Abstract
Ecological studies on marine microbial communities largely focus on fundamental biogeochemical processes or the most abundant constituents, while minor biological fractions are frequently neglected. Youngimonas vesicularis CC-AMW-ET, isolated from coastal surface seawater in Taiwan, is an under-represented marine Paracoccaceae (earlier Rhodobacteraceae) member. The CC-AMW-ET genome was sequenced to gain deeper insights into its role in marine carbon and sulfur cycles. The draft genome (3.7 Mb) contained 63.6% GC, 3773 coding sequences and 51 RNAs, and displayed maximum relatedness (79.06%) to Thalassobius litoralis KU5D5T, a Roseobacteraceae member. While phototrophic genes were absent, genes encoding two distinct subunits of carbon monoxide dehydrogenases (CoxL, BMS/Form II and a novel form III; CoxM and CoxS), and proteins involved in HCO3− uptake and interconversion, and anaplerotic HCO3− fixation were found. In addition, a gene coding for ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO, form II), which fixes atmospheric CO2 was found in CC-AMW-ET. Genes for complete assimilatory sulfate reduction, sulfide oxidation (sulfide:quinone oxidoreductase, SqrA type) and dimethylsulfoniopropionate (DMSP) cleavage (DMSP lyase, DddL) were also identified. Furthermore, genes that degrade aromatic hydrocarbons such as quinate, salicylate, salicylate ester, p-hydroxybenzoate, catechol, gentisate, homogentisate, protocatechuate, 4-hydroxyphenylacetic acid, N-heterocyclic aromatic compounds and aromatic amines were present. Thus, Youngimonas vesicularis CC-AMW-ET is a potential chemolithoautotroph equipped with genetic machinery for the metabolism of aromatics, and predicted to play crucial roles in the biogeochemical cycling of marine carbon and sulfur.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Several members of marine bacterial clades/groups that are numerically important in coastal seawater and sediments have been well characterized for their role in marine carbon and/or sulfur cycle (Swan et al. 2011; Newton et al. 2010; Sorokin 2003; González et al. 1999). Roseobacter genomes include genes dedicated to the oxidation of carbon monoxide, demethylation of dimethylsulfoniopropionate (DMSP), and aromatic compound degradation (Moran et al. 2007). Analysis of culturable representatives in vitro validated the metabolic flexibility of the members of one of the dominant clades (Moran et al. 2004). Nonetheless, the ecological role played by minority bacterial communities inhabiting benthic and pelagic oceans has been often neglected.
The marine bacterium Youngimonas vesicularis CC-AMW-ET, originally classified under the family Rhodobacteraceae, was recently moved along with several other members to the newly established family Paracoccaceae (Liang et al. 2021). While NCBI enlists 96 genera (https://www.ncbi.nlm.nih.gov/datasets/taxonomy/31989/), the List of Prokaryotic names with Standing in Nomenclature (Parte et al. 2020) (LPSN, https://lpsn.dsmz.de/family/paracoccaceae) lists 62 validly published type species under the family Paracoccaceae (https://lpsn.dsmz.de/family/paracoccaceae). Bacterial strains currently classified under Paracoccaceae are widespread in occurrence, and have been isolated from saline terrestrial habitats (Subhash et al. 2013; Wang et al. 2019; Hu et al. 2018), and marine environments such as deep sea (Wei et al. 2023; Kong et al. 2022), shallow sea sediments (Romanenko et al. 2021), seawater (Lim et al. 2008) and sea creatures (Sun et al. 2022; Kim et al. 2021). Few strains were isolated from clinical specimen (Helsel et al. 2007), non-saline aquatic (Li and Zhou 2015) and estuarine (Hameed et al. 2020a, b) habitats. Some strains are inhabitants of extreme environments like sulfidic hydrothermal area (Sorokin et al. 2005), soda lake (Milford et al. 2000) and hotsprings (Albuquerque et al. 2002; Yin et al. 2013). Strains may produce bacteriochlorophyll a (Labrenz et al. 2009; Sorokin et al. 2000), exhibit denitrification (Xu et al. 2021), and metabolize inorganic sulphur/carbon (Hameed et al. 2020a, b; Sorokin et al. 2005; Robertson and Kuenen 1983) and aromatic hydrocarbon (Wang et al. 2019).
At present, the genus Youngimonas accommodates one species (https://lpsn.dsmz.de/species/youngimonas-vesicularis). Similarly, most of the genera that have been classified under Paracoccaceae carry few species. This could be due to the difficulty of growing related strains under laboratory conditions owing to our poor understanding of their ecology and niche (Pohlner et al. 2019). Exploring the ecological functions of this minority population may assist in future taxonomic investigation besides opening new channels for biotechnology and bioremediation. Thus, the genetic makeup of Youngimonas vesicularis CC-AMW-ET was investigated.
Materials and methods
Strain CC-AMW-ET (= JCM 18819 T = BCRC 80549 T) was revived from ‒80 °C and cultured on marine agar (BD Difco 2216) or marine broth (BD Difco 2216) for 48–72 h at 30 °C. Gram staining was performed according to Murray et al. (1994). Fluorescence, transmission electron and scanning electron microscopic analyses were performed as described earlier (Hameed et al. 2020a, b).
