Abstract
Breast cancer (BRCA) is currently the most commonly diagnosed malignancy in women worldwide. Previous studies have demonstrated that mitophagy is important for the prevention and treatment of BRCA. However, few studies have focused on the individual mitochondrial autophagy-related genes (MARG) in human cancers. Based on bioinformatics analyses, TOMM40 was identified as a prognostic DEMARG (PDEMARGs); Kaplan–Meier (KM) survival analysis also indicates that TOMM40 can be useful as a prognostic indicator in BRCAs, with patients in the high expression group having a poorer prognosis. For 20 distinct cancer kinds, there were appreciable differences in the expression of TOMM40 between tumor and normal tissues; in addition, in 21 different cancer types, there were associations between the expression profile of TOMM40 and patient prognosis. Gene Set Enrichment Analysis (GSEA), functional enrichment analysis, and immunological and drug sensitivity analyses of TOMM40 have indicated its biological significance in pan-cancers. Knockdown of TOMM40 in MDA-MB-231 cells inhibited their proliferation, migration, and invasiveness. In conclusion, we found that TOMM40 has prognostic value in 21 cancers, including breast cancer, by bioinformatics analysis. Based on immune correlation analysis, TOMM40 may also be a potential immunotherapeutic target for the treatment of BRCA. Therefore, our results may provide researchers to further explore the role of MARGs, especially TOMM40, in the developmental process of breast cancer, which may provide new directions and targets for the improvement of prognosis of breast cancer patients and their treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is currently the most frequently diagnosed malignancy in women worldwide. According to the latest global statistics (2020), the number of new BRCA cases around the world has risen to 2.26 million and it is also the main cause of cancer mortality for women (Sung et al. 2020). Molecular markers are crucial for BRCA diagnosis and treatment. Tumours expressing estrogen receptors (ERs) and/or progesterone receptors (PRs) are considered hormone receptor-positive, and those expressing the human epidermal growth factor receptor 2 (HER-2) are regarded as HER2-enriched type. Tumours that do not express ER, PR, or HER2 are labelled as triple-negative BRCA (TNBC) (Cheang et al. 2015; Perou et al. 2000). As more and more biomarkers were investigated to predict and guide management, they became increasingly important for the diagnosis and treatment of BRCA.
Mitophagy is a targeted and specific form of autophagy, with a pivotal role in reducing the redundant mitochondria in cells; this allows cells to adapt to hypoxia or nutrient deprivation and limit the generation of reactive oxygen species (ROS) (Mizushima and Komatsu 2011; Mustafa et al. 2021; Palikaras et al. 2018; Kim et al. 2007). Mitophagy also contributes significantly to the development and growth of tumors; however, the effects of mitophagy are complex and are influenced by diverse factors including the tumor microenvironment (Naik et al. 2019). Specific defects of mitophagy related to human cancers were related to the absence of key regulators such as Parkin and BNIP3 (Chourasia et al. 2015). Parkin, as a tumor suppressor (Cesari et al. 2003; Xu et al. 2014), was instrumental in regulating mitochondrial mitophagy, homeostasis, and anti-oxidative stress (Pickrell and Youle 2015). Downregulation and low-frequency mutations of Parkin are apparent in many cancers, including BRCA (Cesari et al. 2003; Tay et al. 2010). The BNIP3 protein targets mitochondria for autophagosomal degradation, which reduces the accumulation of dysfunctional mitochondria and excess ROS production. In genetically engineered mouse models, BNIP3 has been used to restrict the growth and progression of tumors. However, in human TNBCs, BNIP3 is currently being used as a prognostic marker for tumor metastasis as BNIP3 enhances the expression of HIF-1, which promotes metastasis in TNBC (Chourasia and Macleod 2015; Chourasia et al. 2015). Few studies have investigated the relationship between mitophagy and the development and progression of BRCA; however, we have, in a previous study, highlighted the importance of mitophagy in these processes and have proposed a new classification of BRCA based on the expression of 29 mitophagy-related genes.
