Abstract
Objectives
To improve 1,3-propanediol (1,3-PD) production and reduce byproduct concentration during the fermentation of Klebsiella pneumonia.
Results
Klebsiella. pneumonia 2-1ΔldhA, K. pneumonia 2-1ΔaldH and K. pneumonia 2-1ΔldhAΔaldH mutant strains were obtained through deletion of the ldhA gene encoding lactate dehydrogenase required for lactate synthesis and the aldH gene encoding acetaldehyde dehydrogenase involved in the synthesis of ethanol. After fed-batch fermentation, the production of 1,3-PD from glycerol was enhanced and the concentrations of byproducts were reduced compared with the original strain K. pneumonia 2-1. The maximum yields of 1,3-PD were 85.7, 82.5 and 87.5 g/l in the respective mutant strains.
Conclusion
Deletion of either aldH or ldhA promoted 1,3-PD production in K. pneumonia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
1,3-Propanediol (1,3-PD) has applications in polymer synthesis, cosmetics, foods, lubricants, and medicines. Its industrial production is of economic importance particularly as an key monomer in the synthesis of a new type of polyester, polytrimethylene terephthalate (Zeng and Biebl 2002). 1,3-PD is currently produced by chemical processes such as hydroformylation of ethylene oxide and hydration of acrolein. The microbial production of 1,3-PD, a socially beneficial route to obtain chemicals from renewable resources, has been widely researched and is considered as a competitor to the traditional petrochemical synthesis routes (Da Silva et al. 2009). Among the microorganisms that can convert glycerol to 1,3-PD, Klebsiella pneumoniae, Clostridium butyricum and Citrobacter freundii have been the major foci of attention because of their appreciable substrate tolerance, yield, and productivity (Celinska 2010). Biological manipulation including optimization of the natural glycerol utilizing fermentation process and metabolic engineering can contribute to increasing the yield of 1,3-PD (Mu et al. 2006).
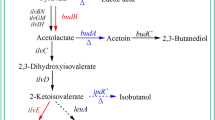
Glycerol metabolism in the model organism K. pneumoniae involves two pathways—the oxidative branch and the reductive branch (Oh et al. 2013a) (Fig. 1). In the oxidative branch, glycerol is transformed to dihydroxyacetone by glycerol dehydrogenase, which then undergoes normal glycolysis to form pyruvate, and finally can be converted into various byproducts such as acids and alcohols. In the reductive pathway, glycerol is converted to 3-hydroxypropionaldehyde (3-HPA) by a coenzyme B12-dependent glycerol dehydratase (GDHt), which is then reduced to 1,3-PD by a NADH-dependent 1,3-PD oxidoreductase (DhaT) (Ji et al. 2010; Jung et al. 2012). Activity of GDHt is the key rate limiting step for this biological process. The second metabolic pathway, which maintains the redox balance of the cell, is essential for the conversion of glycerol to 1,3-PD. However, during the biological production of 1,3-propanediol, a series of byproducts (such as lactate and ethanol) inhibit the pathway and increase the purification cost and time (Oh et al. 2013b). During glycerol metabolism, considerable amounts of metabolites are produced via the oxidative branch. Reduction or elimination of these metabolites is an important strategy to enhance 1,3-PD production during microbial fermentation. In this study, we investigated lactate and ethanol during 1,3-PD production in K. pneumoniae 2-1. We investigated whether direct inactivation of metabolic pathways leading to production of these major byproducts of the oxidative branch would enhance 1,3-PD production. Additionally, fermentation parameters were optimized to achieve this goal (Seo et al. 2009).
Materials and methods
Materials
Klebsiella pneumoniae 2-1 was previously described (Meng et al. 2011). Escherichia coli DH5α was used for DNA manipulation. λRed recombinase (Datsenko and Wanner 2000) and the FLP recombinase (Datsenko and Wanner 2000) were expressed by the helper plasmids pKD46 (Biosci Biotech, Hangzhou, China) and pCP20 (Biosci Biotech, Hangzhou, China), respectively; replication of these plasmids is temperature sensitive, and the plasmids can thus be easily eliminated. The plasmid pMD19-T Simple (TaKaRa Biotech, Dalian, China) was employed for cloning. The primers and restriction endonuclease are shown in Supplementary Table 1. The primers were synthesized by Shanghai Sangon. Restriction endonucleases were purchased in Shanghai Sangon.
The preculture medium contained (g/l)
glycerol, 20.0; KH2PO4, 1.3; K2HPO4, 3.4; (NH4)2SO4, 2.0; MgSO4·7H2O, 0.2; FeSO4·7H2O,5 × 10−2; CaCl2,2 × 10−3; yeast extract, 1; citric acid, 0.42; trace element solution A, 2 ml.
