Abstract
An important feature of EBV-associated gastric cancer (EBVaGC) is extensive methylation of viral and host genomes. This study aims to analyze DNA methylation-driven genes (DMDG) in EBVaGC through bioinformatics methods, providing an important bioinformatics basis for the differential diagnosis and treatment of potential methylation biomarkers in EBVaGC. We downloaded the mRNA expression profiles and methylation datasets of EBVaGC and EBV-negative gastric cancer (EBVnGC) through the TCGA database to screen methylated-differentially expressed genes (MDEGs). DNA methylation-driver genes were identified based on MethylMix algorithm and key genes were further identified by LASSO regression and Random Forest algorithm. Then, we performed gene enrichment analysis for key genes and validated them by GEO database. Gene expression differences in EBVaGC and EBVnGC cell lines was determined by quantitative real-time PCR (qRT-PCR) and western blotting and in GT38 cell and SNU719 cell which all treated by 5-Aza-CdR. Finally, the effect of key gene on the migration and proliferation capacity of EBVaGC cells was determined by Transwells assay and Cell counting Kit-8 (CCK-8) assay. We obtained a total of 687 hypermethylation-low expression genes (Hyper-LGs) and further obtained 53 DNA methylation-driver genes based on the MethylMix algorithm. A total of six key genes (SCIN, ETNK2, PCDH20, PPP1R3C, MATN2, and HOXA5) were identified by LASSO regression and Random Forest algorithm. Among them, SCIN expression was significantly lower in EBVaGC cell lines than in EBVnGC cell lines, and its expression was significantly recovered in EBVaGC cell lines treated with 5-Aza-CdR. Overexpression of SCIN can promote the proliferation and migration capacity of EBVaGC cells. Our study will provide some bioinformatics basis for the study of EBVaGC-related methylation. SCIN may be used as potential methylation biomarkers for the diagnosis and treatment of EBVaGC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epstein-Barr Virus (EBV) is a human gammaherpes virus with lifelong latency in more than 95% of adults (Pagano et al. 2018). As the first human oncovirus to be discovered (Epstein 2015), EBV has been associated with the development of a variety of human malignancies, including Burkitt lymphoma (BL), Hodgkin 's disease, T-cell lymphoma, nasopharyngeal carcinoma (NPC) and approximately 5–10% Epstein-Barr virus-associated gastric cancer (EBVaGC) (Leong and Lung 2021; Shannon-Lowe and Rickinson 2019).
In 2014, the Cancer Genome Atlas (TCGA) classified gastric cancer into four subtypes: EBV-positive, microsatellite unstable (MSI), genomic stable (GS), and chromosomal unstable (CIN) tumors (Bass et al. 2014). Among them, EBV-positive tumors are EBVaGC, which mainly show recurrent PIK3CA mutations, DNA hypermethylation, amplification of JAK2, CD274, and PDCD1LG2 genes (Bass et al. 2014). Alterations in DNA methylation were frequently observed in tumor cells compared to corresponding normal cells (Nishiyama and Nakanishi 2021). In gastric cancer, DNA hypermethylation is more prevalent in EBVaGC than in EBV-negative gastric cancer (EBVnGC) (Kang et al. 2008). Characteristic molecular abnormalities in EBVaGC are caused by global nonrandom CpG island methylation in promoter regions of many cancer-associated genes, which leads to their down-regulation (Chen et al. 2012). It has been shown that EBV infection causes extensive methylation of EBVaGC, and its encoded latent membrane proteins 1 and 2A (LMP1 and LMP2A) can induce the expression of DNA methyltransferases (including DNMT1, DNMT3a, and DNMT3b) and affect the methylation status of the viral and host genomes (Niller et al. 2016).
At present, there are few bioinformatics studies on DNA methylation in EBVaGC, and there is no study on EBVaGC related DNA methylation based on TCGA database. To better understand the role DNA methylation plays in the development of EBVaGC, we screened methylated-differentially expressed genes (MDEGs) in EBVaGC based on the TCGA database to identify DNA methylation-driven genes of EBVaGC, and selected key genes by LASSO regression and Random Forest algorithm. Finally, through experimental verification, Scinderin (SCIN) was identified as a key DNA methylation-driver gene in EBVaGC (Fig. 1). This study will reveal new directions for studying DNA methylation in EBVaGC and provide bioinformatics basis for identifying potential methylation biomarkers for the diagnosis and treatment of EBVaGC.
