also identified two putative PLAG1 consensus binding sites in the p53 promoter (−631 to −644). Co-transfection of PLAG1 expression vector (pCI-neo-PLAG1) with p53 promoter reporter constructs resulted in a dose-dependent increase in luciferase activity (Fig. 4). This suggests that the transcription of p53 is directly activated by PLAG1. This is consistent with previous reports by Van de Ven that described upregulation of p53 in PLAG1-overexpressing pleomorphic adenomas (Voz et al. 2004). Two other reports had described expression of p53 in approx. 75 % of pleomorphic adenomas (Tarakji et al. 2010; Azuma et al. 1992). Tarakji et al. (2010) attributed the high expression of p53 in tumors to the status of p53 mutation, resulting in a more stable form of the transcription factor compared to wild-type protein. However, our study here provides another possible explanation.
We also discovered that PLAG1 induces expression of several genes that promote apoptosis including Bax (Figs. 2, 3). Two putative PLAG1 consensus binding sites were identified in region −158 to −142 and −99 to −82 of the Bax promoter, which was transactivated by PLAG1 (Fig. 4). Additionally, Bax expression can also be directly induced by p53 (Chipuk et al. 2004), indicating that Bax is involved in p53-mediated apoptosis. Low Bax expression is an independent, negative prognostic marker in multivariate regression analysis independent of p53 status(Grabowski et al. 2006; Nehls et al. 2005; Schelwies et al. 2002; Sturm et al. 1999). Therefore, activation of Bax by PLAG1 in tumors is consistent with the pathological and prognostic characteristics of pleomorphic adenoma—a benign (partially borderline) tumor with low recurrence and metastatic potential.
Another two genes identified in our study that promote apoptosis are Fas (also CD95) and TRAIL-R/DR5, both of which belong to the tumor necrosis factor receptor superfamily. Activation of p53 can also induce expression of Fas and TRAIL-R/DR5 (Wu et al. 2000), and the direct transactivation of the promoters of Fas and TRAIL-R/DR5 by PLAG1 has also been described (Fig. 4).
The data we provide here show that PLAG1 functions as both an oncogene and as a tumor suppressor. Indeed, PLAG1 induces expression of some genes that promote cellular growth and others that inhibit proliferation. This may be described as the dual role of oncogene induction. The activation of oncogenes occurs in most tumors and is believed to play an important role in tumorigenesis. However, in many tissues, there are numerous neoplastic lesions that rarely become overt cancers, including established benign tumors such as melanocytic nevi (Pollock et al. 2003). The mechanism was known as oncogene-induced cell senescence involving p53 pathway and pRb pathway. Moreover, apoptotic cell death may be triggered by oncogene-driven cellular proliferation, sometimes directly (Bartkova et al. 2005; Gorgoulis et al. 2005). Thus, tumors resulting from PLAG1 overexpression, including pleomorphic adenomas, lipoblastomas, hepatoblastomas, and leiomyomas, may be described as benign or low-grade malignant tumors because of the dual action of PLAG1 in these diseases. A potential role of PLAG1 was indicated in regulation of oncogene-induced cell senescence.