The fermentation medium contained (g/l)
glycerol, 50; (NH4)2SO4, 6.6; NaH2PO4 1.38; Na2SO4, 0.28; KCl, 0.75; MgCl2·6H2O, 0.26; CaCl2·2H2O, 0.29; yeast extract, 1; citric acid, 0.42; trace element solution B, 5 ml.
Trace element solution A contained (g/l)
CoCl2·6H2O, 0.2; MnCl2·4H2O, 0.1; ZnCl2 0.07; H3BO3, 0.06; Na2MoO4·2H2O, 0.035; CuCl2·2H2O, 0.02; NiCl2·6H2O, 0.025.
Trace element solution B contained (g/l)
CuCl2·2H2O, 0.17; MnCl2·4H2O, 2; H3BO3 0.6; ZnCl2·6H2O, 0.68; Na2MoO4·2H2O, 0.005; CoCl2·6H2O, 0.47; FeCl3·6H2O.
Deletion of ldhA and aldH genes
To make an ldhA deletion mutant, the ldhA gene was cloned using primers A1/A2 (Supplementary Table 1) and the K. pneumoniae 2-1 genome as a template. The recombinant plasmid pMD19-T-ldhA was made from the ldhA gene and the vector pMD19-T. Secondly, in order to obtain linear fragments ldhA was amplified with a 300 bp homologous arm using A3/A4 as primers and pMD19-T-ldhA as template. The Cm r gene was cloned by using L1/L2 as primers and FRT sites in pKD3 (Datsenko and Wanner 2000) as a template. Then the ldhA1-T-ldhA2 and Cm r genes were digested and ligated with KpnI and XhoI to obtain the plasmid pMD19-T-ldhA1-Cm r-ldhA2 which was selected by ampicillin and chloramphenicol resistance. Finally, linear fragments ldhA1-Cm r-ldhA2 with the 300 bp homologous arm were generated using A1/A2 as primers and pMD19-T-ldhA1-Cm r-ldhA2 as a template. The linear fragments were introduced into K. pneumoniae 2-1 by electroporation to induce homologous recombination. Correct integration of the DNA fragment was confirmed by colony PCR with A1/A2 and L1/L2 as primers.
The aldH deletion method was performed in a similar way. First of all, the aldH gene was cloned by using Y1/Y2 as primers and the K. pneumoniae 2-1 genome as a template. Then the recombinant plasmid pMD19-T-aldH was obtained from the aldH gene and pMD19-T. Secondly, the linear fragments with the 300 bp homologous arm were generated by using Y3/Y4 as primers and pMD19-T-aldH as template for the reverse cloning. The Cm r gene was cloned by using L1/L2 as primers and FRT sites with pKD3 as a template. The aldH1-T-aldH2 and Cm r gene were then digested and ligated with KpnI and XhoI to obtain a plasmid with pMD19-T-aldH1-Cm r-aldH2 selected by ampicillin and chloramphenicol resistance. Finally, Y1/Y2 primers were used in order to obtain linear fragments aldH1-Cm r-aldH2 with the 300 bp homologous arm using pMD19-T-aldH1-Cm r-aldH2 as a template for cloning. Linear fragments were then introduced into K. pneumoniae 2-1 by electroporation to induce homologous recombination. Correct integration of the DNA fragment was confirmed by colony PCR with Y1/Y2 and L1/L2 as primers.
To make the ΔaldHΔldhA double mutant, the ldhA mutant linear fragment was PCR-amplified as described above and introduced into K. pneumoniae 2-1ΔaldH by electroporation to induce homologous recombination. Correct integration of the ldhA deletion linear fragment was confirmed as described above.
Fermentation of K. pneumoniae strains
Cells were seeded in a 250 ml shake-flask containing 50 ml medium at 37 °C for 14 h. Fermentations were carried out in a 5 l stirred-vessel system (Kobiotech. Co. Ltd) with 2 l medium. The culture was grown at 37 °C, pH 7, and agitation at 200 rpm. The pH was controlled at 7 by automatic addition of 2.5 M KOH. Anaerobic conditions in the bioreactors were maintained by N2 at 0.4 vvm. In fed-batch fermentation, the limiting substrate, glycerol, was initially at 50 g/l. In the glycerol concentration feedback feeding strategy, after the residual glycerol dropped below 20 g/l, feeding was started to continuously pump concentrated glycerol solution into the bioreactor to maintain it between 15 and 25 g/l.