Mitochondria are encased in two membranes; the inner membrane encloses the mitochondrial matrix, and the outer membrane forms the border of the cytoplasm and the mitochondria (Gabriel et al. 2003). Roughly 99% of all mitochondrial proteins are encoded by nuclear genes and translated into the cytoplasm as precursor proteins (Wiedemann et al. 2004; Endo et al. 2003; Straub et al. 2016). Most precursor proteins (> 90%) are transported into the mitochondria via the translocase outer mitochondrial membrane (TOMM) complex which is composed of seven subunits. These include the channel protein Tom40 and six transmembrane proteins, of which there are three receptor proteins (Tom20, Tom22 and Tom70) and three regulatory proteins (Tom5, Tom6 and Tom7) (Dolezal et al. 2006; Chacinska et al. 2009; Ryan et al. 2000). The Tom40 protein, which is encoded by the TOMM40 gene, forms a channel through which mitochondrial precursor proteins are transported across the outer mitochondrial membrane (Pfanner et al. 2004; Ellenrieder et al. 2015). In addition, Tomm40 also plays an active role in protein sorting for all sub-mitochondrial locations (Gabriel et al. 2003). Many studies have revealed that mutations in TOMM40 may be associated with Alzheimer’s Disease (Bezuch et al. 2021; Lee et al. 2021). Research also suggests that in some cancers, there is an increased expression of the TOMM40 gene; however, the expression levels of TOMM40 in BRCA tissues have not been investigated (Yang et al. 2020).
In this study, we find that TOMM40 can be used as a PDEMARG in BRCA based on the survival analysis of BRCA patients with differentially expressed mitophagy genes. We have also analyzed the expression levels and prognostic value of TOMM40 across different cancers and performed GSEA function enrichment and immunological and drug sensitivity analyses to determine what pathways and potential molecular mechanisms this gene is involved in. The flow chart of this study is shown below.

The flow chart of this research. TCGA: The Cancer Genome Atlas; BRCA: breast cancer; MARGs: mitochondrial autophagy-related genes; DEMARGs: differentially expressed MARGs.
Materials and Methods
Data Collection
The Cancer Genome Atlas (TCGA, xenabrowser.net), in the UCSC Xena database (xenabrowser.net/datapages/), was used to get clinical and patient survival information as well as RNA sequence (RNA-seq) data on 33 different tumors. The Reactome database (reactome.org/) was used to find a total of 28 mitochondrial autophagy-related genes (MARGs), including mitophagy (R-HSA-5205647), pink1 prkn mediated mitophagy (R-HSA-5205685) and receptor mediated mitophagy (R-HSA-8934903) (Wang et al. 2021a).
Identification of Differentially Expressed MARGs (DEMARGs) and Prognostic DEMARGs (PDEMARGs)
The location of MARGs on the chromosomes was drawn using the ‘RCircos’ package in R (version 1.2.2) (Zhang et al. 2013). The expression levels of the 28 MARGs were compared between the BRCA and normal samples using the rank-sum test. The ‘limma’ package in R (version 3.48.3) was then used to screen the DEMARGs between 1,097 BRCA samples and 114 normal samples in the TCGA (|log2FC|> 1, P < 0.05) (Ritchie et al. 2015). The expression levels of the DEMARGs were shown using the R packages ‘ggplot2’(version 3.3.5) and ‘pheatmap’ (version 1.0.12) (Hu 2021). Moreover, Kaplan–Meier (KM) curves were utilized to estimate the prognostic values of the DEMARGs and discover PDEMARGs, including overall survival period (OS) curves and disease-free survival (DFS) curves.
Expression and Clinical Correlation Analysis of PDEMARG in Pan-Cancers
Using the ‘limma’ package (|log2FC|> 1, P < 0.05), the changes in PDEMARG expression levels between tumor and normal samples from 33 different malignancies were examined, as well as the associations between PDEMARGs and clinical parameters (age, sex, tumor grade, and tumor stage).
Functional Enrichment Analysis of PDEMARG
The KM curves of the PDEMARGs across 33 different cancers were plotted to determine the prognostic values of these genes. Based on the results of this analysis, gene set enrichment analysis (GSEA) was used to further explore the pathways and molecular mechanisms that the PDEMARGs are involved in by using the ‘clusterProfiler’ package in R (version 4.0.2) and the ‘org.Hs.eg.db’ package in R (version 3.13.0) (|NES|> 1, NOM P < 0.05, q < 0.25) (Colaprico et al. 2016). Besides, the CancerSEA database (http://biocc.hrbmu.edu.cn/CancerSEA/) was used to evaluate the average correlation between functional status and PDEMARG.
Analysis of the Immune Microenvironment
The tumor microenvironment (TME) is a crucial element that governs the survival, development, and metastasis of tumors (Barriga et al. 2019). In this study, the correlations between the PDEMARGs and the expression levels of 43 immune checkpoints in the 33 cancers included in this study were calculated using the Spearman rank correlation test. We also obtained the RNA-seq data and DTP NCI-60 compound activity data from the CellMiner database (http://discover.nci.nih.gov/cellminer/), to analyze the relationships between the expression levels of the PDEMARGs and drug sensitivity in these cancers (|r|> 0.3 and p < 0.05).
Expression of TOMM40 in BRCA at mRNA and Protein Levels
Bc-GenExMiner (version 4.9) was used to examine the variations in TOMM40 mRNA expression levels between BRCA and normal samples. The differences in the levels of the Tomm40 protein in BRCA and normal breast tissues were analyzed using the UALCAN and the human protein atlas (HPA) databases.