Purified genomic DNA was prepared using the Wizard DNA purification kit (Promega), and sonicated (10 µg) using a Misonix 3000 sonicator to obtain DNA fragments of the size 400‒500 bp. The size of the fragments was checked by the Bioanalyzer DNA 1000 chip (Agilent Technologies). Sonicated DNA (1 µg) was end-repaired, A-tailed and adaptor-ligated according to the Illumina TruSeq DNA preparation protocol. Samples were prepared with the MiSeq Reagent Kit v3 (600-cycle) after library construction and loaded onto a MiSeq cartridge. A 2 × 300 bp paired-end sequencing run was performed using the MiSeq platform (Illumina, San Diego, CA, USA). The raw paired-end reads were trimmed and filtered using Trimmomatic (Bolger et al. 2014) to obtain high-quality reads. The SPAdes genome assembler (Bankevich et al. 2012) was used for de novo genome assembly.
Genes of interest were identified using RAST (Aziz et al. 2008). Genomic relatedness was estimated using the Orthologous Average Nucleotide Identity (OrthoANI) application of EzBioCloud (Lee et al. 2016). Amino acid identity (AAI) was calculated using the enveomics collection, available at http://enve-omics.ce.gatech.edu/aai/ (Rodriguez-R and Konstantinidis 2016). An up-to-date bacterial core gene set (UBCG) analysis, which utilizes a set of 92 single-copy core genes (Na et al. 2018), was conducted for CC-AMW-ET. The core genes were extracted from genomes of interest using Prodigal (Hyatt et al. 2010) and hmmsearch (Eddy 2011), aligned using MAFFT (Katoh and Standley 2013) and concatenated into a single alignment. The core gene tree was constructed using FastTree (Price et al. 2010) and RAxML (Stamatakis 2014) through the built-in pipeline, and visualized through MEGA X software. For this analysis and for genome visualization using Proksee (Grant et al. 2023), nine currently available whole genomes of type strains of Rhodobacterales that shared the highest pair-wise 16S rRNA gene sequence similarity were used in addition to CC-AMW-ET genome. The protein identity was verified through UniProt (UniProt 2023). The carbohydrate active enzymes (CAZymes) were identified through the dbCAN2 Meta server (http://cys.bios.niu.edu/dbCAN2/index.php; Zhang et al. 2018). Sulfatases were screened through SulfAtlas (http://sulfatlas.sb-roscoff.fr/; Barbeyron et al. 2016).
Results and discussion
Morphological characteristics and genomic relatedness
Cells of Youngimonas vesicularis CC-AMW-ET were found to be pleomorphic (Fig. 1a–d). This is in line with the phenotype reported in a closely related strain (Iwaki et al. 2013) and some other Rhodobacterales. A circular map showing genomic features of CC-AMW-ET is depicted in Fig. 2a. The draft genome consists of 47 contigs containing 37,95,539 bp, 63.6% GC content, 3773 coding sequences and 51 RNA genes. Genomic relatedness between CC-AMW-ET and other closely related type strains (based on pairwise 16S rRNA gene sequence similarity) of the order Rhodobacterales was investigated through UBCG and orthologous average nucleotide identity (OrthoANI). Phylogenetic tree based on UBCG data (Fig. S1) showed strong phyletic association of CC-AMW-ET with Thalassobius litoralis (formerly Lutimaribacter litoralis), a marine cyclohexylacetate-degrading pleomorphic bacterium affiliated to the family Roseobacteriaceae isolated from coastal seawater of Japan (Iwaki et al. 2013; Hördt et al. 2020). Furthermore, CC-AMW-ET shared highest OrthoANI value (79.06%, Fig. 2b) and AAI value (81%, Fig. S2) with Thalassobius litoralis (Fig. 2b). These data indicated close genetic relatedness of CC-AMW-ET and Thalassobius litoralis.
Micrographs of cells of Youngimonas vesicularis CC-AMW-ET. Light microscopic image of Gram-stained cells (a); epifluorescence microscopic image (b). transmission electron microscopic image of negatively stained cells (c); scanning microscopic image (d); Cells were grown in marine broth (BD Difco 2216) at 30 °C for 24 h under darkness. Scale bar: 5 μm (a and b); 1 μm (c and d); Arrow, vesicle
Genomic features of Youngimonas vesicularis CC-AMW-ET. Circular genome map of Youngimonas vesicularis CC-AMW-ET showing genome features, base composition, and similarity to closely related type strains (a). From center to the outside: GC content, GC skew, RNA genes on the reverse strand, coding sequences on the reverse strand, contigs (alternating colors), coding sequences on the forward strand, RNA genes on the forward strand, BLAST comparisons with closely related type strains of the order Rhodobacterales. The map was generated using the Proksee (Grant et al. 2023). OrthoANI heatmap showing genomic relatedness between CC-AMW-ET and the same type strains (b)
Carbohydrate-active enzymes and sulfatases
Analysis of the CC-AMW-ET genome in dbCAN2 for genes encoding carbohydrate-active enzymes (CAZymes) revealed maximum genes dedicated to glycosyl transferases (GT, n = 42), followed by glycosyl hydrolases (GH, n = 12), auxiliary activities (AA, n = 10) and carbohydrate esterases (CE, n = 4). Genes coding for polysaccharide lyases and carbohydrate-binding modules were missing. Similarly, no significant hits were found for sulfatases. The CAZymes found in CC-AMW-ET (n = 68) were numerically lower as compared to that of Alteromonas fortis 1 T (n = 130), isolated from marine alga (Rekha et al. 2023). While GH predominated in A. fortis 1 T, GT dominated in CC-AMW-ET. Analysis of the genome at SulfAtlas revealed no significant hits for sulfatases. These data indicated poor biopolymer hydrolytic ability of CC-AMW-ET.