References
Azuma M, Kasai Y, Tamatani T, Sato M (1992) Involvement of p53 mutation in the development of human salivary gland pleomorphic adenomas. Cancer Lett 65(1):61–71
Bartkova J, Horejsí Z, Koed K, Krämer A, Tort F, Zieger K, Guldberg P, Sehested M, Nesland JM, Lukas C, Ørntoft T, Lukas J, Bartek J (2005) DNA damage response as a candidate anticancer barrier in early human tumorigenesis. Nature 434:864–870
Chipuk JE, Kuwana T, Bouchier-Hayes L, Droin NM, Newmeyer DD, Schuler M, Green DR (2004) Direct activation of Bax by p53 mediates mitochondrial membrane permeabilization and apoptosis. Science 303(5660):1010–1014
Enlund F, Nordkvist A, Sahlin P, Mark J, Stenman G (2002) Expression of PLAG1 and HMGIC proteins and fusion transcripts in radiation-associated pleomorphic adenomas. Int J Oncol 20:713–716
Gorgoulis VG, Vassiliou LV, Karakaidos P, Zacharatos P, Kotsinas A, Liloglou T, Venere M, Ditullio RA Jr, Kastrinakis NG, Levy B, Kletsas D, Yoneta A, Herlyn M, Kittas C, Halazonetis TD (2005) Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 434:907–913
Grabowski P, Sturm I, Schelwies K, Maaser K, Buhr HJ, Dörken B, Zeitz M, Daniel PT, Scherübl H (2006) Analysis of neuroendocrine differentiation and the p53/BAX pathway in UICC stage III colorectalcarcinoma identifies patients with good prognosis. Int J Colorectal Dis 21(3):221–230
Landrette SF, Kuo YH, Hensen K, Khosrovani B, Perrat PN, Van de Ven WJ, Delwel R, Castilla LH (2005) Plag1 and Plagl2 are oncogenes that induce acute myeloid leukemia in cooperation with Cbfb-MYH11. Blood 105:2900–2907
Nehls O, Okech T, Hsieh CJ, Sarbia M, Borchard F, Gruenagel HH, Gaco V, Porschen R, Gregor M, Klump B (2005) Low BAX protein expression correlates with disease recurrence in preoperatively irradiated rectal carcinoma. Int J Radiat Oncol Biol Phys 61(1):85–91
Pollock PM, Harper UL, Hansen KS, Yudt LM, Stark M, Robbins CM, Moses TY, Hostetter G, Wagner U, Kakareka J, Salem G, Pohida T, Heenan P, Duray P, Kallioniemi O, Hayward NK, Trent JM, Meltzer PS (2003) High frequency of BRAF mutations in nevi. Nat Genet 33:19–20
Schelwies K, Sturm I, Grabowski P, Scherübl H, Schindler I, Hermann S, Stein H, Buhr HJ, Riecken EO, Zeitz M, Dörken B, Daniel PT (2002) Analysis of p53/BAX in primary colorectal carcinoma: low BAX protein expression is a negative prognostic factor in UICC stage III tumors. Int J Cancer 99(4):589–596
Shen S, Yang W, Wang Z, Lei X, Xu L, Wang Y, Wang L, Huang L, Yu Z, Zhang X, Li J, Chen Y, Zhao X, Yin X, Zhang C (2011) Tumor-initiating cells are enriched in CD44(hi) population in murine salivary gland tumor. PLoS ONE 6(8):e23282
Sturm I, Köhne CH, Wolff G, Petrowsky H, Hillebrand T, Hauptmann S, Lorenz M, Dörken B, Daniel PT (1999) Analysis of the p53/BAX pathway in colorectal cancer: low BAX is a negative prognostic factor in patients with resected liver metastases. J Clin Oncol 17(5):1364–1374
Tarakji B, Kujan O, Nassani MZ (2010) Immunohistochemical expression of p53 in pleomorphic adenoma and carcinoma ex pleomorphic adenoma. J Cancer Epidemiol 2010:250606
Voz ML, Agten NS, Van de Ven WJ, Kas K (2000) PLAG1 The main translocation target in pleomorphic adenoma of the salivary glands, is a positive regulator of IGF-II. Cancer Res 60:106–113
Voz ML, Mathys J, Van de Ven WJ et al (2004) Microarray screening for target genes of the proto-oncogene PLAG1. Oncogene 23(1):179–191
Wu GS, Kim K, el-Deiry WS (2000) KILLER/DR5 a novel DNA-damage inducible death receptor gene, links the p53-tumor suppressor to caspase activation and apoptotic death. Adv Exp Med Biol 465:143–151
Zhao X, Ren W, Yang W, Wang Y, Kong H, Wang L, Yan L, Xu G, Fei J, Fu J, Zhang C, Wang Z (2006) Wnt pathway is involved in pleomorphic adenomas induced by overexpression of PLAG1 in transgenic mice. Int J Cancer 118(3):643–648
Acknowledgments
This work was supported by the National Natural Science Foundation of China 30772432 and 30772430.
Author information
Authors and Affiliations
Corresponding author
Additional information
Yang Wang and Wenjun Shang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, Y., Shang, W., Lei, X. et al. Opposing functions of PLAG1 in pleomorphic adenoma: a microarray analysis of PLAG1 transgenic mice. Biotechnol Lett 35, 1377–1385 (2013). https://doi.org/10.1007/s10529-013-1213-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-013-1213-7