Materials and Methods
Data sources
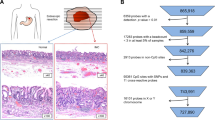
We used the TCGA database (https://portal.gdc.cancer.gov/), RNA-seq data sets (normalized counts) with DNA methylation data sets from gastric cancer patients were downloaded. After excluding cases with missing clinical information and gene expression data, a total of 258 gastric cancer patients (including 23 EBVaGC tissue samples and 235 EBVnGC tissue samples) with complete EBV information were obtained by intersection with EBV information provided in published articles (Rooney et al. 2015). All data used in the study came from the TCGA database and therefore did not involve ethical approval or informed consent.
Data Processing and Screening of MDEGs
DESeq and limma packages of R software were used to identify differentially expressed genes (DEGs) and differentially methylated genes (DMGs) of EBVaGC and EBVnGC. |log2FC|> 1.5 and FDR < 0.05 were used as DEGs threshold, |β|> 0.2 and FDR < 0.05 were used as DMGs threshold.
Functional and Pathway Enrichment Analysis
Based on the DAVID database (Huang et al. 2009), functional enrichment analysis of Hyper-LGs was performed by Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis (P < 0.05 was considered statistically significant).
Identification of DNA Methylation-Driver Genes
MethylMix is an R package for identifying DNA methylation-driver genes (Gevaert 2015), and based on the MethylMix algorithm, the correlation between DNA methylation levels and gene expression is calculated, and genes with significant negative correlations are DNA methylation-driver genes. Hypermethylation-low expression genes (Hyper-LGs) were analyzed using the MethylMix package of R software to identify methylation-driver genes of EBVaGC (|logFC|> 0, P < 0.05, Cor < − 0.3).
LASSO Regression
LASSO algorithm is an estimation method that can realize the reduction of index set. By constructing a penalty function, LASSO regression obtains a more refined model, compresses some regression coefficients, the sum of absolute values of forced coefficients is smaller than a fixed value, and some regression coefficients are reduced to zero, and only variables with non-zero regression coefficients can be retained as part of the model. We performed LASSO regression of 53 methylation-driver genes to select key genes using the glmnet package of R software (McEligot et al. 2020).
Random Forest Algorithm
Build training sample set and prediction sample set. A random forest model was constructed to rank the importance of feature variables using the Random Forest package of R software.
GEO Database Identification
Gene mRNA expression data (GSE51575, including 12 EBVaGC tissue samples versus 14 EBVnGC tissue samples) and gene methylation data (GSE31788, including 11 EBVaGC tissue samples and 43 EBVnGC tissue samples) were obtained from the GEO database (www.ncbi.nlm.nih.gov/geo/). Visualization of the expression and methylation levels of the key gene was performed by Xiantao Academic (https://www.xiantao.love/).
Cell Lines
In this study, we chose EBV-positive gastric cancer cell lines: GT38, GT39, and SNU719, and EBV-negtive gastric cancer cell lines: HGC27 and MGC803. The GT38 and GT39 cell lines was gifted by Prof. Takeshi Sairenji (Tottori University, Japan). SNU719 cell lines were provided by Professor Qian Tao (University of Hong Kong, China). All cell lines were cultured in DMEM (Gibco, USA) with 10% fetal bovine serum (Biological Industries, Israel) and 2% penicillin–streptomycin (Initrogen, USA), 37 °C with 5% CO2.
Treatment of Cell Lines with Decitabine
Decitabine (5-Aza-CdR) is a DNA methyltransferase inhibitor, which can replace DNA incorporation into cytosine allows covalent capture of a DNA methyltransferase into DNA, resulting in irreversible inhibition of the enzyme. When cells were cultivated to 70%-80% confluency, GT38 and SNU719 cells were treated daily with 7.5 μmol/L or 15 μmol/L Decitabine (ApexBio, USA) for three days. Cells treated with the same concentration of DMSO (Solarbio, China) were used as a control.