Cell Culture, Plasmids, shRNA, and Transfection
MDA-MB-231 cells were kindly provided by Dr Shuqun Zhang (Department of Oncology, the Second Affiliated Hospital of Xi’an Jiaotong University, China). The cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM; Gibco Inc.) supplemented with 10% FBS (Gibco Inc.) and 1% penicillin–streptomycin (Sigma Inc.) at 37 ℃ with 5% CO2. For the knockdown assays, 5 × 105 cells were grown in a 6-well plate to 40% confluence before being transfected with Lipofectamine 2000 (5 μl/well; Invitrogen Inc.), and shRNAs targeting human TOMM40 gene (2 μg/well; GeneCopoeia Inc.) to introduce shRNAs that target the human TOMM40 gene.
Real-Time Quantitative Polymerase Chain Reaction (qRT-PCR)
To validate the expression of PDEMARGs, we performed qRT-PCR on tissue samples (mammary cancer tissue and normal breast tissue samples) obtained from patients treated at the Second Affiliated Hospital of Xi’an Jiaotong University; all patients received information about the study and signed consent forms for us to utilize their tissues. Total RNA was extracted using TRIZol (Thermo Fisher Inc.), and the mRNA obtained was reverse transcribed into cDNA, following which qPCR reactions were carried out by the SureScript-First-strand-cDNA-synthesis kit (Servicebio Inc.).
Western Blotting
Cells were lysed with RIPA buffer (Beyotime Inc.) with protease inhibitors. Protein concentrations in the cell lysates were estimated using the BCA Protein Assay Kit (Cwbio Inc.). Protein samples were run on 10% SDS polyacrylamide gels, and then electrotransferred onto PVDF membranes. After blocking, the membranes were incubated with primary antibodies (anti-TOMM40, Affinity Biosciences Inc.) at 4 °C overnight. After immune blotting, the signals were detected using a developing reagent (Beyotime Inc.).
Cell Proliferation Assay
Cells (4–5 × 103 cells/well) were seeded into 96-well plates, six complex wells per set and cultured for 4 days. The number of viable cells were quantified by adding 10 μl of Cell Counting Kit-8 dye (MedChemExpress Inc.) to each well after 4 h of seeding and incubating at 37 °C for 1 h. Following this, the absorbance in each well was measured at 450 nm. The above steps for counting the numbers of viable cells were repeated every 24 h.
Assays for Cell Migration and Invasion
Cells (5 × 104) were added to the upper chamber of a Transwell assay system containing 100 μl of opti-MEM (Gibco Inc.); the lower chamber (Corning Inc.) was filled with 600 μl of DMEM medium with 10% FBS. The cells that had penetrated the membrane dividing the upper and lower chambers were incubated for 24 h before being fixed with 4-paraformaldehyde for 20 min and stained with crystal violet dye for 5 min. The invasion assay was performed in the same way as the migration assay above, with the exception that the upper chamber was precoated with 100 μl Matrigel (300 μg/ml; Corning Inc.). All experiments were performed in triplicate.
Wound-Healing Assay
A sterile 200 μl tip was used to make a straight scratch on a layer of confluent cells in a 6-well plate and washed twice with PBS. Then changed to opti-MEM. Images of the cells were taken at 0 h, 12 h, and 24 h.
Statistical Analysis
Statistical analysis was performed by GraphPad Prism v8.0. Data are presented as the mean ± SEM. Differences between the two groups were compared by t-tests. P value less than 0.05 was considered statistically significant.
Results
TOMM40 was Found to be a PDEMARG for BRCA
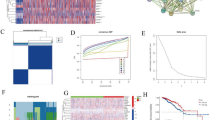
The locations of 28 MARGs on chromosomes was shown in Fig. 1A. Totals of 22 MARGs showed significant differences in expression levels between BRCA and normal samples (p < 0.05) (Fig. 1B). Based on these findings, two DEMARGs were chosen for further analysis. They included PRKN, which was down-regulated in BRCA tissues relative to normal tissues, and TOMM40, which was up-regulated in BRCA tissues (Fig. 1C). The KM curves for the two genes also indicate that TOMM40 is a likely PDEMARG for BRCA (pOS = 0.0021, pDFS = 0.016) (Fig. 1D).