Photosynthesis and phototrophy
The CC-AMW-ET genome was screened for signature genes involved in photosynthesis. CC-AMW-ET lacked genes for the photosynthetic reaction centre, bacteriochlorophyll synthesis, light-harvesting complexes, opsin aproprotein and 15,15'-β-carotene dioxygenase (codes for retinal), confirming the absence of both photosynthesis and rhodopsin-based phototrophy that could complement the heterotrophic lifestyle of CC-AMW-ET (Table 1). The absence of genes coding for bacteriochlorophyll synthesis was in line with the UV‒visible spectroscopy (Hameed et al. 2014).
Inorganic carbon concentration, interconversion and metabolism
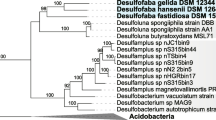
The CC-AMW-ET genome was screened for genes involved in inorganic carbon sequestration. First, genes involved in the metabolism of carbon monoxide (CO), a molecule that participates in a broader range of processes ranging from subcellular to planetary scales (King and Weber 2007), were considered. We found potential genes encoding for a smaller (CoxS, WP_136340385.1), a medium (CoxM, WP_136340383.1) and two larger subunits of CO dehydrogenases (CoxL1 and CoxL2; WP_136340384.1 and WP_136338228.1, respectively) (Table 1). In the UniProt survey, CoxL1 shared the highest amino acid sequence similarity with Actibacterium lipilyticus (90.1%), and formed a tight phylogenetic cluster with BMS/Form II of the CoxL clade in the phylogenetic analysis (Fig. 3). In contrast, CoxL2 formed a separate cluster, distantly associated with BMS/Form II and OMP/Form I. Earlier studies on a subset of nine marine Roseobacter clade (MRC) strains revealed that only MRC strains with both CoxL forms can oxidize CO (Cunliffe 2011). BMS sequences represent functional CODH proteins that are related to but distinct from previously characterized aerobic CODH as evident through a study on Mesorhizobium loti (King 2003). In line with this, the abundance of genes encoding type 1 CODH was used as a marker to quantify soil CO sequestration (Quiza et al. 2014). Thus, CC-AMW-ET is possibly a marine carboxydovore.
Neighbor-joining tree of larger subunits of carbon monoxide dehydrogenase (CoxL) detected in Youngimonas vesicularis CC-AMW-ET (highlighted in bold-phase letters) and other related CoxL homologs. The classification BMS/Form II (green fonts) and OMP/Form I (red fonts) are according to King (2003). Bootstrap values (> 70%) based on 1000 replications are shown at the nodes. The accession number of each sequence is shown in parentheses. The strain name followed by a superscript ‘T’ indicates type strain. Bar, 0.2 substitutions per position. Sequences of xanthine dehydrogenase of CC-AMW-ET (WP_136339391.1) and four additional bacterial strains (A0A1H9GGY6, A0A0P1G2A4, A0A1M4Y7A1, A0A0X3TKM6) were used as an outgroup
The CC-AMW-ET genome was examined for genes involved in HCO3− transport and sequestration. CC-AMW-ET has three copies of the gene encoding BicA (SulP-type Na+-dependent HCO3− transporter) (Table 1). BicA reportedly has a low affinity for the substrate but has a high flux rate (Price et al. 2004). In contrast, the genome lacked a Na+-dependent SbtA type HCO3− transporter that displays a high affinity towards HCO3− (Shibata et al. 2002). Phylogenetic analysis revealed three distinct clusters of CC-AMW-ET BicA (Fig. 4). These HCO3− importer proteins are complemented by a gene coding for monomeric carbonic anhydrase that catalyzes reversible interconversion of CO2 and HCO3− (Guilloton et al. 1992; González et al. 2008). Phylogenetic analysis of carbonic anhydrases showed clustering of CC-AMW-ET within the clade that heterogeneously accommodated carbonic anhydrases from Paracoccaceae and Roseobacteraceae (Fig. S3).
Neighbor-joining tree showing phylogenetic relatedness of SulP-type HCO3− transporters detected in Youngimonas vesicularis CC-AMW-ET (highlighted in bold-phase letters) and other selected bacterial strains. Low-affinity but high flux rate HCO3− transport activity (BicA) has been experimentally demonstrated in a Na+-dependent SulP-type transporter of Synechococcus sp. PCC7002 (*Price et al. 2004). Bootstrap values (> 70%) based on 1000 replications are shown at the nodes. The accession number of each sequence is shown in parentheses. Bar, 0.2 substitutions per position. Photosynthetic cyanobacteria are highlighted in green; Ruegeria pomeroyi, a well-studied Roseobacteraceae member for carbon and sulfur metabolism is shown in blue. Amino acid sequences of Na+-dependent SbtA-type high-affinity HCO3− transporter of Polaribacter sp. MED152 (highlighted in red), Ruegeria pomeroyi and photosynthetic cyanobacteria were used as outgroup
A critical part of CO2 fixation in autotrophs is concentrating carbonate, which could also be an essential step for anaplerotic CO2 fixation in heterotrophs (González et al. 2008). The CC-AMW-ET genome harbored a gene encoding pyruvate carboxylase involved in the ATP-dependent oxaloacetate formation from HCO3− and pyruvate. In addition, CC-AMW-ET also possessed a gene encoding ribulose bisphosphate carboxylase (RuBisCO), involved in atmospheric CO2 fixation directly into organic biomass through the Calvin-Benson-Basham pentose phosphate pathway. Phylogenetic analysis of the protein sequences indicated that CC-AMW-ET RuBisCO belongs to form II reported in the photosynthetic purple non-sulfur bacteria Rhodopseudomonas palustris and R. pentothenatexigens (Fig. 5).