RNA Extraction and Quantitative Real-Time PCR (qRT-PCR)
Total RNA was extracted from cells using TransZol Up Plus RNA Kit (TransGen, China). The extracted RNA was reverse transcribed to cDNA by Evo M-MLV RT Mix Kit (Accurate Biology, China). Using cDNA as a template, the transcription level of mRNA was detected by a LightCycler 96 System with a FastStart DNA Master SYBR Green Kit (Roche, Germany). The relationship between target genes expression and internal reference gene expression β-actin was analyzed by a comparative Ct method based on SYBR Green (relative fold-change = 2 − ΔΔCt). The sequences of the specific forward and reverse primers: SCIN-F (TCTACACGTGGCAAGGAGCAA), SCIN-R(TCTTTGCCTTGGGAGACTCGG); β-actin-F (TCCTGTGGCATCCACGAAACT), β-actin-R (GAAGCATTTGCGGTGGACGAT).
Protein Extraction and Western Blotting Assay
All cells were washed by cold phosphate-buffered saline (PBS) and lysed with RIPA buffer mixture (RIPA: PMSF: phosphatase inhibitors, 100:1:1) on ice for 20 min. Then, the cell lysate was centrifuged at 12,000 rpm at 4 °C for 20 min. The centrifuged supernatant was mixed with 5 × loading buffer and boiled for 5 min to denature the proteins. The proteins were run on 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS–PAGE) gels and were transferred onto PVDF membranes (Millipore, USA). After transfer, the PVDF membranes were blocked in 5% non-fat milk for 2 h at room temperature. Next, the primary antibody was incubated overnight at 4 °C, and the second antibody was incubated for 2 h at room temperature the next day. The protein expression level was detected by enhanced chemiluminescence detection system (Fusion Fx, VILBER LOURMAT). Primary antibody: Anti-DNMT1 antibody (abcam, UK, ab188453, 1:1000 dilution), Anti-SCIN antibody (abcam, ab191396, 1:1000 dilution) Anti-N-cadherin (abcam, ab76011, 1:1000 dilution), E-cadherin (abmart, PA6382, 1:1000 dilution), Anti-β-catenin (abmart, PK02151, 1:1000 dilution), Anti-Vimentin (abmart, T55134, 1000 dilution), and Anti-β-actin antibody (ABclonal, China, AC006, 1:5000 dilution). Second antibody: Goat Anti-Rabbit IgG H&L (HRP) (abcam, ab205718, 1:2000 dilution) and Goat Anti-Mouse IgG H&L (HRP) (Abcam, ab205719, 1:5000 dilution). All antibodies were diluted using the Western-specific primary and secondary antibody dilution (Boster, China, AR1017).
Plasmid Transfection
The pCMV-EGFP-SCIN (human)-Puro plasmid was purchased from Miaoling Biology (China), and the SCIN plasmid and its negative control were transfected into GT38 and SNU719 cells using Lipofectamine 2000 reagent (Invitrogen, USA).
Transwell Migration Assay
The migration ability of the GC cells was measured using the transwell inserts with 8 μm pores (Corning, New York, USA). 8 × 104 cells were seeded into the upper chamber with a serum-free medium and the medium was supplemented with 20% FBS in the lower chamber. Cells transfected with plasmid were incubated for 48 h. Then the cells in the upper compartment were wiped off, and the cells attached to the bottom of the compartment membrane were fixed and stained for observation under a microscope.
Cell Counting Kit-8 (CCK-8) Assay
The Cell proliferation was detected using CCK-8 kits (CCK-8, Boster Biological Technology, China). The inoculation amount of GT38 and SNU719 was 104 cells per well, and six repeat wells were set for each sample. 10 μL CCK-8 (by 1:9 in CCK-8 to medium) were added to each well within the specified time period (24 h, 48 h, 72 h, 96 h). The cells were then incubated for an additional 1 h at 37 °C, and the absorbance was measured using Soft-Max apparatus (Bio-Tek ELx808, USA) at a wavelength of 450 nm.
Statistical Analysis
Data were analyzed with using Students t-test. Analyses were performed using GraphPad Prism 8.0.2 (GraphPad Software, USA. All data were expressed as the mean ± standard error of the mean (SEM)), and all experiments were repeated at least three times.
Results
Identification of MDEGs in EBVaGC
Based on the DESeq2 algorithm, 1031 genes were found to be down-regulated and 118 genes were up-regulated in the expression profiling dataset (Fig. 2A, ). In addition, we found that 7404 genes were hypermethylated and 10 were hypomethylated in the methylation dataset (Fig. 2A). Hypermethylated genes overlapped with down-regulated genes, resulting in 687 Hyper-LGs (Fig. 2C).While upregulated genes did not intersect with hypomethylated genes (Fig. 2D).