TOMM40 was identified as a PDEMARG. A Locations of the 28 MARs on chromosomes. B Differences in the expression profiles of the 28 MARs between tumor and normal samples from the TCGA-BRCA database. *P < 0.05, **P < 0.01, ***P < 0.001. C The identification of DEMARGs from the TCGA-BRCA database and the expression heatmap of two of the identified DEMARGs (TOMM40 and PRKN). D Kaplan–Meier curves showing the overall survival time (left) and disease-free survival time (right) in the high-expression and low-expression groups of TOMM40 and PRKN for BRCA
Clinical Correlation and Prognosis Analysis of TOMM40 in Pan-Cancers
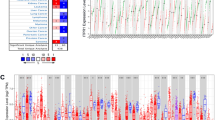
Analysis of the TOMM40 expression in tumors and normal samples from the 33 malignancies revealed that TOMM40 had differential expressions in 20 cancers (p < 0.05). Especially, TOMM40 was highly expressed in all cancers, except for pheochromocytoma and paraganglioma (PCPG) (Fig. 2A). Additionally, the relationship between TOMM40 and different clinical characteristics of cancers was linked to different clinical traits; four of these differences were statistically significant in kidney renal clear cell carcinoma (KIRC) (p < 0.01) (Fig. 2B). Also, we discovered that in 21 malignancies, the expression of TOMM40 was substantially linked with patient prognosis. The curves demonstrated a relationship between overall survival (OS) and TOMM40 expression levels, but these relationships were different for different tumors. In most cases, increased expression levels of TOMM40 were significantly linked to poor prognosis. (Fig. 2C, others in Supplementary Fig. 1).
The expression levels of TOMM40 in different cancers. A The differences in the expression levels of TOMM40 between tumor and normal samples in 33 cancers. *P < 0.05, **P < 0.01, ***P < 0.001. B The correlations between the expression levels of TOMM40 and the clinical features of different types of cancers. *P < 0.05, **P < 0.01, ***P < 0.001. C Kaplan–Meier curves of TOMM40 in different types of cancers
Evaluation of Potential Therapeutic Effects for TOMM40 in Pan-Cancers
To further explore the pertinent pathways and mechanisms of TOMM40 in various cancers, we took GSEA (Gene Set Enrichment Analysis) of it, and the functional analysis of TOMM40 in cancers was shown in Fig. 3A (others in Supplementary Fig. 2). In breast cancer, it was associated with the ribosome and oxidative phosphorylation, etc. Moreover, the average correlation of TOMM40 and functional status (invasion, metastasis, proliferation, epithelial-mesenchymal transformation (EMT), angiogenesis, apoptosis, cell cycle, differentiation, DNA damage, DNA repair, hypoxia, inflammatory) was shown in Fig. 3B, which suggested that TOMM40 was significantly associated with the function of the cell cycle, DNA damage and repair, among other things. TOMM40 was related to angiogenesis and DNA repair in BRCA. Tumor microenvironment (TME) is important to tumorigenesis and progression, which has a predicting value in prognosis and treatment. The majority of immunological checkpoints and TOMM40 have a strong negative connection in pan-cancers. Among these, TOMM40 exhibits a strong negative association with CD200R1 in 24 cancer types and a noteworthy positive correlation with CD200 in 26 cancer types. Notably, TOMM40 and all immunological checkpoints in LIHC showed favorable associations (Fig. 3C). Among 860 anticancer medicines, 5 were selected out because their sensitivities were strongly correlated with TOMM40 (|r|> 0.3 and p < 0.05) (Fig. 3D). Moreover, Fig. 3E displayed the relationship between medication sensitivity and TOMM40.
The immune function and drug sensitivity analyses of TOMM40. A KEGG enrichment analysis of TOMM40 in different types of cancers. If the enrichment score is > 0, the pathway is upregulated and if the score is < 0, the pathway is down-regulated. B Functional enrichment analysis of TOMM40 in cancers, *P < 0.05, **P < 0.01. C Correlation analysis between TOMM40 expression levels and immune checkpoints in cancers. D Drug sensitivity analyses for TOMM40. E Correlation between TOMM40 expression levels and drug susceptibility
Silencing of TOMM40 Expression Suppressed the Proliferation, Invasion and Migration of TNBC Cells In Vitro
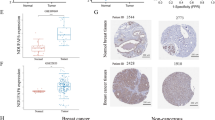
According to the qRT-PCR data, there was a substantial difference between the expression levels of TOMM40 in cancer samples and normal tissue samples (p < 0.0001) (Fig. 4A). We also found that the levels of the TOMM40 protein in BRCA tissues were considerably higher in BRCA tissues compared to normal tissues (Fig. 4B, C). According to the PAM50 subtype classification system, the higher expression levels of TOMM40 in TNBC might indicate that this gene plays a crucial function in the progression of TNBC(Fig. 4D).