Neighbor-joining tree of RuBisCO detected in Youngimonas vesicularis CC-AMW-ET (highlighted in bold-phase letters) as compared to other RuBisCO homologs. The classification I, II, III and IV are according to Tabita et al. (2008). Phylogenetic positions of ‘Green-like’ and ‘red-like’ RuBisCO (Uchino and Yokota, 2003) affiliated to Form I are highlighted in green and red fonts, respectively. Bootstrap values (> 70%) based on 1000 replications are shown at the nodes. The accession number of each sequence is shown in parentheses. The strain name followed by a superscript ‘T’ indicates the type strain. Bar, 0.2 substitutions per position. The GyrB sequence of Rhodobacter azotoformans IAM 14814 T (BAB83770.1) was used as an outgroup
Sulfur metabolism
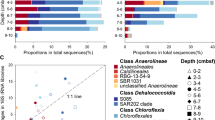
The CC-AMW-ET genome was mined for the genes involved in the metabolism of sulfur-containing osmolyte dimethylsulfoniopropionate (DMSP). CC-AMW-ET harbored a gene encoding DMSP lyase that shared 84.7% sequence similarity with the DddL form of DMSP lyase of Thalassobius litoralis. DddL catalyzes the transformation of DMSP to dimethylsulfide (DMS) (Curson et al. 2008), a climate-changing gas in the ocean. We further evaluated the inorganic sulfur oxidation ability of CC-AMW-ET. A complete set of genes involved in the assimilatory sulfate reduction to sulfide were found in CC-AMW-ET (Table 1). We also found a gene encoding sulfide:quinone oxidoreductase (SQR) that shared 83.4% amino acid similarity with the SQR of Thalassobius teanensis. SQR is essential for photoautotrophic growth on sulfide as determined by the analysis of deletion and interruption strains (Schütz et al. 1999). Bacterial SQR oxidize sulfide during sulfide-dependent chemo- and phototrophic growth (Chan et al. 2009). The detection of genes involved in assimilatory sulfate reduction and sulfide oxidation is in line with Siansivirga zeaxanthinifaciens CC-SAMT-1 T, a marine flavobacterium isolated from coastal seawater (Hameed et al. 2018). Our phylogenetic analysis revealed that the SQR of CC-AMW-ET occupied the SqrA cluster (Fig. 6). Purple non-sulfur bacteria and Cyanobacteria usually harbor SqrA in addition to some Proteobacteria and Aquificaceae (Gregerson et al. 2011). SqrA includes the functionally well-characterized SQRs from Oscillatoria limnetica (Bronstein et al. 2000), Rhodobacter capsulatus (Schütz et al. 1999) and Aquifex aeolicus (Nübel et al. 2000; Marcia et al. 2009).
Neighbor-joining tree of sulfide:quinone oxidoreductase (Sqr) detected in Youngimonas vesicularis CC-AMW-ET (highlighted in bold-phase letters) and other related Sqr homologs. The classification SqrA (red fonts), SqrB, SqrC, SqrD, SqrE, SqrF and SqrX are according to Gregersen et al. (2011). Sequences from phylum Chlorobi (green sulfur bacteria) are shown in green. Bootstrap values (> 70%) based on 1000 replications are shown at the nodes. The accession number of each sequence is shown in parentheses. The strain name followed by a superscript ‘T’ indicates the type strain. Bar, 0.2 substitutions per position. Uncharacterized membrane protein YadS from Chlorobium tepidum TLST (NP_661739.1) was used as an outgroup
Aromatic hydrocarbon metabolism
The genes involved in the aromatic hydrocarbon degradation found in CC-AMW-ET are summarized in Table S1. Key genes dedicated to the degradation of aromatic hydrocarbons such as quinate (3-dehydroquinate dehydratase), salicylate/salicylate ester (salicylate esterase), p-hydroxybenzoate (P-hydroxybenzoate hydroxylase), gentisate (gentisate 1,2-dioxygenase), homogentisate (homogentisate 1,2-dioxygenase), protocatechuate (protocatechuate 3,4-dioxygenase), N-heterocyclic aromatic compounds (isoquinoline 1-oxidoreductase) and aromatic amines (3,4-dihydroxyphenylacetate 2,3-dioxygenase) were found in CC-AMW-ET. These data suggested that Youngimonas vesicularis CC-AMW-ET is capable of metabolizing aromatic hydrocarbons in marine environments.