Identification of MDEGs in EBVaGC. AHistograms of up-regulated genes, down-regulated genes, hypermethylated genes, and hypomethylated genes. B Volcano plot of DMGs in EBVaGC and EBVnGC. C Venn diagram of down-regulated genes and hypermethylated genes. D Venn diagram of up-regulated genes and hypomethylated genes
Functional Enrichment and Pathway Analysis of Hyper-LGs
To further investigate the molecular role of MDEGs in the development of EBVaGC, GO enrichment analysis of these genes was performed and classified according to their function (Fig. 3). The results were divided into three categories: Hyper-LGs were mainly enriched in hormone transport and secretion, potassium ion migration and transmembrane transport, and embryonic organ morphogenesis. For molecular function, these genes are particularly abundant in ion channel activity and ion transmembrane transporters. In the cellular component module, these genes are involved in the assembly of voltage-gated potassium channel complexes, potassium channel complexes, ion channel complexes and transmembrane transport complexes.
Pathway analysis of genes with low expression and hypermethylation in EBVaGC based on the KEGG database revealed that (Fig. 4) the above genes were closely related to neuroactive ligand-receptor interaction, Wnt signaling pathway, Hippo signaling pathway, insulin secretion, fat digestion and absorption, and signaling pathways that regulate stem cell pluripotent.
Identification of DNA Methylation-Driver Genes
Hyper-LGs in EBVaGC were analyzed based on MethylMix algorithm to further identify DNA methylation-driver genes in EBVaGC. Screening criteria for DNA methylation-driver genes were |logFC|> 0, FDR < 0.05, and Cor < -0.3. Eventually, a total of 53 DNA methylation-driver genes were screened (Table 1).
Screening Key Genes
DNA methylation-driver genes were screened by machine learning algorithm LASSO regression (Fig. 5A, B). Subsequently, we construct training and prediction sample sets and do feature importance ranking of methylation-driver genes by Random Forest algorithm (Fig. 5C). A total of 6 key genes were obtained by intersecting genes with importance greater than 0.5 with the screening results of LASSO regression: SCIN, ETNK2, PCDH20, PPP1R3C, MATN2, and HOXA5 (Fig. 5D).
LASSO regression and Random Forest algorithm screen for key genes. A LASSO regression cross validation curve. The parameter corresponding to the dashed line on the left (lambda, min) has the smallest error and determines the number of genes finally screened. B LASSO coefficient path diagram. 53 curves of different colors represent 53 genes, each curve represents the change trajectory of each independent variable coefficient, and the end of each curve points to the respective coefficient. C Random Forest algorithm for feature importance ranking of methylation-driver genes (based on Gini coefficients). D Venn diagram of intersection of LASSO regression and Random Forest algorithm
GEO Database Verification
To confirm our results, we further validated key genes by GEO database (GSE51575 and GSE31788). As verified by dataset GSE51575, the mRNA expression levels of genes ETNK2 and HOXA5 in EBVaGC were significantly higher than those in EBVnGC (Fig. 6A). Because of the GSE31788 dataset is based on the early platform GPL8490 Illumina Human Methylation 27 Bead Chip, probes do not cover genes SCIN, ETNK2, and PCDH20, so only the remaining key genes were identified. It was verified that the methylation levels of genes PPP1R3C, MATN2, and HOXA5 in EBVaGC were significantly higher than those in EBVnGC (Fig. 6B).
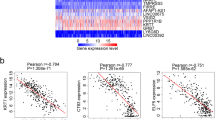
In GEO database, the differences in mRNA expression level and DNA methylation level between key genes in EBVaGC and EBVnGC. A Box plots of mRNA expression level differences between SCIN, ETNK2, PCDH20, HOXA5, MATN2, and PPP1R3C in EBVaGC and EBVnGC. B Box plots of DNA methylation level differences between HOXA5, MATN2, and PPP1R3C in EBVaGC and EBVnGC. (**P < 0.01, ***, P < 0.001)
Gene Expression Verification
To further validate the analysis results, we designed primers for six key genes to verify gene expression differences in EBVaGC and EBVnGC cell lines and examined the changes in gene expression levels in EBVaGC cell lines treated with 5-Aza-CdR by qRT-PCR. After excluding genes that were inconsistent with the analysis of the database and had low expression levels in cells, we found that SCIN expression was significantly different in EBVaGC and EBVnGC gastric cancer cell lines and significantly higher in EBVaGC gastric cancer cell lines treated with 5-Aza-CdR (Fig. 7A, B). Subsequently, we further verified that the expression of SCIN in EBVaGC cell lines was significantly lower by Western blotting than in EBVnGC cell lines. SCIN expression was significantly recovered in EBVaGC cell lines treated with 5-Aza-CdR at 7.5 μmol/L or 15 μmol/L for 3 days (Fig. 7C–F).