The expression levels of TOMM40 in BRCA and its functional effects on TNBC cells. A The expression of TOMM40 was confirmed by qRT-PCR with clinical samples; the results were the same as those of the bioinformatics analysis. B The presence of the Tomm40 protein in BRCAs indicated that primary tumors had higher levels of Tomm40; similarly, the mRNA levels of TOMM40 were higher in primary tumors. C The results of immunohistochemistry to detect TOMM40 in BRCA and normal tissues acquired from HPA. D The classification of TOMM40 according to the PAM50 classification system. E GFP expression was observed by fluorescence microscopy after 24 h of knockdown of TOMM40; the transfection efficiency was evaluated via western blotting 48 h after transfection. F The proliferation of sh-NC and sh-TOMM40 cells was measured by CCK8 assays. G, H The effects of TOMM40 knockdown on the migration and invasiveness of sh-NC and sh-TOMM40 cells were evaluated using Transwell assays and wound-healing assays (G, H, respectively). Magnification: 200× (G) and 100× (H). All experiments were performed thrice. Quantitative data from the images were obtained using ImageJ. (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; ns non-signifificant)
We successfully knocked down the TOMM40 gene in MDA-MB-231 cells (Fig. 4E). The results of the CCK8 assay demonstrate that the deletion of TOMM40 can substantially suppress TNBC cell proliferation (Fig. 4F). Transwell and wound-healing assay results show that TOMM40 knockdown significantly decreased the invasiveness and migration capacity of the MDA-MB-231 cells (Figs. 4G, H). Overall, these results reveal that the down-regulation of TOMM40 expression significantly suppresses the invasiveness and migration potential of TNBC cells.
Discussion
Tumor heterogeneity in BRCA is very high. Although most patients receiving aggressive and systematic treatment usually have good prognoses, many BRCA tumors, especially TNBCs, develop resistance to anticancer therapies. Therefore, new biomarkers and potential therapeutic targets to treat TNBC are necessary. Our previous study proposed a new classification system for BRCAs based on the expression of mitophagy genes (MARGs). We believe that these genes would be very useful as novel biomarkers and therapeutic targets to treat BRCA, especially TNBC. The importance of TOMM40, a MARG that is also a biomarker associated with neurodegenerative processes and healthy aging, has not been studied in detail in cancers. Therefore, in this study, we have investigated the expression levels and prognostic value of TOMM40 in 33 different cancers. We have also, for the first time, investigated the function played by TOMM40 in the progression of BRCA using bioinformatic analyses and functional assays.
We first compared the expression profiles of 28 MARGs between BRCA and normal tissues; of these, 22 MARGs showed significant differences in expression between the two types of tissues. This clearly shows that dysregulation of mitophagy may be strongly associated with the development and progression of BRCA. With further analyses, we have identified TOMM40 as a PDEMARG. A pan-cancer analysis of the expression patterns of TOMM40 indicates that this gene is significantly associated with tumor malignancy. To explore the function of TOMM40 in the progression of BRCA, we knocked down this gene in a BRCA cell line, MDA-MB-231, and found that lower TOMM40 expression suppresses the proliferation of MDA-MB-231, as well as migration and invasiveness of these cells.
TOMM40 was differentially expressed in 20 different types of tumors and was also associated with certain clinical traits in most cancers, specifically in KIRC. we also found that TOMM40 was overexpressed in all the cancers analyzed in this study, except for PCPG (Dahia 2014). However, fewer studies reported TOMM40’s prognostic value in cancer patients (Yang et al. 2020). Based on the KM survival analyses, TOMM40 was significantly negatively linked to patient survival in 21 different types of cancer. This study has shown that TOMM40 could be of prognostic value in several cancers as we have found associations between high expression levels of TOMM40 and better OS in CESC, COAD, LUSC, PAAD, PCPG and STAD; however, higher expression levels of TOMM40 are associated with poor OS in ACC, BRCA, and KIRC. Overall, these results suggested that the prognostic value of TOMM40 is dependent on the type of cancer and needs to be further investigated. Our analyses of the associations between the clinical parameters of cancer patients and TOMM40 expression levels also show that TOMM40 can be useful as a novel, independent prognostic biomarker for KIRC. Functional enrichment analysis indicates that TOMM40 is associated with pathways involved in the cell cycle, DNA damage and repair, etc.