Analytical methods
Cell concentration was measured as the OD650 value (Oh et al. 2013a). The determination of 1,3-PD and ethanol levels was by GC using a Chromosorb 101 column 2 m × 0.5 mm with N2 as a carrier gas at 40 ml/min. The detector was at 240 °C and column at 210 °C) (Wang et al. 2010). Glycerol was measured by a modified titration method (Meng et al. 2011). Lactate was determined by HPLC with an XDB-C18 column (250 mm × 4.6 mm, 5 µm; Agilent, USA) (Saxena et al. 2009). The mobile phase was of 2 % (v/v) H3PO4/acetonitrile (96.5:3.5, v/v), at 1 ml/min. Detection was at 214 nm. The column was held at 35 °C (Xiu and Zeng 2008). All samples tested were initially filtered through a membrane filter (0.45 µm pore size).
Results and discussion
Construction of ΔldhA, ΔaldH and ΔaldH ΔldhA mutant strains
To examine the effect of elimination of the major glycerol oxidative metabolic pathways involved in the synthesis of lactate or ethanol on 1,3-PD production, K. pneumoniae 2-1 was used as the starting strain for the construction of derivatives that were lactate dehydrogenase-deficient, acetaldehyde dehydrogenase-deficient, or deficient in both enzymes. K. pneumoniae 2-1ΔldhA, K. pneumoniae 2-1ΔaldH and K. pneumoniae 2-1ΔaldHΔldhA strains were constructed via homologous recombination with the λRed recombination system (see Materials and methods section). The mutants were identified by colony PCR identification as shown in Figs. 2, 3, and 4 respectively.
Confirmation of construction of the ΔldhA of K. pneumonia 2-1 by colony PCR identification. Lane M 5000 bp DNA Ladder Marker; Lane 1 ldhA gene of K. pneumonia 2-1; Lane 2 Cm r gene of K. pneumonia 2-1; Lane 3 linear fragments ldhA1-Cm r-ldhA2 of K. pneumonia 2-1ΔldhA; Lane 4 Cm r gene of K. pneumonia 2-1ΔldhA
Confirmation of construction of the ΔaldH of K. pneumonia 2-1 by colony PCR identification. Lane M 5000 bp DNA Ladder Marker; Lane 1 aldH gene of K. pneumonia 2-1; Lane 2 Cm r gene of K. pneumonia 2-1; Lane 3 linear fragments aldH1-Cm r-aldH2 of K. pneumonia 2-1ΔaldH; Lane 4, Cm r gene of K. pneumonia 2-1ΔaldH
Confirmation of construction of the ΔldhAΔaldH of K. pneumonia 2-1 by colony PCR identification. Lane M 5000 bp DNA Ladder Marker; Lane 1 ldhA gene of K. pneumonia 2-1; Lane 2 aldH gene of K. pneumonia 2-1; Lane 3 Cm r gene of K. pneumonia 2-1; Lane 4 linear fragments ldhA1-Cm r-ldhA2 of K. pneumonia 2-1ΔldhAΔaldH; Lane 5 linear fragments aldH1-Cm r-aldH2 of K. pneumonia 2-1ΔldhAΔaldH; Lane 6 Cm r gene of K. pneumonia 2-1ΔldhAΔaldH
The glycerol metabolism properties of ΔldhA, ΔaldH and ΔaldH ΔldhA strains
A large amount of metabolites, ethanol and lactate, are co-produced with 1,3-PD from glycerol as byproducts via oxidative metabolism (Oh et al. 2012). Fed-batch fermentation of K. pneumoniae 2-1 resulted in the production of 1,3-PD to a maximum of 78.8 g/l (Fig. 5a). The concentration of lactate and ethanol were 10.2 and 7.2 g/l, respectively. In the ldhA-mutant strain, the synthesis of lactate was decreased, as expected, to 0.5 g/l. 1,3-PD increased to 85.7 g/l (Fig. 5b). The results indicate that the yield of 1,3-PD was increased by deleting the ldhA gene.
In the aldH-mutant strain, the synthesis of ethanol was also decreased, as expected, to 0.5 g/l. 1,3-PD increased to 82.5 g/l (Fig. 5c). Thus the yield of 1,3-PD was increased by deleting the aldH gene. In the K. pneumonia 2-1ΔldhAΔaldH, the synthesis of both lactate and ethanol were decreased, as expected. Lactate and ethanol were 0.5 and 0.4 g/l, respectively. 1,3-PD increased to 87.5 g/l (Fig. 5d). This shows that the yield of 1,3-PD was increased by co-deletion of the aldH and ldhA genes. The figures in Table 1 also demonstrate that the concentrations of formate, acetate and succinate are increased in the mutant strains compared with the parental strain. This shows that the metabolic flux of carbon and hydrogen reduction are transferred to other pathways owing to the deletion of the ldhA and aldH (Seo et al. 2009).