A SCIN expression in EBVaGC and EBVnGC cell lines was determined by qRT-PCR. B SCIN expression in GT38 and SNU719 which all treated with 5-Aza-CdR at 7.5 μmol/L or 15 μmol/L was determined by qRT-PCR. C, D SCIN expression in EBVaGC and EBVnGC cell lines was determined by western blotting. (E–F) SCIN expression in GT38 and SNU719 which all treated with 5-Aza-CdR at 7.5 μmol/L or 15 μmol/L was determined by western blotting. (**P < 0.01, ***P < 0.001, ****P < 0.0001)
SCIN Promotes the Proliferation and Migration Ability of EBVaGC Cells
To further explore the role of SCIN on the development of EBVaGC, we transfected a plasmid for SCIN in GT38 and SNU719, two EBV positive gastric cancer cell lines (Fig. 8A). The results of the CCK-8 assay found that SCIN promotes the proliferative capacity of GT38 and SNU719 cell lines (Fig. 8B). The results in Fig. 8C, D indicated that the overexpression of SCIN was able to promote the expression of N-cadherin, Vimentin, and β-catentin, and to inhibit the expression of E-cadherin. Meanwhile, we also found that the overexpression of SCIN promoted the invasive ability of GT38 and SNU719 cells by transwell assay (Fig. 8E, F).
A CCK-8 assays were performed to determine GT38 and SNU719 cell proliferation ability after transfection with SCIN plasmids. B, C The expression of EMT-related markers was detected by Western blot in GT38 and SNU719 cell line. D Transwell migration assays were performed to determine GT38 and SNU719 cell migration ability after transfection with SCIN plasmids. (**P < 0.01, ***P < 0.001, ****P < 0.0001)
Discussion
In this study, we identified a total of 687 Hyper-LGs and no Hypo-HGs. It is again strongly demonstrated that DNA hypermethylation is more prevalent in EBVaGC than EBVnGC.
GO functional analysis of Hyper-LGs showed that these genes mainly involved ion channels and ion transport across membranes, and the results of KEGG pathway enrichment also confirmed this analysis again. Transporters and ion channels play important roles in regulating ion transport, mucus barrier function, signaling pathways, and gastric acid secretion in the stomach (Yuan et al. 2020). The dysfunction of ion transport mechanism (ITM), such as ion channels and transporters, can participate in cancer cell proliferation, apoptosis, differentiation and other processes through their different signaling pathways (Xie et al. 2018).In addition to this, the KEGG enrichment results also suggested that these genes involved the Wnt signaling pathway and the Hippo signaling pathway. As three repressors in the Wnt signaling pathway, WIF (Hsieh et al. 1999), NLK (Ishitani et al. 1999), and APC (Aoki and Taketo 2007) are all hypermethylated in EBV-positive nasopharyngeal carcinoma and gastric cancer cell lines (Zhao et al. 2017), EBV encoded miR-BART-9-3p may be involved in the Wnt signaling pathway by targeting WIF, NLK, and APC (Zhao et al. 2017). Genomic stable (GS) gastric cancer, on the other hand, has the potential to have unique features in the DNA hypomethylation profile of the Wnt signaling pathway (Toshima et al. 2023). The Hippo signaling effectors YAP and TAZ induce EBV lytic reactivation via TEAD in epithelial cells (Sciver et al. 2021). However, the effect of EBV on the methylation status of genes involved in the Hippo signaling pathway remains unknown and warrants further in-depth study.
In this study, we identified DNA methylation-driver genes based on transcriptome profiling and DNA methylation sequencing data in gastric cancer tissue samples in the TCGA database and screened out six potential key genes by Random Forest algorithm and LASSO regression. We subsequently performed qRT-PCR and western blotting experiments in three EBVaGC cell lines, GT38, GT39, SNU719 and two EBVnGC gastric cancer cell lines, HGC27 and MGC803. Due to the inherent differences in cellular and tissue levels, we only identified SCIN from six key genes as the key DNA methylation-driver gene in EBVaGC. However, this does not negate the potential of other selected genes as key DNA methylation-driver genes for EBVaGC, and relevant experiments should be verified in gastric cancer tissues in the future.