The TME is a crucial part of cancer, cancer cells have a close interaction with stromal cells (Hanahan and Coussens 2012). Accordingly, novel targets associated with TME can be used to develop better therapies, specifically immunotherapies, for treating cancers (Pitt et al. 2016). Immune checkpoints are inhibitory and stimulatory pathways essential for maintaining self-tolerance and regulating immune reactions (Marin-Acevedo et al. 2018). We find that TOMM40 was significantly and negatively correlated with most immune checkpoints in several cancers (Sharma and Allison 2015). Among these are the levels of CD40 and its ligand CD40L, both of which were negatively associated with TOMM40 levels in many tumors such as BLCA, LUSC, PRAD, and SKCM. The interactions between CD40 and CD40L stimulate B cells to secret cytokines and also activate T cells to attack and kill malignant cells (Kooten and Banchereau 2000; Armitage et al. 1992). In addition, levels of CD28 (which is critical for the costimulation of naive T lymphocytes and regulation of T cells) were also negatively associated with TOMM40 levels (Lenschow et al. 1996; Bour-Jordan et al. 2011; Salomon and Bluestone 2001). The levels of the inducible costimulatory molecule, ICOS, which is mainly expressed on activated CD4 cells, were also negatively associated with TOMM40 levels (Hutloff et al. 1999).
CD27 acted as a costimulatory molecule, enhancing T- and B-cell responses. When CD70 binds to CD27, NF-B is activated, which helps cells survive, boosts proliferative signals and improves effector functions(Borst et al. 2005). Modulating the CD70–CD27 interactions to activate the immune system against cancer cells is an attractive strategy to treat solid tumors or directly target malignant stem cells in leukemia (Riether et al. 2012), However, other studies have shown that the expression of CD70 on tumor cells can be a negative prognostic factor (Yang et al. 2007). Contrary to CD27, our findings revealed that CD70 exhibited a favorable connection with TOMM40 in pan-cancers. Cheng’s research found that CSCs utilized CD276 for immune escape in NHSCC and targeted CD276 may reduce CSCs in a CD8 T cell-dependent manner (Wang et al. 2021b). Our result also found that CD276 was positively associated with TOMM40 in many cancers. These results pointed out that TOMM40 may influence the activation and regulation of T cells in malignance to promote tumorigenesis, which needed further research in various tumors. Further, five potential drugs significantly correlated with TOMM40 were screened out, including 5-Fluoro deoxy uridine 10mer, Cladribine, SNS-314, PF-06873600 and LY-2606368, which provided a theoretical basis for the treatment of pan-cancers.
Based on the above research, the expression of TOMM40 in BRCA was further analyzed and found that it was highly expressed in TNBCs. Given the high recurrence and refractory treatment of TNBC, TOMM40 was knocked down in MDA-MB-231. The results showed that TOMM40 can inhibit the proliferation, migration and invasion of TNBC cells.
In summary, our study first identified TOMM40 as a PDEMARG and has found that this molecule could be a potential immunotherapeutic target for treating BRCAs. Few studies have examined the relationships between TOMM40 expression levels and tumors; however, based on the importance of MARGs in cancer development and progression, we have shown that TOMM40 could be of immense prognostic value and may be an important therapeutic target for cancer immunotherapies. We believe that our work will provide data and support the undertaking of more research in this area. Crucially, the function of TOMM40 in BRCAs, and especially in TNBCs, indicates that it is a promising target for anticancer therapy, especially immunotherapy. Although our conclusions need further experimental validation and clinical data support, we believe that it adds value to this field.
Data Availability
The datasets generated during the current study were available in The Cancer Genome Atlas (TCGA, xenabrowser.net) and The Reactome database (reactome.org/). The UCSC Xena database (xenabrowser.net/datapages/) was used to get clinical and patient survival information.
References
Armitage RJ, Fanslow WC, Strockbine L, Sato TA, Clifford KN, Macduff BM et al (1992) Molecular and biological characterization of a murine ligand for CD40. Nature. https://doi.org/10.1038/357080a0
Barriga V, Kuol N, Nurgali K, Apostolopoulos V (2019) The complex interaction between the tumor micro-environment and immune checkpoints in breast cancer. Cancers. https://doi.org/10.3390/cancers11081205