Conclusion
The presence of genes encoding all subunits of carbon monoxide dehydrogenase (CoxS, CoxM and CoxL), RuBisCO (atmospheric CO2 fixation), HCO3− transporter (BicA), carbonic anhydrase (catalyzes the reversible interconversion of CO2 and HCO3−) and anaplerotic inorganic carbon fixation enzymes (malic enzyme and pyruvate carboxylase) indicates a definite role played by CC-AMW-ET in marine carbon cycling. Similarly, the detection of genes involved in assimilatory sulfate reduction, sulfide oxidation (SqrA) and DMSP metabolism reflects a possible role played by CC-AMW-ET in marine sulfur cycling. Furthermore, the strain harbored genomic signatures for the degradation of xenobiotic aromatic organic compounds besides having the ability to utilize sole organic carbons in vitro (Hameed et al. 2014). Thus, Youngimonas vesicularis CC-AMW-ET is a potential chemolithoautotroph adapted to metabolize inorganic compounds (carbon monoxide, carbon dioxide and sulfide) to complement heterotrophy. Heterotrophic and lithoautotrophic dual-life strategies are likely to assist cells of CC-AMW-ET in copiotrophic coastal waters and oligotrophic open oceans.
Data availability
This whole-genome shotgun project for Youngimonas vesicularis CC-AMW-ET has been deposited at DDBJ/ENA/GenBank under BioProject no. PRJNA531816, and the accession no. is SSMD00000000. The version described in this paper is the first version.
References
Albuquerque L, Santos J, Travassos P, Nobre MF, Rainey FA, Wait R, Empadinhas N, Silva MT, da Costa MS (2002) Albidovulum inexpectatum gen. nov., sp. nov., a nonphotosynthetic and slightly thermophilic bacterium from a marine hot spring that is very closely related to members of the photosynthetic genus Rhodovulum. Appl Environ Microbiol 68:4266–4273. https://doi.org/10.1128/AEM.68.9.4266-4273.2002
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Barbeyron T, Brillet-Guéguen L, Carré W, Carrière C, Caron C, Czjzek M, Hoebeke M, Michel G (2016) Matching the diversity of sulfated biomolecules: creation of a classification database for sulfatases reflecting their substrate specificity. PLoS One 11:e0164846. https://doi.org/10.1371/journal.pone.0164846
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bronstein M, Schütz M, Hauska G, Padan E, Shahak Y (2000) Cyanobacterial sulfide-quinone reductase: cloning and heterologous expression. J Bacteriol 182:3336–3344. https://doi.org/10.1128/jb.182.12.3336-3344.2000
Chan LK, Morgan-Kiss RM, Hanson TE (2009) Functional analysis of three sulfide:quinone oxidoreductase homologs in Chlorobaculum tepidum. J Bacteriol 191:1026–1034. https://doi.org/10.1128/JB.01154-08
Cunliffe M (2011) Correlating carbon monoxide oxidation with cox genes in the abundant Marine Roseobacter Clade. The ISME J 5(4):685–691. https://doi.org/10.1038/ismej.2010.170
Curson AR, Rogers R, Todd JD, Brearley CA, Johnston AW (2008) Molecular genetic analysis of a dimethylsulfoniopropionate lyase that liberates the climate-changing gas dimethylsulfide in several marine alpha-proteobacteria and Rhodobacter sphaeroides. Environ Microbiol 10:757–767. https://doi.org/10.1111/j.1462-2920.2007.01499.x
Eddy SR (2011) Accelerated profile HMM searches. PLoS Comput Biol 7:e10002195. https://doi.org/10.1371/journal.pcbi.1002195
González JM, Kiene RP, Moran MA (1999) Transformation of sulfur compounds by an abundant lineage of marine bacteria in the alpha-subclass of the class Proteobacteria. Appl Environ Microbiol 65:3810–3819. https://doi.org/10.1128/AEM.65.9.3810-3819.1999
González JM, Fernández-Gómez B, Fernàndez-Guerra A, Gómez-Consarnau L, Sánchez O, Coll-Lladó M, Del Campo J, Escudero L, Rodríguez-Martínez R, Alonso-Sáez L, Latasa M, Paulsen I, Nedashkovskaya O, Lekunberri I, Pinhassi J, Pedrós-Alió C (2008) Genome analysis of the proteorhodopsin-containing marine bacterium Polaribacter sp. MED152 (Flavobacteria). Proc Natl Acad Sci U S A 105:8724–8729. https://doi.org/10.1073/pnas.0712027105
Grant JR, Enns E, Marinier E, Mandal A, Herman EK, Chen CY, Graham M, Van Domselaar G, Stothard P (2023) Proksee: in-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. https://doi.org/10.1093/nar/gkad326
Gregersen LH, Bryant DA, Frigaard NU (2011) Mechanisms and evolution of oxidative sulfur metabolism in green sulfur bacteria. Front Microbiol 2:116. https://doi.org/10.3389/fmicb.2011.00116
Guilloton MB, Korte JJ, Lamblin AF, Fuchs JA, Anderson PM (1992) Carbonic anhydrase in Escherichia coli. A product of the cyn operon. J Biol Chem 267:3731–3734. https://doi.org/10.1016/S0021-9258(19)50586-5