SCIN encodes scinderin, which is a major member of the actin-binding family of proteins that can affect cell migration by regulating actin function (Marcu et al. 1994; Zhang et al. 1996). Recently, SCIN has been identified as a new oncogene, associated with malignant phenotypes of various tumors such as hepatocellular carcinoma, lung cancer, prostate cancer and glioma, and high levels of SCIN expression in tumors often predict the poor prognosis of patients (Zhou et al. 2020; Wang et al. 2014, 2022; Liu et al. 2015). In gastric cancer, some studies have found that scinderin can promote the proliferation, migration and metastasis of gastric cancer cells, and can be used as an indicator to predict the prognosis of gastric cancer patients. Our results of Transwell and CCK-8 experiments in EBVaGC cell lines also confirm the previous studies. SCIN may be a novel prognostic marker and a potential therapeutic target in gastric cancer (Liu et al. 2016; Huang et al. 2021; Chen et al. 2014). Here, we discover that EBVaGC has a lower level of SCIN expression in EBVaGC compared to EBVnGC, and find that this may be correlated with the epigenetic feature of hypermethylation in EBVaGC. It is worth mentioning that, EBVaGC is a unique subtype of gastric cancer, a series of clinical studies show that EBVaGC lymph node metastasis has lower rates and longer survival. Compared with other types of gastric cancer, EBVaGC patients often have a better prognosis (Camargo et al. 2011, 2014). Our results may provide new ideas for explaining this view, and the better prognosis of EBVaGC may be associated with the methylation status of a range of oncogenes such as SCIN.
Epigenetic abnormalities are an important mechanism driving tumorigenesis and affect the expression of numerous genes (Lee and Kim 2022) Aberrant methylation of CpG islands in gene promoter regions is an important mechanism. Methylation of the promoter region of a gene usually suppresses gene expression and plays a role in driving tumorigenesis (Nishiyama and Nakanishi 2021). Compared to other gastric cancer subtypes, EBVaGC has molecular signatures of hypermethylation status. The EBV encoding product can regulate the expression of DNMTs (Niller et al. 2016). Song et al. (Song et al. 2022) and Wang et al. (Wang et al. 2019) respectively reported that Epstein-Barr virus nuclear antigen 1 (EBNA1) and LMP2A could upregulate the expression of DNMT3a in EBVaGC. In the following study, we will focus on how the SCIN methylation sites change and how EBV factors indirectly contribute to abnormal SCIN expression by affecting methyltransferase expression, and to deeply confirm SCIN expression in animal experiments and clinical samples.
Conclusion
In this study, we identify DNA methylation-driver genes by bioinformatics analysis of MDEGs in EBVaGC based on the TCGA database. After experimental verification, SCIN was identified as a key DNA methylation-driver genes in EBVaGC, which may be used as potential methylation biomarkers for the diagnosis and treatment of EBVaGC. This study will reveal new directions for studying DNA methylation in EBVaGC, providing some bioinformatics basis for EBVaGC-related methylation research.
Data Availability
The datasets analyzed during the current study are available from the corresponding authors on reasonable request.