Bezuch N, Bradburn S, Robinson AC, Pendleton N, Payton A, Murgatroyd C (2021) Superior frontal gyrus TOMM40-APOE locus DNA methylation in Alzheimer’s disease. J Alzheimers Dis Rep. https://doi.org/10.3233/ADR-201000
Borst J, Hendriks J, Xiao Y (2005) CD27 and CD70 in T cell and B cell activation. Curr Opin Immunol. https://doi.org/10.1016/j.coi.2005.04.004
Bour-Jordan H, Esensten JH, Martinez-Llordella M, Penaranda C, Stumpf M, Bluestone JA (2011) Intrinsic and extrinsic control of peripheral T-cell tolerance by costimulatory molecules of the CD28/ B7 family. Immunol Rev. https://doi.org/10.1111/j.1600-065X.2011.01011.x
Cesari R, Martin ES, Calin GA, Pentimalli F, Bichi R, McAdams H et al (2003) Parkin, a gene implicated in autosomal recessive juvenile parkinsonism, is a candidate tumor suppressor gene on chromosome 6q25-q27. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.0931262100
Chacinska A, Koehler CM, Milenkovic D, Lithgow T, Pfanner N (2009) Importing mitochondrial proteins: machineries and mechanisms. Cell. https://doi.org/10.1016/j.cell.2009.08.005
Cheang MC, Martin M, Nielsen TO, Prat A, Voduc D, Rodriguez-Lescure A et al (2015) Defining breast cancer intrinsic subtypes by quantitative receptor expression. Oncologist. https://doi.org/10.1634/theoncologist.2014-0372
Chourasia AH, Macleod KF (2015) Tumor suppressor functions of BNIP3 and mitophagy. Autophagy. https://doi.org/10.1080/15548627.2015.1085136
Chourasia AH, Boland ML, Macleod KF (2015) Mitophagy and cancer. Cancer Metab. https://doi.org/10.1186/s40170-015-0130-8
Chourasia AH, Tracy K, Frankenberger C, Boland ML, Sharifi MN, Drake LE et al (2015) Mitophagy defects arising from BNi-p3 loss promote mammary tumor progression to metastasis. EMBO Rep. https://doi.org/10.15252/embr.201540759
Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D et al (2016) TCGAbiolinks: an R/bioconductor package fo-r integrative analysis of TCGA data. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv1507
Dahia PLM (2014) Pheochromocytoma and paraganglioma pathogenesis: learning from genetic heterogeneity. Nat Rev Cancer. https://doi.org/10.1038/nrc3648
Dolezal P, Likic V, Tachezy J, Lithgow T (2006) Evolution of the molecular machines for protein import into mitochondria. Science. https://doi.org/10.1126/science.1127895
Ellenrieder L, Mårtensson CU, Becker T (2015) Biogenesis of mitochondrial outer membrane proteins, problems and diseases. Biol Chem. https://doi.org/10.1515/hsz-2015-0170
Endo T, Yamamoto H, Esaki M (2003) Functional cooperation and separation of translocators in protein import into mitochondria, the double-membrane bounded organelles. J Cell Sci. https://doi.org/10.1242/jcs.00667
Gabriel K, Egan B, Lithgow T (2003) Tom40, the import channel of the mitochondrial outer membrane, plays an active role in sorting imported proteins. EMBO J. https://doi.org/10.1093/emboj/cdg229
Hanahan D, Coussens LM (2012) Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell. https://doi.org/10.1016/j.ccr.2012.02.022
Hu K (2021) Become competent in generating RNA-Seq heat maps in one day for novices without prior R experience. Methods Mol Biol Clifton NJ. https://doi.org/10.1007/978-1-0716-1084-8_17
Hutloff A, Dittrich AM, Beier KC, Eljaschewitsch B, Kraft R, Anagnostopoulos I et al (1999) ICOS is an inducible T-cell co-stimulator structurally and functionally related to CD28. Nature. https://doi.org/10.1038/16717
Kim I, Rodriguez-Enriquez S, Lemasters JJ (2007) Selective degradation of mitochondria by mitophagy. Arch Biochem Biophys. https://doi.org/10.1016/j.abb.2007.03.034
Lee EG, Chen S, Leong L, Tulloch J, Yu CE (2021) TOMM40 RNA transcription in Alzheimer’s disease brain and its implication in mitochondrial dysfunction. Genes. https://doi.org/10.3390/genes12060871
Lenschow DJ, Walunas TL, Bluestone JA (1996) CD28/B7 system of T cell costimulation. Annu Rev Immunol. https://doi.org/10.1146/annurev.immunol.14.1.233
Marin-Acevedo JA, Dholaria B, Soyano AE, Knutson KL, Chumsri S, Lou Y (2018) Next generation of immune checkpoint therapy in cancer: new developments and challenges. J Hematol OncolJ Hematol Oncol. https://doi.org/10.1186/s13045-018-0582-8
Mizushima N, Komatsu M (2011) Autophagy: renovation of cells and tissues. Cell. https://doi.org/10.1016/j.cell.2011.10.026
Mustafa MF, Saliluddin SM, Fakurazi S, Tizen Laim NMS, Md Pauzi SH, Nik Yahya NH et al (2021) Expression of autophagy and mitoph-agy markers in breast cancer tissues. Front Oncol. https://doi.org/10.3389/fonc.2021.612009
Naik PP, Birbrair A, Bhutia SK (2019) Mitophagy-driven metabolic switch reprograms stem cell fate. Cell Mol Life Sci CMLS. https://doi.org/10.1007/s00018-018-2922-9