Conclusion
Abolishing the major glycerol oxidative metabolic pathway involved in the synthesis of lactate and ethanol, was studied. Deletion of the ldhA gene encoding lactate dehydrogenase (catalyzing the synthesis of lactate) and the aldH gene encoding acetaldehyde dehydrogenase (catalyzing the synthesis of ethanol) enhanced 1,3-PD production. The K. pneumoniae 2-1ΔldhA, K. pneumoniae 2-1ΔaldH and K. pneumoniae 2-1ΔaldHΔldhA variants were constructed via homologous recombination with the λRed recombination system. After 24 h fermentation by K. pneumoniae 2-1ΔldhA, K. pneumoniae 2-1ΔaldH and K. pneumoniae 2-1ΔaldHΔldhA, the maximum yields of 1,3-PD were increased to 85.7, 82.5, 87.5 g/l respectively, compared with 78.8 g/l in the parental strain.
References
Celińska E (2010) Debottlenecking the 1,3-propanediol pathway by metabolic engineering. Biotechnol Adv 28:519–530
Da Silva GP, Mack M, Contiero J (2009) Glycerol: a promising and abundant carbon source for industrial microbiology. Biotechnol Adv 27:30–39
Datsenko KA, Wanner BL (2000) One-step in activation of chrom os om algenes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Ji XJ, Huang H, Zhu JG et al (2010) Engineering Klebsiella oxtoca for efficient 2,3-butanediol production through insertional inactivation of acetaldehyde dehydrogenase gene. Appl Microbiol Biotechnol 85:1751–1758
Jung MY, Ng CY, Song H et al (2012) Deletion of lactate dehydrogenase in Enterobacter aerogenes to enhance 2,3-butanediol production. Appl Microbiol Biotechnol 95:461–469
Meng DD, Wang RM, Ma CL (2011) Isolation, identification and fermentation characterization of a 1,3-propanediol product bacterial. Microbiol Chin 38:803–808
Mu Y, Teng H, Zhang DJ et al (2006) Microbial production of 1, 3-propanediol by Klebsiella pneumoniae using crude glycerol from biodiesel preparation. Biotechnol Lett 28:1755–1759
Oh BR, Seo JW, Heo SY et al (2012) Optimization of culture conditions for 1,3-propanediol production from glycerol using a mutant strain of Klebsiella pneumoniae. Appl Biochem Biotechnol 166:127–137
Oh BR, Seo JW, Heo SY et al (2013a) Efficient production of 1,3-propanediol from glycerol upon constitutive expression of the 1,3-propanediol oxidoreductase gene in engineered Klebsiella pneumoniae with elimination of by-product formation. Bioproc Biosyst Eng 36:757–763
Oh BR, Hong WK, Heo SY et al (2013b) The production of 1,3-propanediol from mixtures of glycerol and glucose by a Klebsiella pneumoniae mutant deficient in carbon catabolite repression. Bioresour Technol 130:719–724
Saxena RK, Anand P, Saran S et al (2009) Microbial production of 1,3-propanediol: recent developments and emerging opportunities. Biotechnol Adv 27:895–913
Seo MY, Seo JW, Heo SY et al (2009) Elimination of by-product formation during production of 1,3-propanediol in Klebsiella pneumoniae by inactivation of glycerol oxidative pathway. Appl Microbiol Biotechnol 84:527–534
Wang A, Wang Y, Jiang T et al (2010) Production of 2,3-butanediol from corncob molasses, a waste byproduct in xylitol production. Appl Microbiol Biotechnol 87:965–970
Xiu ZL, Zeng AP (2008) Present state and perspective of downstream processing of biologically produced 1,3-propanediol and 2,3-butanediol. Appl Microbiol Biotechnol 78:917–926
Zeng AP, Biebl H (2002) Bulk chemicals from biotechnology: the case of 1, 3-propanediol production and the new trends. Adv Biochem Eng Biotechnol 74:239–259
Acknowledgments
This work was financially supported by a grant from Shandong Province Independent Innovation and Achievement Transformation Project (201422cx02602) and a grant from Taishan scholar Construction Project.
Supporting Information
Supplementary Table 1—Primers used in this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, L., Ma, C., Wang, R. et al. Deletion of ldhA and aldH genes in Klebsiella pneumoniae to enhance 1,3-propanediol production. Biotechnol Lett 38, 1769–1774 (2016). https://doi.org/10.1007/s10529-016-2155-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-016-2155-7