References
Aoki K, Taketo MM (2007) Adenomatous polyposis coli (APC): a multi-functional tumor suppressor gene. J Cell Sci 120:3327–3335. https://doi.org/10.1242/jcs.03485
Bass AJ, Thorsson V, Shmulevich I, Reynolds SM, Miller M, Bernard B et al (2014) Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513:202–209. https://doi.org/10.1038/nature13480
Camargo MC, Murphy G, Koriyama C, Pfeiffer RM, Kim WH, Herrera-Goepfert R et al (2011) Determinants of Epstein-Barr virus-positive gastric cancer: an international pooled analysis. Br J Cancer 105:38–43. https://doi.org/10.1038/bjc.2011.215
Camargo MC, Kim WH, Chiaravalli AM, Kim KM, Corvalan AH, Matsuo K et al (2014) Improved survival of gastric cancer with tumour Epstein-Barr virus positivity: an international pooled analysis. Gut 63:236–243. https://doi.org/10.1136/gutjnl-2013-304531
Chen J-N, He D, Tang F, Shao C-k (2012) Epstein-Barr virus-associated gastric carcinoma: a newly defined entity. J Clin Gastroenterol 46:262–271
Chen XM, Guo JM, Chen P, Mao LG, Feng WY, Le DH et al (2014) Suppression of scinderin modulates epithelial-mesenchymal transition markers in highly metastatic gastric cancer cell line SGC-7901. Mol Med Rep 10:2327–2333. https://doi.org/10.3892/mmr.2014.2523
Epstein A (2015) Why and how Epstein-Barr virus was discovered 50 years ago. Curr Top Microbiol Immunol 390:3–15. https://doi.org/10.1007/978-3-319-22822-8_1
Gevaert O (2015) MethylMix: an R package for identifying DNA methylation-driven genes. Bioinformatics 31:1839–1841. https://doi.org/10.1093/bioinformatics/btv020
Hsieh J-C, Kodjabachian L, Rebbert ML, Rattner A, Smallwood PM, Samos CH et al (1999) A new secreted protein that binds to Wnt proteins and inhibits their activites. Nature 398:431–436. https://doi.org/10.1038/18899
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. https://doi.org/10.1038/nprot.2008.211
Huang Y, Du X, Chen X, Chen C, Wang H, Yang Y et al (2021) MiR-301a-5p/SCIN promotes gastric cancer progression via regulating STAT3 and NF-κB signaling. J Cancer 12:5394–5403. https://doi.org/10.7150/jca.59747
Ishitani T, Ninomiya-Tsuji J, Nagai S-i, Nishita M, Meneghini M, Barker N et al (1999) The TAK1–NLK–MAPK-related pathway antagonizes signalling between β-catenin and transcription factor TCF. Nature 399:798–802. https://doi.org/10.1038/21674
Kang GH, Lee S, Cho N-Y, Gandamihardja T, Long TI, Weisenberger DJ et al (2008) DNA methylation profiles of gastric carcinoma characterized by quantitative DNA methylation analysis. Lab Invest 88:161–170. https://doi.org/10.1038/labinvest.3700707
Lee JE, Kim M-Y (2022) Cancer epigenetics: Past, present and future. Semin Cancer Biol 83:4–14. https://doi.org/10.1016/j.semcancer.2021.03.025
Leong MML, Lung ML (2021) The impact of Epstein-Barr virus infection on epigenetic regulation of host cell gene expression in epithelial and lymphocytic malignancies. Front Oncol 2021:11. https://doi.org/10.3389/fonc.2021.629780
Liu H, Shi D, Liu T, Yu Z, Zhou C (2015) Lentivirus-mediated silencing of SCIN inhibits proliferation of human lung carcinoma cells. Gene 554:32–39. https://doi.org/10.1016/j.gene.2014.10.013
Liu JJ, Liu JY, Chen J, Wu YX, Yan P, Ji CD et al (2016) Scinderin promotes the migration and metastasis of gastric cancer cells and predicts the outcome of patients. Cancer Lett 376:110–117. https://doi.org/10.1016/j.canlet.2016.03.035
Marcu MG, Rodríguez del Castillo A, Vitale ML, Trifaró JM (1994) Molecular cloning and functional expression of chromaffin cell scinderin indicates that it belongs to the family of Ca(2+)-dependent F-actin severing proteins. Mol Cell Biochem 141:153–165. https://doi.org/10.1007/bf00926179
McEligot AJ, Poynor V, Sharma R, Panangadan A (2020) Logistic LASSO regression for dietary intakes and breast cancer. Nutrients. https://doi.org/10.3390/nu12092652
Niller HH, Banati F, Salamon D, Minarovits J (2016) Epigenetic Alterations in Epstein-Barr Virus-Associated Diseases. In: Minarovits J, Niller HH (eds) Patho-Epigenetics of Infectious Disease. Springer International Publishing, Cham, pp 39–69