Palikaras K, Lionaki E, Tavernarakis N (2018) Mechanisms of mitophagy in cellular homeostasis, physiology and pathology. Nat Cell Biol. https://doi.org/10.1038/s41556-018-0176-2
Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA et al (2000) Molecular portraits of human breast tumours. Nature. https://doi.org/10.1038/35021093
Pfanner N, Wiedemann N, Meisinger C, Lithgow T (2004) Assembling the mitochondrial outer membrane. Nat Struct Mol Biol. https://doi.org/10.1038/nsmb852
Pickrell AM, Youle RJ (2015) The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron. https://doi.org/10.1016/j.neuron.2014.12.007
Pitt JM, Marabelle A, Eggermont A, Soria JC, Kroemer G, Zitvogel L (2016) Targeting the tumor microenvironment: removing obstruction to anticancer immune responses and immunotherapy. Ann Oncol off J Eur Soc Med Oncol. https://doi.org/10.1093/annonc/mdw168
Riether C, Schürch C, Ochsenbein AF (2012) Modulating CD27 signaling to treat cancer. Oncoimmunology. https://doi.org/10.4161/onci.21425
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W et al (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv007
Ryan MT, Wagner R, Pfanner N (2000) The transport machinery for the import of preprote-ins across the outer mitochondrial membrane. Int J Biochem Cell Biol. https://doi.org/10.1016/s1357-2725(99)00114-4
Salomon B, Bluestone JA (2001) Complexities of CD28/B7: CTLA-4 costimulatory pathway-s in autoimmunity and transplantation. Annu Rev Immunol. https://doi.org/10.1146/annurev.immunol.19.1.225
Sharma P, Allison JP (2015) Immune checkpoint targeting in cancer therapy: toward combination strategies with curative potential. Cell. https://doi.org/10.1016/j.cell.2015.03.030
Straub SP, Stiller SB, Wiedemann N, Pfanner N (2016) Dynamic organization of the mitochondrial protein import machinery. Biol Chem. https://doi.org/10.1515/hsz-2016-0145
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. https://doi.org/10.3322/caac.21660
Tay SP, Yeo CW, Chai C, Chua PJ, Tan HM, Ang AX et al (2010) Parkin enhances the expression of cyclin-dependent kinase 6 and negatively regulates the proliferation of breast cancer cells. J Biol Chem. https://doi.org/10.1074/jbc.M110.108241
van Kooten C, Banchereau J (2000) CD40-CD40 ligand. J Leukoc Biol. https://doi.org/10.1002/jlb.67.1.2
Wang Y, Wang Z, Sun J, Qian Y (2021a) Identification of HCC subtypes with different prognosis and metabolic patterns based on mitophagy. Front Cell Dev Biol. https://doi.org/10.3389/fcell.2021.799507
Wang C, Li Y, Jia L, Kim JK, Li J, Deng P et al (2021b) CD276 expression enables squamous cell carcinoma stem cells to evade immune surveillance. Cell Stem Cell. https://doi.org/10.1016/j.stem.2021.04.011
Wiedemann N, Frazier AE, Pfanner N (2004) The protein import machinery of mitochondria. J Biol Chem. https://doi.org/10.1074/jbc.R400003200
Xu L, Lin D, Yin D, Koeffler HP (2014) An emerging role of PARK2 in cancer. J Mol Med Berl Ger. https://doi.org/10.1007/s00109-013-1107-0
Yang ZZ, Novak AJ, Ziesmer SC, Witzig TE, Ansell SM (2007) CD70+ non-Hodgkin lymphoma B cells induce Foxp3 expression and regulatory function in intratumoral CD4+CD25 T cells. Blood. https://doi.org/10.1182/blood-2007-03-082578
Yang W, Shin HY, Cho H, Chung JY, Lee EJ, Kim JH et al (2020) TOM40 inhibits ovarian cancer cell growth by modulating mitochondrial function including intracellular ATP and ROS levels. Cancers. https://doi.org/10.3390/cancers12051329
Zhang H, Meltzer P, Davis S (2013) RCircos: an R package for Circos 2D track plots. BMC Bioinformatics. https://doi.org/10.1186/1471-2105-14-244
Funding
No funding was received for conducting this study and authors did not receive support from any organization for the submitted work.
Author information
Authors and Affiliations
Contributions
YZ* initiated and directed the study. XW, LJ and WL contributed to the design and implementation of the research. SZ directed and supervised the experiments. YZ performed the experiments. YZ* and YZ interpreted the experimental results and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. All authors have no potential conflict of interest.
Ethical Approval and Consent to Participate
Ethics approval and consent to participate was not applicable.
Consent for Publication
Consent for publication was not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Y., Wei, X., Jia, L. et al. Pan-Cancer Analysis of the Prognostic and Immunological Role of TOMM40 to Identify Its Function in Breast Cancer. Biochem Genet (2024). https://doi.org/10.1007/s10528-024-10794-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10528-024-10794-6