Hameed A, Shahina M, Lin SY, Nakayan P, Liu YC, Lai WA, Hsu YH (2014) Youngimonas vesicularis gen. nov., sp nov., of the family Rhodobacteraceae, isolated from surface seawater, reclassification of Donghicola xiamenensis Tan et al. 2009 as Pseudodonghicola xiamenensis gen. nov., comb. nov and emended description of the genus Donghicola Yoon et al. 2007. Int J Syst Evol Microbiol 64:2729–2737. https://doi.org/10.1099/ijs.0.060962-0
Hameed A, Shahina M, Huang HC, Lai WA, Lin SY, Stothard P, Young CC (2018) Complete genome sequence of Siansivirga zeaxanthinifaciens CC-SAMT-1T, a flavobacterium isolated from coastal surface seawater. Mar Genomics 37:21–25. https://doi.org/10.1016/j.margen.2017.09.003
Hameed A, Lai WA, Shahina M, Stothard P, Young LS, Lin SY, Sridhar KR, Young CC (2020) Differential visible spectral influence on carbon metabolism in heterotrophic marine flavobacteria. FEMS Microbiol Ecol. https://doi.org/10.1093/femsec/fiaa011
Hameed A, Shahina M, Lin SY, Chen WM, Hsu YH, Lai WA, Young CC (2020) Description of Gemmobacter aestuarii sp. nov., isolated from estuarine surface water and reclassification of Cereibacter changlensis as Gemmobacter changlensis Chen et al. 2013. Arch Microbiol 202:1035–1042. https://doi.org/10.1007/s00203-020-01809-y
Helsel LO, Hollis D, Steigerwalt AG, Morey RE, Jordan J, Aye T, Radosevic J, Jannat-Khah D, Thiry D, Lonsway DR, Patel JB, Daneshvar MI, Levette PN (2007) Identification of Haematobacter, a new genus of aerobic Gram-negative rods isolated from clinical specimens, and reclassification of Rhodobacter massiliensis as Haematobacter massiliensis comb. nov. J Clin Microbiol 45:1238–1243. https://doi.org/10.1128/jcm.01188-06
Hördt A, López MG, Meier-Kolthoff JP, Schleuning M, Weinhold LM, Tindall BJ, Gronow S, Kyrpides NC, Woyke T, Göker M (2020) Analysis of 1,000+ Type-Strain Genomes Substantially Improves Taxonomic Classification of Alphaproteobacteria. Front Microbiol 11:468. https://doi.org/10.3389/fmicb.2020.00468
Hu Q, Zhang L, Hang P, Zhou XY, Jia WB, Li SP, Jiang JD (2018) Xinfangfangia soli gen. nov., sp. nov., isolated from a diuron-polluted soil. Int J Syst Evol Microbiol 68:2622–2626. https://doi.org/10.1099/ijsem.0.002887
Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119. https://doi.org/10.1186/1471-2105-11-119
Iwaki H, Yasukawa N, Fujioka M, Takada K, Hasegawa Y (2013) Isolation and characterization of a marine cyclohexylacetate-degrading bacterium Lutimaribacter litoralis sp. nov., and reclassification of Oceanicola pacificus as Lutimaribacter pacificus comb. nov. Curr Microbiol 66:588–593. https://doi.org/10.1007/s00284-013-0321-x
Katoh K, Standley DM (2013) MAFFT Multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kim YO, Noh JK, Kim DG, Park IS, Park S, Yoon JH (2021) Paenihalocynthiibacter styelae gen. nov., sp. nov., isolated from stalked sea squirt Styela clava. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.005085
King GM (2003) Molecular and culture-based analyses of aerobic carbon monoxide oxidizer diversity. Appl Environ Microbiol 69:7257–7265. https://doi.org/10.1128/aem.69.12.7257-7265.2003
King GM, Weber CF (2007) Distribution, diversity and ecology of aerobic CO-oxidizing bacteria. Nat Rev Microbiol 5:107–118. https://doi.org/10.1038/nrmicro1595
Kong YH, Ren WT, Xu L, Cheng H, Zhou P, Wang CS, Wu YH, Xu XW (2022) Mesobacterium pallidum gen. nov., sp. nov., Heliomarina baculiformis gen. nov., sp. nov. and Oricola indica sp. nov., three novel Alphaproteobacteria members isolated from deep-sea water in the southwest Indian ridge. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.005236
Labrenz M, Lawson PA, Tindall BJ, Hirsch P (2009) Roseibaca ekhonensis gen. nov., sp. nov., an alkalitolerant and aerobic bacteriochlorophyll a-producing alphaproteobacterium from hypersaline Ekho Lake. Int J Syst Evol Microbiol 59:1935–1940. https://doi.org/10.1099/ijs.0.016717-0
Lee I, Ouk Kim Y, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Li AH, Zhou YG (2015) Frigidibacter albus gen. nov., sp. nov., a novel member of the family Rhodobacteraceae isolated from lake water. Int J Syst Evol Microbiol 65:1199–1206. https://doi.org/10.1099/ijs.0.000080
Liang KYH, Orata FD, Boucher YF, Case RJ (2021) Roseobacters in a Sea of Poly- and Paraphyly: Whole Genome-Based Taxonomy of the Family Rhodobacteraceae and the Proposal for the Split of the Roseobacter Clade Into a Novel Family Roseobacteraceae fam. nov. Front Microbiol 12:683109. https://doi.org/10.3389/fmicb.2021.683109
Lim JM, Jeon CO, Jang HH, Park DJ, Shin YK, Yeo SH, Kim CJ (2008) Albimonas donghaensis gen. nov., sp. nov., a non-photosynthetic member of the class Alphaproteobacteria isolated from seawater. Int J Syst Evol Microbiol 58:282–285. https://doi.org/10.1099/ijs.0.65429-0