Nishiyama A, Nakanishi M (2021) Navigating the DNA methylation landscape of cancer. Trends Genet 37:1012–1027. https://doi.org/10.1016/j.tig.2021.05.002
Pagano JS, Whitehurst CB, Andrei G (2018) Antiviral drugs for EBV. Cancers 10:197
Rooney MS, Shukla SA, Wu CJ, Getz G, Hacohen N (2015) Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160:48–61. https://doi.org/10.1016/j.cell.2014.12.033
Shannon-Lowe C, Rickinson A (2019) The global landscape of EBV-associated tumors. Front Oncol. https://doi.org/10.3389/fonc.2019.00713
Song H, Zhang Y, Liu J, Liu W, Luo B (2022) Activation of DNA methyltransferase 3a by Epstein-Barr nuclear antigen 1 in gastric carcinoma. Digest Liver Dis 54:973–983. https://doi.org/10.1016/j.dld.2021.06.004
Toshima T, Tanioka H, Mori Y, Tanaka T, Yasui K, Kimura K et al (2023) Genomically Stable gastric cancer characterized by hypomethylation in Wnt signal cascade. Oncology 101:105–116. https://doi.org/10.1159/000527098
Van Sciver N, Ohashi M, Pauly NP, Bristol JA, Nelson SE, Johannsen EC et al (2021) Hippo signaling effectors YAP and TAZ induce Epstein-Barr Virus (EBV) lytic reactivation through TEADs in epithelial cells. PLOS Pathog 17:e1009783. https://doi.org/10.1371/journal.ppat.1009783
Wang D, Sun SQ, Yu YH, Wu WZ, Yang SL, Tan JM (2014) Suppression of SCIN inhibits human prostate cancer cell proliferation and induces G0/G1 phase arrest. Int J Oncol 44:161–166. https://doi.org/10.3892/ijo.2013.2170
Wang J, Liu W, Zhang X, Zhang Y, Xiao H, Luo B (2019) LMP2A induces DNA methylation and expression repression of AQP3 in EBV-associated gastric carcinoma. Virology 534:87–95. https://doi.org/10.1016/j.virol.2019.06.006
Wang Y, Li Z, Bai L, Zhang D, Zhang T, Ren F (2022) Scinderin is a novel oncogene for its correlates with poor prognosis, immune infiltrates and matrix metalloproteinase-2/9 (MMP2/9) in Glioma. Brain Sci 12:1415
Xie J, Zhu XY, Liu LM, Meng ZQ (2018) Solute carrier transporters: potential targets for digestive system neoplasms. Cancer Manag Res 10:153–166. https://doi.org/10.2147/cmar.S152951
Yuan D, Ma Z, Tuo B, Li T, Liu X (2020) Physiological significance of ion transporters and channels in the stomach and pathophysiological relevance in gastric cancer. Evid-Based Complement Altern Med 2020:2869138. https://doi.org/10.1155/2020/2869138
Zhang L, Marcu MG, Nau-Staudt K, Trifaro JM (1996) Recombinant scinderin enhances exocytosis, an effect blocked by two scinderin-derived actin-binding peptides and PIP2. Neuron 17:287–296. https://doi.org/10.1016/s0896-6273(00)80160-9
Zhao Z, Liu W, Liu J, Wang J, Luo B (2017) The effect of EBV on WIF1, NLK, and APC gene methylation and expression in gastric carcinoma and nasopharyngeal cancer. J Med Virol 89:1844–1851. https://doi.org/10.1002/jmv.24863
Zhou B, Chen TW, Jiang YB, Wei XB, Lu CD, Li JJ et al (2020) Scinderin suppresses cell proliferation and predicts the poor prognosis of hepatocellular carcinoma. Oncol Lett 19:2011–2020. https://doi.org/10.3892/ol.2020.11262
Acknowledgements
The authors would like to apologize to the many researchers who have contributed to this area of research but have not been cited in this review due to space limitations. This work is supported by the Natural Science Foundation of Shandong Province (ZR2020MH302 and ZR2021MC068).
Funding
This work was supported by the Natural Science Foundation of Shandong Province (ZR2020MH302 and ZR2021MC068).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Zhiyuan Gong and Chunxia Bi. The first draft of the manuscript was written by Zhiyuan Gong and Wen Liu. Bing Luo provided the funds and was responsible for the revision of the article. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare that they have no conflict of interest.
Informed Consent
There were no human participants in this study.
Research Involving Human and Animal Participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gong, Z., Bi, C., Liu, W. et al. Comprehensive Analysis Based on the TCGA Database Identified SCIN as a Key DNA Methylation-Driver Gene in Epstein-Barr Virus-Associated Gastric Cancer. Biochem Genet (2024). https://doi.org/10.1007/s10528-024-10702-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10528-024-10702-y