Marcia M, Ermler U, Peng G, Michel H (2009) The structure of Aquifex aeolicus sulfide:quinone oxidoreductase, a basis to understand sulfide detoxification and respiration. Proc Natl Acad Sci U S A 106:9625–9630. https://doi.org/10.1073/pnas.09041651064
Milford AD, Achenbach LA, Jung DO, Madigan MT (2000) Rhodobaca bogoriensis gen. nov. and sp. nov., an alkaliphilic purple nonsulfur bacterium from African Rift Valley soda lakes. Arch Microbiol 174:18–27. https://doi.org/10.1007/s002030000166
Moran MA, Buchan A, González JM, Heidelberg JF, Whitman WB, Kiene RP, Henriksen JR, King GM, Belas R, Fuqua C, Brinkac L, Lewis M, Johri S, Weaver B, Pai G, Eisen JA, Rahe E, Sheldon WM, Ye WY, Miller TR, Carlton J, Rasko DA, Paulsen IT, Ren QH, Daugherty SC, Deboy RT, Dodson RJ, Durkin AS, Madupu R, Nelson WC, Sullivan SA, Rosovitz MJ, Haft DH, Selengut J, Ward N (2004) Genome sequence of Silicibacter pomeroyi reveals adaptations to the marine environment. Nature 432:910–913. https://doi.org/10.1038/nature03170
Moran MA, Belas R, Schell MA, González JM, Sun F, Sun S, Binder BJ, Edmonds J, Ye W, Orcutt B, Howard EC, Meile C, Palefsky W, Goesmann A, Ren Q, Paulsen I, Ulrich LE, Thompson LS, Saunders E, Buchan A (2007) Ecological genomics of marine roseobacters. Appl Environ Microbiol 73:4559–4569. https://doi.org/10.1128/Aem.02580-06
Murray RGE, Doetsch RN, Robinow CF (1994) Determinative and cytological light microscopy. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for General and Molecular Bacteriology. American Society for Microbiology, Washington, DC, pp 21–41
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: Up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285. https://doi.org/10.1007/s12275-018-8014-6
Newton RJ, Griffin LE, Bowles KM, Meile C, Gifford S, Givens CE, Howard EC, King E, Oakley CA, Reisch CR, Rinta-Kanto JM, Sharma S, Sun SL, Varaljay V, Vila-Costa M, Westrich JR, Moran MA (2010) Genome characteristics of a generalist marine bacterial lineage. ISME J 4:784–798. https://doi.org/10.1038/ismej.2009.150
Nübel T, Klughammer C, Huber R, Hauska G, Schütz M (2000) Sulfide:quinone oxidoreductase in membranes of the hyperthermophilic bacterium Aquifex aeolicus (VF5). Arch Microbiol 173:233–244. https://doi.org/10.1007/s002030000135
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Pohlner M, Dlugosch L, Wemheuer B, Mills H, Engelen B, Reese BK (2019) The majority of active Rhodobacteraceae in marine sediments belong to uncultured genera: a molecular approach to link their distribution to environmental conditions. Front Microbiol 10:659. https://doi.org/10.3389/fmicb.2019.00659
Price GD, Woodger FJ, Badger MR, Howitt SM, Tucker L (2004) Identification of a SulP-type bicarbonate transporter in marine cyanobacteria. Proc Natl Acad Sci U S A 101:18228–18233. https://doi.org/10.1073/pnas.0405211101
Price MN, Dehal PS, Arkin AP (2010) FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS One 5:e9490. https://doi.org/10.1371/journal.pone.0009490
Quiza L, Lalonde I, Guertin C, Constant P (2014) Land-use influences the distribution and activity of high affinity CO-oxidizing bacteria associated to type l-coxL genotype in soil. Front Microbiol. https://doi.org/10.3389/Fmicb.2014.00271
Rekha PD, Shastry RP, Hameed A, Ghate SD, Arun AB, Athmika N (2023) Genomic potential for exopolysaccharide production and differential polysaccharide degradation in closely related Alteromonas sp. PRIM-21 and Alteromonas fortis 1T. Antonie Van Leeuwenhoek 116:39–51. https://doi.org/10.1007/s10482-022-01796-8
Robertson LA, Kuenen JG (1983) Thiosphaera pantotropha gen. nov., sp. nov., a facultatively anaerobic, facultatively autotrophic sulphur bacterium. J Gen Microbiol 129:2847–2855. https://doi.org/10.1099/00221287-129-9-2847
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Preprints 4:e1900v1. https://doi.org/10.7287/peerj.preprints.1900v1
Romanenko LA, Kurilenko VV, Chernysheva NY, Tekutyeva LA, Velansky PV, Svetashev VI, Isaeva MP (2021) Harenicola maris gen. nov., sp. nov. isolated from the Sea of Japan shallow sediments. Arch Microbiol 203:3973–3979. https://doi.org/10.1007/s00203-021-02360-0
Schütz M, Maldener I, Griesbeck C, Hauska G (1999) Sulfide-quinone reductase from Rhodobacter capsulatus: requirement for growth, periplasmic localization, and extension of gene sequence analysis. J Bacteriol 181:6516–6523. https://doi.org/10.1128/JB.181.20.6516-6523.1999
Shibata M, Katoh H, Sonoda M, Ohkawa H, Shimoyama M, Fukuzawa H, Kaplan A, Ogawa T (2002) Genes essential to sodium-dependent bicarbonate transport in cyanobacteria - Function and phylogenetic analysis. J Biol Chem 277:18658–18664. https://doi.org/10.1074/jbc.M112468200
Sorokin DY (2003) Oxidation of inorganic sulfur compounds by obligately organotrophic bacteria. Microbiology 72:641–653. https://doi.org/10.1023/B:Mici.0000008363.24128.E5
Sorokin DY, Tourova TP, Kuznetsov BB, Briantseva IA, Gorlenko VM (2000) Roseinatronobacter thiooxidans gen. nov., sp. nov., a new alkaliphilic aerobic bacteriochlorophyll-alpha-containing bacteria from a soda lake. Mikrobiologiia 69:89–97. https://doi.org/10.1007/BF02757261
Sorokin DY, Tourova TP, Spiridonova EM, Rainey FA, Muyzer G (2005) Thioclava pacifica gen. nov., sp. nov., a novel facultatively autotrophic, marine, sulfur-oxidizing bacterium from a near-shore sulfidic hydrothermal area. Int J Syst Evol Microbiol 55:1069–1075. https://doi.org/10.1099/ijs.0.63415-0
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Subhash Y, Tushar L, Sasikala Ch (2013) Falsirhodobacter halotolerans gen. nov., sp. nov., isolated from dry soils of a solar saltern. Int J Syst Evol Microbiol 63:2132–2137. https://doi.org/10.1099/ijs.0.044107-0
Sun H, Zheng H, Liao B, Chen B, Li A, Xiao B (2022) Algiphilus acroporae sp. nov. and Coraliihabitans acroporae gen. nov. sp. nov., isolated from scleractinian coral Acropora digitifera. Int J Syst Evol Microbiol 272:5321. https://doi.org/10.1099/ijsem.0.005321
Swan BK, Martinez-Garcia M, Preston CM, Sczyrba A, Woyke T, Lamy D, Reinthaler T, Poulton NJ, Masland EDP, Gomez ML, Sieracki ME, DeLong EF, Herndl GJ, Stepanauskas R (2011) Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the dark ocean. Science 333:1296–1300. https://doi.org/10.1126/science.1203690
Tabita FR, Satagopan S, Hanson TE, Kreel NE, Scott SS (2008) Distinct form I, II, III, and IV Rubisco proteins from the three kingdoms of life provide clues about Rubisco evolution and structure/function relationships. J Exp Bot 59:1515‒24. https://doi.org/10.1093/jxb/erm361
Uchino Y, Yokota A (2003) “Green-like” and “red-like” RubisCO cbbL genes in Rhodobacter azotoformans. Mol Biol Evol 20:821–830. https://doi.org/10.1093/molbev/msg100
UniProt C (2023) UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res 51:D523–D531. https://doi.org/10.1093/nar/gkac1052
Wang S, Yang R, Xu L, Xing YT, Sun JQ (2019) Qingshengfaniella alkalisoli gen. nov., sp. nov., a p-hydroxybenzoate-degrading strain isolated from saline soil. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.004719
Wei TT, Fan XB, Quan ZX (2023) Abyssibius alkaniclasticus gen. nov., sp. nov., a novel member of the family Rhodobacteraceae, isolated from the Mariana Trench. Int J Syst Evol Microbiol 73:5715. https://doi.org/10.1099/ijsem.0.005715
Xu L, Liu A, Zhang YJ (2021) Zongyanglinia huanghaiensis gen. nov., sp. nov., a novel denitrifying bacterium isolated from the yellow sea, and transfer of Pelagicola marinus to Zongyanglinia gen. nov. as Zongyanglinia marinus comb. nov. Antonie Van Leeuwenhoek 114:137–149. https://doi.org/10.1007/s10482-020-01507-1
Yin D, Chen L, Ao J, Ai C, Chen X (2013) Pleomorphobacterium xiamenense gen. nov., sp. nov., a moderate thermophile isolated from a terrestrial hot spring. Int J Syst Evol Microbiol 63:1868–1873. https://doi.org/10.1099/ijs.0.042713-0
Zhang H, Yohe T, Huang L, Entwistle S, Wu P, Yang Z, Busk PK, Xu Y, Yin Y (2018) dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 46:W95–W101. https://doi.org/10.1093/nar/gky418
Acknowledgements
Authors would like to thank Yu-Ting Hseih for her technical assistance.
Funding
This work was financially supported by the National Science and Technology Council (Taiwan) under Grant No. 111–2313-B-005–050 and by the “Innovation and Development Center of Sustainable Agriculture” from The Featured Areas Research Center Program within the framework of the Higher Education Sprout Project by the Ministry of Education (MOE) in Taiwan.
Author information
Authors and Affiliations
Contributions
CCY and AH conceptualized the work. AH drafted the manuscript. AH, SKV and PS performed genomic data mining, annotation and comparative genomics. AH analyzed the data and restructured the manuscript with scientific input from all the authors. SYL performed AAI, UBCG and microscopic analysis. All the authors discussed the results and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare no competing interests.
Ethical approval
None.
Informed consent
None.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hameed, A., Suchithra, K.V., Lin, SY. et al. Genomic potential for inorganic carbon sequestration and xenobiotic degradation in marine bacterium Youngimonas vesicularis CC-AMW-ET affiliated to family Paracoccaceae. Antonie van Leeuwenhoek 116, 1247–1259 (2023). https://doi.org/10.1007/s10482-023-01881-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-023-01881-6